Abstract
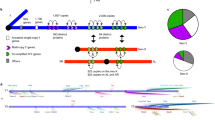
IT is commonly believed1,2 that the mutation rate is much higher in the human male germ line than in the female germ line because the number of germ-cell divisions per generation is much larger in males than in females. But direct estimation of mutation rates is difficult, relying mainly on sex-linked genetic diseases3, so the ratio (αm) of male to female mutation rates is not clear. It has been noted4 that if αm is very large, then the rate of synonymous substitution in X-linked genes should be only 2/3 of that in autosomal genes, and comparison of human and rodent genes supported this prediction4. As the number of X-linked genes used in the study was small and the X-linked and autosomal sequences were non-homologous, and given that the synonymous rate varies among genes5, we sequenced the last intron (~1 kb) of the Y-linked and X-linked zinc-finger-protein genes (ZFY and ZFX) in humans, orang-utans, baboons and squirrel monkeys. The ratio Y/X of the substitution rate in the Y-linked intron to that in the X-linked intron is ~2.3, which is close to that estimated from synonymous rates in the ZFY and ZFX genes6–8 and implies αm≈6. This estimate of αm supports the view that the evolution of DNA sequences in higher primates is male-driven. It is, however, much lower than the previous estimate4 and therefore raises a number of issues.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Haldane, J. B. S. Ann. Eugen. 13, 262–271 (1947).
Penrose, L. S. Lancet II, 312 (1955).
Vogel, F. & Motulsky, A. G. Human Genetics (Springer, Berlin and Heidelberg, Germany, 1986).
Miyata, T., Hayashida, H., Kuma, K., Mitsuyasa, K. & Yasunaga, T. Cold Spring Harbor Symp. quant. Biol. 52, 863–867 (1987).
Li, W.-H. & Graur, D. Fundamentals of Molecular Evolution (Sinauer, Sunderland, MA, 1991).
Lanfear, J. & Holland, P. W. H. J. molec. Evol. 32, 310–315 (1991).
Hayashida, H., Kuma, K. & Miyata, T. J. molec. Evol. 35, 181–183 (1992).
Pamilo, P. & Bianchi, N. O. Molec. Biol. Evol. (in the press).
Bulmer, M. Molec. Biol. Evol. 3, 322–329 (1986).
Winter, R. M., Tuddenham, E. G. D., Goldman, E. & Matthews, K. B. Hum. Genet. 64, 156–159 (1983).
Kohne, D. E. Q. Rev. Biophys. 33, 1–48 (1970).
Tajima, F. & Nei, M. Molec. Biol. Evol. 1, 269–285 (1984).
Schneider-Gadicke, A. et al. Nature 342, 708–711 (1989).
Hattori, M. & Sakaki, Y. Analyt. Biochem. 152, 232–238 (1986).
Sanger, F., Nicklen, S. & Coulsen, A. R. Proc. natn. Acad. Sci. U.S.A. 74, 5463–5467 (1977).
Mizusawa, S., Nishimura, S. & Seela, F. Nucleic Acids Res. 14, 1319–1324 (1986).
Kusukawa, N., Uemori, T., Asada, K. & Kato, I. BioTechniques 9, 66–72 (1990).
Higgins, D. G. & Sharp, P. M. Computer Appl. Biosci. 5, 151–153 (1989).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Shimmin, L., Chang, BJ. & Li, WH. Male-driven evolution of DNA sequences. Nature 362, 745–747 (1993). https://doi.org/10.1038/362745a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/362745a0
This article is cited by
-
Pedigree-based and phylogenetic methods support surprising patterns of mutation rate and spectrum in the gray mouse lemur
Heredity (2021)
-
Rapid functional divergence after small-scale gene duplication in grasses
BMC Evolutionary Biology (2019)
-
The rate of human mutation
Nature (2012)
-
Is the Y chromosome disappearing?—Both sides of the argument
Chromosome Research (2012)
-
Strict evolutionary conservation followed rapid gene loss on human and rhesus Y chromosomes
Nature (2012)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.