Abstract
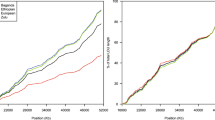
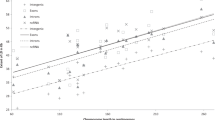
Linkage disequilibrium (LD), or the non-random association of alleles, is poorly understood in the human genome1. Population genetic theory suggests that LD is determined by the age of the markers, population history, recombination rate, selection and genetic drift2. Despite the uncertainties in determining the relative contributions of these factors, some groups have argued that LD is a simple function of distance between markers3,4. Disease-gene mapping studies and a simulation study gave differing predictions on the degree of LD in isolated and general populations5,6. In view of the discrepancies between theory and experimental observations, we constructed a high-density SNP map of the Xq25–Xq28 region7 and analysed the male genotypes and haplotypes across this region for LD in three populations. The populations included an outbred European sample (CEPH males) and isolated population samples from Finland and Sardinia. We found two extended regions of strong LD bracketed by regions with no evidence for LD in all three samples. Haplotype analysis showed a paucity of haplotypes in regions of strong LD. Our results suggest that, in this region of the X chromosome, LD is not a monotonic function of the distance between markers, but is more a property of the particular location in the human genome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Chakravarti, A. Population genetics—making sense out of sequence. Nature Genet. 21 (suppl.), 56–60 (1999).
Hartl, D.L. & Clark, A.G. Principles of Population Genetics (Sinauer Associates, Sunderland, 1989).
Watkins W.S. et al. Linkage disequilibrium patterns vary with chromosomal location: a case study from the von Willebrand factor region. Am. J. Hum. Genet. 55, 348–355 (1994).
Jorde, L.B. et al. Linkage disequilibrium predicts physical distance in the adenomatous polyposis coli region. Am. J. Hum. Genet. 54, 884–898 (1994).
Kruglyak, L. Prospects for whole-genome genome linkage disequilibrium mapping of common disease genes. Nature Genet. 22, 139–144 (1999).
Jorde, L.B., Watkins, W.S., Viskochil, D., O'Connell, P. & Ward, K. Linkage disequilibrium in the neurofibromatosis 1 (NF1) region: implications for gene mapping. Am. J. Hum. Genet. 53, 1038–1050 (1993).
Taillon-Miller, P. & Kwok, P.-Y. A high density single nucleotide polymorphism map of Xq25–Xq28. Genomics 65, 195–202 (2000).
Kere, J. et al. Cystic fibrosis in a low-incidence population: two major mutations in Finland. Hum. Genet. 93, 162–166 (1994).
Tahvanainen, E. et al. The gene for a recessively inherited human childhood progressive epilepsy with mental retardation maps to the distal short arm of chromosome 8. Proc. Natl Acad. Sci. USA 91, 7267–7270 (1994).
Höglund, P. et al. Fine mapping of the congenital chloride diarrhea gene by linkage disequilibrium. Am. J. Hum. Genet. 57, 95–102 (1995).
Huttley, G.A., Smith, M.W., Carrington, M. & O'Brien, S.J. A scan for linkage disequilibrium across the human genome. Genetics 152, 1711–1722 (1999).
Taillon-Miller, P., Piernot, E.E. & Kwok, P.-Y. Efficient approach to unique single nucleotide polymorphism discovery. Genome Res. 9, 499–505 (1999).
Taillon-Miller, P. et al. The homozygous complete hydatidiform mole: a unique resource for genome studies. Genomics 46, 307–310 (1997).
Kwok, P.-Y., Carlson, C., Yager, T., Ankener, W. & Nickerson, D.A. Comparative analysis of human DNA variations by fluorescence-based sequencing of PCR products. Genomics 23, 138–144 (1994).
Chen, X., Zehnbauer, B., Gnirke, A. & Kwok, P.-Y. Fluorescence energy transfer detection as a homogeneous DNA diagnostic method. Proc. Natl Acad. Sci. USA 94, 10756–10761 (1997).
Kwok, P.-Y., Gremaud, M.F., Nickerson, D.A., Hood, L. & Olson, M.V. Automatable screening of yeast artificial-chromosome libraries based on the oligonucleotide-ligation assay. Genomics 13, 935–941 (1992).
Laitinen, T. et al. Genetic control of serum IgE levels and asthma: linkage and linkage disequilibrium studies in an isolated population. Hum. Mol. Genet. 6, 2069–2076 (1997).
Cavalli-Sforza, L.L., Menozzi, P. & Piazza, A. Demic expansions and human evolution. Science 259, 639–646 (1993).
Cao, A., Galanello, R. & Rosatelli, M.C. Genotype-phenotype correlations in β-thalassemias. Blood Rev. 8, 1–12 (1994).
Hedrick, P.W. Gametic disequilibrium measures: proceed with caution. Genetics 117, 331–341 (1987).
Lewontin, R.C. On measures of gametic disequilibrium. Genetics 120, 849–852 (1988).
Acknowledgements
We thank D. Schlessinger for encouragement and support; R.D. Miller for discussions; and P. Buzby and S. Spurgeon for reagents. This work was supported in part by grants from the National Human Genome Research Institute (RO1HG1439, P50HG00201, P50HG00835) and by MH37685 and MH17104.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Taillon-Miller, P., Bauer-Sardiña, I., Saccone, N. et al. Juxtaposed regions of extensive and minimal linkage disequilibrium in human Xq25 and Xq28. Nat Genet 25, 324–328 (2000). https://doi.org/10.1038/77100
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/77100
This article is cited by
-
Combined effects of multiple linked loci on pairwise sibling tests
International Journal of Legal Medicine (2017)
-
Genetic structure of the Newfoundland and Labrador population: founder effects modulate variability
European Journal of Human Genetics (2016)
-
Vascular endothelial growth factor (VEGF) gene polymorphisms and breast cancer risk in Punjabi population from North West India
Tumor Biology (2014)
-
Genetic mapping of high caries experience on human chromosome 13
BMC Medical Genetics (2013)
-
Population-genetic comparison of the Sorbian isolate population in Germany with the German KORA population using genome-wide SNP arrays
BMC Genetics (2011)