Abstract
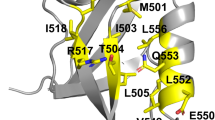
PROTEIN secondary structures have been viewed as fundamental building blocks for protein folding, structure and design. Pre-vious studies indicate that the propensities of individual amino acids to form particular secondary structures are the result of a combination of local conformational preferences1,2 and non-local factors3–7. To examine the extent to which non-local factors influence the formation of secondary structural elements, we have designed an 11-amino-acid sequence (dubbed the 'chameleon' sequence) that folds as an α-helix when in one position but as a β-sheet when in another position of the primary sequence of the IgG-binding domain of protein G (GUI). Both proteins, chameleon-α and chameleon-β, are folded into structures similar to native GB1, as judged by several biophysical criteria. Our results demonstrate that non-local interactions can determine the secondary structure of peptide sequences of substantial length. They also support views of protein folding that favour tertiary interactions as dominant determinants of structure (for example, see refs 8,9).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Chakrabartty, A. & Baldwin, R. L. Adv. Prat. Chem. 46, 141–176 (1995).
Bryson, J. W. et al. Science 270, 935–941 (1995).
Garratt, R. C., Taylor, W. R. & Thornton, J. M. FEBS Lett. 188, 59–62 (1985).
Heinz, D. W., Baase, W. A., Dahlquist, F. W. & Matthews, B. W. Nature 361, 561–564 (1993).
Minor, D. L. Jr & Kim, P. S. Nature 371, 264–267 (1994).
Xiong, H., Buckwalter, B. L., Shieh, H.-M. & Hecht, M. H. Proc. natn. Acad. Sci. U.S.A. 92, 6349–6353 (1995).
Smith, C. K. & Regan, L. Science 270, 980–982 (1995).
Dill, K. A. et al. Prot. Sci. 4, 561–602 (1995).
Lumb, K. J., Carr, C. M. & Kim, P. S. Biochemistry 33, 7361–7367 (1994).
Privalov, P. L. A. Rev. Biophys. biophys. Chem. 18, 47–69 (1989).
Lumb, K. J. & Kim, P. S. Biochemistry 34, 8642–8648 (1995).
Gronenborn, A. M. & Clore, G. M. J. molec. Biol. 223, 331–335 (1993).
Dyson, H. J. & Wright, P. E. A. Rev. Biophys. biophys. Chem. 20, 519–538 (1991).
Carr, C. M. & Kim, P. S. Cell 73, 823–832 (1993).
Bullough, P. A., Hughson, F. M., Skehel, J. J. & Wiley, D. C. Nature 371, 37–43 (1994).
Goldsmith, E. J. & Mottonen, J. Structure 2, 241–244 (1994).
Kabsch, W. & Sander, C. Proc. natn. Acad. Sci. U.S.A. 81, 1075–1078 (1984).
Wilson, I. A., Haft, D. H., Getzoff, E. D., Tainer, J. A. & Lerner, R. A. Proc. natn. Acad. Sci. U.S.A. 82, 5255–5259 (1985).
Argos, P. J. molec. Biol. 197, 331–348 (1987).
Cohen, B. I., Presnell, S. R. & Cohen, F. E. Prot. Sci. 2, 2134–2145 (1993).
Gronenborn, A. M. et al. Science 253, 657–661 (1991).
Minor, D. L. Jr & Kim, P. S. Nature 367, 660–663 (1994).
Achari, A. et al. Biochemistry 31, 10449–10457 (1991).
Alexander, P., Fahnestock, S., Lee, T., Orban, J. & Bryan, P. Biochemistry 31, 3597–3603 (1992).
Wüthrich, K. NMR of Proteins and Nucleic Acids (Wiley, New York, 1986).
Bai, Y., Milne, J. S., Mayne, L & Englander, S. W. Proteins 17, 75–86 (1993).
Orban, J., Alexander, P., Bryan, P. & Khare, D. Biochemistry 34, 15291–15300 (1995).
Fahnestock, S. R., Alexander, P., Filpula, D. & Nagle, J. in Bacterial immunoglobin-binding Proteins (ed. Boyle. M. D. P.) (Academic, San Diego, 1990).
Woody, R. W. in The Peptides (Academic, New York, 1985).
Chou, P. Y. & Fasman, G. D. Biochemistry 13, 211–222 (1973).
Schuster, T. M. & Laue, T. M. Modern Analytical Ultracentrifugation (Birkhäuser, Boston, 1994).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Minor, D., Kim, P. Context-dependent secondary structure formation of a designed protein sequence. Nature 380, 730–734 (1996). https://doi.org/10.1038/380730a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/380730a0
This article is cited by
-
A Monist Proposal: Against Integrative Pluralism About Protein Structure
Erkenntnis (2022)
-
De novo design of modular and tunable protein biosensors
Nature (2021)
-
Auxiliary interfaces support the evolution of specific toxin–antitoxin pairing
Nature Chemical Biology (2021)
-
Environment-transformable sequence–structure relationship: a general mechanism for proteotoxicity
Biophysical Reviews (2018)
-
Dihedral angle preferences of amino acid residues forming various non-local interactions in proteins
Journal of Biological Physics (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.