Abstract
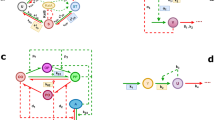
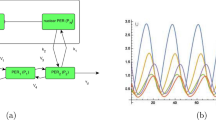
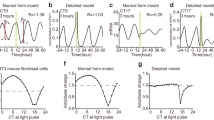
Circadian rhythms, which provide internal daily periodicity, are used by a wide range of organisms to anticipate daily changes in the environment1. It seems that these organisms generate circadian periodicity by similar biochemical networks within a single cell2. A model based on the common features of these biochemical networks shows that a circadian network can oscillate reliably in the presence of stochastic biochemical noise and when cellular conditions are altered. We propose that the ability to resist such perturbations imposes strict constraints on the oscillation mechanisms underlying circadian periodicity in vivo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Edmund, L.N. Cellular and Molecular Basis of Biological Clocks (Springer, New York, 1988).
Dunlap, J. Cell 96, 271–290 (1999).
Suri, V., Lanjuin, A. & Rosbash, M. EMBO J. 18, 675–686 (1999).
Goldbeter, A. Biochemical Oscillations and Cellular Rhythms (Cambridge Univ. Press, 1995).
Murray, J. D. Mathematical Biology (Springer, Berlin, 1994).
Gillespie, D. T. J. Phys. Chem. 81, 2340–2361 (1977).
Ko, M. S. Bioessays 14, 341 (1992).
McAdams, H. N. & Arkin, A. Trends. Genet. 15, 65–69 (1999).
Wodicka, L. et al. Nature Biotechnol. 15, 1359–1367 (1997).
Leloup, J. C. & Goldbeter, A. J. Biol. Rhythms 13, 70–87 (1998).
Kondo, T. et al. Science 275, 224–227 (1997).
Glossop, N. R. J., Lyons, L. C. & Hardin, P. E. Science 286, 766–768 (1999).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Barkai, N., Leibler, S. Circadian clocks limited by noise. Nature 403, 267–268 (2000). https://doi.org/10.1038/35002258
Issue Date:
DOI: https://doi.org/10.1038/35002258
This article is cited by
-
Quorum sensing for population-level control of bacteria and potential therapeutic applications
Cellular and Molecular Life Sciences (2020)
-
Noise control and utility: From regulatory network to spatial patterning
Science China Mathematics (2020)
-
Parameter estimation in models of biological oscillators: an automated regularised estimation approach
BMC Bioinformatics (2019)
-
Stochastic Simulation of Pattern Formation in Growing Tissue: A Multilevel Approach
Bulletin of Mathematical Biology (2019)
-
Algebraic expressions of conditional expectations in gene regulatory networks
Journal of Mathematical Biology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.