Abstract
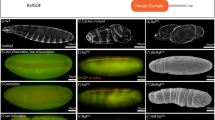
The Krüppel (Kr) locus is a member of the ‘gap’ class of segmentation genes of Drosophila melanogaster. Mutations at the Kr locus cause the deletion of contiguous segments from the embryonic body pattern. We have elucidated the spatial and temporal characteristics of Kr gene expression during early embryo development, the localization of cytoplasmic Kr+ activity and its spatial requirement for normal segmentation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Wieschaus, E. & Gehring, W. J. Devl Biol. 50, 249–263 (1976).
Schubiger, G. Devl Biol. 50, 476–488 (1976).
Szabad, J., Schüpbach, T. & Wieschaus, E. Devl Biol. 73, 265–271 (1979).
Wieschaus, E., Nüsslein-Volhard, C. & Jürgens, G. Wilhelm Roux Arch dev. Biol. 193, 296–307 (1984).
Nüsslein-Volhard, C., Wieschaus, E. & Kluding, H. Wilhelm Roux Arch dev. Biol. 193, 267–282 (1984).
Jürgens, G., Kluding, H., Nüsslein-Volhard, C. & Wieschaus, E. Wilhelm Roux Arch dev. Biol. 193, 283–295 (1984).
Nüsslein-Volhard, C. & Wieschaus, E. Nature 287, 795–801 (1980).
Preiss, A., Rosenberg, U. B., Kienlin, A., Seifert, E. & Jäckle, H. Nature 313, 27–32 (1985).
Gloor, H. Archs Julius Klaus-Stift. 25, 38–44 (1950).
Wieschaus, E., Nüsslein-Volhard, C. & Kluding, H. Devl Biol. 104, 172–186 (1984).
Rosenberg, U. B., Preiss, A., Seifert, E., Jäckle, H. & Knipple, D. C. Nature 313, 703–706 (1985).
Hafen, E., Levine, M., Garber, R. L. & Gehring, W. J. EMBO J. 2, 617–623 (1983).
Akam, M. E. EMBO J. 2, 2075–2084 (1983).
Foe, V. E. & Alberts, B. M. J. Cell Sci. 61, 31–70 (1983).
Fullilove, S. L. & Jacobson, A. G. C. in The Genetics and Biology of Drosophila (eds Ashburner, M. & Wright, T. R. F.) 106–277 (Academic, New York, 1978).
Turner, F. R. & Mahowald, A. P. Devl Biol. 57, 403–416 (1977).
Poulson, D. F. in Biology of Drosophila (ed. Demerec, M.) 168–174 (Wiley, New York, 1950).
Hartenstein, V., Technau, G. M. & Campos-Ortéga, J. A. Wilhelm Roux Arch dev. Biol. 194, 213–216 (1985).
Hafen, E., Kuroiwa, A. & Gehring, W. J. Cell 37, 833–841 (1984).
Fjose, A., McGinnis, W. J. & Gehring, W. J. Nature 313, 284–299 (1985).
Kornberg, T., Sidén, I., O'Farrell, P. O. & Simon, M. Cell 40, 45–53 (1985).
Lohs-Schardin, M., Cremer, C. & Nüsslein-Volhard, C. Devl Biol. 73, 239–255 (1979).
Meinhardt, H. J. Cell Sci. 23, 117–139 (1977).
Anderson, K. V. & Lengyel, J. A. Devl Biol. 70, 217–231 (1979).
McKnight, S. L. & Miller, O. L. Cell 8, 305–319 (1976).
Akam, M. & Martinez-Arias, A. EMBO J. (in the press).
Cox, K. H., DeLeon, D. V., Angerer, L. M. & Anferer, R. C. Devl Biol. 101, 485–502 (1984).
Gall, J. G. & Pardue, M. L. Meth. Enzym. 21, 470–480 (1971).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Knipple, D., Seifert, E., Rosenberg, U. et al. Spatial and temporal patterns of Krüppel gene expression in early Drosophila embryos. Nature 317, 40–44 (1985). https://doi.org/10.1038/317040a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/317040a0
This article is cited by
-
Bayesian model selection for the Drosophila gap gene network
BMC Bioinformatics (2019)
-
Improved readout precision of the Bicoid morphogen gradient by early decoding
Journal of Biological Physics (2012)
-
The gap gene network
Cellular and Molecular Life Sciences (2011)
-
Alterations in the regulation of expression of the αl(I) collagen gene (COL1A1) in systemic sclerosis (scleroderma)
Springer Seminars in Immunopathology (2000)
-
Conservation of wingless patterning functions in the short-germ embryos of Tribolium castaneum
Nature (1994)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.