Abstract
Mutations in the GJB2 (connexin 26, Cx26) gene are the major cause of nonsyndromic hearing impairment in many populations. Genetic testing offers opportunities to determine the cause of deafness and predict the course of hearing, enabling the prognostication of language development. In the current study, we compared severity of hearing impairment in 60 patients associated with biallelic GJB2 mutations and assessed the correlation of genotypes and phenotypes. Within a spectrum of GJB2 mutations found in the Japanese population, the phenotype of the most prevalent mutation, 235delC, was found to show more severe hearing impairment than that of V37I, which is the second most frequent mutation. The results of the present study, taken together with phenotypes caused by other types of mutations, support the general rule that phenotypes caused by the truncating GJB2 mutations are more severe than those caused by missense mutations. The present in vitro study further confirmed that differences in phenotypes could be explained by the protein expression pattern.
Similar content being viewed by others
Introduction
Mutations of the GJB2 (connexin 26, Cx26) gene have recently drawn much attention because they have been recognized as the most prevalent genetic cause of congenital hearing loss. A broad range of phenotypes, from mild to profound hearing loss, is associated with GJB2 mutations (Cryns et al. 2004), and more than 90 different GJB2 mutations are associated with recessive forms of nonsyndromic hearing loss (The Connexins-deafness Homepage: http://www.crg.es/deafness). Universal neonatal hearing screening programs are the current trend and have become popular in many countries (Govaerts et al. 2001; Joint Committee on Infant Hearing 2000; Mehl and Thomson 2002; National Institutes of Health 1993), because it is thought that optimum language development requires early identification of hearing loss and early intervention (Yoshinaga-Itano et al. 1998). Cochlear implantation has resulted in remarkable improvement in auditory skills and development of speech production for patients with profound hearing loss associated with GJB2 mutations (Fukushima et al. 2002; Matsushiro et al. 2002). It is clear that genetic testing to determine the cause of deafness facilitates prediction of the course of hearing loss and prognostication of language development. There is, however, some controversy regarding genotype/phenotype correlation (Cohn et al. 1999; Cryns et al. 2004; Denoyelle et al. 1999; Estivill et al. 1998; Murgia et al. 1999; Orzan et al. 1999). For example, prediction of the degree of hearing loss was difficult, and environmental factors as well as modifier genes may have been involved (Cohn et al. 1999; Murgia et al. 1999; Orzan et al. 1999). On the other hand, a series of reports have indicated that certain phenotypes are dependent on certain genotypes (Denoyelle et al. 1999; Estivill et al. 1998). A recent report of a multi-center-based study in Europe and the United States suggested that inactivating mutations, which include stop or frameshift mutations, show significantly severer phenotypes than those caused by noninactivating mutations (missense mutations) (Cryns et al. 2004).
We have recently shown that mutation spectrums are quite different between the Japanese population and populations with European ancestry and emphasized the importance of specific population-based genetic databases for genetic testing (Ohtsuka et al. 2003). In Japanese (who are one example of Asian populations), the most common mutation was an inactivating mutation, 235delC, which is comparative to the 35delG mutation known as the most prevalent mutation in those with European ancestry. Interestingly, the second most common mutation was the V37I mutation, which has recently been reported as a mild phenotype causative genotype (Cryns et al. 2004). Given this background, we attempted to: (1) compare the differences in phenotypes caused by the 235delC and V37I mutations, (2) test a hypothetical general rule that inactivating mutations show more severe phenotypes than those caused by noninactivating mutations, and (3) test whether the differences in phenotype could be explained by protein expression study.
Materials and methods
Subjects and clinical evaluation
Pure-tone audiometry results were available for 60 individuals from independent families in whom biallelic GJB2 mutations were identified. These patients were from seven university hospitals (Hirosaki, Iwate, Gunma, Shinshu, Kokusai Iryoufukushi, Hamamatsu, and Kyushu) located in different regions in Japan. The age when the patients/parents noticed hearing impairment was from 0 to 49 (mean 8.00, SD 12.51) years of age. None of these patients had any other associated neurological signs. All subjects gave prior informed consent for participation in the project, which was approved by the ethical committee of each hospital.
Severity was classified by using a pure-tone average over 500, 1,000, 2,000, and 4,000 Hz in the better-hearing ear. Hearing impairment was classified as follows: normal hearing, <20 dB; mild hearing loss, 21–40 dB; moderate hearing loss, 41–70 dB; severe hearing loss, 71–95 dB; and profound hearing loss, greater than 95 dB. When the threshold exceeded the output limits of the audiometer, it was recorded as the output limit for air-conducted sounds plus 10 dBHL; i.e., if the output limit of the audiometer was120 dBHL, the threshold was described as 130 dBHL.
Mutation Analysis
To identify GJB2 mutations, a DNA fragment containing the entire coding region was amplified using the primer pair Cx48U/Cx1040L, as described elsewhere (Abe et al. 2000). Polymerase chain reaction (PCR) products were sequenced and analyzed with an ABI sequencer 377XL (Perkin-Elmer, Wellesley, MA, USA). DNA samples from 147 unrelated Japanese who had normal hearing were used as controls.
Reverse transcription (RT)-PCR analysis
Total RNA was extracted from NCTC2544 cells with the Catrimox-14 RNA Isolation Kit Ver.2.11 (Iowa Biotechnology, Urbandale, IA, USA). The yield of total RNA was determined by Agilent 2100 Bioanalyzer RNA 6000 Nano Assay (Agilent Technologies, Palo Alto, CA, USA). Reverse transcription (RT)-PCR assay was performed with the aid of an RNA PCR kit (Takara, Tokyo, Japan). The primers for human GJB2 and the specific sites of restriction enzymes were added with the amplification step. The primers were sense Xho I-Cx26 5′-cccctcgaggatggattggggcacgctgcagacgatcctggg-3′ and antisense Cx26-EcoR I 5′-cccgaattcgttaaactggcttttttgacttcccagaac-3′. These primers yield oligomer products of a distinctive size: 712 bp. PCR steps were denaturing at 94°C for 2 min, followed by 30 cycles of 94°C for 30 s, 60°C for 30 s, and 72°C for 1 min, and then processing with a final extension at 72°C for 5 min. After amplification, expected sizes of PCR products were confirmed on 2% agarose gel, and the bands were visualized by ethidium bromide upon exposure to an ultraviolet transilluminator.
Transformation
Wild-type Cx26 PCR products were inserted into a pEGFP-C2 vector (Clontech, Palo Alto, CA, USA). The PCR products and vector were digested with EcoR I and Xho I. Prepared PCR products were inserted into vector. Ligation reactants were transformed into Escherichia coli DH5α. Positive colonies were incubated in Luria-Bertani (LB) liquid medium containing kanamycin. A QIAprep spin miniprep kit (Qiagen, Valencia, CA, USA) was used for purification of plasmid DNA according to the manufacturer’s protocol. Plasmid DNA was identified by restriction enzyme analysis. Selected constructs were sequenced and analyzed with an ABI sequencer 377XL (Perkin-Elmer).
Mutagenesis of the GJB2 gene
The following primers were used to produce the GJB2 mutations: V27I sense 5′-ctggctcaccatcctcttcatt-3′, V37I sense 5′-tatgatcctcattgtggctgcaa-3′, and 235delC sense 5′-ccggctatgggcctgcagctgatct-3′. First, PCR reactions (100 μl) were prepared containing 4 μg of the plasmid DNA (see above), 1.0 μM of mutation primer, 1.0 μM of Cx26-EcoR I primer, 2.5 U of Takara Ex taq Hot Start Version (Takara, Tokyo, Japan), and Ex-taq buffer (10x) consisting of 100 mM Tris-HCl (pH 8.3), 500 mM KCl, 15 mM MgCl2, and 1 mM deoxynucleoside triphosphate mixture. These PCR reactions were denatured at 94°C for 2 min, followed by 30 cycles of 94°C for 30 s, 60°C for 30 s, and 72°C for 1 min, and then processed with a final extension at 72°C for 5 min. Second PCR reactions (100 μl) were prepared containing 10 μl of first PCR products, 1.0 μM of Xho I-Cx26 primer, 2.5 U of Takara Ex taq Hot Start Version (Takara), and Ex-taq buffer (10x) consisting of 100 mM Tris-HCl (pH 8.3), 500 mM KCl, 15 mM MgCl2, and 1 mM deoxynucleoside triphosphate mixture. Second PCR conditions were the same as above. These PCR products were inserted into a pEGFP-C2 vector with the same techniques as transformation (see above). The plasmid DNA containing Cx26 mutations were sequenced and analyzed with a sequencer and identified by restriction enzyme analysis.
Transfection and visualization
COS-7 cells grown on glass cover slips were transfected with the cloned plasmid vectors using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA). Forty-eight hours after the transfection, cells were fixed by 4% formaldehyde and stained by DAPI and TRITC-conjugated phalloidin (Chemicon, Temecula, CA, USA). Cover slips were mounted onto glass slides and visualized under a Leica confocal microscope TCS SP2 AOBS (Leica Microsystems, Wetzlar, Germany).
Results
Mutation spectrums
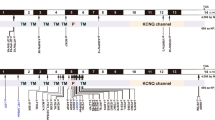
Among thirteen mutations that have been reported in Japanese (Ohtsuka et al. 2003), 11 were identified in our 60 biallelic patients. These biallelic mutations were found to be either four different homozygous or 14 different compound heterozygous mutations. These included five inactivating mutations and six missense mutations. The five inactivating mutations were one stop mutation (Y136X), three deletion frameshift mutations (235delC, 176-191del16, 299-300delAT), and one insertion frameshift mutation (605ins46). The six missense mutations were V37I (109G→A), G45E (134G→A), T86R (257C→G), T123N (368C→A), R143W (427C→T), and F191L (570T→C). T123N and F191L were categorized as changes with unknown relation to disease ( The Connexin-deafness Homepage: http://www.crg.es/deafness); however, we included both mutations as missense mutations in the present report because both were found among the hearing-loss patients in either a homozygous or compound heterozygous state. The nonsense mutation, Y136X (408C→A), converts a tyrosine residue (TAC) at codon 136 to a stop codon (TAA). Three deletion frameshift mutations, 235delC, 176-191del16, and 299-300delAT, and one insertion frameshift mutation, 605ins46, were found. The 235delC mutation causes a frameshift at codon 79 resulting in a truncated polypeptide and was found in two of the 147 controls (294 alleles). The 176-191del16 mutation, present in four subjects, causes a frameshift leading to an altered amino-acid sequence from codon 59 followed by a stop at codon 76. The 299-300delAT deletion, seen in two subjects, causes a frameshift leading to an altered amino-acid sequence from codon 100 followed by a stop at codon 113. The 605ins46 mutation has a tandem repeat of 46 nucleotides (corresponding to the positions 559–604 of the Cx26 DNA sequence) at the position 605. A stop codon (TGA) is produced at the 202nd amino acid, leading to the premature truncation in the series of polypeptide synthesis. Three previously described common sequence changes, V27I (79G→A), E114G (341A→G), and I203T (608T→C), which were thought to be nonpathological polymorphic changes (Abe et al. 2000), were frequently found in patients as well as controls.
Audiometric evaluation of the patients with biallelic GJB2 mutations
Audiometric results were obtained from 60 patients with biallelic GJB2 mutations. Fig. 1 shows a collection of overlapping audiograms from subjects bearing 18 combinations of mutations. Although the severity of hearing impairment in individuals varied according to the combinations of mutations, there seemed to be certain phenotypes determined by each combination. First, the hearing levels of the patients homozygous for 235delC mutations were comparatively severe to profound (Fig. 1). In addition, 235delC/299-300ATdel, G45E/G45E/Y136X/Y136X, G45E/Y136X/R143W, 235delC/R143W, and R143W/T86R also showed severe hearing impairment. In contrast, the patients homozygous for V37I had significantly mild-to-moderate hearing impairment (Fig. 1). Similarly, relatively milder phenotypes were found in the patients with 235delC/V37I, V37I/R143W, F191L/F191L, T123N/176-191del16, and V37I/G45E/Y136X. Hearing of one patient associated with V37I/T123N was within normal range (Fig. 1).
The comparison between patients homozygous for 235delC or V37I, which are the most and the second most prevalent mutations in Japanese (Ohtsuka et al. 2003), showed significant differences in phenotype (Figs. 1, 2). Those homozygous for the 235delC mutation (n=11, mean 100.68 dB, SD 21.25 dB) exhibited a significantly severer phenotype than that caused by V37I (n=5, mean 37.75 dB, SD 23.09 dB) (P=0.003 Fisher’s exact test). Those compound heterozygous for the 235delC mutation (n=19, mean 78.75 dB, SD 27.76 dB) were significantly different from those compound heterozygous for V37I (n=7, mean 47.14 dB, SD 18.35 dB) (P=0.021 Fisher’s exact test). Concerning the comparison between a combination of inactivating mutations and a combination of noninactivating mutations, the former (n=30, mean 88.33 dB, SD 25.67 dB) showed a severer phenotype than that caused by the latter (n=11, mean 47.39 dB, SD 31.19 dB) (P=0.0003 Fisher’s exact test).
Overlapping audiograms caused by 235delC/non 235delC, V37I/non V37I, inactivating mutation/inactivating mutation, and noninactivating mutation/noninactivating mutation. Note that patients associated with 235delC show relatively severer hearing loss whereas V37I-involved patients show a relatively mild phenotype. It is also evident that patients associated with inactivating mutation/inactivating mutation showed a severer phenotype than patients with noninactivating mutation/noninactivating mutation
Localization of Cx26 and its mutants
The inherent fluorescence of GFP determined the intracellular localization of the recombinant fusion proteins. Transfected GFP-Cx26 wt (wild type) were found to be localized as labeled puncta, which may be representative of gap junctions along the plasma membrane. In contrast, GFP- Cx26 235delC was not recognized at the plasma membrane but was retained within the cytoplasm close to the nucleus. Both GFP-Cx26 V27I and GFP- Cx26 V37I were found to be localized along the plasma membrane as well as being dispersed in the cytoplasm, which is a similar pattern to that shown in the wild type. (Fig. 3.)
Protein expression in transfected COS-7 cells. COS-7 cells transfected with GFP-Cx26 wt, GFP-Cx26 V27I, and GFP-Cx26 V37I, which were associated with normal–mild phenotypes, showed a characteristic puncta along the membrane. In contrast, only perinuclear staining was seen in GFP-Cx26 235delC. Red actin filament (TRITC- conjugated phalloidin): cell membrane, Blue DAPI: nucleus, Green Green fluorescent protein: chimeric protein.
Discussion
The present study, using different spectrums of GJB2 mutations (Ohtsuka et al. 2003), confirmed that certain genotypes are correlated with certain phenotypes in GJB2 deafness. The most common mutation, 235delC, exhibited severer hearing impairment whereas V37I, which is the second most common mutation, showed significantly mild hearing impairment. Audiometric data revealed an additional comparatively severe phenotype as well as a relatively mild phenotype.
Among more than 90 different GJB2 mutations, 35delG, accounts for up to 75% of mutated alleles in populations with European ancestry (Estivill et al. 1998; Gasparini et al. 2000; Van Laer et al. 2001). A series of reports has described that patients associated with 35delG exhibit severe-to-profound hearing impairment (Cohn et al. 1999; Cryns et al. 2004; Denoyelle et al. 1997, 1999; Green et al. 1999; Marlin et al. 2001; Wilcox et al. 2000). The status of the 235delC mutation, which seems to be a unique mutation in populations with Asian ancestry, is comparable to the 35delG mutation in Caucasoid populations. High prevalence of 35delG and 235delC mutations in the respective populations are due to a founder effect (Ohtsuka et al. 2003; Van Laer et al. 2001). Patients homozygous or compound heterozygous for the 235delC mutation exhibit a comparatively severer phenotype (Fig. 2), indicating that this frequent mutation should be the first to be considered when genetic screening for congenitally deaf patients is performed in Asian populations.
Several reports have indicated the existence of less-severe phenotypes correlated with certain specific mutations, especially in association with V37I (Bason et al. 2002; Cryns et al. 2004; Marlin et al. 2001; Rabionet et al. 2000; Wilcox et al. 2000). The exact phenotype has been rather difficult to prove because of the relatively small number of patients with V37I. The V37I mutation was originally reported as a polymorphism (Kelley et al. 1998), but the fact that valine 37 residue is highly conserved among different connexins, and that a series of reports identified homozygous or compound heterozygous V37I deafness patients (Abe et al. 2000; Bason et al. 2002; Marlin et al. 2001; Rabionet et al. 2000; Wilcox et al. 2000), indicate that it may be a disease-causing mutation. There seem to be ethnic differences in the allele frequency of V37I, as it was not detected in the control subjects from Italy, Spain, Germany, Greece, Israel, Ghana, or Austria (see Discussion in Bason et al. 2002) in spite of a high prevalence in the Japanese population (Abe et al. 2000; Kudo et al. 2000; Ohtsuka et al. 2003). The reported patients in whom the ethnic background was known were all of eastern-Asian origin (Abe et al. 2000; Bason et al. 2002; Kudo et al. 2000; Ohtsuka et al. 2003). In Japanese, V37I is the second most frequent mutated allele, and in this study, it was possible to collect a significant number of patients, and the present data confirmed a less severe phenotype caused by V37I. Due to such a mild phenotype, timing of presentation at clinics and diagnosis may be comparatively delayed. For patients with V37I/V37I, hearing impairment was noticed at the age of 20.6 (range 7–49, SD 17.08) years of age in contrast with 0.33 (range 0–3, SD 1.00) years for patients with 235delC/235delC. It should therefore be noted that patients with GJB2 mutations can also be found among less-severe hearing-impaired populations.
A recent multi-center-based genotype–phenotype correlation study clearly showed that severity of hearing impairment is correlated with some particular genotype and proposed a hypothetical general rule that inactivating mutations (stop or frameshift mutations) cause more severe phenotypes than those caused by noninactivating mutations (Cryns et al. 2004). Concerning the comparison between combinations of inactivating mutations and combinations of noninactivating mutations, the present study also showed that the former cause a severer phenotype than that caused by the latter. Therefore, our study supports the above hypothetical general rule.
Overlapped audiograms showed high-frequency-predominant sensorineural hearing loss regardless of genotype. Overall, there seemed to be certain rules regarding genotype and phenotype correlations. Particular genotypes tended to have similar audiograms with minor exceptions (Fig. 1). Therefore, genotype is a fundamental factor to predict phenotype. However, variations among the same phenotypes still exist (Fig. 1). These variations may be explained by the following factors involved in phenotypes: (1) alterations in promoter regions, (2) additional genes such as GJB6 (del Castillo et al. 2002), (3) modifier genes (Abe et al. 2001), (4) environmental factors. Concerning patients with G45E/Y136X, there was great variability in their phenotypes, ranging from normal to profound. A segregation study indicated that either G45E or Y136X situated on the same allele or different alleles. Our subcloning experiments confirmed the existence of two types of allele: cis allele and trans allele (data not shown). When two mutations are on different alleles (compound heterozygous state), the patients may exhibit severe-to-profound hearing impairment.
The present study further investigated whether the differences in phenotype could be explained by protein-expression study. In contrast to transfected GFP-Cx26 wt, which were found to be localized as labeled puncta along the plasma membrane (Fig. 3), the localization of transfected GFP-Cx26 235delC was not seen on the cellular membrane but mainly cohered at or around the nucleus. Such abnormal subcellular localization of mutated Cx26 protein with 235delC is consistent with a previous study (Choung et al. 2002). From these results, truncated mutations at the transmembrane domain, such as 235delC, were considered to lead to loss of function, resulting in serious hearing impairment. In the case of V37I, which is categorized as a noninactivating mutation, transfected GFP-Cx26 V37I was found along the membrane as in the wild type, indicating that the V37I protein may retain its function and therefore results in a rather mild phenotype. As expected, V27I, a known polymorphism, showed a similar distribution pattern to the wild type and V37I. To summarize, in the present study, the results indicate that protein expression patterns are well correlated with clinical phenotypes. A series of in vitro studies, including protein expression study, cell-to-cell communication properties, or physiological conductance experiments, sometimes provided discrepant results when compared to the phenotypic results, and limitations have been suggested (see discussion in Cryns et al. 2004). In the case of V37I, a complete loss of junctional properties has been reported (Bruzzone et al. 2003) in spite of a rather mild phenotype shown in a series of studies. The protein expression experiments in the current study, however, were in line with the phenotype associated with this mutation.
In conclusion, the present genotype–phenotype correlation results supported the view that phenotypes caused by the truncating GJB2 mutations are severer than those caused by missense mutations. Anticipating severity of hearing impairment is sometimes difficult, but if such general rules can be drawn with regard to genotype–phenotype correlation, determination of these correlations will facilitate the prediction of the course of hearing and help in making decisions regarding treatment/intervention.
References
Abe S, Usami S, Shinkawa H, Kelley PM, Kimberling WJ (2000) Prevalent connexin 26 gene (GJB2) mutations in Japanese. J Med Genet 37:41–43
Abe S, Kelley PM, Kimberling WJ, Usami SI (2001) Connexin 26 gene (GJB2) mutation modulates the severity of hearing loss associated with the 1555A–>G mitochondrial mutation. Am J Med Genet 103:334–338
Bason L, Dudley T, Lewis K, Shah U, Potsic W, Ferraris A, Fortina P, Rappaport E, Krantz ID (2002) Homozygosity for the V37I Connexin 26 mutation in three unrelated children with sensorineural hearing loss. Clin Genet 61:459–464
Bruzzone R, Veronesi V, Gomes D, Bicego M, Duval N, Marlin S, Petit C, D’Andrea P, White TW (2003) Loss-of-function and residual channel activity of connexin26 mutations associated with non-syndromic deafness. FEBS Lett 533:79–88
Choung YH, Moon SK, Park HJ (2002) Functional study of GJB2 in hereditary hearing loss. Laryngoscope 112:1667–1671
Cohn ES, Kelley PM, Fowler TW, Gorga MP, Lefkowitz DM, Kuehn HJ, Schaefer GB, Gobar LS, Hahn FJ, Harris DJ, Kimberling WJ (1999) Clinical studies of families with hearing loss attributable to mutations in the connexin 26 gene (GJB2/DFNB1). Pediatrics 103:546–550
Cryns K, Orzan E, Murgia A, Huygen PL, Moreno F, del Castillo I, Chamberlin GP, Azaiez H, Prasad S, Cucci RA, Leonardi E, Snoeckx RL, Govaerts PJ, Van de Heyning PH, Van de Heyning CM, Smith RJ, Van Camp G (2004) A genotype-phenotype correlation for GJB2 (connexin 26) deafness. J Med Genet 41:147–154
del Castillo I, Villamar M, Moreno-Pelayo MA, del Castillo FJ, Alvarez A, Telleria D, Menendez I, Moreno F (2002) A deletion involving the connexin 30 gene in nonsyndromic hearing impairment. N Engl J Med 346:243–249
Denoyelle F, Weil D, Maw MA, Wilcox SA, Lench NJ, Allen-Powell DR, Osborn AH, Dahl HH, Middleton A, Houseman MJ, Dode C, Marlin S, Boulila-ElGaied A, Grati M, Ayadi H, BenArab S, Bitoun P, Lina-Granade G, Godet J, Mustapha M, Loiselet J, El-Zir E, Aubois A, Joannard A, Petit C et al. (1997) Prelingual deafness: high prevalence of a 30delG mutation in the connexin 26 gene. Hum Mol Genet 6:2173–2177
Denoyelle F, Marlin S, Weil D, Moatti L, Chauvin P, Garabedian EN, Petit C (1999) Clinical features of the prevalent form of childhood deafness, DFNB1, due to a connexin-26 gene defect: implications for genetic counselling. Lancet 353:1298–1303
Estivill X, Fortina P, Surrey S, Rabionet R, Melchionda S, D’Agruma L, Mansfield E, Rappaport E, Govea N, Mila M, Zelante L, Gasparini P (1998) Connexin-26 mutations in sporadic and inherited sensorineural deafness. Lancet 351:394–398
Fukushima K, Sugata K, Kasai N, Fukuda S, Nagayasu R, Toida N, Kimura N, Takishita T, Gunduz M, Nishizaki K (2002) Better speech performance in cochlear implant patients with GJB2-related deafness. Int J Pediatr Otorhinolaryngol 62:151–157
Gasparini P, Rabionet R, Barbujani G, Melchionda S, Petersen M, Brondum-Nielsen K, Metspalu A, Oitmaa E, Pisano M, Fortina P, Zelante L, Estivill X (2000) High carrier frequency of the 35delG deafness mutation in European populations. Genetic Analysis Consortium of GJB2 35delG. Eur J Hum Genet 8:19–23
Govaerts PJ, Yperman M, De Ceulaer G, Daemers K, Van Driessche K, Somers T, Offeciers FE (2001) A Two-stage bipodal screening model for universal neonatal hearing screening. Otol Neurotol 22:850–854
Green GE, Scott DA, McDonald JM, Woodworth GG, Sheffield VC, Smith RJ (1999) Carrier rates in the midwestern United States for GJB2 mutations causing inherited deafness. JAMA 281:2211–2216
Joint Committee on Infant Hearing (2000) Year 2000 position statement: principles and guidelines for early hearing detection and intervention programs. Joint Committee on Infant Hearing, American Academy of Audiology, American Academy of Pediatrics, American Speech-Language-Hearing Association, and Directors of Speech and Hearing Programs in State Health and Welfare Agencies. Pediatrics 106:798–817
Kelley PM, Harris DJ, Comer BC, Askew JW, Fowler T, Smith SD, Kimberling WJ (1998) Novel mutations in the connexin 26 gene (GJB2) that cause autosomal recessive (DFNB1) hearing loss. Am J Hum Genet 62:792–799
Kenneson A, Van Naarden Braun K, Boyle C (2002) GJB2 (connexin 26) variants and nonsyndromic sensorineural hearing loss: a HuGE review. Genet Med 4:258–274
Kudo T, Ikeda K, Kure S, Matsubara Y, Oshima T, Watanabe K, Kawase T, Narisawa K, Takasaka T (2000) Novel mutations in the connexin 26 gene (GJB2) responsible for childhood deafness in the Japanese population. Am J Med Genet 90:141–145
Marlin S, Garabedian EN, Roger G, Moatti L, Matha N, Lewin P, Petit C, Denoyelle F (2001) Connexin 26 gene mutations in congenitally deaf children: pitfalls for genetic counseling. Arch Otolaryngol Head Neck Surg 127:927–933
Matsushiro N, Doi K, Fuse Y, Nagai K, Yamamoto K, Iwaki T, Kawashima T, Sawada A, Hibino H, Kubo T (2002) Successful cochlear implantation in prelingual profound deafness resulting from the common 233delC mutation of the GJB2 gene in the Japanese. Laryngoscope 112:255–261
Mehl AL, Thomson V (2002) The Colorado newborn hearing screening project, 1992–1999: on the threshold of effective population-based universal newborn hearing screening. Pediatrics 109:E7
Murgia A, Orzan E, Polli R, Martella M, Vinanzi C, Leonardi E, Arslan E, Zacchello F (1999) Cx26 deafness: mutation analysis and clinical variability. J Med Genet 36:829–832
National Institutes of Health (1993) Early identification of hearing impairment in infants and young children. NIH Consens Statement 11:1–24
Ohtsuka A, Yuge I, Kimura S, Namba A, Abe S, Van Laer L, Van Camp G, Usami S (2003) GJB2 deafness gene shows a specific spectrum of mutations in Japan, including a frequent founder mutation. Hum Genet 112:329–333
Orzan E, Polli R, Martella M, Vinanzi C, Leonardi M, Murgia A (1999) Molecular genetics applied to clinical practice: the Cx26 hearing impairment. Br J Audiol 33:291–295
Rabionet R, Zelante L, Lopez-Bigas N, D’Agruma L, Melchionda S, Restagno G, Arbones ML, Gasparini P, Estivill X (2000) Molecular basis of childhood deafness resulting from mutations in the GJB2 (connexin 26) gene. Hum Genet 106:40–44
Van Laer L, Coucke P, Mueller RF, Caethoven G, Flothmann K, Prasad SD, Chamberlin GP, Houseman M, Taylor GR, Van de Heyning CM, Fransen E, Rowland J, Cucci RA, Smith RJ, Van Camp G (2001) A common founder for the 35delG GJB2 gene mutation in connexin 26 hearing impairment. J Med Genet 38:515–518
Wilcox SA, Saunders K, Osborn AH, Arnold A, Wunderlich J, Kelly T, Collins V, Wilcox LJ, McKinlay Gardner RJ, Kamarinos M, Cone-Wesson B, Williamson R, Dahl HH (2000) High frequency hearing loss correlated with mutations in the GJB2 gene. Hum Genet 106:399–405
Yoshinaga-Itano C, Sedey AL, Coulter DK, Mehl AL (1998) Language of early- and later-identified children with hearing loss. Pediatrics 102:1161–1171
Acknowledgements
We thank all subjects who participated in the present project. This work was supported by the Ministry of Health and Welfare, Japan, (S.U.), and a Grant-in-Aid for Scientific Research from the Ministry of Education, Science and Culture of Japan (S.U.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Oguchi, T., Ohtsuka, A., Hashimoto, S. et al. Clinical features of patients with GJB2 (connexin 26) mutations: severity of hearing loss is correlated with genotypes and protein expression patterns. J Hum Genet 50, 76–83 (2005). https://doi.org/10.1007/s10038-004-0223-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-004-0223-7
Keywords
This article is cited by
-
Carrier re-sequencing reveals rare but benign variants in recessive deafness genes
Scientific Reports (2017)
-
Connexinopathies: a structural and functional glimpse
BMC Cell Biology (2016)
-
An effective screening strategy for deafness in combination with a next-generation sequencing platform: a consecutive analysis
Journal of Human Genetics (2016)
-
Characterization of a knock-in mouse model of the homozygous p.V37I variant in Gjb2
Scientific Reports (2016)
-
Carrier frequency of the GJB2 mutations that cause hereditary hearing loss in the Japanese population
Journal of Human Genetics (2015)