Abstract
The tumor suppressor p53 is essential for several cellular processes that are involved in the response to diverse genotoxic stress, including cell cycle arrest, DNA repair, apoptosis and senescence. Studies of the regulation of p53 have mostly focused on its stability and transactivation; however, new regulatory molecules for p53 have also been frequently identified. Here, we report that human ssDNA binding protein SSB1 (hSSB1), a novel DNA damage-associated protein, can interact with p53 and protect p53 from ubiquitin-mediated degradation. Furthermore, hSSB1 also associates with the acetyltransferase p300 and is required for efficient transcriptional activation of the p53 target gene p21 by affecting the acetylation of p53 at lysine382. Functionally, the hSSB1 knockdown-induced abrogation of the G2/M checkpoint is partially dependent on p53 or p300. Collectively, our results indicate that hSSB1 may regulate DNA damage checkpoints by positively modulating p53 and its downstream target p21.
Similar content being viewed by others
Introduction
The tumor suppressor p53 plays a critical role in maintaining genomic stability and preventing tumorigenesis1,2. Much evidence suggests that p53 is a multifunctional protein that is involved in diverse biological processes including cell cycle arrest, DNA repair, apoptosis and senescence3. The biological effects of p53 can in part be explained by its ability to activate or suppress the expression of a wide range of downstream target genes4. It is generally believed that tight regulation of p53, for example by post-translational modification, is essential for its stability and transcriptional activity5,6,7. It has become increasingly clear that a plethora of redundant post-translational modifications can occur in p53, including ubiquitination, acetylation, methylation, phosphorylation, neddylation and sumoylation, all of which significantly affects its stability and functions8,9,10. For example, E3 ligase MDM2 binds to and ubiquitinates p53, thereby regulating the stability and subcellular localization of p53. Acetyltransferase p300 functions as a co-activator and acetylates p53 at K382. This acetylation of p53 can enhance the stability of p53 by excluding ubiquitination or recruit some cofactors for the activation of p53 transcriptional activity. Moreover, increasing numbers of reports suggest that p53 can interact with several cellular proteins, including MDM2, NPM/B23 and NUMB11,12,13,14, which in turn modulate both the stability and the transcriptional activity of p53. In addition, some p53 coactivators, such as ADA3, VHL and BRD7, have also been shown to bind p53 and regulate p53-dependent transcriptional activity15,16,17.
In eukaryotic cells, RPA (human replication protein) is a key single-stranded DNA (ssDNA)-binding protein with three subunits: RPA70, RPA32 and RPA1418. RPA plays essential roles in cell cycle regulation and DNA replication and repair19,20. Recently, the novel human ssDNA binding protein SSB1 (hSSB1; OBFC2B) was identified as a key player in the cellular response to DNA damage21. Cells deficient in hSSB1 exhibit increased radiosensitivity, defective checkpoint activation and genomic instability21. Furthermore, the SOSS complex (INTS3-hSSB1-C9orf80) has been shown to play key roles in DNA damage response and DNA repair22,23,24,25. Mechanistically, hSSB1 has recently been shown to (1) be required for the efficient recruitment of the MRN complex, (2) bind directly with NBS1 and (3) stimulate the endonuclease activity of the MRN complex26,27. Moreover, we have recently shown that hSSB1 regulates cell cycle progression and DNA damage checkpoints by modulating the stability of p21 in cancer cells28. In this study, we report that hSSB1 not only physically interacts with p53 and affects its stability, but also associates with the acetyltransferase p300 to regulate the acetylation of p53 at lysine 382, a key event for the p53-mediated expression of p21.
Results
RNAi-mediated depletion of hSSB1 reduces the protein levels of endogenous p53 in p53 wild-type cancer cells
We have recently shown that p21 protein was markedly reduced in multiple cancer cell lines when hSSB1 was knocked down28. However, the regulation of p21 by hSSB1 is different among those cell lines. When hSSB1 was depleted, both the mRNA and protein levels of p21 were decreased in p53 wild-type cancer cell lines such as HepG2, SMMC-7721 and Bel7402 (Figure 1A), whereas in the p53-null Hep3B cells, the mRNA level of p21 remained constant while its protein level was downregulated (Figure 1A). Given that p53 is the major regulator of p21 at the transcriptional level, this observation raises the possibility that p53 may also be regulated by hSSB1. To examine this possibility, we used three different siRNA duplexes specifically targeting different hSSB1-coding regions. As shown in Figure 1B, p53 protein level was markedly reduced in HepG2 cells depleted of hSSB1, while overexpression of hSSB1 did not affect the protein level of p53 in HepG2 cells. This result indicates that endogenous hSSB1 is necessary and sufficient to execute its function on the stability of p53. Interestingly, our quantitative real-time PCR data showed that the level of p53 mRNA was not altered after depletion of hSSB1 in HepG2 cells, indicating that hSSB1 regulates p53 at the post-translational level rather than at the transcriptional level (Figure 1C). This positive regulation of p53 by hSSB1 is further confirmed in multiple p53 wild-type cancer cell lines (Figure 1D and data not shown). Because the depletion of hSSB1 can cause cell cycle redistribution and p53 expression is cell cycle dependent, we used synchronized cells depleted of hSSB1 to rule out the possibility that cell cycle redistribution results in altered expression of p53 protein. As shown in Figure 1E, knockdown of hSSB1 resulted in a decrease of p53 protein level in each phase, although the level of p53 protein varied during the cell cycle. This indicates that the positive relationship between the p53 protein level and hSSB1 is independent of the cell cycle. In other words, p53 stability is positively regulated by hSSB1.
The hSSB1 knockdown results in decreased p53 protein level in p53 wild-type cancer cells. (A) HepG2, SMMC-7721, Bel-7402 and Hep3B cells were transfected with scrambled or hSSB1 siRNAs as indicated for 48 h, the indicated mRNA and protein levels were detected by real time-PCR and western blotting with the indicated antibodies. (B) Left: HepG2 cells were transfected with the indicated siRNAs for 48 h (P: a pool mixed with equal amount of three individual hSSB1 siRNAs) and were analyzed as in A. Right: HepG2 cells were transfected with vector or Flag-hSSB1 for 24 h, and protein was detected by western blotting with the indicated antibodies. (C) HepG2 cells were transfected with scrambled or hSSB1 siRNAs as indicated for 48 h, and the indicated mRNA levels were determined by real time-PCR. (D) SMMC-7721 and Bel-7402 cells were transfected with scrambled or hSSB1 siRNAs as indicated for 48 h, and were analyzed as A. (E) HepG2 cells transfected with the indicated siRNAs were synchronized using a double thymidine block and release. Cells were collected for G1, S, G2 and M phase as described in Materials and Methods. Some cells were collected for flow cytometry (top panel), and the rest of the cells were analyzed by western blotting with the indicated antibodies (bottom panel).
hSSB1 interacts with p53 both in vitro and in vivo
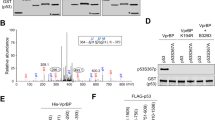
Given that hSSB1 regulates p53 stability and that the hSSB1 homolog RPA has been shown to interact with p5329,30,31,32, we examined whether hSSB1 also associates with p53. Following co-immunoprecipitation (co-IP) using Flag-agarose or Myc-agarose, a complex containing hSSB1 and p53 was clearly detected in HEK293T cells overexpressing both exogenous hSSB1 and p53 (Figure 2A). Since p53 is a very sticky protein that interacts with a number of protein partners, we used Flag-AEG1 and Flag-ERK1 as the negative and positive controls, respectively, and found that Flag-hSSB1 and Flag-ERK1 could co-immunoprecipitate HA-p53, whereas Flag-AEG1 nearly have no interaction with p53 (Supplementary information, Figure S1). To further confirm this interaction, we performed endogenous co-IP and found that a complex containing hSSB1 and p53 was detectable at endogenous levels, as shown in Figure 2B. To determine whether the interaction between hSSB1 and p53 is direct, we expressed and purified MBP-tagged p53 and GST-tagged hSSB1. Pull-down assay revealed that hSSB1 binds with p53 directly in vitro (Figure 2C). Collectively, these results suggest that hSSB1 associates with p53 both in vitro and in vivo.
The hSSB1 interacts with p53. (A) HEK293T cells transfected with the indicated plasmids for 24 h were lysed with MCLB. Immunoprecipitation (IP) using anti-Myc or anti-Flag antibodies and western blotting with the indicated antibody were performed. WCL, whole cell lysate. (B) HepG2 cells were lysed with MCLB, and lysates were subjected to immunoprecipitation using anti-IgG or anti-hSSB1 as indicated, and were analyzed by western blotting. (C) Beads coated with bacterially expressed GST or GST-hSSB1 were incubated with purified MBP-p53 protein overnight at 4 °C. Beads were washed, and the bound proteins were analyzed by western blotting using antibodies as indicated. (D) Upper: Schematic description of the domains of p53. Lower: HEK293T cells co-transfected with Flag-hSSB1 with various p53 mutants as indicated for 24 h were lysed with MCLB, and were analyzed as A. (E) Upper: Schematic description of the domains of hSSB1. Lower: HEK293T cells co-transfected HA-p53 with various hSSB1 mutants as indicated for 24 h were lysed with MCLB, and were analyzed as A.
Next, we attempted to determine the regions responsible for the interaction between hSSB1 and p53. A series of deletions of hSSB1 and p53 were generated (Figure 2D and 2E), and co-IP was performed to map the binding domains of these two proteins. As shown in Figure 2D, the core domain of p53 (residues 100-300) was necessary and sufficient to bind hSSB1. Interestingly, as shown in Figure 2E, both the N-terminal oligosaccharide-binding fold domain (residues 1-113) and the C-terminus (114-211) fragment of hSSB1 were capable of binding p53. It is noteworthy that the mobility of C-terminal fragment of hSSB1 (114-211, 98 aa) is much slower than the N-terminal fragment hSSB1 (1-113, 113 aa). We speculate that the C-terminal fragment of hSSB1 (114-211) contains a higher frequency (20/98 = 20.4%) of proline, as compared with the N-terminal fragment (1-113, 113 aa) that has a much lower frequency (5/113 = 4.4%) of proline. Consistent with this result (Figure 2E), Skaar et al.24 also found that the N-terminal hSSB1 (1-113, 113 aa) fragment was less than 17 KD in the SDS-PAGE gel. Together, these results reinforce the notion that hSSB1 physically and specifically interacts with p53 in cells.
hSSB1 protects p53 from ubiquitin-mediated degradation
Because hSSB1 binds to and positively regulates p53, which is well known to be labile in cells, we speculated that hSSB1 might affect the proteolysis of p53. Indeed, MG132, a specific proteasome inhibitor, substantially rescued the decline of p53 protein level caused by hSSB1 knockdown in HepG2 cells (Figure 3A), suggesting that hSSB1 prevents p53 from degradation by the proteasome. Furthermore, as shown in Figure 3B, poly-ubiquitinated p53 was upregulated in the hSSB1-knockdown HepG2 cells, indicating that the degradation of p53 prevented by hSSB1 is ubiquitination dependent. If the ubiquitination of p53 increases, a shorter half-life of endogenous p53 protein is expected in cells depleted of hSSB1. This is indeed the case, as shown in Figure 3C. Collectively, these results indicate that knockdown of hSSB1 increases the ubiquitination of p53, which in turn promotes the degradation of p53 by the proteasome.
The hSSB1 protects p53 from ubiquitin-mediated degradation. (A) HepG2 cells were transfected with the indicated siRNA for 48 h, incubated with dimethyl sulfoxide (DMSO) or 10 μM MG132 for 4 h, and probed for the indicated proteins by western blotting. (B) HepG2 cells transfected with hSSB1 or control siRNA for 24 h were co-transfected with Myc-p53 and HA-ubiquitin for another 24 h, and MG132 (10 μM) was added for 4 h and lysed with MCLB, followed by anti-Myc IP and western blotting with the indicated antibody. (C) HepG2 cells transfected with hSSB1 or scrambled siRNA for 48 h were incubated with 20 mg/ml cycloheximide (CHX) for the indicated periods of time. Lysates were harvested from the cells and analyzed by western blotting (left panel). Quantitation of p53 protein levels were shown in right pane l (n = 3).
hSSB1 regulates the acetylation of p53 at K382, which in turn modulates the transcriptional activity of p53
Given that hSSB1 contributes to p53 stability, we subsequently investigated whether hSSB1 affects the transcriptional activity of p53. Genomic expression profiles of HepG2 cells treated with si-scramble or si-hSSB1 showed that p21, a main downstream target of p53, was the second most downregulated gene when hSSB1 was depleted (Supplementary information, Figure S2). In addition, the newly identified p53 target gene SULF2 was also downregulated (Supplementary information, Figure S2). These findings were further validated by quantitative real-time PCR. As shown in Figure 4A, hSSB1 knockdown clearly reduced the basal expression levels of p21 and SULF2 by over twofold in HepG2 cells. Furthermore, in HepG2 cells treated with ionizing radiation (IR), hSSB1 knockdown clearly decreased the expression levels of p21, SULF2, MDM2 and BAX by over twofold. These results strongly suggest that hSSB1 is capable of regulating the transcriptional activity of p53 in normal conditions or in response to DNA damage.
hSSB1 regulates the transcriptional activity of p53 by controlling the K382 acetylation. (A) HepG2 cells transfected with scrambled or hSSB1 siRNA for 48 h were either untreated or irradiated (6Gy). After 8 h, cells were harvested and the indicated mRNA levels were determined by real time-PCR. (B) HepG2 cells transfected with scrambled or hSSB1 siRNA for 24 h were either treated with or without doxorubicin (DOX, 1 μM). After 24 h, cells were then harvested in MCLB and analyzed by immunoblotting. The number presented the ratio of acetylated p53 (K382) and phosphorylated p53 (Ser15) to the corresponding total p53 protein in doxorubicin-treated experiments (control, set at 1). The quantitation of the intensity of the acetylated p53 and the total p53 signals were shown in right panel. (C) HepG2 cells transfected with scrambled or hSSB1 siRNA for 24 h were either untreated or treated with doxorubicin (DOX, 1 μM) for another 24 h. Cells were then harvested, and equivalent amounts of total cell lysates (note: the loaded protein level in lane 4 was higher than that in other lanes in order to normalize the total-p53 protein level) were subjected to western blotting with the indicated antibodies. (D) Upper: HepG2 cells transfected with the indicated plasmids for 24 h were lysed with MCLB. Immunoprecipitation (IP) using anti-Flag antibodies and western blotting with the indicated antibody were performed. Lower: HepG2 cells were lysed with MCLB, and lysates were subjected to immunoprecipitation using anti-IgG or anti-hSSB1 as indicated, and were analyzed by western blotting. (E) HepG2 cells transfected with scrambled or hSSB1 siRNA for 24 h, and then transfected with the empty vector or HA-p300 plasmid. After 12h, cells were either untreated or treated with doxorubicin (DOX, 1 μM) for 12 h before harvesting. Equivalent amounts of total cell lysates were subjected to western blotting with the indicated antibodies. (F) HepG2 cells transfected with scrambled or hSSB1 siRNA for 24 h were transiently transfected with p21-Luc promoter plasmid along with vector, p53 or p300 expression plasmid. After 6 h, cells were treated with or without doxorubicin (DOX, 1 μM) for 24 h. Then cells were harvested and p21 luciferase activities were measured (n = 3).
The transactivation of p53, for instance in response to DNA damage, is believed to be regulated through protein accumulation and through post-translational modifications such as phosphorylation and acetylation33. We therefore determined whether depletion of hSSB1 affects the phosphorylation or acetylation of p53 in response to DNA damage. In HepG2 cells depleted of hSSB1 after doxorubicin treatment, the ratio of p53 acetylated at K382 (Ac-p53) to total p53 was consistently decreased by half, whereas the ratio of p53 phosphorylated at Ser15 (Phos-p53) to total p53 was slightly decreased (Figure 4B). Furthermore, when the total p53 protein level was normalized, Ac-p53, but not Phos-p53, was clearly decreased in HepG2 cells depleted of hSSB1 after doxorubicin treatment (Figure 4C). This finding indicates that hSSB1 may be required for effective acetylation of p53 at K382, especially in response to DNA damage.
Acetyltransferase p300 is a cofactor that is mainly responsible for the acetylation of p53 at K382, which modulates the transcriptional activity of p5334,35,36,37. Activated p53 induces expression of p21 mainly by recruiting p30038,39. In addition, it has been reported that SKP2 and BRD7 can control the acetylation of p53 at K382 by binding and regulating p30017,40. Thus, we surmised that acetyltransferase p300 might play a key role in the hSSB1-mediated regulation of p53 acetylation at K382. Indeed, as shown in Figure 4D, a complex including hSSB1 and p300 was clearly detected at both exogeous and endogenous levels. More importantly, the decreased p53 acetylation at K382 induced by knockdown of hSSB1 was completely rescued when etopic p300 was introduced into the cells (Figure 4E). In addition, as shown in Figure 4F, a luciferase assay also showed that overexpression of either p53 or p300 can partially rescue the decreased activity of p21 promoter in response to DNA damage in cells depleted of hSSB1. This is consistent with the result that the p21 mRNA level can be regulated by hSSB1 in p53 wild-type cancer cells (Figure 1A, 4A and Supplementary information, Figure S2).
hSSB1 is involved in the p53-dependent G2/M checkpoint in response to DNA damage
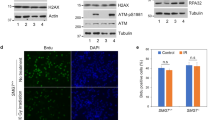
To determine the biological functions and significance of the regulation of p53 by hSSB1 described above, apoptosis, the G1/S transition and DNA damage checkpoints were examined, as both hSSB1 and p53 are key components in these processes. First, we tested whether the regulation of p53 by hSSB1 is involved in the G1/S transition. As shown in Supplementary information, Figure S3, the promotion of the G1/S transition by hSSB1 knockdown was mostly rescued by overexpression of p21, but neither p53 nor p300, in HepG2 cells, indicating that the G1/S transition regulated by hSSB1 is directly through p21 protein, consistent with our recent report28. Second, we tested whether hSSB1 affects apoptosis induced by doxorubicin. As shown in Supplementary information, Figure S4, the doxorubicin-induced apoptosis was not altered in hSSB1-knockdown cells, indicating that the regulation of p53 by hSSB1 is not involved in apoptosis induced by doxorubicin in HepG2 cells. Third, we tested whether hSSB1 is required for stabilization of p53 in response to DNA damage. As shown in Figure 5A, the IR-induced increase of p53 protein level was completely suppressed in hSSB1-knockdown cells, and as expected, the p53 target p21 was also suppressed. Based on the fact that p53 protein level is increased in response to DNA damage, we were curious to determine whether the interaction between hSSB1 and p53 is altered in such a response. Interestingly, as shown in Figure 5B, the binding between hSSB1 and p53 was obviously enhanced upon DNA damage induced by doxorubicin. Fourth, as expected, IR-induced cell cycle arrest in G2/M is dramatically diminished by the depletion of hSSB1 (Figure 5C). To discriminate between cells in G2 and M phases, phospho-Ser10-histone3 (pH3), an indicator of cells in M phase, was used to monitor the G2/M checkpoint in HepG2 (p53 wild type) cells exposed to IR. As shown in Figure 5D, the percentage of cells in M phase was markedly increased (1.0% vs 3.1%) by knockdown of hSSB1 in p53 wild-type HepG2 cells (Figure 5D). Indeed, as shown in Figure 5E, the abolishment of the G2/M checkpoint by hSSB1 knockdown was partially rescued by overexpression of p53 or p300 in HepG2 cells. Taken together, these results strongly suggest that hSSB1 regulates the p53-dependent G2/M checkpoint in response to DNA damage.
hSSB1 is involved in p53-dependent G2/M checkpoint in response to DNA damage. (A) Left: HepG2 cells transfected with scrambled or hSSB1 siRNA for 48 h were either untreated or irradiated (6 Gy). After 24 h, cells were then harvested in MCLB and analyzed by immunoblotting. Right: HepG2 cells transfected with scrambled or hSSB1 siRNA for 48 h were either untreated or irradiated (6 Gy). After 4h, 12h and 24 h, cells were then harvested in MCLB and analyzed by immunoblotting. (B) HepG2 cells were treated with or without doxorubicin (DOX, 1 μM) for 24 h before lysed with MCLB, and lysates were subjected to immunoprecipitation using anti-IgG or anti-hSSB1 as indicated, and were analyzed by western blotting. Note: hSSB1 immunoprecipitated with p53 in DNA damage induced by DOX was twofold compared with the control group, as shown in the inserted numbers, which were measured by the intensities of the immunoprecipitated p53 and the total p53 protein levels. (C) HepG2 cells were transfected by scrambled or hSSB1 siRNA for 48 h and then exposed to IR (6 Gy). After 24 h, cells were then harvested and analyzed using flow cytometry (n = 3). (D) HepG2 cells were transfected by scrambled or hSSB1 siRNA for 48 h and then exposed to IR (6 Gy). They were then incubated with 100 ng/ml of nocodazole for 10 h, harvested and analyzed using flow cytometry. The percentage of cells positive for phosphohistone H3 (pH3) is indicated (n = 3). (E) HepG2 cells transfected with scramble or hSSB1 siRNA for 24 h were transfected with vector, p53 or p300 plasmid. After 24 h, cells were exposed to IR (6 Gy), incubated with 100 ng/ml of nocodazole for 10 h, and analyzed as in C. All results shown are representative of three independent experiments (n = 3). (F) Schematic model showing how hSSB1 modulates the G2/M checkpoint.
Discussion
In this report, we have shown that hSSB1 binds to and protects p53 from degradation by the proteasome. hSSB1 also contributes to the regulation of DNA damage checkpoints in p53 wild-type cancer cells by regulating the acetylation of p53, which in turn modulates the transcriptional activity of p53. To our knowledge, this is the first evidence that hSSB1 may (1) positively regulate p53 at the protein level and (2) bind to p300 and bring it to the promoter of p21 to modulate p53-induced p21 expression in cells.
p53 is the most frequently mutated gene in human tumors, over 50% of which harbor p53 mutations41. As a key tumor suppressor and transcription factor, p53 is often degraded by the ubiquitin-proteasome system. MDM2 is a major E3 ubiquitin ligase that can bind to p53 and catalyze either mono- or poly-ubiquitination of p53, leading to nuclear export or proteasomal degradation42. Furthermore, post-translational modifications such as acetylation may have a crucial role in the stabilization of p5343. For example, acetylation may regulate the stability of p53 by inhibiting MDM2-induced ubiquitination44. Here, we found that depletion of hSSB1 reduced the acetylation of p53 at lysine 382 and increased the ubiquitination of p53 (Figure 4B, 4C and 3B), indicating that hSSB1 might regulate MDM2-mediated ubiquitination by affecting the acetylation of p53. Taken together, these results highlight hSSB1 as a novel component in the regulation of p53 stability. In addition, activated p53 induces the expression of p21 mainly by recruiting p300, which is primarily responsible for the acetylation of p53 at lysine 38234,35,36,37. This observation is consistent with our results showing that depletion of hSSB1 can downregulate p21 expression in p53 wild-type cancer cells, probably by decreasing the acetylation of p53 (Figure 1A, 4A-4C and Supplementary information, Figure S2), although the mechanisms need to be investigated in further detail. We can speculate that hSSB1 may be required for p53-induced p21 expression mainly by binding and bringing p300 to the promoter of p21 in cells, because hSSB1 can interact with p300. Furthermore, overexpression of p300 can completely rescue the decreased acetylation of p53 at lysine 382 in the cells depleted of hSSB1 (Figure 4E), and overexpression of either p53 or p300 can partially rescue the decline in p21 promoter activity induced by depletion of hSSB1 (Figure 4F). These functions of hSSB1 are similar to those of BRD717, indicating that hSSB1 interacts with p53 and p300, which in turn affect p53 acetylation and p21 promoter activity.
It has been reported that INTS3-hSSB1-C9orf80 forms a subunit complex22,23,24,25, and that the protein level of hSSB1 was dramatically reduced in the cells of depleted of INTS322,23,24. In addition, Skaar et al.24 also demonstrated that knockdown of INTS3 resulted in the decrease of hSSB1 mRNA levels, indicating that INTS3 may regulate the transcription of hSSB1. Therefore, we believe that the INTS3 subunit in the complex of INTS3-hSSB1-C9orf80 may have the same function on p53 as hSSB1 does, although we did not perform such an experiment. As regard to another component, C9orf80, we have no clue to its effect on p53, since its function is unclear so far. In addition, the subunits in a complex may function differently and independently. For instance, we have recently showed that damaged DNA binding protein 1 (DDB1) regulates the activity of APC/CCdh1 independent of the Cul4-DDB1 complex45. Here, we have mainly focused on hSSB1, showing that hSSB1 binds to and regulates p53 stability and transcriptional activity, indicating that p53 may be one of the downstream molecules of hSSB1 to control the cell cycle checkpoint and DNA damage response. Interestingly, both hSSB1 and p53 are DNA-binding proteins, and hSSB1 binds to the core domain, which is also known as DNA-binding domain, of p53. Therefore, we could not rule out the possibility that DNA may be also involved in the regulation of p53 by hSSB1, and this desires to be further investigated in future.
We have recently revealed that hSSB1 binds p21 and modulates its degradation in both p53 wild-type and p53-deficient cell lines to regulate cell cycle and DNA damage checkpoint28. In current study, when hSSB1 was depleted, p21 was decreased at both the mRNA and protein levels in p53 wild-type cancer cell lines such as HepG2, SMMC-7721 and Bel7402 (Figure 1A), whereas in the p53-null Hep3B cells, the p21 mRNA level remained constant while the protein level was downregulated (Figure 1A). This regulatory pathway is also involved in the cell cycle and DNA damage checkpoint. Thus, we speculate that hSSB1 can regulate p21 at both protein and mRNA levels in the p53 wild-type cells whereas hSSB1 mainly stabilizes p21 protein in p53-deficient cells to regulate cell cycle and DNA damage checkpoint. In other words, the regulation of p21 by hSSB1 can be occurred in p53-dependent and -independent manner. Therefore, like NPM/B2346, hSSB1 regulates p21 at both the transcriptional and post-translational levels, as shown in the proposed mechanism in Figure 5F, in order to ultimately regulate DNA damage checkpoints.
The protein p53 is involved in multiple biological processes that act to prevent tumorigenesis, including reversible cell cycle arrest, apoptosis, DNA repair and senescence. p21 is one of the critical target genes that mediates p53-induced cell cycle arrest. Much evidence has been presented showing that both p53 and p21 participate in arrest at the G1/S and G2/M checkpoints after DNA damage47,48,49,50. Our results show that hSSB1 is required to stabilize both p53 and p21 in response to IR (Figure 5A)28, and that hSSB1 knockdown allows bypass of the G2/M checkpoint response to IR. The effects of hSSB1 knockdown were mostly rescued by overexpressing p2128, and mildly by overexpressing p53 or p300 in cells (Figure 5E). This phenomenon can be explained by the fact that p21, which can also be positively regulated by hSSB128, is an important effecter for inhibiting Cdks in order to arrest at the checkpoints of the cell cycle. However, p53 and p300 must execute their functions through p21 and p53, respectively. In addition to p53 and p21, other unknown molecules involved in the G2/M checkpoint may also be regulated by hSSB1.
In conclusion, we have demonstrated that hSSB1 can participate in the G2/M checkpoint by positively regulating both p21 and p53, as proposed in Figure 5F. This may have an important impact on our understanding of molecular pathways related to DNA damage checkpoints and may also partially explain why cells deficient in hSSB1 exhibit increased radiosensitivity, defective checkpoint activation and genomic instability21. Most importantly, based on the fact that bypass of the G2/M checkpoint is widely used for clinical trials on cancers51,52, hSSB1 may be a promising therapeutic target for cancer radiotherapy, especially for cancer cells with high levels of hSSB1 expression.
Materials and Methods
Cell lines
Cells (HepG2, Hep3B, Bel-7402, SMMC-7721, HEK293, HEK293T) were cultured in Dulbecco's modified Eagle's medium (DMEM, Invitrogen) supplemented with 10% fetal bovine serum (FBS, Hyclone), 1 mM glutamine and 100 U/ml each of penicillin and streptomycin.
Plasmids
Full-length cDNAs coding for hSSB1, p21, p53, AEG1 and ERK1 were obtained from human HEK293 cell cDNA by PCR and were then cloned into pcDNA3.1 with a FLAG, HA or Myc tag sequence. GST and GST-hSSB1 were used previously28. MBP-tagged p53 was a gift from Dr Junjie Chen (The University of Texas MD Anderson Cancer Center, USA). HA-tagged ubiquitin was a gift from Dr Helen Piwnica-Worms (Washington University, St. Louis, USA). HA-tagged p300 was a gift from Dr Binhua P Zhou (University of Kentucky, USA). Mutations were introduced using the Quik-Change Site-Directed Mutagenesis Kit (Stratagene), and all mutations were verified by DNA sequencing.
Antibodies and reagents
Human anti-hSSB1 antibody was from Bethyl Laboratories. Human anti-p53, anti-p300, anti-Hsp70 and anti-GST antibodies were from Santa Cruz Biotechnology. Human anti-MBP antibody was a gift from Dr Junjie Chen. Human anti-FLAG and anti-actin antibodies were from Sigma Chemical Co. Human anti-GAPDH was from Bioworld Technology. Other primary antibodies used for western blotting (anti-acetyl-p53 Lys382, anti-phospho-p53 Ser15, anti-HA, anti-p21 and anti-Myc) were from Cell Signaling Technology. Bound primary antibodies were detected with either horseradish peroxidase (HRP)-conjugated goat anti-mouse antibody or HRP-conjugated goat anti-rabbit antibody (Sigma Chemical Co.). MG132, protease inhibitor cocktail and phosphatase inhibitor cocktail were purchased from Calbiochem. Doxorubin (ADR) was purchased from Sigma Chemical Co.
RNAi treatment
The sequences of oligonucleotides targeting hSSB1 mRNA were as follows: (1) CCCUGUUAGUAACGGCAAA, (2) GGAGAAUUCUGUAUGGUUU and (3) CGACGGAGACCUUUGUGAA. Approximately 2 × 105 HepG2 cells were seeded into each well of a 6-well tissue culture dish the day before transfection. Transfection was performed according to the manufacturer's instructions using LipofectamineTM RNAiMAX transfection reagent (Invitrogen) and 50 nM siRNA. 48 h post-transfection, cells were incubated in the presence or absence of 25 μg/ml cycloheximide for the indicated times and harvested in mammalian cell lysis buffer (MCLB) (50 mM Tris-HCl pH 8.0, 2 mM DTT, 5 mM EDTA, 0.5% Nonidet P-40, 100 mM NaCl, 1 mM microcystin, 1 mM sodium orthovanadate, 2 mM PMSF, protease inhibitor cocktail (Calbiochem) and phosphatase inhibitor cocktail (Calbiochem)). Alternatively, cells were exposed to 6 Gy IR and harvested in MCLB after 24 h.
Western blotting and immunoprecipitation
Western blotting and immunoprecipitation were performed as described previously53. Briefly, cells were lysed in MCLB, and the clarified lysates were resolved by SDS-PAGE and transferred to PVDF membranes for western blotting using ECL detection reagents (Beyotime Co.). Alternatively, the clarified supernatants were first incubated with anti-Myc-agarose (Santa Cruz), anti-FLAG-agarose (Sigma Chemical Co.), or anti-HA-agarose (Sigma Chemical Co.) for 2 h to overnight at 4 °C, and the precipitates were washed five times with MCLB. To investigate the interaction between hSSB1 and p53 or p300 at the endogenous level, the clarified supernatants were first incubated with anti-hSSB1 for 2 h at 4 °C. Protein A/G-agaroses were then added for 2 h to overnight, and the precipitates were washed five times with MCLB and analyzed by western blotting.
In vitro GST pull-down assay
The GST, GST-hSSB1 and MBP-p53 were expressed in E. coli BL21 cells. These fusion proteins were purified using glutathione-Sepharose 4B beads (GE Life Sciences) or amylase-Sepharose beads (New England Biolabs). GST-tagged hSSB1 or GST alone was immobilized on glutathione-Sepharose 4B beads and incubated with purified MBP-tagged p53 protein overnight at 4 °C. Beads were washed four times and these bound proteins were subjected to western blotting with anti-MBP or anti-GST antibodies.
Real-time quantitative PCR
Total RNA was extracted from HepG2 cells using Trizol reagent (Invitrogen), and 1 μg DNase-treated RNA was reverse transcribed using the Revert AidTM First Strand cDNA Synthesis Kit (MBI Fermentas, USA) according to the manufacturer's instructions. The threshold cycle (Ct) value of each sample was determined using Platinum SYBR Green qPCR SuperMix-UDG with ROX (Invitrogen) in an ABI 7900HT Real-Time PCR System (Applied Biosystems, Foster City, CA). Primer sequences are shown in Supplementary information, Table S1. The relative mRNA levels of each gene shown were normalized to the expression of the housekeeping gene β-actin. The relative level of a transcript was calculated as two power values of ΔCt (Ct of β-Actin minus the Ct of the target cDNA).
mRNA expression array analysis
Total RNA was extracted with Trizol reagent (Invitrogen), and an aliquot of 2 μg of total RNA was used to synthesize double-stranded cDNA, which in turn was used to produce biotin-tagged cRNA. The resulting biotin-tagged cRNA was fragmented to strands of 35-200 bases in length according to the protocols from Affymetrix. The fragmented cRNA was hybridized to an mRNA array. The hybridization data were analyzed using GeneChip Operating software (GCOS 1.4). The scanned images were first assessed by visual inspection and then analyzed to generate raw data files, which were saved as CEL files using the default setting of GCOS 1.4. In a comparison analysis, we applied a two-class unpaired method in the Significant Analysis of Microarray software (SAM) to identify significantly differentially expressed genes between test (B-1 and B-2) and control (S1 and S2) groups. Genes were determined to be significantly differentially expressed with a selection threshold of false discovery rate (FDR) < 5% and fold change > 2.0 in the SAM output result. The results from this calculation were then sorted; the 11 most upregulated (indicated as Red color) and 9 most downregulated (indicated as Green color) genes are presented in Supplementary information, Table S2 as heat maps.
Luciferase reporter assay
HepG2 cells were transiently transfected with hSSB1 siRNA oligonucleotides or scrambled oligonucleotides. About 24 h after transfection, cells were cotransfected with the indicated expression plasmids and reporter constructs. After 6 h, the cells were treated with doxorubin (ADR, 1 μM) for another 24 h. Whole cell lysates were then prepared and assayed using a dual luciferase assay system (Promega) according to the manufacturer's instructions. The intensity of the firefly luciferase signals generated was normalized against an internal control (Renilla luciferase). The luciferase reporter plasmid PGL3-p21 was a gift from Wei-Guo Zhu (Peking University, China). The results were obtained from three sets of transfections.
In vivo ubiquitination assay
This assay was performed as described previously28. Briefly, HepG2 cells were transfected with scrambled or hSSB1 siRNA. About 24 h after transfection, cells were cotransfected with the indicated Myc-p53 and HA-ubiquitin constructs for 24 h. MG132 (10 μM) was added for 4 h prior to collection, and cells were lysed in MCLB. The clarified supernatants were first incubated with anti-Myc-agarose (Santa Cruz) and then subjected to western blotting.
Synchronization of HepG2 cells
HepG2 cells synchronized at the G1/S boundary using a double thymidine block-and-release protocol54 were released into culture media for 3 h (S-phase cells), 6 h (G2-phase cells), 9 h with 100 ng/ml of nocodazole (M-phase cells) or 11 h (G1-phase cells). Cells were collected and analyzed by flow cytometry and western blotting.
G2/M checkpoint assay
The G2/M checkpoint assay was performed essentially as described previously28. Cells were first exposed to IR (6 Gy) and incubated with 100 ng/ml of nocodazole for 10 h, then were counted by flow cytometry following fixation and staining with phosphohistone H3 antibody (Upstate Biotechnology, Lake Placid, NY, USA) and PI.
Apoptosis assay
HepG2 cells were transfected with scrambled or hSSB1 siRNAs for 6 h, and then treated with doxorubicin (DOX, 1 μM), after 48 h the cells were collected by trypsin without EDTA,washed with PBS, then subjected to annexin V-EGFP and propidium iodide staining as the manufacturer's recommendations (KeyGen Biotec),and analyzed by flow cytometry.
G1/S transition assay
As detailed previously28, HepG2 cells were transfected with a control siRNAs and hSSB1 siRNAs. After 24 h, p21-WT and control plasmid then were transfected. Nocodazole (100 ng/ml; Sigma Aldrich) was added 8 h after the second transfection for 16 h. Cells were harvested by trypsinization and fixed with 70% ethanol at −20 °C overnight. Cell cycle distributions were measured by staining with PI, followed by analysis on a flow cytometer (Beckman).
References
Levine AJ, Hu W, Feng Z . The P53 pathway: what questions remain to be explored? Cell Death Differ 2006; 13:1027–1036.
Vousden KH, Prives C . Blinded by the light: The growing complexity of p53. Cell 2009; 137:413–431.
Vousden KH, Lu X . Live or let die: the cell's response to p53. Nat Rev Cancer 2002; 2:594–604.
Riley T, Sontag E, Chen P, Levine A . Transcriptional control of human p53-regulated genes. Nat Rev Mol Cell Biol 2008; 9:402–412.
Bode AM, Dong Z . Post-translational modification of p53 in tumorigenesis. Nat Rev Cancer 2004; 4:793–805.
Dai C, Gu W . p53 post-translational modification: deregulated in tumorigenesis. Trends Mol Med 2010; 16:528–536.
Brooks CL, Gu W . New insights into p53 activation. Cell Res 2010; 20:614–621.
Kruse JP, Gu W . Modes of p53 regulation. Cell 2009; 137:609–622.
Brooks CL, Gu W . The impact of acetylation and deacetylation on the p53 pathway. Protein Cell 2011; 2:456–462.
Taira N, Yoshida K . Post-translational modifications of p53 tumor suppressor: determinants of its functional targets. Histol Histopathol 2012; 27:437–443.
Haupt Y, Maya R, Kazaz A, Oren M . Mdm2 promotes the rapid degradation of p53. Nature 1997; 387:296–299.
Kubbutat MH, Jones SN, Vousden KH . Regulation of p53 stability by Mdm2. Nature 1997; 387:299–303.
Colombo E, Marine JC, Danovi D, Falini B, Pelicci PG . Nucleophosmin regulates the stability and transcriptional activity of p53. Nat Cell Biol 2002; 4:529–533.
Colaluca IN, Tosoni D, Nuciforo P, et al. NUMB controls p53 tumour suppressor activity. Nature 2008; 451:76–80.
Kuo PC, Tsao YP, Chang HW, et al. Breast cancer amplified sequence 2, a novel negative regulator of the p53 tumor suppressor. Cancer Res 2009; 69:8877–8885.
Lukas J, Lukas C, Bartek J . More than just a focus: The chromatin response to DNA damage and its role in genome integrity maintenance. Nat Cell Biol 2011; 13:1161–1169.
Drost J, Mantovani F, Tocco F, et al. BRD7 is a candidate tumour suppressor gene required for p53 function. Nat Cell Biol 2010; 12:380–389.
Wold MS . Replication protein A: a heterotrimeric, single-stranded DNA-binding protein required for eukaryotic DNA metabolism. Annu Rev Biochem 1997; 66:61–92.
Richard DJ, Bolderson E, Khanna KK . Multiple human single-stranded DNA binding proteins function in genome maintenance: structural, biochemical and functional analysis. Crit Rev Biochem Mol Biol 2009; 44:98–116.
Oakley GG, Patrick SM . Replication protein A: directing traffic at the intersection of replication and repair. Front Biosci 2010; 15:883–900.
Richard DJ, Bolderson E, Cubeddu L, et al. Single-stranded DNA-binding protein hSSB1 is critical for genomic stability. Nature 2008; 453:677–681.
Li Y, Bolderson E, Kumar R, et al. HSSB1 and hSSB2 form similar multiprotein complexes that participate in DNA damage response. J Biol Chem 2009; 284:23525–23531.
Huang J, Gong Z, Ghosal G, Chen J . SOSS complexes participate in the maintenance of genomic stability. Mol Cell 2009; 35:384–393.
Skaar JR, Richard DJ, Saraf A, et al. INTS3 controls the hSSB1-mediated DNA damage response. J Cell Biol 2009; 187:25–32.
Zhang F, Wu J, Yu X . Integrator3, a partner of single-stranded DNA-binding protein 1, participates in the DNA damage response. J Biol Chem 2009; 284:30408–30415.
Richard DJ, Cubeddu L, Urquhart AJ, et al. hSSB1 interacts directly with the MRN complex stimulating its recruitment to DNA double-strand breaks and its endo-nuclease activity. Nucleic Acids Res 2011; 39:3643–3651.
Richard DJ, Savage K, Bolderson E, et al. hSSB1 rapidly binds at the sites of DNA double-strand breaks and is required for the efficient recruitment of the MRN complex. Nucleic Acids Res 2011; 39:1692–1702.
Xu S, Feng Z, Zhang M, et al. hSSB1 binds and protects p21 from ubiquitin-mediated degradation and positively correlates with p21 in human hepatocellular carcinomas. Oncogene 2011; 30:2219–2229.
Dutta A, Ruppert JM, Aster JC, Winchester E . Inhibition of DNA replication factor RPA by p53. Nature 1993; 365:79–82.
Abramova NA, Russell J, Botchan M, Li R . Interaction between replication protein A and p53 is disrupted after UV damage in a DNA repair-dependent manner. Proc Natl Acad Sci USA 1997; 94:7186–7191.
Romanova LY, Willers H, Blagosklonny MV, Powell SN . The interaction of p53 with replication protein A mediates suppression of homologous recombination. Oncogene 2004; 23:9025–9033.
Bochkareva E, Kaustov L, Ayed A, et al. Single-stranded DNA mimicry in the p53 transactivation domain interaction with replication protein A. Proc Natl Acad Sci USA 2005; 102:15412–15417.
Olsson A, Manzl C, Strasser A, Villunger A . How important are post-translational modifications in p53 for selectivity in target-gene transcription and tumour suppression? Cell Death Differ 2007; 14:1561–1575.
Liu L, Scolnick DM, Trievel RC, et al. p53 sites acetylated in vitro by PCAF and p300 are acetylated in vivo in response to DNA damage. Mol Cell Biol 1999; 19:1202–1209.
Lill NL, Grossman SR, Ginsberg D, DeCaprio J, Livingston DM . Binding and modulation of p53 by p300/CBP coactivators. Nature 1997; 387:823–827.
Avantaggiati ML, Ogryzko V, Gardner K, Giordano A, Levine AS, Kelly K . Recruitment of p300/CBP in p53-dependent signal pathways. Cell 1997; 89:1175–1184.
Gu W, Roeder RG . Activation of p53 sequence-specific DNA binding by acetylation of the p53 C-terminal domain. Cell 1997; 90:595–606.
Barlev NA, Liu L, Chehab NH, et al. Acetylation of p53 activates transcription through recruitment of coactivators/histone acetyltransferases. Mol Cell 2001; 8:1243–1254.
Zhao Y, Lu S, Wu L, et al. Acetylation of p53 at lysine 373/382 by the histone deacetylase inhibitor depsipeptide induces expression of p21(Waf1/Cip1). Mol Cell Biol 2006; 26:2782–2790.
Kitagawa M, Lee SH, McCormick F . Skp2 suppresses p53-dependent apoptosis by inhibiting p300. Mol Cell 2008; 29:217–231.
Hollstein M, Sidransky D, Vogelstein B, Harris CC . p53 mutations in human cancers. Science 1991; 253:49–53.
Li M, Brooks CL, Wu-Baer F, Chen D, Baer R, Gu W . Mono- versus polyubiquitination: differential control of p53 fate by Mdm2. Science 2003; 302:1972–1975.
Rodriguez MS, Desterro JM, Lain S, Lane DP, Hay RT . Multiple C-terminal lysine residues target p53 for ubiquitin-proteasome-mediated degradation. Mol Cell Biol 2000; 20:8458–8467.
Li M, Luo J, Brooks CL, Gu W . Acetylation of p53 inhibits its ubiquitination by Mdm2. J Biol Chem 2002; 277:50607–50611.
Lv XB, Xie F, Hu K, et al. Damaged DNA-binding protein 1 (DDB1) interacts with Cdh1 and modulates the function of APC/CCdh1. J Biol Chem 2010; 285:18234–18240.
Xiao J, Zhang Z, Chen GG, et al. Nucleophosmin/B23 interacts with p21WAF1/CIP1 and contributes to its stability. Cell Cycle 2009; 8:889–895.
el-Deiry WS, Tokino T, Velculescu VE, et al. WAF1, a potential mediator of p53 tumor suppression. Cell 1993; 75:817–825.
Ceraline J, Deplanque G, Duclos B, et al. Inactivation of p53 in normal human cells increases G2/M arrest and sensitivity to DNA-damaging agents. Int J Cancer 1998; 75:432–438.
Bunz F, Dutriaux A, Lengauer C, et al. Requirement for p53 and p21 to sustain G2 arrest after DNA damage. Science 1998; 282:1497–1501.
Ando T, Kawabe T, Ohara H, Ducommun B, Itoh M, Okamoto T . Involvement of the interaction between p21 and proliferating cell nuclear antigen for the maintenance of G2/M arrest after DNA damage. J Biol Chem 2001; 276:42971–42977.
Lobrich M, Jeggo PA . The impact of a negligent G2/M checkpoint on genomic instability and cancer induction. Nat Rev Cancer 2007; 7:861–869.
Bucher N, Britten CD . G2 checkpoint abrogation and checkpoint kinase-1 targeting in the treatment of cancer. Br J Cancer 2008; 98:523–528.
Kang T, Wei Y, Honaker Y, et al. GSK-3 beta targets Cdc25A for ubiquitin-mediated proteolysis, and GSK-3 beta inactivation correlates with Cdc25A overproduction in human cancers. Cancer Cell 2008; 13:36–47.
Chen MS, Ryan CE, Piwnica-Worms H . Chk1 kinase negatively regulates mitotic function of Cdc25A phosphatase through 14-3-3 binding. Mol Cell Biol 2003; 23:7488–7497.
Acknowledgements
We thank the members of the laboratory for their helpful comments on the manuscript. This work was supported by the National Basic Research Program of China (2010CB912201 and 2012CB967000 to TK), and the National Natural Science Foundation of China (81171890 to SL, 81125015 and 30930045 to TK).
Author information
Authors and Affiliations
Corresponding author
Additional information
( Supplementary information is linked to the online version of the paper on the Cell Research website.)
Supplementary information
Supplementary information, Figure S1
HEK293T cells transfected with the indicated plasmids for 24 h were lysed with MCLB. (PDF 67 kb)
Supplementary information, Figure S2
HepG2 cells were transfected with scrambled or hSSB1 siRNAs as indicated for 48 h, and mRNA was extracted and detected by microarray chip. (PDF 38 kb)
Supplementary information, Figure S3
HepG2 cells transfected with scrambled or hSSB1 siRNA for 24 h were transfected with p21, p53, p300 plasmids or control as indicated for 8 h, and then nocodazole (100 ng/ml) was added for another 16 h, followed by flow cytometry (n = 3). (PDF 209 kb)
Supplementary information, Figure S4
HepG2 cells were transfected with scrambled or hSSB1 siRNAs for 6 h, and then treated with doxorubicin (DOX, 1 μM), after 48h the cells were subjected to annexin V-EGFP/propidium iodide staining, and analyzed by flow cytometry (n = 3). (PDF 172 kb)
Supplementary information, Table S1
Sequences of primers used for Real-time quantitative PCR (PDF 9 kb)
Supplementary information, Table S2
Ratio=B / S, significantly differentially expressed genes:20 (XLS 7967 kb)
Rights and permissions
About this article
Cite this article
Xu, S., Wu, Y., Chen, Q. et al. hSSB1 regulates both the stability and the transcriptional activity of p53. Cell Res 23, 423–435 (2013). https://doi.org/10.1038/cr.2012.162
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2012.162
Keywords
This article is cited by
-
KLF12 promotes the proliferation of breast cancer cells by reducing the transcription of p21 in a p53-dependent and p53-independent manner
Cell Death & Disease (2023)
-
SOSSB1 and SOSSB2 mutually regulate protein stability through competitive binding of SOSSA
Cell Death Discovery (2023)
-
UBE2O targets Mxi1 for ubiquitination and degradation to promote lung cancer progression and radioresistance
Cell Death & Differentiation (2021)
-
Forcing ATGL expression in hepatocarcinoma cells imposes glycolytic rewiring through PPAR-α/p300-mediated acetylation of p53
Oncogene (2019)
-
RETRACTED ARTICLE: MNAT1 is overexpressed in colorectal cancer and mediates p53 ubiquitin-degradation to promote colorectal cancer malignance
Journal of Experimental & Clinical Cancer Research (2018)