Abstract
Interleukin-2 (IL)-2 signaling plays a pivotal role in the activation of immune responses, and drugs that block this pathway have been shown to be effective for the immunosuppression in patients with organ transplantation to alleviate/eliminate allograft rejection. The first humanized monoclonal antibody (mAb) daclizumab falls into this category and shows high specificity and affinity against a key component of the IL-2 receptor complex, namely IL-2Rα. To reveal the molecular mechanism of the inhibition of the IL-2 signaling pathway by daclizumab, we determined the crystal structures of the daclizumab Fab in free form and in complex with the IL-2Rα ectodomain at 2.6 and 2.8 Å resolution, respectively. The daclizumab Fab adopts a similar conformation in the presence or absence of the IL-2Rα ectodomain. The antigen-binding site of daclizumab is mainly composed of five complementarity determining regions (CDRs) that form a large positively charged surface depression and two flanking patches that are generally hydrophobic. The conformational epitope consists of several discontinuous segments of the IL-2Rα ectodomain, a large portion of which overlaps with the regions that interact with IL-2, suggesting that the binding of daclizumab to IL-2Rα would prevent the IL-2 binding to IL-2Rα and the subsequent formation of the IL-2/IL-2Rαβγc complex, and therefore block the IL-2 signaling pathway. These results also have implications for the design and development of improved mAb drugs targeting IL-2Rα.
Similar content being viewed by others
Introduction
The cytokine interleukin-2 (IL-2) plays a central role in immune responses by promoting proliferation, differentiation, and survival of mature T and B cells, and the cytolytic activity of natural killer cells 1, 2, 3, 4. The in vivo biological effects of IL-2 are mediated mainly through the high-affinity IL-2 receptor (IL-2R) complex 2 composed of the alpha chain IL-2Rα (p55, CD25, or Tac Ag), the beta chain IL-2Rβ (p75 or CD122), and the common gamma chain γc (p65 or CD132). In the structure of the IL-2/IL-2Rαβγc complex, IL-2Rα interacts extensively with two dominantly hydrophobic regions of IL-2 largely via its D1 domain, and IL-2Rβ and the γc interact with IL-2 from the opposite side with less contacts and have no interaction with IL-2Rα 5, 6, 7. The structural and biochemical data suggest a sequential assembly of the IL-2/IL-2Rαβγc complex: (1) IL-2Rα that expresses abundantly on the activated T cells acts as a ligand carrier to facilitate the secretion of IL-2 and then binds it, leading to conformational changes of IL-2 to a favorable IL-2Rβ binding state; (2) the IL-2/IL-2Rα complex is delivered to IL-2Rβ through two-dimensional cell surface diffusion to form the IL-2/IL-2Rαβ complex; and (3) the γc is recruited to form the IL-2/IL-2Rαβγc complex that triggers the signaling pathways 6, 7, 8, 9.
Because the interactions of IL-2 with IL-2Rs play a pivotal role in the activation of immune defense, inhibition of the interactions can suppress the immune responses and provide therapeutic benefits 1. IL-2Rα is considered as a good target in clinical treatment, particularly for the specific immunosuppression in organ transplantation, due to several reasons: (1) the binding of IL-2Rα to IL-2 initiates the sequential assembly of the IL-2/IL-2Rαβγc complex and then activates the IL-2 signaling pathway; (2) IL-2Rα is specific for IL-2 binding 10, 11, whereas IL-2Rβ is shared by IL-15 12, 13, 14, 15, and the γc is a common receptor component for a variety of cytokines 15, 16, 17; and (3) IL-2Rα is not expressed on resting T and B cells but is abundantly expressed on activated T cells and abnormal T cells of patients suffering from allograft rejection 18, 19, some autoimmune diseases 20, 21, and T cell leukemia 20, 22. Currently there are two monoclonal antibody (mAb) drugs targeting at IL-2Rα, namely daclizumab (Zenapax®, Roche) and basiliximab (Simulect®, Novartis), which were approved by the US Food and Drug Administration (FDA) for prevention of allograft rejection in organ transplantation. Basiliximab is a chimeric mAb with the variable domains of murine anti-IL-2Rα mAb RTF5 and the constant domains of human IgG1 (κ). Daclizumab is the first humanized mAb drug approved by FDA in 1997. It is a humanized IgG1 form of murine anti-IL-2Rα mAb anti-Tac and retains the murine complementarity determining regions (CDRs) and some key residues of the framework regions (FWRs) for high specificity and high affinity 23. Daclizumab has been successfully used in the treatment of renal transplantation 24, 25, 26, 27, 28 and liver transplantation 29, 30, 31, 32, and is promising for the treatment of acute graft-versus-host disease 33, 34, multiple sclerosis 35, and malignancies such as T cell leukemia 19. The non-randomized large multi-center trials in phase I/II showed that the safe and efficacious dosages of basiliximab and daclizumab are quite different (basiliximab in two 20 mg/kg doses and daclizumab in five 2 mg/kg doses) 36. However, the randomized, open-label, single-center studies found no significant differences in the efficacy and safety of the two mAbs 36, 37. Basiliximab and daclizumab have similarly high binding affinities to the IL-2Rα ectodomain with KD values of 0.14 nM 38 and 0.3 nM 39, respectively.
The IL-2Rα ectodomain contains two β-strand-swapped “sushi-like” domains D1 (residues 1-64) and D2 (residues 101-164) 5, 6, 7, 38. Previously, by phage display library screening, Binder et al. 40 suggested that residues 116-120 of the D2 domain of IL-2Rα form the epitope recognized by both basiliximab and daclizumab. Recently, our structural study of the basiliximab Fab in complex with IL-2Rα has shown that basiliximab binds IL-2Rα via a discontinuous epitope composed of several segments (residues 1-6, 21-29, 38-48, 56-57) of the D1 domain and a minor segment (residues 118-120) of the D2 domain. As the epitope covers most of the regions responsible for the interactions of IL-2Rα with IL-2, our results indicate that the binding of basiliximab to IL-2Rα blocks the interactions of IL-2 with IL-2Rα and thus inhibits the activation of the IL-2 signaling pathway 38.
To understand the inhibition mechanism of daclizumab on the IL-2 signaling pathway and to compare the respective interactions of IL-2R with daclizumab and basiliximab, we determined the crystal structures of the daclizumab Fab both in free form and in complex with the IL-2Rα ectodomain. Structural analysis of the daclizumab Fab/IL-2Rα complex and its comparisons with the IL-2/IL-2Rαβγc and basiliximab Fab/IL-2Rα complexes reveal the molecular basis of the high affinity of daclizumab to IL-2Rα, the inhibition mechanism of daclizumab on the IL-2 signaling pathway, and the differences in the two mAbs. Our results also have implications for further improvement of mAb drugs targeting IL-2Rα.
Results and Discussion
Structures of the daclizumab Fab in free form and in complex with the IL-2Rα ectodomain
The crystal structures of the daclizumab Fab in free form and in complex with the IL-2Rα ectodomain were determined at 2.6 and 2.8 Å resolution, respectively (Figure 1A and Table 1). In the free-form daclizumab Fab structure, there is one Fab molecule in an asymmetric unit. In the structure of the daclizumab Fab/IL-2Rα complex, there are two complexes in an asymmetric unit with almost identical overall structures (superposition of all Cα atoms yields a root-mean-square deviation of 1.2 Å for IL-2Rα, 0.6 Å for the Fab, and 0.9 Å for the complex, respectively), and the one with more detectable residues and better electron densities was chosen for further structural analysis and discussion.
Structure of the daclizumab Fab/IL-2Rα complex. (A) A stereoview of the overall structure of the daclizumab Fab/IL-2Rα complex. The Fab is colored with V and C domains of the light chain in yellow and orange, and those of the heavy chain in green and pink, respectively. IL-2Rα is colored with the D1 domain in magenta and the D2 domain in cyan. (B) A stereoview of a representative SIGMAA-weighted 2Fo - Fc map (1σ contour level) showing part of the binding interface in IL-2Rα D1 domain (residues 1-7) and D2 domain (residues 147-154), both of which are involved in interactions with the daclizumab Fab. The atomic coordinates of the residues are shown with ball-and-stick models. The disulfide bond between CysI3 and CysI147 is clearly defined. (C) Structural comparison of IL-2Rα among different complexes. IL-2Rα structures from the complexes with the daclizumab Fab and the basiliximab Fab (PDB code: 3U3I) are colored in green and cyan, respectively, and those from the IL-2/IL-2Rα complex (PDB code: 1Z92) and the IL-2/IL-2Rαβγc complex in two different crystal forms (PDB codes: 2B5I and 2ERJ) are colored in red, yellow and magenta, respectively. Residues 1-64 and 101-164 of the IL-2Rα ectodomain from the complex with the daclizumab Fab can be traced, whereas the two linker regions are disordered due to their inherent flexibility. IL-2Rα in the complex with the daclizumab Fab adopts a conformation very similar to that observed in the other IL-2Rα complexes except that some conformational changes occur at three flexible regions of residues 36-41, 109-117, and 132-144 as indicated.
In the free form or in complex with IL-2Rα, the daclizumab Fab exhibits a canonical β-sandwich immunoglobulin fold consisting of the heavy-chain VH and CH1 domains (residues 1-216) and the light-chain VL and CL domains (residues 1-212) (Figure 1A). In both structures, the daclizumab Fab has well-defined electron density, except for a small segment (SerH130 to GlyH136 in the free-form Fab or SerH130 and SerH131 in the complex) of the conserved loop of the CH1 domain, which is often disordered in many other Fab structures 41. Both structure models have good stereochemical geometry, except that residue ThrL50 resides in a classic γ-turn and hence is located in the disallowed region of the Ramachandran plot, which is commonly seen in other Fab structures 42. Superposition of the structures based on all Cα atoms yields a root-mean-square deviation of 0.8 Å for the V domains and 0.8 Å for the C domains, indicating that the V and C domains of the free-form daclizumab Fab resemble those in the complex. In particular, the six CDRs of the Fab adopt very similar conformations in the two structures. The elbow angle of the Fab, defined as the subtended angle by the two pseudo two-fold axes relating VH to VL and CH to CL, is 164.6° in the free-form Fab and 169.0° in the complex, which is in agreement with the notion that daclizumab has a κ light chain 43. Crystal packing analysis reveals that both V and C domains participate in the interactions with symmetry-related molecules. Thus, the slight change of the elbow angle in the two structures is probably caused by crystal packing and/or the intrinsic flexibility of the elbow.
The IL-2Rα ectodomain in the complex assumes an atypical cytokine receptor structure as reported previously 5, 6, 7, 38 (Figure 1A). The D1 (residues 1-64) and D2 (residues 101-164) domains have well-defined electron density, especially in the regions interacting with the Fab (Figure 1B). The linker between the D1 and D2 domains (residues 65-100) and the C-terminal region connecting the D2 domain to the trans-membrane domain (residues 165-217) cannot be observed as in all other IL-2Rα structures 5, 6, 7, 38, and hence may not participate in the binding of the Fab. Thus, only 129 out of 217 residues of the IL-2Rα ectodomain are structured. Comparison with the previously reported structures 5, 6, 7, 38 shows that the IL-2Rα ectodomain adopts a similar overall structure (superposition of all Cα atoms yields a root-mean-square deviation of 1.4–1.7 Å for the D1 domain, 1.5 Å for the D2 domain, and 1.4–1.7 Å for the whole molecule). The major conformational differences occur in three solvent-exposed regions (residues 36-41 of the D1 domain and residues 109-117 and 132-144 of the D2 domain) (Figure 1C). These regions are adjacent to the epitope, but have no interactions with the Fab. Nevertheless, we cannot completely exclude the possibility that the conformational changes might have resulted from the Fab binding.
Interactions between the daclizumab Fab and the IL-2Rα ectodomain
The epitope of IL-2Rα is composed of several discontinuous segments, including residues 1-6, 25-27, and 42-43 of the D1 domain, and residues 118-120 and 149-155 of the D2 domain (Figure 2A, panel a). The antigen-binding site of the daclizumab Fab consists of a large, positively charged surface depression flanked by two generally hydrophobic patches, which involves CDRs L2, L3, H1, H2, and H3 and FWR L2 of the Fab (Figure 2A, panel b). Formation of the complex buries a total of 2078.6 Å2 solvent-accessible surface area (1053.0 Å2 on the Fab and 1025.6 Å2 on IL-2Rα), which is much higher than the common range in the other antibody-antigen complexes 44, 45. The Fab heavy chain contributes 555.2 Å2 of the buried solvent-accessible surface area, whereas the light chain contributes 497.8 Å2. The interaction interface has a shape complementarity (Sc) value of 0.58, which is slightly lower than the average range of 0.64–0.68 for the other antibody-antigen complexes 46. In total, there are 12 hydrogen bonds, 2 salt bridges, and 91 van der Waals contacts at the interface (Figure 2, Tables 2 and 3). The extensive interactions at the interface result in a high binding affinity of daclizumab to the IL-2Rα ectodomain with an apparent disassociation constant (KD) of 0.27 nM as measured by the SPR analysis. The KD value is similar to that of daclizumab to human T lymphoma cell line HuT-102, as determined by the Scatchard plot analysis (0.3 nM) 39.
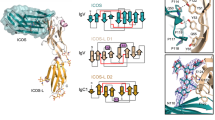
Interactions between the daclizumab Fab and IL-2Rα. (A) Molecular surfaces of the daclizumab Fab and IL-2Rα showing the residues at the interface. Panel a: the daclizumab Fab is shown with an electrostatic potential surface with some of the residues involved in the interactions labeled in black, while IL-2Rα is shown with a stick model with the D1 domain in magenta and the D2 domain in cyan. In panel b, IL-2Rα is shown with an electrostatic potential surface with some of the residues involved in the interactions labeled in black, while the Fab is shown as a stick model with CDR loops L2 and L3 in yellow, and CDR loops H1, H2, and H3 in green, respectively. The residues of IL-2Rα and the Fab involved in the interactions are shown with ball-and-stick models. (B-D) The hydrophilic interactions between IL-2Rα and the daclizumab Fab. IL-2Rα forms hydrophilic interactions with the Fab mainly in three regions, which are illustrated in panels B-D, respectively. The hydrogen bonds are indicated by black dashes and the salt bridges with gray dashes. The color coding of the structural elements is the same as in Figure 1A.
Specifically, residues 1-6 of IL-2Rα (GluI1 to AspI6) interact with the central positively charged surface depression of the Fab formed mainly by residues ArgL90, TyrH32, ArgH33, HisH35, GlyH100, and ValH102 (hereafter residues of the Fab heavy chain and light chain and IL-2Rα are designated by superscripted chain identifiers H, L, and I, respectively) (Figure 2A and 2B and Tables 2 and 3). This part of the interface makes up more than half of the hydrophilic interactions and one-third of the hydrophobic contacts (8 hydrogen bonds, 2 salt bridges, and 38 van der Waals contacts). In particular, the side chain of AspI4 of IL-2Rα inserts into a small cavity formed by ArgH33, HisH35, GlyH100, and GlyH101 and forms a number of hydrophilic and hydrophobic interactions. Residues of CDRs L3 and H1 contribute most of the interactions at this region. Particularly, ArgH33 of CDR H1 contributes two salt bridges, one hydrogen bond, and 13 van der Waals contacts.
In addition, residues 25-27, 42-43, and 118-120 of IL-2Rα interact with a generally hydrophobic surface patch composed of several residues of CDRs L3 and H2, including TyrL93, TyrH50, TyrH57, GluH59, and GlnH62 (Figure 2A and 2C and Tables 2 and 3). In this region, a total of 4 hydrogen bonds and 24 van der Waals contacts are formed, and in particular, GluH59 of CDR H2 makes a major contribution by interacting with AsnI27, TyrI43, and HisI120 of IL-2Rα. Moreover, residues 149-155 of IL-2Rα interact with another hydrophobic surface patch formed by CDRs L2, L3, H1, and H3 (including residues ThrL49, SerH31, ArgL90, TyrH32, GlyH100, GlyH101, and ValH102), and FWR L2 (TyrL48) (Figure 2A and 2D and Tables 2 and 3). Nearly half of the interactions are formed by residues GlyH100 and TyrH32, including 2 hydrogen bonds and 12 van der Waals contacts. It is noteworthy that TyrL48 of FWR L2 makes 11 van der Waals contacts with residues ThrI150, HisI151, and GlyI152.
Previously, the phage display library screening results suggested that the daclizumab epitope is located in the region of residues 116-122 of IL-2Rα 40. In the structure of the daclizumab Fab/IL-2Rα complex, only two residues of this region participate in the interactions with the daclizumab Fab. Specifically, IleI118 forms two van der Waals contacts with TyrH57, and HisI120 makes one hydrogen bond and eight van der Waals contacts with CDRs H2 and L3 (Tables 2 and 3). Similarly, the basiliximab epitope defined in our previous study 38 contains several discontinuous segments, of which the linear epitope identified by the phage display method 40 constitutes only a small portion. Also, the 182YCYSI186 fragment of CD20 suggested to be part of the rituximab epitope 47 does not bind to rituximab directly; instead, our structural study shows that Cys183 of this fragment may contribute to maintaining a unique cyclic conformation of the CD20 epitope by forming a disulfide bond with Cys167 48. In addition, in a search for peptide mimics of the epitope recognized by trastuzumab, a therapeutic antibody specific to Her2/neu, the five mimotopes identified by the phage display screening 49 just mimic one of the three regions of the Her-2/neu epitope determined by the structural study 50. These results indicate that the potential epitope identified by the phage display method is very likely incomplete and needs to be treated cautiously in the studies of antibody-antigen interactions and in the development of improved antibodies.
Comparison of the epitopes of daclizumab and basiliximab
Our biochemical data show that the basiliximab mAb has a binding affinity to IL-2Rα (0.14 nM) 38 similar to that of the daclizumab mAb (0.27 nM, in this study), consistent with the similarity of the two antibodies in the Sc value (0.58 vs. 0.57) and the buried solvent-accessible surface area (1108.6 Å2 vs. 1053.0 Å2). On the Fab side, the daclizumab paratope consists of five CDRs L2, L3, H1, H2, and H3, whereas the basiliximab paratope is composed of all of the six CDRs. On the IL-2Rα side, the epitopes of both daclizumab and basiliximab consist of several discontinuous segments of IL-2Rα and a large portion of the two epitopes overlap with each other. Nevertheless, the two epitopes differ to some extent. Residues 149-155 of the D2 domain form numerous interactions with the daclizumab Fab (3 hydrogen bonds and 29 van der Waals contacts), but have no direct contact with the basiliximab Fab. On the other hand, residues 56-57 of the D1 domain interact with the basiliximab Fab, but do not interact with the daclizumab Fab. In general, the basiliximab epitope relies more on the D1 domain whereas the daclizumab epitope relies evenly on both the D1 and D2 domains. Moreover, AspI4, which has intensive interactions (1 salt bridge, 3 hydrogen bonds, and 20 van der Waals contacts) and hence is a key residue for the recognition and binding of daclizumab, plays a less important role in the binding of basiliximab due to a smaller number of the interactions (1 hydrogen bond and 17 van der Waals contacts).
Inhibition mechanism of daclizumab on the IL-2 signaling pathway
The structural studies of the IL-2/IL-2Rαβγc complex have shown that the interactions between IL-2 and IL-2Rα involve 20 residues of IL-2 and 21 residues of IL-2Rα, forming 8 hydrogen bonds, 2 salt bridges, and 100 van der Waals contacts 5. In particular, residues 1-6, 25-29, 35-43, 57, and 64 of the D1 domain of IL-2Rα make a major contribution to the interactions, whereas residues 118-120 of the D2 domain make a minor contribution. Structural comparisons of the daclizumab Fab/IL-2Rα complex with the basiliximab Fab/IL-2Rα complex and the IL-2/IL-2Rαβγc complex reveal that the daclizumab epitope is largely shared by the basiliximab epitope and both epitopes overlap extensively with the regions of IL-2Rα that interact with IL-2 (Figure 3). Specifically, 11 out of 21 residues (52.4%) of IL-2Rα that interact with IL-2 are involved in the interactions with the daclizumab Fab, which account for 11 out of 18 residues and 722.6 Å2 (70.4%) of the buried solvent-accessible surface area at the interface. Two regions, including residues 4-6 of IL-2Rα that form a negatively charged surface patch and residues LeuI2, MetI25, LeuI42, TyrI43, and HisI120 that form a hydrophobic surface patch, are shared by both IL-2 and the daclizumab Fab (Figure 3B). In contrast, residues 149-155 of the D2 domain that contribute about one-third of the interactions between the daclizumab Fab and IL-2Rα have no contact with IL-2. Given that the binding affinity of IL-2Rα to IL-2 is 10 nM, which is significantly lower than that of IL-2Rα to daclizumab (0.27 nM), our structural and biochemical data suggest that, like basiliximab, daclizumab would competitively bind to the IL-2-binding site of IL-2Rα, and hence block the binding of IL-2Rα by IL-2 and the subsequent formation of the IL-2/IL-2Rαβγc complex, and inhibit the IL-2 signaling pathway when applied with a sufficient dose.
Mechanism of the inhibition of the IL-2 signaling pathway by daclizumab. (A) Structural comparison of the daclizumab Fab/IL-2Rα complex with the IL-2/IL-2Rαβγc complex. The daclizumab Fab is shown with a molecular surface with the light chain in yellow and the heavy chain in green. The IL-2/IL-2Rαβγc complex is superposed to the daclizumab Fab/IL-2Rα complex based on the position of IL-2Rα and shown with a stick model with IL-2 colored in blue, IL-2Rβ in red, and γc in orange, and the D1 and D2 domains of IL-2Rα in magenta and cyan, respectively. The occupation of daclizumab prevents the IL-2 binding by IL-2Rα and thus inhibits IL-2 signaling through the IL-2/IL-2Rαβγc complex. (B) Overlapping among the IL-Rα regions that bind daclizumab, basiliximab, and IL-2, respectively. The daclizumab epitope overlaps largely with the basiliximab epitope and the binding regions for IL-2. IL-2Rα is shown with a molecular surface. The binding regions and the overlapping areas are colored and marked differently.
Implications for therapeutic applications
The activation of T cells plays a central role in allograft rejection in organ transplantation. Although drugs such as corticosteroids, tacrolimus, and cyclosporine can suppress the immune responses successfully, the lack of specificity of these drugs for activated T cells leads to severe side effects, including nephrotoxicity (tacrolimus), hypertension, and metabolic effects (corticosteroids) 51. Thus, more specific immunosuppressive regimens are in demand. IL-2 signaling is pivotal for the activation and survival of T cells in allograft rejection, and IL-2Rα is critical for the activation of this pathway in T cells as the binding of IL-2 by IL-2Rα initiates the sequential assembly of the receptor 6, 7, 8, 9. Given that IL-2Rα is also the only IL-2-sepcific component of the IL-2Rαβγc receptor complex, blockage of IL-2Rα would efficiently and specifically interfere with the interactions between IL-2 and IL-2R on T cells, and thus provide therapeutic benefits of specific immunosuppression in organ transplantation.
Daclizumab is one such drug and improvement of the efficacy of the drug would be clinically beneficial. As the interface between the Fab and IL-2Rα has a relatively low Sc value of 0.58, the paratope of daclizumab may be modified to have a higher shape and chemical complementarity with IL-2Rα. In our previous work, it has been shown that modification of the rituximab paratope based on the structural information substantially enhances its binding affinity, and a rituximab variant displays an 18-fold higher affinity compared with the wild-type antibody and exhibits potent apoptosis-inducing activity 52. Similarly, structural analysis of the daclizumab Fab/IL-2Rα complex could provide some valuable information for the improvement of this mAb. For example, the side chain of GluI1 of IL-2Rα forms only one and four van der Waals contacts with SerL30 and SerL91, respectively. Substitution of SerL30 and SerL91 with positively charged residues such as arginine or lysine might provide favorable salt bridges. In addition, AspI6 stretches its side chain near the positively charged surface and forms only one van der Waals contact with AsnH52. Change of AsnH52 to a residue with a longer polar side chain might generate favorable hydrogen bonds. Together, mutations on CDR loops as mentioned above would form favorable interactions and hence result in a higher affinity for IL-2Rα.
In several clinical studies, murine mAbs and chimeric mAbs could induce human anti-mouse antibody and human anti-chimeric antibody responses, respectively, and the resulting neutralization would lead to limitation in efficacy. To date, a large number of humanized mAbs have been generated based on the computer-guided humanization technology to reduce immunogenicity, including daclizumab 53. It has been proposed that key framework residues that interact with CDRs are important for the integral capacity of antigen binding and should be retained during humanization 23. Based on the computer-generated three-dimensional model of the V domain of anti-Tac, nine amino acids were identified as key framework residues and retained as mouse-specific sequence in humanization, three of which are the same as human consensus sequence 39. All of these residues of daclizumab form extensive hydrophobic and hydrophilic interactions with nearby CDRs in our structure. It is worth noting that TyrL48 of FWR L2, which is the same as the mouse-specific amino acid and is next to the key framework residue IleL47, not only has extensive contacts with the nearby CDRs, but also makes 11 van der Waals contacts with the D2 domain of IL-2Rα. These results strengthen the notion that key FWR residues should be retained and also indicate that structural information would provide a guide in the humanization process for maintaining the binding affinity of parental murine antibodies and in further improvement of the humanized antibodies.
Materials and Methods
Protein preparation
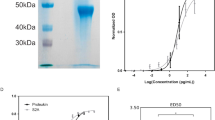
The Fab fragment was obtained by digestion of the mAb daclizumab (Roche Pharmaceuticals) with papain (Sigma) and further purification with exchange chromatography using a Q-Sepharose Fast-Flow column (GE Healthcare) and gel filtration chromatography using a Superdex G200 16/60 column (GE Healthcare). The protein fractions were pooled and concentrated to 15 mg/ml and then exchanged to a stock buffer of 10 mM HEPES (pH 7.2), and 50 mM NaCl. The recombinant human IL-2Rα ectodomain (residues 1-217) was expressed using a modified Bac-to-Bac baculovirus expression system (Invitrogen) and purified as described previously 38.
To obtain the daclizumab Fab/IL-2Rα ectodomain complex, the daclizumab Fab was mixed with the IL-2Rα ectodomain at a molar ratio of 1.5:1 at 4 °C for 12 h and then loaded onto a Superdex G200 16/60 column to remove excessive daclizumab Fab. The protein complex was eluted in the stock buffer and concentrated to 8 mg/ml for crystallization. The concentrations of the proteins were measured using a Bio-Rad protein assay kit and the purity and homogeneity were confirmed by SDS-PAGE and dynamic light-scattering analyses.
Crystallization and diffraction data collection
Crystallization was carried out using the hanging drop vapor diffusion method at 20 °C by mixing equal volumes (0.8 μl) of the protein solution and the reservoir solution and then equilibrated against 400 μl of the reservoir solution. Crystals of the free-form daclizumab Fab grew under the conditions of 0.1 M HEPES (pH 7.5), 0.1 M ammonium sulfate, and 2% PEG 400, and those of the Fab/IL-2Rα complex were formed under the conditions of 0.1 M Tris-HCl (pH 7.0), 0.2 M MgCl2, and 10% PEG 8000. Crystals of the free-form Fab were cryoprotected using the reservoir solution supplemented with 30% glycerol and then flash-cooled into liquid N2 stream. Diffraction data were collected using an in-house Rigaku R-AXIS IV ++ diffractometer and processed with the program suite CrystalClear 54. Crystals of the Fab/IL-2Rα complex were cryoprotected using the reservoir solution supplemented with 25% PEG 400 and then flash-cooled into liquid N2 stream. Diffraction data were collected at the Shanghai Synchrotron Radiation Facility beamline BL-17U1 and processed with the program suite HKL2000 55. The statistics of the diffraction data are summarized in Table 1.
Structure determination and refinement
The structure of the free-form daclizumab Fab was determined by the molecular replacement method implemented in the program Phaser 56, with the structure of the anti-CD20 antibody ofatumumab Fab (PDB code 3GIZ) 57 as the search model. There is one Fab molecule in the asymmetric unit. The structure of the daclizumab Fab/IL-2Rα ectodomain complex was determined by molecular replacement with the free-form daclizumab Fab structure as the search model, which revealed two Fab molecules in the asymmetric unit with outstanding solutions in both rotation function and translation function searches. In the initial electron density map, the daclizumab Fab had well-defined electron density but the IL-2Rα ectodomain had very poor density. Further trials using the molecular replacement method implemented in Phaser 56 and other commonly used programs with all of the four available structures of IL-2Rα (PDB codes 1Z92, 2B5I, 2ERJ, 3IU3) 5, 6, 7, 38 as templates failed to locate the correct positions of the IL-2Rα ectodomain. After several cycles of structure refinement of the Fab using the program Refmac5 58 and model building using the program Coot 59, the R factor and free R factor decreased from 43.9% and 44.3% to 34.7% and 41.2%, respectively, and the electron density for the IL-2Rα ectodomain developed gradually, which allowed us to manually fit the D1 and D2 domains of IL-2Rα (PDB code 3IU3) 38 into the electron density. The final structure model contains 129 of 217 residues of the IL-2Rα ectodomain; similar to other previously reported structures 5, 6, 7, 38, several flexible regions including the linker region between domains D1 and D2 (residues 65-100) and the C-terminal region (residues 166-217) have no defined electron density and could not be modeled. The stereochemical geometry of the structures was analyzed using the program Procheck 60. Structure analysis was carried out using the programs in the CCP4 61 and the PISA server 62. The figures were generated using the program Pymol (http://www.pymol.org). The statistics of the refinement and structure models are summarized in Table 1.
Surface plasmon resonance measurements
The kinetic studies of the interaction between daclizumab and the IL-2Rα ectodomain were performed with the surface plasmon resonance method using a Biacore 3000 instrument (GE Healthcare) at 25 °C. The daclizumab mAb was covalently coupled onto a CM5 sensor chip (GE Healthcare) using an amine coupling kit (GE Healthcare). The IL-2Rα ectodomain was dialyzed against the HBS buffer (GE Healthcare) and used as the analyte. The association was monitored for a 240-s period and the dissociation was monitored by flowing the HBS buffer for a 600-s period subsequently. An irrelevant mAb (Rituximab, a chimeric anti-CD20 mAb, Roche) was used as reference. The experimental data were analyzed with a 1:1 Langmuir model using the program BIAevaluation (GE Healthcare).
Accession codes
The coordinates and structure factors of the daclizumab Fab in free form and in complex with the IL-2Rα ectodomain have been deposited in the RCSB Protein Data Bank with the accession codes 3NFS and 3NFP, respectively.
Abbreviations
- (mAb):
-
monoclonal antibody
- (CDR):
-
complementarity determining region
- (FWR):
-
framework region
- (IL-2R):
-
interleukin-2 receptor
References
Smith KA . Interleukin-2: inception, impact, and implications. Science 1988; 240:1169–1176.
Nelson BH, Willerford DM . Biology of the interleukin-2 receptor. Adv Immunol 1998; 70:1–81.
Klebb G, Autenrieth IB, Haber H, et al. Interleukin-2 is indispensable for development of immunological self-tolerance. Clin Immunol Immunopathol 1996; 81:282–286.
Robb RJ, Munck A, Smith KA . T cell growth factor receptors. Quantitation, specificity, and biological relevance. J Exp Med 1981; 154:1455–1474.
Rickert M, Wang X, Boulanger MJ, Goriatcheva N, Garcia KC . The structure of interleukin-2 complexed with its alpha receptor. Science 2005; 308:1477–1480.
Stauber DJ, Debler EW, Horton PA, Smith KA, Wilson IA . Crystal structure of the IL-2 signaling complex: paradigm for a heterotrimeric cytokine receptor. Proc Natl Acad Sci USA 2006; 103:2788–2793.
Wang X, Rickert M, Garcia KC . Structure of the quaternary complex of interleukin-2 with its alpha, beta, and gammac receptors. Science 2005; 310:1159–1163.
Forsten KE, Lauffenburger DA . The role of low-affinity interleukin-2 receptors in autocrine ligand binding: alternative mechanisms for enhanced binding effect. Mol Immunol 1994; 31:739–751.
Rickert M, Boulanger MJ, Goriatcheva N, Garcia KC . Compensatory energetic mechanisms mediating the assembly of signaling complexes between interleukin-2 and its alpha, beta, and gamma(c) receptors. J Mol Biol 2004; 339:1115–1128.
Bazan JF . Structural design and molecular evolution of a cytokine receptor superfamily. Proc Natl Acad Sci USA 1990; 87:6934–6938.
Uchiyama T, Hori T, Tsudo M, et al. Interleukin-2 receptor (Tac antigen) expressed on adult T cell leukemia cells. J Clin Invest 1985; 76:446–453.
Sharon M, Klausner RD, Cullen BR, Chizzonite R, Leonard WJ . Novel interleukin-2 receptor subunit detected by cross-linking under high-affinity conditions. Science 1986; 234:859–863.
Teshigawara K, Wang HM, Kato K, Smith KA . Interleukin 2 high-affinity receptor expression requires two distinct binding proteins. J Exp Med 1987; 165:223–238.
Tsudo M, Kozak RW, Goldman CK, Waldmann TA . Demonstration of a non-Tac peptide that binds interleukin 2: a potential participant in a multichain interleukin 2 receptor complex. Proc Natl Acad Sci USA 1986; 83:9694–9698.
Ozaki K, Leonard WJ . Cytokine and cytokine receptor pleiotropy and redundancy. J Biol Chem 2002; 277:29355–29358.
Takeshita T, Asao H, Ohtani K, et al. Cloning of the gamma chain of the human IL-2 receptor. Science 1992; 257:379–382.
He YW, Malek TR . The structure and function of gamma c-dependent cytokines and receptors: regulation of T lymphocyte development and homeostasis. Crit Rev Immunol 1998; 18:503–524.
Rubin LA, Nelson DL . The soluble interleukin-2 receptor: biology, function, and clinical application. Ann Intern Med 1990; 113:619–627.
Waldmann TA . Daclizumab (anti-Tac, Zenapax) in the treatment of leukemia/lymphoma. Oncogene 2007; 26:3699–3703.
Waldmann TA . The multi-subunit interleukin-2 receptor. Annu Rev Biochem 1989; 58:875–911.
Waldmann TA . The meandering 45-year odyssey of a clinical immunologist. Annu Rev Immunol 2003; 21:1–27.
Waldmann TA . The structure, function, and expression of interleukin-2 receptors on normal and malignant lymphocytes. Science 1986; 232:727–732.
Queen C, Schneider WP, Selick HE, et al. A humanized antibody that binds to the interleukin 2 receptor. Proc Natl Acad Sci USA 1989; 86:10029–10033.
Ciancio G, Mattiazzi A, Roth D, Kupin W, Miller J, Burke GW . The use of daclizumab as induction therapy in combination with tacrolimus and mycophenolate mofetil in recipients with previous transplants. Clin Transplant 2003; 17:428–432.
Nashan B, Light S, Hardie IR, Lin A, Johnson JR . Reduction of acute renal allograft rejection by daclizumab. Daclizumab Double Therapy Study Group. Transplantation 1999; 67:110–115.
Pescovitz MD, Bumgardner G, Gaston RS, et al. Pharmacokinetics of daclizumab and mycophenolate mofetil with cyclosporine and steroids in renal transplantation. Clin Transplant 2003; 17:511–517.
Vincenti F, Kirkman R, Light S, et al. Interleukin-2-receptor blockade with daclizumab to prevent acute rejection in renal transplantation. Daclizumab Triple Therapy Study Group. N Engl J Med 1998; 338:161–165.
Vincenti F, Nashan B, Light S . Daclizumab: outcome of phase III trials and mechanism of action. Double Therapy and the Triple Therapy Study Groups. Transplant Proc 1998; 30:2155–2158.
Eckhoff DE, McGuire B, Sellers M, et al. The safety and efficacy of a two-dose daclizumab (zenapax) induction therapy in liver transplant recipients. Transplantation 2000; 69:1867–1872.
Emre S, Gondolesi G, Polat K, et al. Use of daclizumab as initial immunosuppression in liver transplant recipients with impaired renal function. Liver Transpl 2001; 7:220–225.
Innocenti F, Humeres R, Zamboni M, et al. IL-2 receptor blockers in liver transplantation: initial experience with daclizumab in Chile. Transplant Proc 2003; 35:2520–2521.
Koch M, Niemeyer G, Patel I, Light S, Nashan B . Pharmacokinetics, pharmacodynamics, and immunodynamics of daclizumab in a two-dose regimen in liver transplantation. Transplantation 2002; 73:1640–1646.
Anasetti C, Hansen JA, Waldmann TA, et al. Treatment of acute graft-versus-host disease with humanized anti-Tac: an antibody that binds to the interleukin-2 receptor. Blood 1994; 84:1320–1327.
Przepiorka D, Kernan NA, Ippoliti C, et al. Daclizumab, a humanized anti-interleukin-2 receptor alpha chain antibody, for treatment of acute graft-versus-host disease. Blood 2000; 95:83–89.
Rose JW, Watt HE, White AT, Carlson NG . Treatment of multiple sclerosis with an anti-interleukin-2 receptor monoclonal antibody. Ann Neurol 2004; 56:864–867.
Pascual J, Marcen R, Ortuno J . Anti-interleukin-2 receptor antibodies: basiliximab and daclizumab. Nephrol Dial Transplant 2001; 16:1756–1760.
Kandus A, Arnol M, Omahen K, et al. Basiliximab versus daclizumab combined with triple immunosuppression in deceased donor renal transplantation: a prospective, randomized study. Transplantation 2010; 89:1022–1027.
Du J, Yang H, Zhang D, et al. Structural basis for the blockage of IL-2 signaling by therapeutic antibody basiliximab. J Immunol 2010; 184:1361–1368.
Tsurushita N, Hinton PR, Kumar S . Design of humanized antibodies: from anti-Tac to Zenapax. Methods 2005; 36:69–83.
Binder M, Vogtle FN, Michelfelder S, et al. Identification of their epitope reveals the structural basis for the mechanism of action of the immunosuppressive antibodies basiliximab and daclizumab. Cancer Res 2007; 67:3518–3523.
Sheriff S, Jeffrey PD, Bajorath J . Comparison of CH1 domains in different classes of murine antibodies. J Mol Biol 1996; 263:385–389.
Al-Lazikani B, Lesk AM, Chothia C . Standard conformations for the canonical structures of immunoglobulins. J Mol Biol 1997; 273:927–948.
Gaboriaud C, Juanhuix J, Gruez A, et al. The crystal structure of the globular head of complement protein C1q provides a basis for its versatile recognition properties. J Biol Chem 2003; 278:46974–46982.
Davies DR, Cohen GH . Interactions of protein antigens with antibodies. Proc Natl Acad Sci USA 1996; 93:7–12.
Wilson IA, Stanfield RL . Antibody-antigen interactions. Curr Opin Struct Biol 1993; 234:946–950.
Lawrence MC, Colman PM . Shape complementarity at protein/protein interfaces. J Mol Biol 1993; 234:946–950.
Binder M, Otto F, Mertelsmann R, Veelken H, Trepel M . The epitope recognized by rituximab. Blood 2006; 108:1975–1978.
Du J, Wang H, Zhong C, et al. Structural basis for recognition of CD20 by therapeutic antibody Rituximab. J Biol Chem 2007; 282:15073–15080.
Riemer AB, Klinger M, Wagner S, et al. Generation of peptide mimics of the epitope recognized by trastuzumab on the oncogenic protein Her-2/neu. J Immunol 2004; 173:394–401.
Cho HS, Mason K, Ramyar KX, et al. Structure of the extracellular region of HER2 alone and in complex with the Herceptin Fab. Nature 2003; 421:756–760.
Perry I, Neuberger J . Immunosuppression: towards a logical approach in liver transplantation. Clin Exp Immunol 2005; 139:2–10.
Li B, Zhao L, Guo H, et al. Characterization of a rituximab variant with potent antitumor activity against rituximab-resistant B-cell lymphoma. Blood 2009; 114:5007–5015.
Reichert JM . Monoclonal antibodies in the clinic. Nat Biotechnol 2001; 19:819–822.
Pflugrath JW . The finer things in X-ray diffraction data collection. Acta Crystallogr 1999; D55:1718–1725.
Otwinowski ZMW . Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol 1997; 276:307–326.
McCoy AJ, Grosse-Kunstleve RW, Storoni LC, Read RJ . Likelihood-enhanced fast translation functions. Acta Crystallogr 2005; D61:458–464.
Du J, Yang H, Guo Y, Ding J . Structure of the Fab fragment of therapeutic antibody Ofatumumab provides insights into the recognition mechanism with CD20. Mol Immunol 2009; 46:2419–2423.
Murshudov GN, Vagin AA, Dodson EJ . Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr 1997; D53:240–255.
Emsley P, Cowtan K . Coot: model-building tools for molecular graphics. Acta Crystallogr 2004; D60:2126–2132.
Laskowski RA, Macarthur MW, Moss DS, Thornton JM . PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr 1993; 26:283–291.
CCP4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr 1994; D50:760–763.
Krissinel E, Henrick K . Inference of macromolecular assemblies from crystalline state. J Mol Biol 2007; 372:774–797.
Acknowledgements
We are grateful to the staff members at Shanghai Synchrotron Radiation Facility for support in diffraction data collection and other members of our group for helpful discussion. This work was supported by grants from the Ministry of Science and Technology of China (2010CB833601, 2006AA02A313, and 2009ZX09503-009), the National Natural Science Foundation of China (30730028 and 90713046), and the Chinese Academy of Sciences (KSCX2-YW-R-107 and SIBS2008002).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yang, H., Wang, J., Du, J. et al. Structural basis of immunosuppression by the therapeutic antibody daclizumab. Cell Res 20, 1361–1371 (2010). https://doi.org/10.1038/cr.2010.130
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2010.130
Keywords
This article is cited by
-
Antibody-guided design and identification of CD25-binding small antibody mimetics using mammalian cell surface display
Scientific Reports (2021)
-
An update on the use of biologic therapies in the management of uveitis in Behçet’s disease: a comprehensive review
Orphanet Journal of Rare Diseases (2017)
-
Human antibody 3E1 targets the HA stem region of H1N1 and H5N6 influenza A viruses
Nature Communications (2016)
-
Translational medicine in China I: Perspectives from Chinese physicians and scientists
Science China Life Sciences (2011)