Abstract
The role of Yamanaka factors as the core regulators in the induction of pluripotency during somatic cell reprogramming has been discovered recently. Our previous study found that Yamanaka factors regulate a developmental signaling network in maintaining embryonic stem (ES) cell pluripotency. Here, we established completely reprogrammed induced pluripotent stem (iPS) cells and analyzed the global promoter occupancy of Yamanaka factors in these cells by ChIP-chip assays. We found that promoters of 565 genes were co-bound by four Yamanaka factors in iPS cells, a 10-fold increase when compared with their binding in ES cells. The promoters occupied by a single Yamanaka factor distributed equally in activated and repressed genes in iPS cells, while in ES cells Oct4, Sox2, or Klf4 distributed mostly in repressed genes and c-Myc in activated ones. Pathway analysis of the ChIP-chip data revealed that Yamanaka factors regulated 16 developmental signaling pathways in iPS cells, among which 12 were common and 4 were unique compared to pathways regulated in ES cells. We further analyzed another recently published ChIP-chip dataset in iPS cells and observed similar results, showing the power of ChIP-chip plus pathway analysis for revealing the nature of pluripotency maintenance and regeneration. Next, we experimentally tested one of the repressive signaling pathways and found that its inhibition indeed improved efficiency of cell reprogramming. Taken together, we proposed that there is a core developmental signaling network necessary for pluripotency, with TGF-β, Hedgehog, Wnt, p53 as repressive (Yin) regulators and Jak-STAT, cell cycle, focal adhesion, adherens junction as active (Yang) ones; and Yamanaka factors synergistically regulate them in a Yin-Yang balanced way to induce pluripotency.
Similar content being viewed by others
Introduction
Induced pluripotent stem (iPS) cells have been obtained from various somatic cell types by forced expression of four transcription factors, Oct4, Sox2, Klf4, and c-Myc, which are subsequently referred to as “Yamanaka factors” 1, 2, 3, 4, 5, 6, 7. iPS cells are thought to have great potential in cell transplantation therapeutics, because of their embryonic stem (ES) cell-like properties namely self-renewal and capacity for differentiation into almost all cell types. Furthermore, the iPS technology, especially the recent patient-specific and virus-free iPS cell generation technique overcomes both the ethical issues concerning the use of human embryos and the clinical risk of immune rejection that hinders human ES cell research and application 8, 9, 10, 11. However, despite several proof-of-concept studies suggesting the potential to use iPS to cure diseases such as Parkinson's disease and sickle cell anemia 12, 13, the therapeutic application of iPS cells remains a long-term goal because of the limited understanding of reprogramming process at the molecular level 14.
Studies have shown that signaling network is critical to maintain the ground state of pluripotency as well as to control lineage-specific ES-cell differentiation. Mouse ES cells are commonly cultured in medium containing the cytokine leukemia inhibitory factor (LIF) and serum bone morphogenetic proteins (BMPs). LIF is believed to activate the STAT3 signaling pathway to maintain pluripotency while BMP signals act to inhibit differentiation 15, 16. When cultured in the absence of LIF and BMPs, ES cells can be maintained in a pluripotent state by inhibition of GSK3β signaling together with the elimination of the differentiation-inducing signaling from ERK1/2 mitogen-activated protein kinases 17. Lineage commitment of ES cells is also regulated by signals either from the environment or intrinsically within the cells. Wnt/β-catenin signaling is involved in the cardiomyocyte differentiation of mouse ES cells, while induction of ERK1/2 signaling triggers the differentiation of mouse ES cells towards the primitive endoderm lineage 18, 19. We thus speculate that the signal transduction network must play critical roles during pluripotency induction from somatic cells. Given the diverse origins of iPS cells, systematically studying the fundamental signaling network during reprogramming is of great interest.
Consistent with the critical roles of Yamanaka factors in ES cell pluripotency and iPS cell induction, our previous work reveals that Yamanaka factors regulate a crucial developmental signaling network comprised of 16 signaling pathways, including 9 with previously unknown roles in the maintenance of ES cell pluripotency 20. In the present study, we identified the promoters occupied by endogenous Yamanaka factors in iPS cells, analyzed their regulated signaling network, and tested the effect of several signaling pathways on Yamanaka factors-induced iPS generation. We also analyzed published data from another iPS cell study 21, and propose a core developmental signaling network underlying the induction of pluripotency.
Results
Establishment of a completely reprogrammed iPS cell line using an inducible system
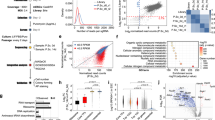
We generated an inducible lentivirus expression system for Yamanaka factors (Figure 1A) to obtain completely reprogrammed iPS cells. The inducible lentivirus vectors for Yamanaka factors were obtained by separately cloning the coding regions of Oct4, Sox2, c-Myc, and Klf4 into a lentivirus backbone under the control of a TetO-MinCMV promoter (Figure 1B), and the viruses were packaged in 293T cells using a second generation of lentivirus recombination system 22. Mouse embryonic fibroblast (MEF) cells from 129/C57 F1 mice E13.5 embryos were infected overnight with virus containing Oct4, Sox2, c-Myc and Klf4 together with a lentivirus constitutively expressing the reverse tetracycline transactivator (rtTA). The infected MEF cells were cultured first for 2 days in Dulbecco's modified Eagle's medium (DMEM) until they reached confluency and were then reseeded on feeder cell layers in standard mES medium. The medium was changed daily and Dox was added to the medium on day 2 post-infection, and the treatment continued until day 12. Emerged colonies were picked on day 12 and for several subsequent days until enough clones were collected. Finally, eight clones with homogenous ES-like morphology were further expanded in standard mES culture conditions until they formed stable ES-like cell lines. Four of these iPS cell lines, iPS-tet-A1, iPS-tet-B1, iPS-tet-B3, and iPS-tet-F1, were then analyzed for their property of pluripotency. All of them showed an ES-like morphology, were positive for alkaline phosphatase (AP) activity, and endogenously expressed ES pluripotency marker, Nanog (Figure 1C, shown are the representative data of iPS-tet-B3). We further evaluated the developmental potency of these iPS cell lines by injecting them into the ICR mice blastocysts. Remarkably, all four iPS cell lines tested gave rise to live chimeras with high coat color chimerism (Figure 1D, Supplementary information, Table S1). To test for germline transmission, chimeras derived from iPS-tet-B3 were mated with normal females. Live offsprings with agouti coat color were obtained, showing the origin from iPS cells (Figure 1D). iPS cells from clone iPS-tet-B3 were thus used in the following experiments.
Generation of completely reprogrammed iPS cell lines using an inducible system. (A) Schematic design of the inducible system by Yamanaka-factor-induced reprogramming. (B) The coding regions of Oct4, Sox2, c-Myc, and Klf4 were cloned into a lentivirus backbone under the control of a TetO-MinCMV promoter. (C) Established iPS cells show an ES-like morphology (Phase), express alkaline phosphotase (AP) and endogenous Nanog. (D) Established iPS cells give rise to live chimera (left panel) and contribute to germline (right panel).
Global mapping of promoter occupancy of Yamanaka factors in iPS cells
To characterize the global binding pattern of endogenous Yamanaka factors in iPS cells, iPS-Tet-B3 cells were maintained and expanded in normal mES medium without Dox, and ChIP-chip experiments were performed using the ChIP-grade antibodies against Oct4, Sox2, c-Myc, or Klf4 protein and a Mouse Promoter Tiling Microarray Set covering [−8 kb, +2.5 kb] (from transcriptional start sites, TSS) promoters of about 17 kb RefSeq transcripts 20. Target genes for Yamanaka factors were identified using Agilent G4477AA ChIP Analytics 1.3.1 software, where the “binding peak” in promoter regions and the binding activity of the four factors were inferred from Agilent's error modeling. The binding positions of each Yamanaka factor relative to the TSS of the corresponding genes are shown in Supplementary information, Table S2. Similar to what we previously performed in ES cells 20, the high quality nature of the ChIP-chip data is further validated using ChIP-qPCR assays with primer sets for several well-known target loci of each of the Yamanaka factors (data not shown).
The number of target genes of Oct4, Sox2, Klf4, and c-Myc is 1 388, 1 372, 1 832, and 2 531, respectively (Figure 2A), which is similar to the data that we obtained in ES cells 20 as well as the data published recently 21. Most target sites of these four factors are enriched in proximity to TSS (approximately ±2 kb from TSS) (Figure 2B), which is also observed in ES cells 20, confirming the reproducibility of our ChIP-chip experiment. To gain further insights into the binding pattern of the Yamanaka factors, we analyzed the number of target genes for each individual factor and different combinations of the four factors, and compared them with those in ES cells (Figure 2C, Supplementary information, Figure S1). Surprisingly, although the total number of target genes for each individual factor is largely the same within these two datasets, the number of target genes for different combinations of factors is quite different between iPS and ES cells. The total number of target genes occupied by only one factor is 4 000 in ES cells (Supplementary information, Figure S1), whereas the number in iPS cells is only 1 584. The number of target genes co-occupied by two factors only is also reduced from 1 083 in ES cells to 733 in iPS cells. In contrast, the number of target genes co-occupied by at least three factors or four factors is significantly increased. In iPS cells, the number of genes co-occupied by at least three factors is 1 104, a threefold increase compared to that in ES cells. More strikingly, there is a 10-fold increase in the number of target genes shared by all four factors, from 58 in ES cells to 565 in iPS cells. This may result largely from the changed binding pattern of c-Myc in iPS cells. In iPS cells, only one-third (717 of 2 531) of the c-Myc target genes are bound by c-Myc alone (Figure 2C), while in ES cells, over 70% (2 714 of 3 869) of c-Myc target genes are lone c-Myc targets. A similar iPS data set published recently corroborated these results (Supplementary information, Figure S2).
Summary of the endogenous Yamanaka factor occupancy in mouse iPS. (A) Number of target promoters occupied by each Yamanaka factor. (B) Chromosomal target loci distribution of each Yamanaka factor to the TSS. The -axis represents the relative distance to the TSS of target genes. (C) Number of target promoters occupied by multiple Yamanaka factors. Red line represents the accumulated number of target promoters by at least 1, 2, 3, or 4 factors. The bars represent the targets occupied by only 1, 2, 3, or 4 factors. In each bar, different color means different combination of Yamanaka factors. The 1 factor only bar means that the targets are occupied by only one of c-Myc, Klf4, Oct4, or Sox2 factors.
We then analyzed the possible effects of differential binding of the Yamanaka factors on the expression pattern of the target genes by supervised clustering and gene set enrichment analysis (GSEA) (Figure 3A). In ES cells, previous studies showed that genes occupied by only one of the non-c-Myc factors (Oct4, Sox2, or Klf4) were largely repressed, while genes occupied by two or more factors were activated 20. However, in iPS cells, we found that the expression of target genes bound by non-c-Myc factors showed a balanced pattern, with almost equal number of activated and repressed genes, no matter whether they were targeted by only one single factor or by multiple factors (Figure 3B–3D). Notably, while c-Myc mainly functions as an activator of gene expression 20 in ES cells, c-Myc target genes in iPS cells show a balanced expression pattern, regardless of being targeted by c-Myc alone or together with other factors (Figure 3B, 3C). Coincidently, in another recently published iPS ChIP-chip dataset, the targets of Oct4, Sox2, and Klf4 also show a nearly balanced expression pattern when bound by a single factor alone or by two factors together, while targets co-bound by Oct4, Sox2, and Klf4 together are largely repressed, and c-Myc mainly functions as a suppressor of gene expression (Supplementary information, Figure S3). Collectively, these data indicate that Yamanaka factors play some distinct roles in ES and iPS cells, with more synergetic effects on regulation of gene transcription in iPS cells.
Transcriptional profile regulated by Yamanaka factors in mouse iPS cells. (A) A supervised clustering showing different occupancy of the target genes by endogenous Yamanaka factors (left panel). Bar “a” represents the target genes occupied by three or four Yamanaka factors; bar “b” represents the target genes occupied by two factors; and bar “c” represents the targets occupied by only one factor. The moving window average (bin size 25 and step size 1) of the expression values (Log2) of the target genes in control mouse iPS cells and MEF is shown in the right panel. (B-D) Analysis of the relationship between Yamanaka factors occupancy and target gene expression profile using Gene Set Enrichment Analysis (GSEA). (B) The four kinds of target genes were almost equally distributed in active and repressed genes in iPS cells. (C) Co-occupancy with c-Myc did not affect the regulation of Oct4, Sox2, or Klf4 on the target gene expression. (D) Without c-Myc, the targets were distributed equally in active and repressed genes when occupied by Oct4, Sox2, or Klf4 alone, mostly in active genes when occupied by any of two factors, or mostly in repressed genes when occupied by all three factors.
Characterization of signaling pathways regulated by Yamanaka factors in iPS cells
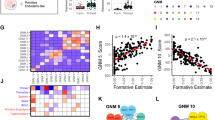
To investigate the signaling network regulated by Yamanaka factors in iPS cells, we analyzed our iPS data using PANTHER (Protein ANalysis Through Evolutionary Relationships, http://www.pantherdb.org/), as we did in ES cells 20, and compared the two cases. The primary categorization of KEGG pathways is shown in Supplementary information, Table S3 and the pathways regulated by each and multiple Yamanaka factors are listed in Supplementary information, Tables S3,S4,S5,S6,S7. In iPS cells, the genes only bound by one of the three non-c-Myc factors (Oct4, Sox2, or Klf4) are enriched mainly in developmental processes and mRNA transcription processes, while genes bound by c-Myc alone are mainly enriched in the protein metabolism and modification systems (Figure 4A). These observations are consistent with that in ES cells 20. However, when the total targets of c-Myc were analyzed, they were found to be enriched significantly in metabolism system, as well as in developmental processes and mRNA transcription processes (Figure 4B), which is not the case in ES cells 20. Interestingly, in iPS cells the union target genes of Oct4, Sox2, and Klf4 are also enriched in the metabolism process (Figure 4B). Further analysis of target genes in different combinations of the four factors revealed that the targets of Oct4 and Sox2 are enriched in both developmental and metabolism processes either with or without c-Myc and Klf4 (Figure 4C, 4D). This suggests that in iPS cells, in addition to their role in developmental signaling, Oct4 and Sox2 may act together to function as metabolism regulators, which is not observed in ES cells 20. Similarly, PANTHER analysis of c-Myc target genes gave similar results (Supplementary information, Figure S4). To investigate the synergistic effect of Yamanaka factors in iPS cells, we analyzed the hierarchical clustering of the four factors on the basis of their targets correlations. Interestingly, although the correlation between Oct4 and Sox2 remains similarly highest in ES and iPS cells (0.521 and 0.626, respectively), the correlation between other factors becomes closer in iPS than in ES cells (Figure 4E). The most striking difference is again in c-Myc. In ES cells, our previous analysis shows that c-Myc correlated poorly to the other three factors, with the correlation coefficients with Oct4, Sox2, and Klf4 being −0.223, −0.249, −0.294, respectively 20. However, in iPS cells, the correlation coefficients of c-Myc with Oct4, Sox2, and Klf4 are 0.356, 0.337, and 0.282, respectively (Figure 4E). Our analysis of another recently published iPS dataset 21 shows similar results (Supplementary information, Figure S5) and the enriched pathways identified using these data are listed in Supplementary information, Table S8. All together, these data indicate that in iPS cells, the four Yamanaka factors function more synergistically with each other in regulating developmental signaling as well as other processes.
Functional analysis of cell signaling role of Yamanaka factors' targeted genes using PANTHER classification system. The y-axis represents the relative enrichment calculated as the following: the obtained number of genes in our binding lists for “developmental process”, “mRNA transcription”, “protein metabolism and modification”, and “cell cycle” is divided by the expected number of genes calculated for all mouse genes. The values above and below one indicate the enrichment or depletion of the target genes in the category, respectively. The numbers above each bar represents “the genes included in the category”/“the genes expected in the category”. All targets occupied by Oct4, Sox2, Klf4, or c-Myc, and the targets occupied by Oct4, Sox2, Klf4, or c-Myc only were separately analyzed in (A) and (B). Target genes were mainly enriched during developmental process and mRNA transcription. (C) “Oct4 plus Sox2” represents the core cluster genes that are occupied by the factors containing Oct4 or Sox2; “Oct4 plus Sox2 only” represents the genes occupied by the factors containing Oct4 or Sox2, but not Klf4 and c-Myc; “Oct4 plus Sox2 and Klf4” represents the “Oct4 plus Sox2” genes co-occupied by Klf4; “Oct4 plus Sox2 and c-Myc” represents the “Oct4 plus Sox2” genes co-occupied by c-Myc; “Oct4 plus Sox2 and Klf4 and c-Myc” represents the “Oct4 plus Sox2” genes co-occupied by Klf4 and c-Myc together. The “Oct4 plus Sox2 and c-Myc only” core genes were enriched during the developmental processes, mRNA transcription, and cell cycle, but not during the protein metabolism. Each of the combination has lower relative enrichment in protein metabolism. (D) “Oct4 plus Sox2 plus Klf4” represents the core cluster of genes that are occupied by the factors containing Oct4, Sox2, or Klf4. “Oct4 plus Sox2 plus Klf4 only” represents the genes occupied by the factors containing Oct4, Sox2, or Klf4, but not c-Myc; “Oct4 plus Sox2 plus Klf4 and c-Myc” represents the “Oct4 plus Sox2 plus Klf4” genes co-occupied by c-Myc. The “Oct4 plus Sox2 plus Klf4” core genes were highly enriched during developmental processes, mRNA transcription, and cell cycle. In protein metabolism, each of the combination still has lower relative enrichment. (E) Hierarchical clustering of Yamanaka factors based on their target correlations reveals the overall target similarity among Oct4, Sox2, and Klf4.
Defining a signaling network controlling pluripotency in iPS cells
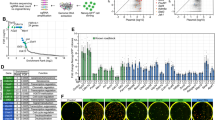
To further explore how the Yamanaka factors are involved in regulating signaling pathways in iPS cells, we used KOBAS 23, 24 to clarify the pathways regulated by the four factors into subcategories of development, cancer, metabolism, and other signaling pathways (Figure 5A). Consistent with the results from PANTHER analysis, the distributions of pathways regulated by Yamanaka factors between different subcategories in iPS cells also differ from those in ES cells. In iPS cells, c-myc regulates 38 signaling pathways. Out of them, 16 are developmental signaling pathways. However, in ES cells, c-myc only regulates 9 develpomental signaling pathways out of 47 total regulated pathways. The number of developmental signaling pathways related to Klf4 is 13 out of 29 in iPS cells, but 6 out of 32 in ES cells. Moreover, although ES and iPS cells share most of the developmental signaling pathways identified (12 out of 16 pathways overlap), they do have some differences (Figure 5B). Four signaling pathways we previously identified in ES cells 20, axon guidance, cytokine-cytokine receptor interaction, dorsal-ventral axis formation, and Notch signaling pathways, are not identified in iPS cells, while VEGF signaling pathway, calcium signaling, tight junction, and mTOR signaling pathways are unique to iPS cells (Figure 5B). We also similarly analyzed another published ChIP-chip data on iPS cells and partially reprogrammed iPS (piPS) cells 21 for signaling pathways regulated by Yamanaka factors and obtained similar results (Supplementary information, Figure S6). In this set of data, 15 and 14 developmental signaling pathways are regulated in ES and iPS cells, respectively, among which 12 overlap. Ten developmental signaling pathways are identical in iPS cells of the two datasets, further validating the power of ChIP-chip and pathway analysis. As expected, some signaling pathways observed in iPS cells are different from those in piPS cells (Supplementary information, Figure S6). On the basis of these observations, we experimentally tested the effect of MAPK signaling, one repressive regulator in iPS cell induction identified by us, and found that inhibition of the JNK and Erk1/2 MAPK pathways indeed significantly improves iPS cell generation efficiency (Figure 6). Finally, we compared these four groups of pathways found in ES and iPS cells, and identified a core developmental signaling network regulated by Yamanaka factors (Figure 5C). Totally, eight pathways are core pathways in iPS and ES cells, among them pathways of TGF-β, Hedgehog, Wnt, and p53 are repressed (Yin), while those of Jak-STAT, cell-cycle, focal adhesion, adherens junction are activated (Yang) in pluripotent state. Other developmental signaling pathways, including MAPK, VEGF, Notch, gap junction, axon guidance, and tight junction pathways, were identified as Yin pathways, while calcium signaling, cytokine-cytokine receptor interaction, ErbB signaling, mTOR signaling, and dorso-ventral axis formation signaling pathways were identified as Yang pathways supporting the core network.
Developmental signaling pathways regulated by Yamanka factors in iPS cells. (A) Pathway classification of the target genes of Yamanka factors in iPS cells using KOBAS. The numbers represent the number of pathways that are categorized into the indicated process. (B) Developmental signaling pathways regulated by Yamanaka factors in mES and iPS cells. +/−: Signaling pathways regulated or not regulated by Yamanaka factors. (C) Core developmental signaling network regulated by Yamanaka factors. The developmental signaling pathways located in the pink circle are enriched in both ES and iPS data. The round rectangle shapes are the developmental signaling pathways only enriched in ES, while the octagon shapes are the pathways only enriched in iPS. (Red: the activated pluripotency-associated pathways; green: the repressed pluripotency-associated pathways).
Role of MAPK signaling in Yamanaka-factors-induced reprogramming. (A) Representative morphology of GFP-positive colonies emerged on twelfth day post-infection. Left panel: phase contrast; right panel: GFP. (B) The numbers of GFP-positive colonies on the twelfth day post-infection upon four-factor induction (Ctrl), four-factor induction plus VPA (2 mM), four-factor induction plus p38 inhibitor SB203580 (1 μM), four-factor induction plus JNK inhibitor SP600125 (10 μM), and two ERK inhibitors, PD98059 (10 μM) and U0126 (10 μM), were counted and normalized as fold increase relative to Ctrl. Statistics were carried out by one-way ANOVA followed by LSD post hoc test. *P < 0.05, **P < 0.01, ***P < 0.001.
Discussion
In the present study we focused on the signaling network regulated by the Yamanaka factors during the reprogramming process. By analyzing the genome-wide promoter occupancy of the four factors, we provide a set of data about different signaling pathways that may regulate pluripotency, which may be a useful resource for researchers in both ES and iPS cell community. We also carried out signaling pathway analysis from another recently published ChIP-chip dataset on the promoter occupancy of the four factors in ES and iPS cells 21. We identified a core developmental signaling network regulated by Yamanaka factors through analyzing and comparing the developmental signaling pathways from the four ChIP-chip datasets (Figure 5, Supplementary information, Figure S6). We found that 12 developmental signaling pathways overlap between the 16 pathways obtained each from our iPS cell data and those from our ES cell data, while a recently published ChIP-chip dataset 21 identified 15 and 14 developmental signaling pathways in ES and iPS cells, respectively, among which 12 are identical. Furthermore, between the two iPS datasets, 10 pathways are identical, including Wnt, Jak-STAT, MAPK, p53, TGF-β, VEGF, Hedgehog, adherens junction, cell cycle, and focal adhesion signaling pathways. Collectively, despite the differential binding of Yamanaka factors in two different ES cells 20, 21 and different iPS cells, the signaling pathways revealed by these datasets are highly overlapped. ChIP-chip plus pathway analysis is thus a powerful tool for discovering the molecular mechanisms of the reprogramming process.
In Yamanaka factor-regulated core signaling pathways, the role of some pathways in cell reprogramming has been confirmed. Wnt signaling and p53 signaling pathways were known to be regulated by Yamanaka factors in both iPS and ES cells, and their role in Yamanaka factors-induced reprogramming or in cell fusion-mediated reprogramming has been proven recently 25, 26. Calcium signaling pathway is not a previously reported pathway related with pluripotency, however, it was found to be an active regulator in iPS cells by our data. Interestingly, the improvement of reprogramming efficiency through activation of calcium signaling has been recently discovered through a large-scale screening of small chemicals 27. PI3K signaling pathway was found in piPS cells, while the MAPK signaling pathways, including JNK, ERK1/2 and p38 MAPK pathways were identified as repressive regulators in iPS cells, and the functions of JNK, ERK1/2, and PI3K signaling pathways in induction of pluripotency were also confirmed by our experiments (Figure 6 and data not shown). These results provide further evidence for the quality of data and significance of our ChIP-chip plus pathway analysis, suggesting that our study not only reveals knowledge about the signaling nature of iPS and ES cells, but also provides useful insights for the development of new reprogramming methods. Further exploration of the roles of pathways with previously unknown functions in cell pluripotency maintenance such as mTOR signaling, tight junction signaling and GnRH signaling pathways in the reprogramming process will be of great interest to reveal the specific roles of these pathways and Yamanaka factors during cell reprogramming and development.
Consistent with the specific identity of different ES and iPS cell lines 20, we noticed that there are some differences between the pathways obtained from using our data and those from another published data (Figure 5B, Supplementary information, Figure S6C). In our data, there are 16 developmental signaling pathways regulated by Yamanaka factors in iPS cells, while 14 enriched pathways are identified in another recently published iPS dataset. Interestingly, among these two datasets, 10 pathways are overlapped. When the data from ES and iPS cells are analyzed, eight pathways including Jak-STAT, Wnt signaling, TGF-β, p53, Hedgehog, cell cycle, adherens junction, and focal adhesion pathways are identified as core pathways that associate with pluripotency. VEGF signaling is not found in our ES data (Figure 5B) and MAPK signaling is not found in the published ES data (Supplementary information, Figure S6C). Meanwhile, four pathways, axon guidance, CAM ligands, ECM-receptor interaction and Notch signaling are enriched in the published iPS and ES data, while six pathways, apoptosis, calcium signaling, ErbB, gap junction, mTOR and tight junction are not enriched in the recently published iPS data 21. These observations indicate that despite the many similarities, different iPS cell lines may possess some specific characteristics. Thus, properties of ES and iPS cells should be fully investigated and optimized before the application of specific iPS cells.
In addition to the developmental and signaling pathways discussed above, we also investigated other pathways such as metabolism pathways. We found that the following pathways were highly enriched in ES and iPS cells, but not in piPS cells. These include hypoxia signaling, VDR/RXR activation, CCR3 signaling, cell cycle G1/S checkpoint regulation, calcium-induced apoptosis, IGF-1 signaling, Aryl hydrocarbon receptor signaling, ephrin receptor signaling, protein ubiquitination pathway, mitochondria dysfunction, etc. In contrast, pathways such as clathrin-mediated endocytosis and actin cytoskeleton signaling were enriched in piPS cells but not in iPS cells or ES cells. Further exploration of these unexpected pathways may give us insights about pluripotency and cell reprogramming.
In summary, through ChIP-chip plus pathway analysis of promoter occupancy of Yamanaka factors, we found that Yamanaka factors function more synergistically in iPS cell induction, and identified Yin-Yang-balanced developmental signaling pathways in maintaining and regenerating cell pluripotency. Our study thus not only provides a useful resource and a powerful methodology to understand the nature of induced pluripotency, but also provides a variety of potential choices to improve the iPS cell induction efficiency through specific signaling pathway activators or inhibitors. Further exploration of the similarity and difference between piPS, iPS, and ES cells revealed here will open new doors to understand the molecular nature of cell reprogramming and development.
Materials and Methods
Virus production and iPS cell generation
The coding region of Oct4, Sox2, c-Myc, and Klf4 were amplified from pMXs-Oct4, pMXs-Sox2, pMXs-c-Myc, and pMXs-Klf4 plasmids (purchased from Addgene), respectively, and inserted between the BamHI and SalI site in the Lv-Tre lentivirus vector following BamHI/XhoI double digestion. The constitutively expressed rtTA lentivirus vector was a kind gift of Dr Lei Xiao. All plasmids were sequenced before being used in our experiments.
Viruses were produced in 293T cells using a second generation of lentivirus packaging system. Briefly, 4.0×106 293T cells were transfected with 10 μg of the lentivirus vector together with 7.5 μg of the delta-8.91 vector and 5 μg of the VSVG vector following the standard calcium transfection procedure. Viruses were harvested 48 h after transfection, filtered through a 0.45 μM filter, and stored at 4 °C before use.
MEF cells were generated from the E13.5 129/C57 F1 mice embryos, as previously described 1. For iPS cell induction, 8×104 MEF cells were seeded in a 35-mm plate. At 12 h after seeding, 1 ml each of the Oct4, Sox2, c-Myc, Klf4, and rtTA virus was mixed and added to the MEF culture. Polybrene was added to the culture in a final concentration of 8 μg/ml. At 24 h after infection, the viruses were removed. Infected MEF cells were then cultured in DMEM for 2 days and reseeded on irradiated MEFs at a density of 1.0×104 cells/cm2 in typical mES cell media consisting of DMEM (GIBCO) supplemented with 15% heat-inactivated fetal bovine serum (GIBCO), 0.055 mM beta-mercaptoethanol (GIBCO), 2 mM L-glutamine, 0.1 mM MEM non-essential amino acid, and 1 000 U/ml LIF (Chemicon). Potential iPS colonies began to appear 12 days after infection. These colonies were individually picked and expanded for further analysis.
Immunostaining and AP staining
iPS cells were seeded at a density of 1.0×104 cells/cm2 on irradiated MEFs in 24-well plates and immunostained for Nanog expression 2 days later. Briefly, cells were washed twice with PBS, fixed with 4% PFA/PBS for 30 min, permeablized with 0.2% Triton X-100 for 10 min, and blocked with 1% BSA/PBS for 2 h. Cells were then incubated with a primary antibody against Nanog (Santa Cruz, Catalog number sc-33760) diluted 1:100 in 1% BSA/PBS for 2 h, washed twice with 1% BSA/PBS, incubated with a Texas-red conjugated secondary antibody diluted 1:100 in 1% BSA/PBS for 2 h, washed twice with 1% BSA/PBS, covered with PBS, and scanned for fluorescence under a microscope.
For AP staining, iPS cells were seeded on irradiated MEFs in 24-well plates at a density of 1.0×104 cells/cm2. At 2 days after seeding, AP staining was carried out using an Alkaline Phosphatase Detection Kit (Chemicon, Catalog number SCR004), following the manufacturers instructions.
Blastocyst injections
Blastocysts derived from ICR mice were placed in a drop of DMEM with 15% FCS under mineral oil. A flat tip microinjection pipette was used for iPS cell injection. After injection, blastocysts were transferred to each uterine horn of 2.5-day post-coitum pseudo-pregnant females. For test of germline transmission, chimeras derived from iPS cells were mated with normal ICR females.
Chromatin immunoprecipitation and hybridization
Chromatin immunoprecipitation and hybridization were carried out following the standard Agilent mammalian ChIP-chip protocol 9.1 (available online: http://www.chem.agilent.com/scripts/generic.asp?lpage=11617&indcol=N&prodcol=Y), as previously described 20. Briefly, iPS-tet-B3 cells were grown to a final count of 1×108 cells for each ChIP-chip experiment. Cells were harvested with diastase and chemically cross-linked in 50 ml of 0.5% formaldehyde for 20 min at room temperature. Cells were then rinsed twice with 50 ml of 1× PBS and stored at −80 °C prior to use. Cells were resuspended and lysed in lysis buffers, and then sonicated to shear the cross-linked DNA to an average length of 500 bp. Since the sonication conditions vary mainly depending on cell line, cell number, degree of cross-linking, and equipment, we used a Bioruptor sonicator (Diagenode) and sonicated the 1×108 iPS cells with intensity set at high for 26×15-s pulse (30-s pause between each pulse) at 4 °C, while the samples were kept immersed in an ice-water bath. In total, 50 μl of the sonicated lysate was saved for whole-cell extract (WCE) DNA extraction. The remaining lysate was incubated overnight at 4 °C with 100 μl of Dynal Protein A magnetic beads pre-incubated with 10 μg of specific antibody. The Dynal beads were washed five times with RIPA buffer and once with TE containing 50 mM NaCl. To elute the bound complex from beads, 210 μl of the elution buffer was added and the beads were incubated for 15 min at 65 °C with interval vortexing at every 2 min. As for the WCE DNA extraction, 150 μl of elution buffer was added to 50 μl of the sonicated lysate. Both the immunoprecipitated DNA sample eluted from the Dynal beads and the WCE DNA sample were incubated overnight at 65 °C to reverse the chemical cross-linking between protein and DNA. The DNA samples were then purified by treatment with RNaseA, proteinase K, and phenol:chloroform:isoamyl alcohol extraction. Purified DNA was treated with Whole Genome Amplification Kit (WGA2, Sigma) 28, which allowed more linear amplification of a small amount of DNA than traditional ligation-mediated PCR 29. We used a standard protocol for modified WGA amplification (http://www.genomecenter.ucdavis.edu/farnham/protocol.html). Amplified DNA was labeled and purified using Invitrogen Bioprime random primer labeling kits (immunoprecipitated DNA was labeled with Cy5 fluorophore; WCE DNA was label with Cy3 fluorophore). The labeled IP and WCE DNA were combined in equal amounts (5 μg) and hybridized to mouse promoter arrays in Agilent hybridization chambers for 40 h at 65 °C. After washing, slides were scanned and analyzed. At least three biological replicates of hybridization were performed for each sample.
Array design and data extraction
The Mouse Promoter ChIP-on-chip Microarray Set used in this study was manufactured by Agilent Technologies (http://www.agilent.com). Each microarray set (product number: G4490A) contains two slides (design number: slide 1=014716 and slide 2=014717) that cover [−8 kb, +2.5 kb] (from transcriptional start sites, TSS) promoters of ∼17 000 mouse RefSeq transcripts. These arrays were designed on the basis of the UCSC mm8, NCBI build 36 mouse genome (February 2006). Data were extracted from image files by Feature Extraction V9.5.3.1, and peak analysis was performed using Agilent ChIP Analytics V1.3.1.
Microarray data processing
The gene expression data were retrieved from a previous study 21, in which RNA was extracted from duplicate samples of V6.5 and E14 ES cells, 1D4 and 2D4 iPS cells, 1A2 and a single sample of 1B3 partially reprogrammed cells, and four samples of three different MEF lines. The gene expression data were then analyzed by GSEA, which is a computational method that determines whether an a priori defined set of genes shows statistically significant, concordant difference between two biological states or phenotypes 30, 31. Experimental expression values were analyzed by GSEA.
In order to produce the supervised clustering image given in Figure 3A (left panel), we first performed unsupervised hierarchical clustering of the transcription factors on the basis of the correlation of their target genes, using Cluster and Treeview software 32. To produce a cluster view of common targets of multiple factors and unique targets of single factor, we first randomized the order of target genes and then sequentially sorted targets of each factor. For the target gene expression profile image given in Figure 3A (right panel), the expression value of each gene on iPS was measured by duplicate 1D4 and 2D4, and four samples of three different MEF lines were averaged. We then applied moving window average on the expression values of target genes in the order determined in Figure 3A (left panel).
PANTHER analysis
Gene ontology analysis was performed with the Panther Classification System. The analysis procedure consists of uploading Yamanaka target gene list into the system and submitting the list. The map lists genes to a PANTHER ontology. The binomial statistics tool is used to compare classifications of multiple clusters of lists to a reference list to statistically determine over- or underrepresentation of PANTHER classification categories. The results of the target genes of each transcription factor were combined and compared to produce Figure 4.
Visualization of signaling pathway network
Cytoscape software version 2.6.2 36 was used to visualize the signaling regulatory networks shown in Figure 5C.
( Supplementary information is linked to the online version of the paper on the Cell Research website.)
References
Takahashi K, Yamanaka S . Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006; 126:663–676.
Takahashi K, Tanabe K, Ohnuki M, et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 2007; 131:861–872.
Okita K, Ichisaka T, Yamanaka S . Generation of germline-competent induced pluripotent stem cells. Nature 2007; 448:313–317.
Wernig M, Meissner A, Foreman R, et al. In vitro reprogramming of fibroblasts into a pluripotent ES-cell-like state. Nature 2007; 448:318–324.
Nakagawa M, Koyanagi M, Tanabe K, et al. Generation of induced pluripotent stem cells without Myc from mouse and human fibroblasts. Nat Biotechnol 2008; 26:101–106.
Aoi T, Yae K, Nakagawa M, et al. Generation of pluripotent stem cells from adult mouse liver and stomach cells. Science 2008; 321:699–702.
Yu J, Vodyanik MA, Smuga-Otto K, et al. Induced pluripotent stem cell lines derived from human somatic cells. Science 2007; 318:1917–1920.
Kaji K, Norrby K, Paca A, et al. Virus-free induction of pluripotency and subsequent excision of reprogramming factors. Nature 2009; 458:771–775.
Woltjen K, Michael IP, Mohseni P, et al. piggyBac transposition reprograms fibroblasts to induced pluripotent stem cells. Nature 2009; 458:766–770.
Park I-H, Arora N, Huo H, et al. Disease-specific induced pluripotent stem cells. Cell 2008; 134:877–886.
Soldner F, Hockemeyer D, Beard C, et al. Parkinson's disease patient-derived induced pluripotent stem cells free of viral reprogramming factors. Cell 2009; 136:964–977.
Hanna J, Wernig M, Markoulaki S, et al. Treatment of sickle cell anemia mouse model with iPS cells generated from autologous skin. Science 2007; 318:1920–1923.
Wernig M, Zhao J-P, Pruszak J, et al. Neurons derived from reprogrammed fibroblasts functionally integrate into the fetal brain and improve symptoms of rats with Parkinson's disease. Proc Natl Acad Sci USA 2008; 105:5856–5861.
Yamanaka S . A fresh look at iPS cells. Cell 2009; 137:13–17.
Niwa H, Burdon T, Chambers I, Smith A . Self-renewal of pluripotent embryonic stem cells is mediated via activation of STAT3. Genes Dev 1998; 12:2048–2060.
Ying QL, Nichols J, Chambers I, Smith A . BMP induction of Id proteins suppresses differentiation and sustains embryonic stem cell self-renewal in collaboration with STAT3. Cell 2003; 115:281–292.
Ying Q-L, Wray J, Nichols J, et al. The ground state of embryonic stem cell self-renewal. Nature 2008; 453:519–523.
Naito AT, Shiojima I, Akazawa H, et al. Developmental stage-specific biphasic roles of Wnt/beta-catenin signaling in cardiomyogenesis and hematopoiesis. Proc Natl Acad Sci USA 2006; 103:19812–19817.
Chazaud C, Yamanaka Y, Pawson T, Rossant J . Early lineage segregation between epiblast and primitive endoderm in mouse blastocysts through the Grb2-MAPK pathway. Dev Cell 2006; 10:615–624.
Liu X, Huang J, Chen T, et al. Yamanaka factors critically regulate the developmental signaling network in mouse embryonic stem cells. Cell Res 2008; 18:1177–1189.
Sridharan R, Tchieu J, Mason MJ, et al. Role of the murine reprogramming factors in the induction of pluripotency. Cell 2009; 136:364–377.
Zufferey R, Dull T, Mandel RJ, et al. Self-inactivating lentivirus vector for safe and efficient in vivo gene delivery. J Virol 1998; 72:9873–9880.
Wu J, Mao X, Cai T, Luo J, Wei L . KOBAS server: a web-based platform for automated annotation and pathway identification. Nucleic Acids Res 2006; 34:W720–W724.
Mao X, Cai T, Olyarchuk JG, Wei L . Automated genome annotation and pathway identification using the KEGG Orthology (KO) as a controlled vocabulary. Bioinformatics 2005; 21:3787–3793.
Marson A, Foreman R, Chevalier B, et al. Wnt signaling promotes reprogramming of somatic cells to pluripotency. Cell Stem Cell 2008; 3:132–135.
Zhao Y, Yin X, Qin H, et al. Two supporting factors greatly improve the efficiency of human iPSC generation. Cell Stem Cell 2008; 3:475–479.
Shi Y, Desponts C, Do JT, Hahm HS, Scholer HR, Ding S . Induction of pluripotent stem cells from mouse embryonic fibroblasts by Oct4 and Klf4 with small-molecule compounds. Cell Stem Cell 2008; 3:568–574.
Jin VX, O'Geen H, Iyengar S, Green R, Farnham PJ . Identification of an OCT4 and SRY regulatory module using integrated computational and experimental genomics approaches. Genome Res 2007; 17:807–817.
O'Geen H, Nicolet CM, Blahnik K, Green R, Farnham PJ . Comparison of sample preparation methods for ChIP-chip assays. Biotechniques 2006; 41:577–580.
Subramanian A, Tamayo P, Mootha VK . Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 2005; 102:15545–15550.
Mootha VK, Lindgren CM, Eriksson KF, et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet 2003; 34:267–273.
Eisen MB, Spellman PT, Brown PO, Botstein D . Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 1998; 95:14863–14868.
Huang da W, Sherman BT, Lempicki RA . Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 2009; 4:44–57.
Huang da W, Sherman BT, Tan Q, et al. DAVID Bioinformatics Resources: expanded annotation database and novel algorithms to better extract biology from large gene lists. Nucleic Acids Res 2007; 35:W169–W175.
Huang da W, Sherman BT, Tan Q, et al. The DAVID Gene Functional Classification Tool: a novel biological module-centric algorithm to functionally analyze large gene lists. Genome Biol 2007; 8:R183.
Shannon P, Markiel A, Ozier O, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 2003; 13:2498–2504.
Acknowledgements
This research was supported by grants from the Ministry of Science and Technology (2005CB522406, 2007CB947904, 2007CB947100, 2007CB948000, 2010CB944900, and 2009CB941100), the National Natural Science Foundation of China (30621091, 30625014, 30623003, 30871285, 90713047, and 90919028), the Shanghai Municipal Commission for Science and Technology (07PJ14096, 08dj1400500, 09DZ2260100, and 06DZ22032), and the Chinese Academy of Sciences (KSCX2-YW-R-56, 2007KIP204, and SIBS2008001). We thank Dr Jian Zhao, Dr Lei Xiao and Shunmei Xin for their helpful discussions and technical assistance.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary information Table S1
Chimera formation and germline transmission of tested iPS cell lines (XLS 15 kb)
Supplementary information Table S3
Primary Categorization of KEGG pathways. (PDF 20 kb)
Supplementary information Table S4
Significantly enriched developmental signaling pathways by the target genes of endogenous Yamanaka factors in iPS and E14.1 mouse ES cells. (PDF 7 kb)
Supplementary information Table S5
All the significantly enriched KEGG pathways for each endogenous Yamanaka factor (PDF 14 kb)
Supplementary information Table S6
Comparison of the developmental signaling pathways significantly enriched by the target genes of Yamanaka factors in E14.1 mES cells and iPS cells. (PDF 9 kb)
Supplementary information Table S7
Significantly enriched KEGG pathways by Yamanaka factors (PDF 19 kb)
Supplementary information Table S8
c-Myc targeted Gene Ontology(biological process) common enrichment in piPS, iPS and ES. (XLS 818 kb)
Supplementary information Figure S1
Summary of the Yamanaka factors occupancy in mouse ES cells. (PDF 105 kb)
Supplementary information Figure S2
Summary of the Yamanaka factors occupancy in the published iPS ChIP-chip data. (PDF 104 kb)
Supplementary information Figure S3
Transcriptional profile regulated by Yamanaka factors in published data iPS ChIP-chip data. (PDF 202 kb)
Supplementary information Figure S4
Effects of Oct4, Sox2 and Klf4 on the Signaling Regulated by c-Myc. (PDF 117 kb)
Supplementary information Figure S5
Functional analysis of cell signaling role of Yamanaka factors targeted genes in published iPS ChIP-chip data using PANTHER classification system. (PDF 120 kb)
Supplementary information Figure S6
Signaling pathways enrichment analysis by endogenous Yamanaka factors targeted genes in published piPS, iPS and ES data. (PDF 100 kb)
Rights and permissions
About this article
Cite this article
Huang, J., Chen, T., Liu, X. et al. More synergetic cooperation of Yamanaka factors in induced pluripotent stem cells than in embryonic stem cells. Cell Res 19, 1127–1138 (2009). https://doi.org/10.1038/cr.2009.106
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2009.106
Keywords
This article is cited by
-
A revisiting of “the hallmarks of aging” in domestic dogs: current status of the literature
GeroScience (2023)
-
Identification of New Transcription Factors that Can Promote Pluripotent Reprogramming
Stem Cell Reviews and Reports (2021)
-
MicroRNA and transcription factor co-regulatory networks and subtype classification of seminoma and non-seminoma in testicular germ cell tumors
Scientific Reports (2020)
-
Enhanced generation of human induced pluripotent stem cells by ectopic expression of Connexin 45
Scientific Reports (2017)
-
Oct4 links multiple epigenetic pathways to the pluripotency network
Cell Research (2012)