Abstract
A major feature of apoptotic cell death is gross structural changes, one of which is the loss of cell–cell contacts. The caspases, executioners of apoptosis, were shown to cleave several proteins involved in the formation of cell junctions. The membrane-associated guanylate kinases (MAGUKs), which are typically associated with cell junctions, have a major role in the organization of protein–protein complexes at plasma membranes and are therefore potentially important caspase targets during apoptosis. We report here that MAGUKs are cleaved and/or degraded by executioner caspases, granzyme B and several cysteine cathepsins in vitro. When apoptosis was induced by UV-irradiation and staurosporine in different epithelial cell lines, caspases were found to efficiently cleave MAGUKs in these cell models, as the cleavages could be prevented by a pan-caspase inhibitor N-benzyloxycarbonyl-Val-Ala-Asp(OMe)fluoromethylketone. Using a selective lysosomal disrupting agent L-leucyl-L-leucine methyl ester, which induces apoptosis through the lysosomal pathway, it was further shown that MAGUKs are also cleaved by the cathepsins in HaCaT and CaCo-2 cells. Immunohistological data showed rapid loss of MAGUKs at the sites of cell–cell contacts, preceding actual cell detachment, suggesting that cleavage of MAGUKs is an important step in fast and efficient cell detachment.
Similar content being viewed by others
Main
Sites of cell–cell contacts, specialized regions for signal transduction and cellular adhesion, have crucial roles in tissue development, differentiation and physiology of multicellular organisms. Along the membrane, protein complexes are targeted to spatially and functionally distinct cell junctions, assembled by the integral and submembranous proteins. Among the latter, a family of proteins termed MAGUKs (membrane-associated guanylate kinases) has been extensively characterized. They are often found in different cell junction structures, where they have a general role in creating and maintaining specialized membrane domains and act as molecular scaffolds for the signalling pathway components.1, 2 The MAGUKs have a common multidomain organization including several PDZ domains, an SH3 or a WW domain, and an inactive guanylate kinase (GUK) domain. All these domains act as protein–protein interacting domains enabling MAGUKs to assemble a combination of cell adhesion molecules, cytoskeletal proteins, receptors, ion channels and their associate signaling components at the specific membrane sites.2, 3, 4, 5
The loss of cell–cell contacts and isolation of a dying cell is one of the important morphological features of apoptosis. The main proteases involved in the execution of apoptosis are the caspases, which reside in the cytosol as inactive proenzymes and are activated in response to cellular damage.6 During the last decade, a number of caspase substrates have been identified, among them the components of cell–cell contacts including MAGUKs.7, 8, 9, 10, 11
Beside caspases, other proteases have been implicated in apoptosis, such as cathepsins,12 which on release from the lysosomes by a number of apoptotic stimuli13, 14 indirectly activate caspases through the processing of the Bcl-2 homologue Bid and/or degradation of the antiapoptotic Bcl-2 proteins,15, 16 or through the activation of Bax independently of Bid cleavage.17 In addition, the serine protease Omi/HtrA2, which is released from the mitochondria together with cytochrome c and Smac/Diablo, is also involved in apoptosis, although it was found to process only a very limited number of substrates.18, 19, 20 Moreover, cytotoxic T lymphocytes (CTLs) and natural killer cells (NK) can induce apoptosis of target cells through a pseudo-receptor pathway, that is, through exocytosis of cytotoxic granules, also known as secretory lysosomes.21, 22 Granzyme B, a major apoptotic serine protease from the cytotoxic granules, has a substrate specificity similar but not identical to those of caspases and cleaves substrates exclusively after aspartate residues.23
The aim of this study was to elucidate the role of MAGUKs in dismantling cell–cell contacts during apoptosis and to identify the proteases that cleave them in this process. We show here that Dlg1, ZO-1 and ZO-3 are cleaved in different epithelial cell lines on induction of apoptosis by several apoptotic stimuli, suggesting that this effect is neither cell nor pathway specific. In addition, caspases were identified as the major proteases responsible for MAGUK cleavage in all models used, whereas cathepsins, and possibly granzymes, could contribute to MAGUK degradation when released from conventional or secretory lysosomes.
Results
Dlg1, ZO-1 and ZO-3 are cleaved during apoptosis
Previously, some members of the MAGUK family were found to be cleaved during apoptosis,7, 8, 9, 10, 11 which prompted us to ask whether MAGUKs are general protease targets during apoptosis. To do this, MDCK and HaCaT cells that contain high levels of endogenous epithelial MAGUKs, were used as model systems, with the major focus being on ZO-1, ZO-3 and Dlg1. Apoptosis was induced using two well-established methods, staurosporine (STS) and UV-irradiation (Figures 1a and b) and confirmed by measurement of phosphatydilserine (PS) exposure (Supplementary Figures S1A and S1B), DEVD-ase activity (Supplementary Figures S1C and S1D) and also by the detection of the caspase-generated 85 kDa N-terminal poly(ADP-ribose)polymerase (PARP) fragment; the last two were abolished using the pan-caspase inhibitor N-benzyloxycarbonyl-Val-Ala-Asp(OMe)fluoromethylketone (zVAD-fmk) (Figure 1d).
Cleavage of Dlg1, ZO-1 and ZO-3 in MDCK and HaCaT cells after induction of apoptosis with UV-irradiation and STS. (a) MDCK cells 12 h after UV irradiation and HaCaT cells 12 h after STS treatment were viewed under light microscopy. zVAD-fmk was added 2 h before induction of apoptosis. (b) The percentage of detached MDCK cells 12 and 16 h after UV irradiation and HaCaT cells after 12 h of 500 nM and 1 μM STS treatment was determined as described in Materials and methods section. zVAD-fmk was added 2 h before induction of apoptosis. (c and d) Western blot analysis of total cell lysates from MDCK and HaCaT cells treated with UV light or STS in presence or absence of z-VAD-fmk. Cells were harvested and lysed 12 or 16 h after the treatment (same conditions as in b and c). The total lysates were subjected to western blotting with antibodies indicated in the figure
Next, MDCK cells were harvested 12 and 16 h after UV irradiation and Dlg1, ZO-1 and ZO-3 proteins were followed by immunodetection. A concomitant decrease in full-length proteins can be observed, which is more pronounced after 16 than 12 h, consistent with apoptosis progression (Figure 1c). Addition of zVAD-fmk completely abolishes the cleavage, suggesting that caspases are responsible for the cleavages in this cell model. Several cleavage products of ZO-1 (∼120, 115 and 90 kDa) could be detected (Figure 1c). In contrast, no cleavage products of Dlg1 and ZO-3 were detected, suggesting degradation of the proteins, which was more pronounced in the case of ZO-3. Similar results were observed in MDCK cells treated with STS (Figure 1c) and in HaCaT cells treated with either UV irradiation or STS (Figure 1d). It can therefore be suggested that Dlg1, ZO-1 and ZO-3 are substrates of the caspases during UV- and STS-induced apoptosis in MDCK and HaCaT cells. There are a number of consensus caspase cleavage sequences (DXXD) found in ZO-1, ZO-3, and in Dlg, mostly in the linker regions between the protein–protein interaction domains (Supplementary Figure S2A). As proteolytic cleavage fragments of MAGI-1 have no active role in apoptosis and such cleavage products of MAGUKs generally result in a loss of protein scaffolding function,11, 24 we decided not to pursue their identification in ZO-1. However, the presence of several DXXD sites at the C-terminus of ZO-3 could explain its apparent degradation during apoptosis as the antibodies used were generated against the C-terminus of the protein, which also precluded us from further analysis.
Changes in localization of MAGUKs during apoptosis
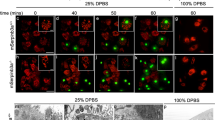
As MAGUKs are normally localized at sites of cell junctions in confluent cells, we asked whether this localization would change during apoptosis. Immunohistological staining of MDCK and HaCaT cells showed that ZO-1 and Dlg1 are found at the regions of cell–cell contacts (Figure 2a and Supplementary Figure S3, controls). Six hours after UV irradiation of MDCK cells, the membrane localization of ZO-1 was partially lost and a small fraction of ZO-1 appeared to be present in both perinuclear and nuclear compartments (Figure 2a). This is similar to the situation seen in non-confluent cells, where ZO-1 is mainly localized in the nucleus with its binding partner, the transcription factor ZONAB. On confluency, both ZO-1 and ZONAB are translocated to the cell junctions at the membrane.25 Therefore, the conclusion that after induction of apoptosis a fraction of ZO-1 changes its localization to a subnuclear compartment relies solely on the fragment that is recognized by the antibody used in these experiments. It is possible that some of the cleavage product(s) could be still at the membrane or in some other compartment. Furthermore, cell–cell contacts and nuclei were largely preserved at the 3- and 6-h time points (Figures 2a–c), suggesting that ZO-1 cleavage and its removal from the tight junctions precedes cell–cell detachment and nuclear fragmentation (Figures 2c and d). Nuclear condensation and fragmentation as well as cell detachment were observed in a higher percentage of cells at the 9- and 12-h time points where ZO-1 localization was almost completely lost from the membrane (Figures 2b–d). Addition of zVAD-fmk mainly preserved the ZO-1 membrane localization, implying the involvement of caspases in the localization changes (Figure 2a). In a similar experiment, apoptosis was induced with STS in MDCK cells and in HaCaT cells with STS and UV-irradiation with essentially the same result (Supplementary Figure S3, reference Gregorc et al.10 and data not shown).
Loss of ZO-1 from the membrane during UV irradiation-induced apoptosis. (a) MDCK cells were subjected to UV irradiation in the presence or absence of z-VAD-fmk. Cells were fixed 6 h later and ZO-1 localization was analyzed by immunofluorescence (green). DAPI was used to stain the nuclei. White arrows (middle panel) point to loss of ZO-1 from the membrane and red arrows (middle panel) to possible subnuclear or nuclear localization of cleavage fragment(s) of ZO-1. Forty-fold magnification was used. (b and c) MDCK cells were subjected to UV irradiation and fixed at indicated times. After staining with ZO-1 antibody and DAPI, cells were visualized under fluorescent microscope (b) and the percentage of condensed and fragmented nuclei was determined (c). (d) At indicated times after UV irradiation, detached and attached cells were collected separately, counted and the percentage of detached cells was calculated
ZO-1 is cleaved early during apoptosis
To confirm that cleavage of MAGUKs is indeed an early event during apoptosis, preceding the actual cell detachment, we followed the cleavage of ZO-1 with time. ZO is the closest to the membrane among all the MAGUKs as it directly binds to the transmembrane proteins of the cell junctions,26, 27 and is therefore probably one of the last MAGUKs to be cleaved during apoptosis in the process of cell detachment. The cleavage products of ZO-1 can be seen by 3 h after induction of apoptosis by STS as well as PARP cleavage (Figure 3b); at the same time the DEVDase activity increased fourfold compared with the control (Figure 3a). By 24 h after induction of apoptosis, we could not detect the native form of ZO-1. E-cadherin appears to be cleaved later than ZO-1, since very little cleavage could be observed at the 6-h time point, and significant amounts of the protein could still be detected at the 24- and 36-h time points (Figure 3b). zVAD-fmk almost completely blocked the cleavage of ZO-1, but not the cleavage of E-cadherin (Figure 3b), suggesting that some other proteases besides caspases can cleave it, most likely the extracellular matrix metaloprotease (MMPs).28 Similarly, ZO-1 was also cleaved early (∼ at 4 h) in UV-induced apoptosis in HaCaT cells (Supplementary Figure S2B).
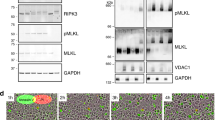
Time-dependent cleavage of ZO-1 during apoptosis in HaCaT cells. (a) HaCaT cells were treated with 1 μM STS in the presence or absence of zVAD-fmk and collected at indicated times. DEVDase activity was quantified by fluorogenic caspase substrate DEVD-afc. (b) Time-dependent cleavage of ZO-1, PARP and E-cadherin during STS-induced apoptosis in HaCaT cells was analyzed by western blot assays using specific antibodies. E-cadherin densiometry is indicated. (c) HaCaT cells were UV irradiated in the presence or absence of zVAD-fmk and detached and attached cells were collected separately 12 and 24 h later and the percentage of detached cells was determined. (d) HaCaT cells were UV irradiated in the presence or absence of zVAD-fmk. After 12 h, cells were collected either all together or detached and attached separately and stained with annexin V-PE and PI for flow cytometry analysis. In control experiment treatment was omitted. Each plot represents a typical experiment of 10 000 cells. Almost all detached HaCaT cells were annexin V-positive, while in the attached cells the percentage of the annexin V-positive cells was low. (e and f) Detached and attached HaCaT cells were collected separately at indicated times after UV-induced apoptosis. DEVD-ase activity of attached and detached cells was determined with fluorogenic caspase substrate Ac-DEVD-afc (e). In parallel, total cell lysates were subjected to western blotting with anti-ZO-1 and E-cadherin (f). Owing to the low number of detached cells remaining after 12 h in the presence of zVAD-fmk, we followed the cleavage of ZO-1 and E-cadherin 24 h after UV irradiation, when the number of detached cells is slightly lower compared with the 12-h time point without the inhibitor. zVAD-fmk almost completely abolished the cleavage of ZO-1 in both detached and attached cells, but not the cleavage of E-cadherin
Furthermore, we focused on the different properties of the attached and detached cells after induction of apoptosis (Figures 3c–f). Our results indicate that ZO-1 is cleaved before E-cadherin in apoptosis and before PS externalization (Figures 3d and f); and complete cleavage of ZO-1 and E-cadherin is not necessary for cell detachment (Figure 3f). Collectively, these results suggest that for a fast and efficient cell detachment during apoptosis the components from the both sides of the membrane have to be cleaved, including the MAGUKs, which appear to be cleaved before the exposure of PS on the outer leaflet of plasma membrane, and before detachment of the cells from the substrate.
MAGUKs are cleaved by caspases in vitro
To identify the caspases responsible for the cleavages, the MAGUK proteins were in vitro translated and incubated with recombinant caspases-3, -6 -7 and -8 for 1 h. Only caspase-3 cleaved all the MAGUKs (Figure 4a). Interestingly, caspase-6 also cleaved several MAGUKs in vitro (Figure 4a). However, all of them were substantially more efficiently cleaved by caspase-3, suggesting a minor, if any, physiological importance of caspase-6. Caspase-3 and -7 exhibited different cleavage patterns for ZO-3, which is consistent with the slightly different cleavage profile of the two proteases on protein substrates as revealed by proteomic studies.29 Additional in vitro studies confirmed that ZO-3 is a better substrate for caspase-3 than ZO-1 (Supplementary Figure S3), as seen in cell models (Figures 1c and d).
In vitro cleavage of MAGUK proteins with different proteases involved in apoptosis. (a) In vitro translated MAGUK proteins were incubated with recombinant executioner caspases-3, -6 and -7 for 1 h at 37 °C. Samples were resolved on 10 or 12.5% SDS-PAGE and visualized by autoradiography. (b) In vitro translated MAGUK proteins were incubated with recombinant cathepsins as indicated in the figure. Cathepsins were activated at pH 6 and added to the translated proteins and buffer with pH 7 for 45 min at 37 °C. (c) Radiolabelled MAGUK proteins were incubated with recombinant granzyme B (GrB) or inactive form of recombinant granzyme B (NA) for 4 h at 37 °C. (d) In vitro translated wt, D761A and DD543AA mutants of MAGI-1 were incubated with granzyme B (GrB) or inactive form of granzyme B (NA) for 4 h at 37 °C. (e) In vitro translated MAGUK proteins and mutants of MAGI-1 (D761A and DD543AA) were incubated with recombinant caspase-8 for 1 h at 37 °C. All samples were resolved on 10 or 12.5% SDS-PAGE gels and visualized by autoradiography. In control experiments proteases were omitted
Caspases could cleave MAGUKs at multiple sites in vitro (Figure 4a), although probably not all of these cleavages occur in vivo owing to conformational changes and/or unavailability of the cleavage sites in MAGUKs complexed with their cognate ligands. Taken together, these results demonstrate that MAGUKs are cleaved by at least one of the executioner caspases in vitro and that probably caspase-3 is the main executioner caspase responsible for the cleavages.
MAGUKs are cleaved by the cathepsins and granzyme B in vitro, but not by the serine protease Omi/HtrA2
As release of the lysosomal proteases into the cytosol was shown to be an early event in apoptosis induced by various apoptotic stimuli,13 we first examined whether cathepsins B, K, L, S and D are able to cleave MAGUKs in vitro. All the MAGUKs tested were degraded by all four cysteine cathepsins, but not by cathepsin D (Figure 4b). Even at high concentration cathepsin D could only partially cleave ZO-3 and SAP-102, suggesting that the enzyme is not really important as a MAGUK-processing protease. However, the efficient degradation of MAGUKs by all the cysteine cathepsins at neutral pH suggests that they can degrade MAGUKs in vivo after translocation from the lysosomes into the cytosol.
CTLs induce apoptosis of target cells through the action of granzymes. Like the caspases, granzyme B specifically cleaves after aspartate residues.22 In this respect, we have previously shown that endothelial tight junctions are disrupted in cells treated with recombinant purified granzyme B.30 This was the main rationale for testing whether granzyme B can also cleave MAGUKs, thus enhancing disruption of the cell contacts and facilitating phagocytosis of the target cells. Granzyme B cleaved all the MAGUKs tested in vitro, however, the cleavage of Dlg1, MAGI-1, MAGI-3 and ZO-1 was more efficient than the cleavage of other MAGUKs (Figure 4c). These proteins are predominantly found in epithelial cells, where CTLs have access, while PSD-95, PSD-93, SAP102 and MAGI-2 are almost exclusively localized in the brain, where under normal conditions CTLs are not found. In addition, all the MAGUKs tested were only marginally cleaved by caspase-8 (Figure 4e), questioning its relevance in vivo, where the concentration of caspase-8 is significantly lower from the one used in these assays. Furthermore, the major caspase cleavage site in MAGI-1 (DQTD761S; 11) is also the major cleavage site for granzyme B (Figure 4d) and caspase-8 (Figure 4e), suggesting that the apical caspase-8, the executioner caspases (namely caspase-3 and -7) and granzyme B share the same cleavage site for MAGI-1, Asp761.
Finally, recombinant serine protease Omi/HtrA2 was found not to cleave any of the MAGUKs (Supplementary Figure S4), indicating that this protease cleaves a limited set of proteins during apoptosis.18, 19, 20
Cathepsins cleave Dlg1, ZO-1 and ZO-3 during LeuLeuOMe-induced cell death
Next, cysteine cathepsins were examined for their ability to cleave Dlg1, ZO-1 and ZO-3 in a cellular model. Apoptosis was induced by the lysosomotropic reagent L-leucyl-L-leucine methyl ester (LeuLeuOMe), which is known to critically involve cysteine cathepsins upstream of caspase activation.15 Twenty hours after induction of apoptosis, HaCaT and CaCo-2 cells exhibited several characteristics of apoptosis; that is, morphological changes, annexin V-positive cells, DEVDase activity and PARP cleavage (Supplementary Figure S5 and Figure 5).
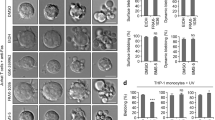
Cleavage of Dlg1, ZO-1 and ZO-3 in HaCaT and CaCo-2 cells after induction of apoptosis with LeuLeuOMe. (a) HaCaT and CaCo-2 cells were incubated with LeuLeuOMe for 20 h in the presence or absence of cysteine cathepsin inhibitor E64d or pan-caspase inhibitor zVAD-fmk. Morphology of cells was visualized under light microscopy. (b) HaCaT and CaCo-2 cells were treated with LeuLeuOMe in presence or absence of E64d or zVAD-fmk. Detached and attached cells were collected separately and percentage of detached cells was determined. (c and d) HaCaT (c) and CaCo-2 (d) cells were harvested 20 h after induction of apoptosis with LeuLeuOMe in presence or absence of inhibitors. Total cell lysates were subjected to western blotting with anti-Dlg1, anti-ZO-1, anti-ZO-3 and anti-PARP. (e) CaCo-2 cells were treated with LeuLeuOMe in the presence or absence of E64d and zVAD-fmk, and analyzed 12 h after induction of apoptosis by immunofluorescence. DAPI was used as a counterstain to visualize the nuclei. Forty-fold magnification was used
Dlg1, ZO-1 and ZO-3 were cleaved in HaCaT and CaCo-2 cells in LeuLeuOMe-induced apoptosis, and these cleavages could be partially prevented by L-trans-epoxysuccinyl(OEt)-Leu-3-methylbutylamide (E64d), but not by zVAD-fmk (Figures 5c and d). These data suggest that, in this model, both caspases and cathepsins are responsible for the processing of MAGUKs, and that the degradation observed in the cells pretreated with zVAD-fmk is probably because of cathepsin activity. Furthermore, cathepsins can also disrupt the membrane localization of ZO-1 and Dlg1 in a similar way as caspases (Figure 5e and Supplementary Figure S5C).
Discussion
During apoptotic cell death, a fraction of the cellular proteome is targeted by caspases with several hundred substrates identified so far.31, 32 In addition to caspases, other proteases have also been found to participate in cell death, often in concert with them.13, 14, 21 As a consequence of protease action, a number of vital functions of the cell are disabled and there is increasing evidence that the disruption of cell–cell contacts and isolation of a dying cell is one of them. Here, we have shown that ZO-1, ZO-3 and Dlg1, the components of the cellular junctions from the MAGUK family, are cleaved by the caspases and cysteine cathepsins in several epithelial cell lines during apoptosis induced by STS, by UV light and by the lysosomotropic compound LeuLeuOMe. Their cleavage was accompanied by their dissociation from the membrane, which was shown to precede the actual disruption of the cell–cell contacts and fragmentation of the nuclei.
MAGUKs are composed of a number of protein-binding domains, such as PDZ domains, SH3 domains or WW domains, which are critical for their function as scaffolding proteins mediating protein–protein interaction at the membrane; thus, such cleavages usually inactivate the MAGUKs. This is based on previous findings demonstrating that proteases, including trypsin and chymotrypsin as model proteases,33 and caspases, process MAGUKs by cleaving them exclusively in the interdomain linker regions24 and thereby inactivate them by separating the protein-interacting domains. These results are therefore consistent with the idea that inactivation of the MAGUKs is required to ensure a complete, fast and efficient cell detachment during apoptosis. In addition to the MAGUKs,9, 10, 11 several other components of different cell junctions have been identified as the caspase targets during apoptosis. These include β- and γ-catenin, E- and P-cadherin, and several desmosomal proteins.7, 8, 9, 32 This large number is consistent with the disruption of cell contacts being an important step in apoptosis. However, the multiple components involved in cell–cell contact also suggest that cleaving a single protein is not sufficient for efficient cell detachment. This is consistent with our data which show that the loss of ZO-1 and Dlg1 from the area of cell–cell contacts precedes the actual disruption of these contacts, fragmentation of the nuclei and PS exposure, suggesting that cleavage of the MAGUKs is an earlier event (Figures 1, 2 and 3). In addition, MAGI-1 was previously found to be cleaved by caspases during Fas-, UV- and STS-induced apoptosis. Moreover, a dominant-negative mutant of MAGI-1 delayed cell–cell detachment but not PS exposure and nuclear fragmentation, further supporting the idea that MAGUKs are general caspase targets and that their cleavage is a conserved process found both in different cells and in response to different apoptotic stimuli.11 Although the C-terminal caspase cleavage fragment of MAGI-1 was found to translocate to the nucleus, its function there is currently unclear. However, this does not affect apoptosis progression, suggesting that the physiological function may be questionable.24
In vitro experiments showed that caspases, primarily caspase-3, cysteine cathepsins B, K, L and S, and granzyme B can cleave nine different MAGUKs, including MAGI-1, MAGI-2, MAGI-3, Dlg1, PSD-95, PSD-93, SAP102, ZO-1 and ZO-3. These results, combined with the cellular data, suggest that caspase-3 is the major protease responsible for cleaving MAGUKs. However, it seems that a complete inactivation of the proteins is not necessary for the efficient disruption of cell contacts. Thus, Dlg1, ZO-1 and ZO-3 were only partially cleaved in the cellular models, which is consistent with previous data on Dlg1 degradation in UV- and STS-induced apoptosis in HaCaT cells.10 Although the cathepsins were found to also degrade Dlg1, ZO-1 and ZO-3 in LeuLeuOMe-induced lysosomal membrane permabilization (LMP) in HaCaT and CaCo-2 cells; the cleavage efficiency was very weak, pointing to caspases as the major proteases responsible for MAGUK inactivation during apoptosis. In addition, based on the data in a model of LMP-induced apoptosis, it can be suggested that cysteine cathepsins can assist the caspases in the inactivation of MAGUKs. However, this is probably limited to apoptosis triggered through the LMP, as the caspases seem to be by far more efficient. Although cathepsin D, the major aspartic lysosomal protease, was often found to be associated with apoptosis,34 it is probably not involved in inactivation of MAGUKs. This is based on the finding that pepstatin A, an inhibitor of cathepsin D, had no effect on the cleavage of ZO-3 in LeuLeuOMe-induced apoptosis in HaCaT cells (data not shown) and inhibition of caspases was sufficient to completely prevent ZO-3 cleavage in STS-induced apoptosis in MDCK cells, although cathepsin D has been implicated in STS-induced apoptosis in human foreskin fibroblasts.34 Although the cathepsins are relatively inefficient in inactivating the MAGUKs, they might be sufficient to remove MAGUKs, and possibly other components of the cell junctions, from the membrane on release into the cytosol (Figure 5) in tumor cells, where the absence of junctions is a major contributing factor to the loss of contact inhibition of growth and tumorigenesis.32 Moreover, both MAGUKs and cathepsins have been implicated in tumor biology,12, 35, 36, 37, 38 further supporting the idea. MAGUKs are also cleaved by granzyme B in vitro suggesting that granzyme B cleaves MAGUKs either directly or indirectly via previous proteolysis of pro-caspase-322 in vivo during NK- or CTL-mediated cell death.
On this basis, we propose a model for cell–cell detachment during apoptosis, in which, depending on the type of cell junction and the type of cell, several MAGUKs must be cleaved by caspases or other proteases, such as granzymes or cysteine cathepsins to allow the protease access to other components of cell junctions located closer to the membrane, such as β-catenin and E-cadherin, a known cathepsin target during tumorigenesis,38 in the case of adherent junctions. In the final stages of cell–cell detachment, MMPs probably shed the extracellular part of the adhesion molecules, such as E-cadherin and claudins,9, 28 thus completing the process (Figure 6). Therefore, it can be suggested that, besides ensuring fast and efficient cell detachment, the cleavage of MAGUKs is probably important also for shutting off or disrupting the communication between the dying cell and its neighbors, as the MAGUKs are important scaffolding proteins for the different signaling pathways.1, 2 However, none of the MAGUKs alone seems to be a critical protease target. Instead, multiple MAGUKs are probably cleaved in parallel to ensure a really rapid cellular detachment.
Model of cell–cell detachment during apoptosis. (a) Schematic representation of adherens junction complex. Adhesion molecule E-cadherin is connected to actin cytoskeleton via β- and α-catenin. MAGUK proteins strengthen the complex and also bind other adaptor and signalling molecules (not shown). (b) After caspase activation, MAGUK proteins are the first proteins cleaved in this complex thus allowing access to other components of cell junction closer to the membrane, such as β-catenin and E-cadherin. On LMP cathepsins translocate to the cytosol and can cleave MAGUKs. MMPs probably in later stages of apoptosis shed the extracellular part of E-cadherin, thus completing the disruption of cell–cell contact
In conclusion, our results show that MAGUKs, key scaffolding proteins at cell junctions, are targets of caspases during STS-, UV irradiation- and LeuLeuOMe-induced apoptosis in different epithelial cell lines, indicating that their cleavage is neither cell nor pathway specific and is probably a general mechanism required for dismantling cell–cell contacts. Our data further suggest that the MAGUK cleavage occurs early in the course of apoptosis, that is, before exposure of PS and nuclear fragmentation, and before the actual detachment of the dying cell from the surface. In addition, we showed that cathepsins are capable of cleaving MAGUKs in the apoptotic model following LMP and removing them from the membrane in a similar way to the caspases. However, the most potent executioners are still the caspases, particularly caspase-3. In addition, granzyme B cleaves MAGUKs in vitro, thus increasing the number of proteases that can cleave them in apoptosis, and further strengthening the conclusion of their importance in the process of fast and proper disruption of cell–cell contacts and the isolation of a dying cell during apoptosis.
Materials and Methods
Plasmid constructs
For our study we used following constructs: pCDNA3-MAGI-2, pCDNA-MAGI-3, pCDNA3-PSD-95, pCDNA3-PSD-93, pCDNA-SAP102 and pCMV-BK-ZO-1. For in vitro expression the MAGI-1 gene was cloned into HindIII/EcoRI and Dlg1 gene into EcoRI/KpnI sites of pCDNA3, ZO-1 gene was cloned into pGEM according to the manufacturer's instructions (Promega, Madison, WI, USA).
The sequences were verified by DNA sequencing using an Abi Prism 310 automated DNA sequencer (PerkinElmer, Boston, MA, USA).
Protein expression
Recombinant caspases-3, -6 and -7 were expressed in E. coli and purified as described previously.39 Recombinant cathepsins B, K, L, S and D were prepared according to published procedures.15, 16 Recombinant serine protease Omi/HtrA2 and granzyme B were expressed as described.20, 30
Caspase, cathepsin, granzyme B and Omi/HtrA2 cleavage assays
In vitro transcription/translation of the MAGUKs was performed with 35S-methionine labelling mix (Amersham, Piscataway, WI, USA) using rabbit reticulosyte lysate – TNT7 Coupled Extract System (Promega). Caspase cleavage assays were performed in 20 mM Hepes buffer, pH 7.2, containing 100 mM NaCl, 10 mM dithiothreitol, 1 mM EDTA, 0.1 (w/v) CHAPS and 10 % (w/v) sucrose at 37 °C. Briefly, caspases were incubated for 5 min in the reaction buffer at 37 °C before the addition of 5 μl of translation product to the final volume of 25 μl. Final concentration of the caspases was 1 μM. Cathepsins were activated for 5 min at 37 °C in their respective buffers containing 2 mM dithiothreitol (pH 6.0) before the addition to the 5 μl of each MAGUK protein to the final active concentration of 300 nM (pH 7.2) and volume of 25 μl. Omi/HtrA2 was incubated for 5 min in 50 mM Tris, 1 mM dithiothreitol and 150 mM NaCl (pH 7.5) before addition of the 5 μl of translation product to the final concentration of 400 nM. Granzyme B was added to the buffer D (50 mM Tris/HCl and 100 mM NaCl; pH 8.5) with 5 μl of translation product to the final concentration of 10 μg/ml. After 1-h incubation with caspases and Omi/HtrA2, 45 min with cathepsins and 4 h with granzyme B at 37 °C the reactions were terminated by the addition of l × SDS loading buffer and boiling. The reaction mixtures were analyzed by 8.5 or 10% SDS-PAGE gels and autoradiography.
Cell culture
Human HaCaT cells (from ATCC, Wesel, Germany) were grown in DMEM (Invitrogen, Camarillo, CA, USA) supplemented with 10% fetal bovine serum (FBS) (Sigma, St. Louis, MO, USA). Human CaCo-2 cells and canine kidney epithelial cells (MDCK) (both cell lines purchased from (ECACC, Salisbury, UK)) were cultured in EMEM (Invitrogen) with 10% FBS (Sigma). Staurosporine (Sigma) was dissolved in dimethylsulfoxide (DMSO). Lysosomotropic agent LeuLeuOMe (Sigma) was dissolved in methanol and used at final concentrations of 1.5 mM for HaCaT and 5 mM for CaCo-2 cells. UV irradiation-induced apoptosis was initiated by irradiating the cells with UV light for 45 s. Inhibitors zVAD-fmk (Bachem, Bubendorf, Switzerland) and E64d (Peptide Institute, Inc., Osaka, Japan) were dissolved in DMSO and used at a final concentrations of 20 μM. All inhibitors were added 2 h before induction of apoptosis.
For calculating the percentage of detached cells after induction of apoptosis, we separately collected detached and attached cells and counted them using Neubauer improved bright-line counting chamber (Brand, Wertheim, Germany).
Immunoblotting
Cells were collected at the times indicated in the text post induction of apoptosis and incubated in RIPA buffer (50 mM Tris (pH 8.0), 100 mM NaCl, 0.1% (w/v) SDS, 1% (v/v) Nonident P-40, 0.5% (w/v) deoxyholic acid, 1 mM EDTA) for 10 min on ice. Insoluble material was removed by centrifugation at 14 000 r.p.m. for 10 min. Bradford assay (Bio-Rad, Hercules, CA, USA) was used to determine protein concentration and 75 μg of proteins were resolved in 8.5 or 10% SDS-PAGE gels. After transfer to nitrocellulose membrane (Serva, Heidelberg, Germany), blots were probed with antibodies against Dlg1 (Santa Cruz, Santa Cruz, CA, USA), ZO-1 (Zymed Laboratories, San Francisco, CA, USA), ZO-3 (Zymed Laboratories), E-cadherin (Invitrogen), 85 kDa cleaved form of PARP (Promega), actin (Sigma), appropriate HRP-conjugated secondary antibodies (Sigma) and visualized with ECL according to the manufacturer's instructions (Amersham). E-cadherin densiometry was analyzed using SynGene Laboratories software (Cambridge, UK).
DEVDase activity and FACS analysis
In all, 30 μg of protein of untreated and treated cells in the presence or absence of inhibitors were used to determine caspase activity by measuring the cleavage of fluorogenic substrate acetyl-Asp-Glu-Val-Asp-7-amino-4-trifluoromethylcoumarin (Ac-DEVD-AFC) (Bachem).
Cells were harvested at indicated times in text after treatment. Culture medium supernatant and PBS washes were retained to ensure that both floating and adherent cells were analyzed. After being incubated for 15 min with PE-conjugated annexin V (Becton Dickinson, Erembodegem-Aalst, Belgium) according to the manufacturer's instructions, cells were subjected to FACS analysis using a FACScalibur cytometer (Becton Dickinson, San Jose, CA, USA) and CellQuest software (San Jose, CA, USA).
Fluorescence microscopy
Cells were grown overnight on coverslips before induction of apoptosis and incubation for times indicated in text. Cells were then washed with PBS and fixed in 4% paraformaldehyde in PBS for 10 min, followed by permeabilization with 0.2% (v/v) Triton X-100 in PBS for 15 min. After extensive washing with PBS, cells were incubated with hDlg, ZO-1 or ZO-3 antibody diluted in PBS overnight at 4 °C. Following an additional round of extensive washing with PBS, cells were incubated either with goat anti-rabbit rhodamine-conjugated antibody or with goat anti-mouse fluorescein-conjugated antibody (Molecular Probes, Eugene, OR, USA) for 60 min at 37 °C. After incubation, cells were washed with PBS, mounted on slides and visualized by fluorescence microscopy (Olympus IX71, Hamburg, Germany). DAPI (Sigma) was used to stain the nuclei. Nuclear changes were counted in five randomly chosen fields at 40-fold magnification with at least 300 cells per field.
Abbreviations
- Ac-DEVD-AFC:
-
acetyl-Asp-Glu-Val-Asp-7-amino-4-trifluoromethylcoumarin
- E64d:
-
L-trans-epoxysuccinyl(OEt)-Leu-3-methylbutylamide
- LeuLeuOMe:
-
L-leucyl-L-leucine methyl ester
- LMP:
-
lysosomal membrane permabilization
- MMP:
-
matrix metaloprotease
- PARP:
-
poly(ADP-ribose)polymerase
- PS:
-
phosphatydilserine
- STS:
-
staurosporine
- zVAD-fmk:
-
N-benzyloxycarbonyl-Val-Ala-Asp(OMe)fluoromethylketone
References
Anderson JM . Cell signalling: MAGUK magic. Curr Biol 1996; 6: 382–384.
Funke L, Dakoji S, Bredt DS . Membrane-associated guanylate kinases regulate adhesion and plasticity at cell junctions. Annu Rev Biochem 2005; 74: 219–245.
Gonzales-Mariscal L, Betanzos A, Avila-Flores A . MAGUK proteins: structure and role in the tight junction. Semin Cell Dev Biol 2000; 11: 315–324.
Caruana G . Genetic studies define MAGUK proteins as regulators of epithelial cell polarity. Int J Dev Biol 2002; 46: 511–518.
Hata Y, Nakanishi H, Takai Y . Synaptic PDZ domain-containing proteins. Neurosci Res 1998; 32: 1–7.
Vaux DL, Korsmeyer SJ . Cell death in development. Cell 1999; 96: 245–254.
Schmeiser K, Grand RJ . The fate of E- and P-cadherin during the early stages of apoptosis. Cell Death Differ 1999; 6: 377–386.
Brancolini C, Sgorbissa A, Schneider C . Proteolytic processing of the adherens junctions components β-catenin and γ-catenin/plakoglobin during apoptosis. Cell Death Differ 1998; 5: 1042–1050.
Bojarski C, Weiske J, Schoneberg T, Schroder W, Mankertz J, Schulzke JD et al. The specific fates of tight junction proteins in apoptotic epithelial cells. J Cell Sci 2004; 117: 2097–2107.
Gregorc U, Ivanova S, Thomas M, Turk V, Banks L, Turk B . hDLG, a member of MAGUK family proteins, is a novel caspase target during cell-cell detachment in apoptosis. Biol Chem 2005; 386: 705–710.
Gregorc U, Ivanova S, Thomas M, Guccione E, Glaunsinger B, Javier R et al. Cleavage of MAGI-1, a tight junction PDZ protein, by caspases is an important step for cell-cell detachment in apoptosis. Apoptosis 2007; 12: 343–354.
Turk V, Turk B, Turk D . Lysosome cysteine proteases: facts and opportunities. EMBO J 2001; 20: 4629–4633.
Turk B, Turk V . Lysosomes as ‘suicide bags’ in cell death: myth or reality? J Biol Chem 2009; 284: 21783–21787.
Stoka V, Turk B, Turk V . Lysosomal cysteine protease : structural features and their role in apoptosis. IUBMB Life 2005; 57: 347–353.
Cirman T, Orešić K, Droga Mazovec G, Turk V, Reed JC, Myers RM et al. Selective disruption of lysosomes in HeLa cells triggers apoptosis mediated by cleavage of Bid by multiple papain-like lysosomal cathepsins. J Biol Chem 2004; 279: 3578–3587.
Droga-Mazovec G, Bojič L, Petelin A, Ivanova S, Romih R, Repnik U et al. Cysteine cathepsins trigger caspase-dependent death through cleavage of bid and antiapoptotic Bcl-2 homologues. J Biol Chem 2008; 283: 19140–19150.
Bidere N, Lorenzo HK, Carmona S, Laforge M, Harper F, Dumont C et al. Cathepsin D triggers Bax activation, resulting in selective apoptosis-inducing factor (AIF) relocation in T lymphocytes entering the early commitment phase to apoptosis. J Biol Chem 2003; 278: 31401–31411.
Van Loo G, Van Gurp M, Depuydt B, Srinivasula SM, Rodriguez M, Alnemri ES et al. The serine protease Omi/HtrA2 is released from mitochondria during apoptosis. Omi interacts with caspase-inhibitor XIAP and induces enhanced caspases activity. Cell Death Differ 2002; 9: 20–26.
Yang QH, Church-Hajduk R, Ren J, Newton ML, Du C . Omi/HtrA2 catalytic cleavage of inhibitor of apoptosis (IAP) irreversibly inactivates IAPs and facilitates caspase activity in apoptosis. Gen Dev 2003; 17: 1487–1496.
Vande Walle L, Van Damme P, Lamkanfi M, Saelens X, Vandekerckhove J, Gevaert K et al. Proteome-wide indentification of HtrA2/Omi substrates. J Proteom Res 2007; 6: 1006–1015.
Trapani JA, Smyth MJ . Functional significance of the perforin/granzyme cell death pathway. Nat Rev Immunol 2002; 2: 735–747.
Pardo J, Aguilo JI, Anel A, Martin P, Joeckel L, Borner C et al. The biology of cytotoxic cell granule exocytosis pathway: granzymes have evolved to induce cell death and inflammation. Microbes Infect 2009; 11: 462–469.
Thornberry NA, Rano TA, Peterson EP, Rasper DM, Timkey T, Garcia-Calvo M et al. A combinatorial approach defines specificities of members of the caspase family and granzyme B. J Biol Chem 1997; 272: 17907–17911.
Ivanova S, Repnik U, Banks L, Turk V, Turk B . Cellular localization of MAGI-1 caspase cleavage products and their role in apoptosis. Biol Chem 2007; 388: 1195–1198.
Balda MS, Garret MD, Matter K . The ZO-1-associated Y-box factor ZONAB regulates epithelial cell proliferation and cell density. J Cell Biol 2003; 160: 423–432.
Furuse M, Itoh M, Hirase T, Nagafuchi A, Yonemura S, Tsukita S et al. Direct association of occludin with ZO-1 and its possible involvement in the localization of occludin at tight junctions. J Cell Biol 1994; 127: 1617–1626.
Itoh M, Furuse M, Morita K, Kubota K, Saitou M, Tsukita S . Direct binding of three tight junction-associated MAGUKs, ZO-1, ZO-2, and ZO-3, with the COOH termini of claudins. J Cell Biol 1999; 147: 1351–1363.
Herren B, Levkau B, Raines EW, Ross R . Cleavage of beta-catenin and plakoglobin and shedding of VE-cadherin during endothelial apoptosis: evidence for a role for caspases and metalloproteinases. Mol Biol Cell 1998; 9: 1589–1601.
Demon D, Van Damme P, Vanden Berghe T, Deceuninck A, Van Durme J, Verspurten J et al. Proteome-wide substrate analysis indicates substrate exclusion as a mechanism to generate caspase-7 versus caspase-3 specificity. Mol Cell Proteomics 2009; 8: 2700–2714.
Pardo J, Wallich R, Ebnet K, Iden S, Zentgraf H, Martin P et al. Granzyme B is expressed in mouse mast cells in vivo and in vitro and causes delayed cell death independent of perforin. Cell Death Differ 2007; 14: 1768–1779.
Fischer U, Janicke RU, Schulze-Osthoff K . Many cuts to ruin: a comprehensive update of caspase substrates. Cell Death Differ 2003; 10: 76–100.
Timmer JC, Salvesen GS . Caspase substrates. Cell Death Differ 2007; 14: 66–72.
Marfatia SM, Morais Cabral JH, Lin L, Hough C, Bryant PJ, Stolz L et al. Modular organization of the PDZ domains in the human discs-large protein suggests a mechanism for coupling PDZ domain-binding proteins to ATP and the membrane cytoskeleton. J Cell Biol 1996; 135: 753–766.
Johansson AC, Steen H, Ollinger K, Roberg K . Cathepsin D mediates cytochrome c release and caspase activation in human fibroblast apoptosis induced by staurosporine. Cell Death Differ 2003; 10: 1253–1259.
Woods DF, Bryant PJ . The discs-large tumor suppressor gene of Drosophila encodes a guanylate kinase homolog localized at septate junctions. Cell 1991; 66: 451–464.
Gardiol D, Kühne C, Glaunsinger B, Lee SS, Javier R, Banks L . Oncogenic human papillomavirus E6 proteins target the discs large tumour suppressor for proteasome-mediated degradation. Oncogene 1999; 18: 5487–5496.
Thomas M, Laura R, Hepner K, Guccione E, Sawyers C, Lasky L et al. Oncogenic human papillomavirus E6 proteins target the MAGI-2 and MAGI-3 proteins for degradation. Oncogene 2002; 21: 5088–5096.
Gocheva V, Zeng W, Ke D, Klimstra D, Reinheckel T, Peters C et al. Distinct roles for cysteine cathepsin genes in multistage tumorigenesis. Gen Dev 2006; 20: 543–556.
Stennicke HR, Salvesen GS . Caspases: preparation and characterization. Methods 1999; 17: 313–319.
Acknowledgements
The caspase-3, -6 and -7 clones in pET expression vectors were a generous gift of Guy S Salvesen (La Jolla, CA, USA). Recombinant caspase-8 was a generous gift of Mauro Degli Esposti (Manchester, UK). This work was supported in part by research grants from the Slovenian Research Agency (P1-0140V T and BT), by the European Commission Framework VI Program (CAMP project, LSHG-2006-018830 BT), a research grant from the Associazione Italiana per la Ricerca sul Cancro (LB), Public Health Service grants CA-58541 (RTJ). PV was supported by the Methusalem grant (BOF09/01M00709) from the Flemish government and by VIB, IAP6/18, FWO 3G.0218.06, Apo-Sys FP7-200767 and FP6nApopTrain, MRTN-CT-035624 grants. JP was supported by a research grant SAF2008-02139 from Ministerio de Ciencia e Innovacion (MICINN) (Spain), Fundacion Aragon I+D (ARAID) and the Alexander von Humboldt foundation. We would also like to thank Urška Repnik for help with flow cytometry measurements, and Miha Renko, Primož Meh, Mojca Prebanda, Dejan Caglič and Vida Puizdar for recombinant cathepsins.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by M Piacentini
Supplementary Information accompanies the paper on Cell Death and Disease website
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Ivanova, S., Gregorc, U., Vidergar, N. et al. MAGUKs, scaffolding proteins at cell junctions, are substrates of different proteases during apoptosis. Cell Death Dis 2, e116 (2011). https://doi.org/10.1038/cddis.2010.92
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2010.92
Keywords
This article is cited by
-
The SF3B1R625H mutation promotes prolactinoma tumor progression through aberrant splicing of DLG1
Journal of Experimental & Clinical Cancer Research (2022)
-
Loss of the cell polarity determinant human Discs-large is a novel molecular marker of nodal involvement and poor prognosis in endometrial cancer
British Journal of Cancer (2016)
-
Analysis of apoptosis methods recently used in Cancer Research and Cell Death & Disease publications
Cell Death & Disease (2012)