Abstract
Deregulation of signaling pathways that control differentiation, expansion and migration of neural crest-derived melanoblasts during normal development contributes also to melanoma progression and metastasis. Although several epithelial-to-mesenchymal (EMT) transcription factors, such as zinc finger E-box binding protein 1 (ZEB1) and ZEB2, have been implicated in neural crest cell biology, little is known about their role in melanocyte homeostasis and melanoma. Here we show that mice lacking Zeb2 in the melanocyte lineage exhibit a melanoblast migration defect and, unexpectedly, a severe melanocyte differentiation defect. Loss of Zeb2 in the melanocyte lineage results in a downregulation of the Microphthalmia-associated transcription factor (Mitf) and melanocyte differentiation markers concomitant with an upregulation of Zeb1. We identify a transcriptional signaling network in which the EMT transcription factor ZEB2 regulates MITF levels to control melanocyte differentiation. Moreover, our data are also relevant for human melanomagenesis as loss of ZEB2 expression is associated with reduced patient survival.
Similar content being viewed by others
Main
Melanocytes are specialized cells in the skin that produce melanin, a pigment that is responsible for skin and hair color and that provides protection against ultraviolet (UV) radiation. During mouse embryogenesis, melanoblasts originate from the neural crest and migrate along a dorsolateral pathway from the neural tube to the developing dermis.1 Around embryonic day (E) E11 they move into the epidermis and eventually populate the developing hair follicle.2 Here they separate into two distinct populations: the differentiated pigmented melanocytes, which reside in the hair matrix, and the non-pigmented melanocyte stem cells (MSC) in the bulge. The latter cells are responsible for replenishing the hair follicle with new melanocytes during each hair cycle. Genetic studies in mice demonstrated the importance of several key players (such as sex-determining region Y (SRY)-box 10 (Sox10), paired-box 3 (Pax3), microphthalmia-associated transcription factor (Mitf), endothelin 3/endothelin receptor B (Edn3/Ednrb), Kitl/Kit, Slug, cellular myelocytomatosis oncogene cellular homolog (cMyc) and β-Catenin (β-Cat)) for melanoblast cell fate specification, proliferation, migration and survival.2, 3, 4 The master regulator of the melanocyte development is MITF, which is spatio-temporally controlled by several key transcription factors such as SOX10, PAX3 and β-catenin.5, 6, 7 Fundamentally, MITF induces gene expression patterns that prompt a melanocyte to differentiate and initiate pigment production by activating genes important for melanin biosynthesis (such as Tyrosinase (Tyr), Dopachrome tautomerase (Dct), Tyrosinase-related protein 1 (Tyrp1) and melanocortin 1 receptor (Mc1r)) and melanosome formation (such as the premelanosome protein (Pmel)).8, 9 Maintenance of the melanocyte stem cell compartment is mainly determined by transforming growth factor β (TGF-β), NOTCH signaling and MITF-dependent BCL-2 activity.10, 11 Deficiency in melanoblast development and migration generally result in white spotting phenotypes, whereas defects in melanocyte stem cell maintenance result in progressive hair graying phenomena.3
Melanoma is a malignant tumor arising from the melanocyte lineage and is the most therapy-resistant and deadly form of skin cancer. Several pathways have been identified to have a major role in the development of malignant melanoma, among which the RAS/RAF/MEK/ERK pathway is probably the most studied and has recently become the target for anti-melanoma therapy.12 Activating mutations in RAS (Q61K), with NRAS being the most frequent, occur in 15% of cases of human melanoma.12 Melanocyte-specific expression of NRASQ61K together with cyclin-dependent kinase inhibitor 2A (Cdkn2a) or p53 deficiency leads to accelerated melanoma formation in mice.13, 14 BRAF activating mutations (V600E) occur at high frequencies (50–60%) in human melanoma. Specific inhibitors of BRAFV600E are being successfully used in melanoma therapy.15 However, patients become rapidly resistant to this therapy, and combination therapies with MEK inhibitors are being investigated.16, 17 Drug resistance of malignant melanoma cells has been associated with the expression of the drug transporter P-glycoprotein (PGP) and increased migratory and invasive behavior.18 EMT (Epithelial-to-mesenchymal transition) has a key role in the collective movement of normal cells during different stages of embryonic development, whereas in adults it has been associated with several pathologies, including fibrosis and cancer progression.19, 20 EMT is a process in which epithelial cells lose their epithelial characteristics, gain mesenchymal features and become migratory. Several pathways have been shown to induce EMT, all of which function through the induction of three families of transcription factors, the Snail (SNAIL, SLUG), Zeb (ZEB1, ZEB2) and basic Helix Loop Helix (bHLH) (E47, TWIST and others) families.21 Melanoblasts undergo a migration process during embryogenesis and mature melanocytes are periodically replenished from the stem cells and undergo a constant cycling and migration process. The EMT transcription factor SLUG, has been shown to have an important role in the melanocyte lineage, as Slug-deficient mice have pigmentation abnormalities due to melanoblast migration defects and SLUG mutations have also been found in some cases of human piebaldism.22, 23, 24 Furthermore, homozygous deletions of the SLUG gene were found in patients with type 2D of Waardenburg syndrome, an auditory-pigmentary syndrome caused by a migration deficiency of melanocytes and other neural crest-derived cells.25, 26 However, although it has been shown that siRNA-mediated inhibition of Slug leads to the suppression of metastasis in an orthotopic mouse model of melanoma, SLUG expression has not been correlated with melanoma metastasis.27 Recently, it became clear that in vitro activation of the MEK-ERK signaling drives the reversion of the EMT transcription factor expression pattern in melanocytes with the downregulation of SLUG and ZEB2 and the induction of TWIST1 and ZEB1.28
ZEB2 is a transcription factor required for neural crest cell development. It belongs to the Zeb family of zinc-finger-homeodomain transcription factors and binds the E-box sequences CACCT(G).29, 30 We previously showed that conditional Zeb2 expression in epithelial cells results in specific downregulation of E-cadherin and gain of malignant features characterized by a mesenchymal gene expression profile, such as N-cadherin and Vimentin.29, 31 Furthermore, it has recently become clear that during EMT not only epithelial characteristics are abolished, but that cells are reprogrammed to different extents.32 Mouse embryos deficient in Zeb2 exhibit defects from E8.5 onwards, with early arrest of cranial neural crest cell migration and absence of neural crest cells at the postotic vagal level.33 Tissue specific disruption of Zeb2 very early in neural crest cell development using a Wnt1-Cre-based approach, results in abnormalities in craniofacial and heart development, as well as defects in the peripheral nervous system.34
On the basis of these observations we hypothesized that ZEB2 may have a key role in the process of melanogenesis. To test this possibility we used a melanocyte-specific loss of function approach in mice. Unexpectedly we find that in addition to a role in proper melanoblast development, ZEB2 is required for the correct differentiation of melanocytes through its ability to act as an upstream regulator of Mitf. Our results propose a model whereby in melanocytes ZEB2 is necessary for Mitf expression and activity which in its turn is responsible for Zeb1 repression. Measurement of nuclear ZEB2 levels shows a significantly shorter melanoma-specific survival in those patients with low ZEB2 expression.
Results
Melanocyte-specific ZEB2 deficiency causes congenital loss of hair pigmentation
We show for the first time that ZEB2 is expressed in the melanocytes of human skin epidermis, as well as in the differentiated melanocytes of mouse hair follicles (Figure 1a). ZEB2 is also expressed in migrating melanoblasts of the mouse embryo and their precursors, the neural crest cells.33, 35 As both complete knockout and Wnt1-Cre-mediated neural-crest-specific deletion of Zeb2 are embryonic lethal (E9.5 and E12.5-postnatal (P) 0, respectively),33, 34 we conditionally deleted Zeb2 in the melanocyte lineage by using the tyrosinase(Tyr)::Cre mouse line, which allows CRE-mediated Zeb2 deletion under the control of the Tyrosinase promoter starting at E10.5 (Supplementary Figure S1a).34, 36 Homozygous melanocyte-specific deletion of Zeb2 (ZEB2MCKO) caused a severe loss of hair pigmentation compared with wild-type littermates (ZEB2MCWT). This hair pigmentation loss was present from the first hair cycle and continued through adulthood (Supplementary Figure S1b and Figure 1b). The melanin content of the dorsal and ventral hair were reduced by 93 and 91%, respectively (Figure 1c). Light microscopic analysis confirmed the disappearance of melanin pigment from the hair shafts (Supplementary Figure S1c), but hair with mixed pigmentation were also observed (data not shown).
Zeb2 loss in the melanocyte lineage causes congenital loss of pigmentation and impairs melanoblast development. (a) ZEB2 is expressed in differentiated melanocytes in the human epidermis, and in the hair follicles of mouse skin. (b) Melanocyte-specific deletion of Zeb2 (ZEB2MCKO) causes congenital loss of hair pigmentation in homozygous knockout mice (2 months old). This loss of pigmentation was more pronounced on the belly than on the back. (c) Determination of melanin pigment (μg/ml) in the dorsal and ventral hair of ZEB2MCWT and ZEB2MCKO mice (n=7–8 for each group) at the age of 4–6 months. (d) ZEB2 is required for proper melanoblast development in homozygous embryos at E15.5. Whole-mount LacZ staining of E15.5 ZEB2MCWT;Dct-LacZ and ZEB2MCKO;Dct-LacZ embryos. (e) Comparison of melanoblast migration in the epidermis and dermis of dorsal and ventral areas of ZEB2MCWT;Dct-LacZ and ZEB2MCKO;Dct-LacZ embryos (n=4 and 5, respectively). Results represent the number of melanoblasts per area analyzed. (f) Nuclear fast-red staining of LacZ-stained skin sections of ZEB2MCWT;Dct-LacZ and ZEB2MCKO;Dct-LacZ mice to visualize hair follicles (HF) (arrow: pigmented HF, arrowhead: LacZ-positive HF, *: LacZ-negative HF). (g) Quantitative analysis of the pigmented, LacZ-positive, non-pigmented LacZ-negative HF and LacZ-positive stem cells in the bulge area of ZEB2MCWT (n=7) and ZEB2MCKO (n=8) mice per 0.5 mm analyzed. Data of ZEB2MCWT and ZEB2MCKOmice were compared by using unpaired Student’s t-test and are presented as means±95% CI. P-values are indicated with (P<0.001). Micrograph images were taken with a × 60/0.8 objective (a) or a × 10/0.25 objective (e and f)
The absence of ZEB2 impairs melanoblast development
Congenital loss of hair pigmentation indicates a disturbance of melanoblast migration or proliferation due to the absence of ZEB2. To further analyze this phenotype in-depth, we crossed conditional melanocyte-specific Zeb2 knockout mice with Dct::LacZ melanoblast reporter mice37 and analyzed the LacZ-positive melanoblast population in ZEB2MCWT and ZEB2MCKO embryos at E15.5. Absence of Zeb2 severely impaired melanoblast development (Figure 1d) and quantification of the LacZ-positive melanoblasts on the back and belly of embryos further confirmed a significant reduction of melanoblast numbers in both areas (Supplementary Figure S1d). In agreement with an expected defect in migration, melanoblast numbers decreased more on the belly (95%) than on the back (70%) of the ZEB2-deficient embryos. To visualize the distribution of melanoblasts in the dermis and their subsequent migration to the epidermis, we made transversal sections of E15.5 LacZ embryos and quantified the melanoblasts in both areas. There was no difference in the number of melanoblasts present in the dermis between ZEB2MCKO compared with control embryos (Figure 1e). However, in the epidermis of both the belly and back there was a significant reduction of melanoblasts in the ZEB2MCKO embryos, suggesting an important role for ZEB2 in this process. In addition, melanoblast of ZEB2MCKO embryos had fewer cell protrusions, compatible with their altered characteristics (Supplementary Figure S1e).
ZEB2 is necessary for proper melanocyte differentiation
Although melanoblast development was severely affected, 30% of them reached the dorsal area at E15.5. This raised the question whether the remaining melanoblasts could still populate the hair follicles of ZEB2MCKO mice. Histological analysis of ZEB2MCWT;Dct-LacZ and ZEB2MCKO;Dct-LacZ skin from 5.5-day-old (P5) mice showed that some hair follicles in ZEB2MCKO mice are still LacZ positive and/or (hypo-)pigmented, indicating that ZEB2-deficient melanoblasts can migrate and populate the bulb area of the hair follicles (Figure 1f). However, quantitative analysis of the hair follicles showed that there were significantly fewer pigmented or LacZ-positive hair follicles present in ZEB2MCKO mice (Figure 1g). Furthermore, ZEB2MCKO skin contained some completely undifferentiated hair follicles that were neither pigmented nor LacZ positive, which was not the case for ZEB2MCWT skin (Figures 1f and g). Immunohistochemical analysis of depigmented ZEB2-stained sections confirmed nuclear ZEB2 presence in the differentiated melanocytes of the bulb area, the migrating melanocytes in the epidermis and importantly also in the melanocyte stem cells of ZEB2MCWT mice, whereas ZEB2 was absent in all cells of the melanocyte lineage of ZEB2MCKO skin (Supplementary Figure S2a). Quantitative analysis of the melanocyte stem cells revealed a reduction in LacZ-positive stem cells in ZEB2MCKO mice compared with ZEB2MCWT mice (Figure 1g), which agrees with the presence of fewer melanoblasts in the epidermis in ZEB2MCKO embryos. To visualize melanocytes independently of Dct promoter reporter activity, we analyzed the expression of the general melanocyte marker S100 calcium-binding protein B (S100b), which does not depend on the differentiation status of the melanocytes. S100b staining showed that melanocytes were present in most of ZEB2MCKO hair follicles (Figure 2a). Moreover, quantifying S100-positive melanocytes per hair follicle showed that the number of S100-positive ZEB2MCKO melanocytes was reduced by 40% per hair follicle, which further points to a role of ZEB2 in migration and/or survival of melanocytes (Supplementary Figures S2b and c). Further immunohistochemical analysis and quantification of positive cells of the melanocyte-specific transcription factor MITF and one of its regulators (i.e., PAX3) showed that in the absence of ZEB2 the amount of MITF-positive melanocytes was strongly reduced, when compared with the total amount of melanocytes per hair follicle (S100-positive melanocytes), whereas the amount of PAX3-positive melanocytes was not affected (Figure 2a, Supplementary Figures S2b and c). The intensity of MITF staining was also lower, indicating a decrease in both the number of MITF-positive cells and in MITF protein levels in still MITF-positive cells. These observations indicate that ZEB2 is an important regulator of MITF-dependent melanocyte differentiation. We analyzed the presence of other terminal differentiation markers, such as TYRP1 and tyrosinase enzymatic activity. Most ZEB2MCKO hair follicles were indeed negative for these markers (Figure 2a and data not shown). mRNA expression analysis of several melanocyte differentiation markers (Tyrp1, Tyr, Dct, Pmel, Mc1R and Mitf) on whole skin mRNA of ZEB2MCKO and ZEB2MCWT mice further showed that the differentiation status of ZEB2MCKO melanocytes was affected, as all of these markers are significantly downregulated in ZEB2MCKO skin (Supplementary Figure S2d). Close examination of the melanosomes in ZEB2-stained sections of ZEB2MCWT and ZEB2MCKO skin confirmed the absence of melanin in most melanocytes of the bulb area and in hair shafts of ZEB2MCKO skin, although in some hair follicles a few remaining melanosomes were visible (Figure 2b). Electron microscopy analysis showed that these melanosomes were spherical with irregular borders, in contrast to the rod-shaped melanosomes of ZEB2MCWT hair follicles (Figure 2c). These results collectively show that melanocytes form but do not differentiate properly, pointing to a crucial role for ZEB2 in melanocyte differentiation. These effects were strictly ZEB2-dependent, as in a complementation experiment in which we crossed the ZEB2MCKO mice with a conditional Zeb2 transgenic strain (Rosa26-ZEB2TG/TG-IRES-GFP) activated by the same Tyr-Cre as for the deletion of Zeb2, melanogenesis was completely rescued (Supplementary Figure S3a and Figure 2d). Such Zeb2 complemented mice (ZEB2MCKO; ZEB2MCTG) regained pigmentation, and histological sections demonstrated that ZEB2-positive melanocytes are pigmented (Figure 2d), confirming the cell autonomous action of ZEB2. Furthermore, in vivo mRNA expression analysis of the melanocyte differentiation markers (Tyrp1, Tyr, Dct, PmeL, Mc1R and Mitf) confirmed the full functional complementation of Zeb2 deficiency with transgenic Zeb2 (data not shown).
Melanocyte-specific Zeb2 deficiency causes the formation of undifferentiated melanocytes in the bulge area of hair follicles. (a) Immunohistochemical staining of sections of ZEB2MCWT and ZEB2MCKO;Dct-LacZ-positive skin sections with S100b, PAX3, MITF and TYRP1 antibodies. (b) Detailed microscopic analysis of melanosomes in the hair shafts and the bulb area of the hair follicles, combined with immunohistochemical staining of ZEB2 on LacZ-positive skin sections of ZEB2MCWT and ZEB2MCKO; Dct-LacZ-positive mice. Insets show the altered morphology of melanosomes in the ZEB2MCKO sections compared with ZEB2MCWT. (c) Electron microscopic analysis of melanosomes in the bulb area of ZEB2MCWT and ZEB2MCKO mice demonstrates the absence or irregular morphology of melanosomes in the ZEB2MCKO hair follicles. (d) Genetic compensation of the loss of Zeb2 in the ZEB2MCKO;Dct-LacZ mice with melanocyte-specific overexpression of ZEB2 (ZEB2MCTG). Left panel: complete compensation of pigmentation in ZEB2MCKO ZEB2MCTG;Dct-LacZ mice compared with ZEB2MCKO;Dct-LacZ mice. Right panels: reappearance of pigmented melanosomes and ZEB2 expression in the ZEB2MCKO ZEB2MCTG;Dct-LacZ hair follicles. All microscopic analyses were done on skin sections of 5.5-day-old (a–c) or 13.5-day-old mice (d) and Immunohistochemical micrograph images were taken with a × 60/0.8 objective (a and d) or a × 100/1.25 objective (b)
ZEB2 controls an MITF-ZEB1-dependent transcriptional network essential for melanocyte differentiation
The melanocyte differentiation block observed upon Zeb2 ablation could also be due to a change in the functionality of the melanoblasts or melanocyte stem cells. We therefore investigated whether ZEB2 is required for the maintenance of the differentiation program in melanocytes. To this end, we transfected primary mouse melanocytes with an siRNA pool targeting Zeb2 (siZEB2) or with a control siRNA pool (siCTRL). Significant knockdown of Zeb2 caused nearly complete loss of differentiation of the melanocytes (Figure 3a) and a concomitant decrease in the steady-state mRNA levels of Mitf and several genes of the melanin synthesis pathway (Figure 3b). To examine whether EMT was also affected, we assessed the expression levels of two representative EMT target genes, E-cadherin (Cdh1; epithelial marker) and Vimentin (Vim; mesenchymal marker). Unexpectedly, Cdh1 was downregulated, whereas Vim was upregulated (Figure 3b). Interestingly we found that Zeb2 knockdown caused transcriptional upregulation of Zeb1 (Figure 3b). The latter results were confirmed at the protein level in the immortalized melanocyte cell line Melan-a in which Zeb2 knockdown caused a strong downregulation of MITF and concomitant upregulation of ZEB1, and the mesenchymal markers Vimentin (VIM) and Fibronectin (FN1) (Figure 3c). These observations support the hypothetical model of ZEB2 dictating MITF expression and activity. In the absence of Zeb2, MITF expression is lost, coinciding with ZEB1 upregulation (Figure 3d). This model is confirmed in vivo as ZEB1 staining intensity was clearly increased in a fraction of the undifferentiated ZEB2-deficient melanocytes (Figure 3e), whereas ZEB1 could not be detected in differentiated melanocytes of ZEB2MCWT mice. ZEB1 protein was also undetectable in primary melanocytes of human origin (Supplementary Figure S3b and see below). On the other hand, ZEB1 was readily observed in the stem cells of both ZEB2MCWT and ZEB2MCKO mice (Figure 3e). Thus, an intricate balance between these two ZEB-family members appears to be important in the transcriptional regulation of melanocyte differentiation.
ZEB2 is necessary for proper differentiation of primary melanocytes. (a and b) Morphology of primary melanocytes after knockdown of Zeb2 by siRNA transfection and relative mRNA expression of Zeb2, melanocytes-specific differentiation genes Tyrp1, Tyr, Dct, PmeL and Mc1R, the epithelial/mesenchymal markers E-cadherin/Vimentin and Zeb1. Scrambled siRNA was used as a control (siCTRL). (c) Western blot analysis after siRNA knockdown of Zeb2 in the primary Melan a cell line. (d) Hypothetical model of the ZEB2-MITF-ZEB1 balance in melanocytes (e) ZEB1 expression in dedifferentiated LacZ-positive melanocytes of the hair follicles of ZEB2MCKO mice and in the melanocyte stem cells of both ZEB2MCWT and ZEB2MCKO mice. QPCR data were compared by using unpaired Student’s t-test (n=3) and are presented as averages±S.D. P-values are indicated with (**P<0.005 and *P<0.05). Micrograph images were taken with a × 20/0.2 objective (a) or a × 60/0.8 objective (e)
Strong nuclear ZEB2 expression in primary melanomas is associated with better survival
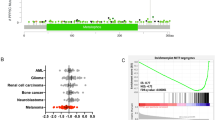
A role for ZEB2 in the regulation of MITF levels and/or activity, melanoblast development and melanocyte differentiation raises the possibility that deregulation of ZEB2 expression may contribute to melanoma progression/metastasis. Consistently, immunohistochemical analyses of ZEB2 in a cohort of human benign naevi, primary and metastatic melanoma samples revealed a very heterogeneous pattern of expression particularly in primary melanoma, with areas of high expression next to strong local downregulation (Supplementary Figure S3c and Figure 4a). This heterogeneous downregulation of ZEB2 expression is reminiscent to the profile of MITF expression in human melanoma.38 Interestingly we found that human melanoma metastases express high nuclear ZEB2 levels suggesting reversible loss of expression during melanoma dissemination (Supplementary Figure S3c). We extended this study by analyzing 178 primary melanoma samples on a tissue array. This study showed that ZEB2 has significant prognostic relevance in melanoma, as strong nuclear ZEB2 expression is beneficial for the patients in terms of melanoma-specific survival and recurrence-free survival (Figure 4b). In contrast, weak nuclear ZEB2 expression was associated with a worse patient outcome (Figure 4b). Therefore, ZEB2 might serve as a gatekeeper controlling human melanoma progression and provide a novel prognostic marker for melanoma.
ZEB2 expression in melanoma. (a) Heterogeneous ZEB2 expression in primary human melanoma with vertical growth phase. (b) Kaplan–Meier curves for melanoma recurrence-free survival, comparing human melanoma samples with strong nuclear ZEB2 expression (n=83) and weak nuclear ZEB2 expression (n=82) from a melanoma tissue array. Data were compared using the log-rank test and **P=0.0065. A representative ZEB2 staining of both groups is shown in the right panel (*** very high nuclear ZEB2, ** high nuclear ZEB2, * low nuclear ZEB2). ZEB2 protein expression values were determined using an automated image analysis approach (IHC-MARK, Oncomark, Dublin, Ireland) designed to quantify immunohistochemically stained slides. Micrograph images were taken with a × 4/0.1 and × 20/0.4 objective (a) or a × 10/0.25 objective (b)
ZEB2 loss results in reduced MITF expression, and is associated with melanoma progression
We have provided genetic evidence that ZEB2 regulates Mitf expression levels in primary melanocytes in vivo (Figure 2a and Supplementary Figure S2d). This was confirmed in melanoma cells, as Zeb2 knockdown in the B16 mouse melanoma cell line caused a transcriptional decrease in expression of Mitf and its well-established target genes and a concomitant increase in ZEB1 expression (Figure 5a). Importantly, transfection of the Zeb2 KD cells with an Mitf expression vector rescued the differentiation defect indicating that this phenotype is MITF-dependent (Figure 5b). This expression switching of ZEBs was confirmed in short-term culture human melanoma cells, in which we observed this particular ZEB heterogeneity in NRAS- or BRAF-mutated samples: 67% (8 out of 12) express both ZEB1 and ZEB2 whereas the remaining express either ZEB1 (2 out of 12) or ZEB2 (2 out of 12) (Figure 5c and Supplementary Figure S3d). Melanoma cells that are wild type for NRAS or BRAF mainly express ZEB2 only (67%, 6 out of 9), resembling the expression pattern of normal melanocytes which also only express ZEB2. In addition, the analysis of a cohort of human melanoma cell lines supported the tendency of an inverse correlation between ZEB2 and MITF versus ZEB1 expression (Figure 5d). These results were further confirmed on the human SKMel28 melanoma cell line in which we performed siRNA-mediated KD of ZEB2, which caused a strong downregulation of MITF and concomitant upregulation of ZEB1 (Supplementary Figure S3e). Furthermore, knockdown of MITF in human melanoma 501Mel cells results in upregulation of ZEB1 expression suggesting that MITF contributes to the direct repression of ZEB1 (Figure 5e). Interestingly, MITF-ChIP-seq analysis recently identified the ZEB1 locus as a potential MITF target in human melanoma 501Mel cells (Figure 5f).39 The specificity of this binding to the ZEB1 locus was confirmed by anti-HA-ChIP-qRT-PCR for both the 5′- and 3′-end of the gene (Figure 5g).
The ZEB2-MITF-ZEB1 transcriptional network. (a) Knockdown of Zeb2 by siRNA transfection of the B16 melanoma cell line. Scrambled siRNA was used as a control (siCTRL). Relative mRNA expression of endogenous Zeb2, the melanocyte-specific differentiation genes Mitf, Tyrp1, Tyr, Dct, Mc1R and Zeb1 after Zeb2 knockdown are shown. (b) Relative mRNA expression of subset of MITF target genes after siRNA knockdown of Zeb2 in the B16 melanoma cell line after compensation with Mitf. (c) Western blot analysis of ZEB2 and ZEB1 expression in short-term culture human melanoma cell lines with different NRAS (blue circles) or BRAF (red circles) mutated or WT (green circles) status. Primary human melanocytes were used as a reference. (→) specific ZEB1 band, (*) aspecific band. (d) Western blot analysis of ZEB2, MITF and ZEB1 in a panel of human melanoma cell lines. Primary human melanocytes were used as a reference. (e) 501Mel cells were infected with lentiviral vectors expressing control shRNA (shCTRL) or shRNA directed against MITF (shMITF). Left panel: western blot analysis shows strongly downregulated MITF expression after shMITF infection. Right panel: shMITF knockdown leads to increased ZEB1 mRNA expression. Two independent primer pairs located in exons 1–2 and 4–5 for the ZEB1 gene were used. (f) UCSC screenshot of a wig file of an HA-ChIP-seq from 501Mel cells expressing a 3HA-Control vector or 3HA-tagged MITF at the ZEB1 locus. Several MITF binding sites are observed (shown by arrows) both upstream and downstream of the ZEB1 gene. (g) Anti-HA-ChIP-qPCR from native 501Mel cells (CTRL) or 501MEL cells stably expressing 3HA-tagged MITF (HA-MITF). qPCR was performed with primers that amplify a region 1 kb upstream of the TYR transcription start site (TSS) as negative control, the TYR TSS as positive control and two sites from the ZEB1 locus 5′ and 3′ of the gene as indicated in f. QPCR data were compared by using unpaired Student’s t-test (n=3, for a, e and g) or paired Student’s t-test (n=5, for b) and are presented as averages±S.D. or are presented as log-transformed relative expression values with 95% CI. P-values are indicated with ***P<0.0005, **P<0.005 and *P<0.05
Discussion
In this work we have identified an important novel function for ZEB2 in melanocytes: ZEB2 regulates the expression of MITF and thereby coordinates the development and differentiation of the melanocyte lineage. Moreover we provide evidence that ZEB2 is expressed in a heterogeneous manner in human melanoma. These observations have important implications for our understanding of melanoma biology as well as for anti-melanoma therapy.
Emerging evidence indicates that the transcriptional pathways critical for melanogenesis, like those controlled by MITF, could be disturbed in pigmentation-associated diseases and melanoma progression and metastasis.40 MITF is considered to be the master regulator of melanocytes as it is essential for melanoblast survival and melanocyte-specific lineage differentiation. On the basis of the proven role of Zeb2 in neural crest cell delamination, we expected a major defect in melanoblast development.34 However, although mice deficient for Zeb2 in the melanocyte lineage have an impaired melanoblast development; a significant fraction of ZEB2-deficient melanoblasts was still able to migrate to the epidermis where they populate the hair follicles. Unexpectedly, the ZEB2-deficient melanocytes in the hair follicle bulb area remain undifferentiated, causing a congenital loss of pigmentation. These melanocytes have a very low MITF expression, supporting the role of ZEB2 as a driver for melanocyte differentiation. In vitro data further confirm the crucial role of ZEB2 as a regulator of MITF and melanocyte differentiation coinciding with an upregulation of ZEB1. In this context, it is noteworthy that the loss of Zeb2 in non-transformed melanocytes in vivo can result in ZEB1 misexpression. Furthermore the strongly reduced expression of the melanocortin receptor upon Zeb2 loss potentially contributes to the maintenance of the observed low Mitf transcriptional levels.
EMT has a key role during different stages of embryonic development including the formation and migration of the neural crest cells. Neural crest cells are a multipotent, migratory, transient cell population that migrate through the vertebrate embryo to infiltrate different organs and differentiate in various cell lineages including melanocytes.41 During the progression to melanoma, melanocytes may acquire multiple traits of high grade malignancy in context of additional oncogenic changes by reactivating this silenced embryonic program.25 In contrast to epithelial cells where EMT-inducing transcription factors favor dedifferentiation and dissemination, we shedded light on a more complex interaction between ZEB2 and MITF in the melanocyte lineage. As ZEB2 expression favors MITF expression and thus differentiation in melanocytes, the loss of Zeb2 in the melanocyte lineage results in the formation of undifferentiated melanocytes with loss of Mitf and upregulation of Zeb1. Together with our finding that ZEB1 is expressed in melanocyte stem cells, it seems that ZEB switching is a naturally occurring transcription factor reprogramming that also can be facilitated by activated oncogenic RAS/RAF signaling.28
Examination of human melanoma samples and cell lines assessed the relevance of the identified effects upon in vivo loss of ZEB2 in a clinical context. It seems that ZEB2, particularly in the highly invasive vertical growth phase of human melanoma, gets downregulated in a very heterogeneous manner. Importantly, nuclear ZEB2 expression in melanomas is linked with good prognostic markers whereas loss of ZEB2 is associated with poor melanoma-specific survival. The partial or total clearance of ZEB2 in human melanoma could be caused by the well-known negative control of ZEB2 expression by the miR200 family. However this reasoning is not supported by the observation that expression of the miR200 family is lost during melanoma progression ruling out their potential dominant action of ZEB2 repression, at least during melanoma progression.42, 43
As MITF depletion forces melanocytes to senescence,39 ZEB1 may be a failsafe program for a melanoma cell to overcome the senescent state, exemplifying why ZEB1 expression in human melanoma is inversely correlated with MITF status. In this context, it is noteworthy that loss of ZEB2 in non-transformed melanocytes in vivo can result in ZEB1 misexpression, which potentially takes over in an alternative way the function of ZEB2 on the expense of differentiation. Our novel finding demonstrates that the switch from ZEB2 to ZEB1 is associated with a reduction of MITF, the master controller of differentiation, growth and migration in melanocytes. In melanoma, MITF has been described as a lineage addiction oncogene that is controlling melanoma progression following a rheostat model, in which high MITF expression is required for proliferation whereas its reduced level results in invasive behavior. We postulate that low MITF expression favors melanoma invasion and metastasis, which corresponds with the poor patient prognosis predicted by the loss of ZEB2, as shown in our study.
Materials and Methods
Mice
Mice were kept in accordance with the institutional guidelines regarding the care and use of laboratory animals and all procedures were approved by the institutional ethics committee. Mice with the following genotypes have been described elsewhere: conditional Zeb2fl/fl,44 Tyr::Cre36 and Dct::LacZ.37 Conditionally, Rosa-Zeb2TG/TG-IRES-GFP overexpressing mice were generated using G4 hybrid ES cells and the Gateway-compatible Rosa26 locus targeting vector, as previously described.45, 46
Melanin content determination
Melanin was extracted with an alkaline solution as previously described.47
Whole-mount LacZ staining of embryos and skin
To obtain E15.5 embryos, timed matings were set up, and the day on which a vaginal plug was detected was set as E0.5. For skin samples, back skin of 5.5-day-old mice was isolated. Embryos and back skin were washed in PBS and fixed in 4% paraformaldehyde for 3 h at room temperature (RT), after which they were washed with permeabilization solution (0.1 M phosphate buffer pH 7.4, 2 mM MgCl2, 2.5 mM EGTA, 0.01% sodium deoxycholate, 0.02% NP40, 0.005% BSA) and stained overnight at RT with permeabilization buffer containing 5 mM potassium ferricyanide, 5 mM potassium ferrocyanide and 0.1% LacZ solution. The samples were washed in PBS and postfixed for 2 h in 4% paraformaldehyde at RT. To count embryonic melanoblasts externally, photos were taken with a Nikon AZ100 Multizoom microscope (Nikon, Tokyo, Japan) and defined regions were analyzed for LacZ-positive melanoblasts with Volocity software. For further immunohistochemical analysis of the back skin, samples were dehydrated, embedded in paraffin and sectioned at 6 μm.
Human tissues
Tissue samples were obtained from the Department of Dermatology, Universitair Ziekenhuis Gent, Ghent, Belgium, from the Department of Pathology, University Hospital Leuven, KU Leuven, Leuven, Belgium and from archival paraffin-embedded patient samples from St. Vincent’s University Hospital, Dublin, Ireland. All patient specimens were used in accordance with institutional and national policies at the respective locations, with appropriate approval provided by the relevant Ethics committees at the respective institutions. All patient-related information was anonymized.
Immunohistochemistry
Tumors, organs and skin were isolated and fixed overnight in 4% paraformaldehyde solution, dehydrated, embedded in paraffin and cut into 5-μm sections. For histology, samples were stained with hematoxylin and eosin, or with nuclear fast red. For immunohistochemical staining, antigen retrieval was done in citrate buffer and endogenous peroxidases were blocked with 3% H2O2 in methanol. The sections were incubated with primary antibodies and stained with biotin-conjugated secondary antibodies followed by Streptavidin-HRP based development (substrate development with DAB or AEC). When necessary, the signal was amplified using the Tyramide Signal Amplification (TSA) kit (Perkin Elmer Applied Biosystems, Zaventem, Belgium), indicated with asterisk. The following antibodies were used: mouse anti-ZEB2 (1/300*) (in-house monoclonal antibody, clone 7F7) for normal mouse sections, rabbit anti-ZEB2 (1/500*) (HPA003456, Sigma, Diegem, Belgium) for human sections, goat anti-ZEB1 (1/50) (sc-10572, Santa Cruz, Heidelberg, Germany) for mouse sections, rabbit anti-ZEB1 (1/10 000*) (gift from Professor D Darling) for human sections; rabbit anti-S100b (1/1000*) (ZO311, Dakopatts, Leuven, Belgium), rabbit TYRP1 (1/1000*) (gift received kindly from Professor V Hearing); mouse anti-MITF (1/300) (Ab 12039, Abcam, Cambridge, UK), mouse anti-PAX3 (1/200) (DSHB, Iowa city, IA, USA).
Tyrosinase assay
The Tyrosinase assay was performed as described previously.47
Transmission electron microscopy
Back skin from 5.5-day-old ZEB2MCWT and ZEB2MCKO mice was immersed in a fixative solution of 2.5% glutaraldehyde, 3% formaldehyde and 0.02% CaCl2 in 0.1 M Na-cacodylate buffer, and processed as previously described.48 Ultrathin sections of a gold interference color were cut using an ultra microtome (Leica EM UC6, Diegem, Belgium), post-stained with uranyl acetate and lead citrate in a Leica EM AC20, and collected on Formvar-coated copper slot grids. They were viewed with a JEOL 1010 transmission electron microscope ( × 10 000 images) (JEOL, Tokyo, Japan).
Primary melanocyte cultures
Primary melanocytes were isolated from 5.5-day-old pups as previously described.47
siRNA and plasmid transfections
To knock down Zeb2 in primary NRAS transformed melanocytes, primary Melan a mouse melanocytes, B16-BL6 mouse melanoma and SKMel28 human melanoma, we used siRNA pools for mouse Zeb2 (Dharmacon, St. Leon Rot, Germany) and a scrambled siRNA pool (Dharmacon) as a control. Transfections were performed with HiPerfect (Qiagen, Antwerpen, Belgium) according to the manufacturer’s instructions. Five days after siRNA transfection, RNA and protein lysates were prepared. To overexpress mouse Mitf we used a mouse Mitf expression vector and an empty vector as a control. Transfections were performed with GenJet (Stratagene, Amsterdam, The Netherlands) according to the manufacturer’s instructions. Mitf transfections were done 1 day after siRNA knockdown of Zeb2.
Lentiviral transfections
To knock down MITF in 501Mel cells we used an shRNA for human MITF and a scrambled shRNA (shCTRL) as a control. The shRNA vectors are pLKO based from Sigma, the Mission shRNA series. 501Mel cells were infected with shRNA directed against MITF or the Mission shRNA control sequence and selected for 4 days with 3.0 μg/ml of puromycin.
ChIP analysis
ChIP and ChIP-Seq experiments were performed on chromatin from native 501Mel and on 501Mel cells stably expressing 3HA-tagged MITF to standard protocols as previously described.39
Real-time qPCR analysis
RNA was extracted from cell cultures or ears of 15.5-day-old ZEB2MCWT and ZEB2MCKO mice by using RNeasy extraction columns (Qiagen). RNA was treated with 1 U of RNAse-free DNase RQ1 (Promega, Leiden, The Netherlands) per μg RNA for 30 min at 37 °C in appropriate buffer. DNAse was inactivated by incubation in Promega stop solution for 10 min at 65 °C. Bulk Mg2+ was removed by using Amicon ultra 0.5-ml centrifugal filters (Millipore, Brussels, Belgium) in two consecutive diluting washes. cDNA synthesis was performed with iScript (Bio-Rad, Hercules, CA, USA) following the manufacturer’s instructions. Quantitative PCR was done using the Fast SYBR master mix kit (Applied Biosystems, Gent, Belgium) or SensiFastTM SYBR No-Rox kit (Bioline, Alpen aan de Rijn, The Netherlands) for the genes of interest and reference genes. Primers were designed using Primer Express 1.0 Software (Perkin Elmer Applied Biosystems) or obtained from the literature.47, 49 Plates were run on the LightCycler 480 (Roche, Vilvoorde, Belgium). The average threshold cycle of triplicate reactions was used for all subsequent calculations using the deltaCT method. Graphs represent the average normalized relative expression values of ZEB2MCWT and ZEB2MCKO mice, with the WT average normalized relative to the expression value at 4 days set to 1.
Western blot analysis
Cells were lysed in Laemli-lysis buffer (50 mMTris-HCl pH 6.8, 10% glycerol, 2% SDS). After sonicating and centrifuging the samples, 20 μg of protein was separated on gel and transferred to a PVDF membrane. Membranes were incubated with primary antibodies and appropriate HRP-labeled secondary antibodies (GE Healthcare, Diegem, Belgium). Detection was performed with the Western LightningT chemiluminescence reagent plus kit (Perkin Elmer, Waltham, MA, USA) or the Immobilion western HRP substrate (Millipore). The following antibodies were used: rabbit anti-ZEB2 (1/1000) (HPA003456, Sigma); rabbit anti-ZEB1 (1/1000) (Gift from Professor Darling); mouse anti-β-Tubulin (1/1000) (T4026, Sigma); mouse anti-MITF (Ab 12039 Abcam); mouse anti-Vinculin (1/3000) (V9131, Sigma); mouse anti-Fibronectin (1/1000) (Abcam, ab23750).
Statistical analysis
Data were analyzed with GraphPad Prism 5 (GraphPad Software Inc., La Jolla, CA, USA) and R (The Comprehensive R Archive Network, http://www.cran.r-project.org).
Abbreviations
- β-Cat:
-
β-Catenin
- BCL-2:
-
B-Cell CLL/Lymphoma 2
- bHLH:
-
basic Helix Loop Helix
- BRAF:
-
viral RAF murine sarcoma viral oncogene homolog B1
- Cdkn2a:
-
cyclin-dependent kinase inhibitor 2A
- ChIP:
-
chromatin immunoprecipitation
- c-MYC:
-
cellular myelocytomatosis oncogene cellular homolog
- CRE:
-
cyclisation recombination
- CTRL:
-
control
- DCT:
-
dopachrome tautomerase
- E:
-
embryonic day
- EDN3/EDNRB:
-
endothelin 3/endothelin receptor B
- ERK:
-
extracellular signal-regulated kinase
- EMT:
-
epithelial-to-mesenchymal transition
- FL:
-
floxed
- HF:
-
hair follicle
- KD:
-
knockdown
- LacZ:
-
β-D-galactosidase
- MC1R:
-
melanocortin 1 receptor
- MCKO:
-
melanocyte-specific knockout
- MCWT:
-
melanocyte-specific wild-type
- MEK:
-
MAPK/ERK kinase
- MITF:
-
microphthalmia-associated transcription factor
- P:
-
postnatal day
- PAX3:
-
paired-box 3
- PGP:
-
P-glycoprotein
- PMEL:
-
premelanosome protein
- NHM:
-
normal human primary melanocytes
- NRAS:
-
neuroblastoma RAS viral oncogene homolog
- qRT-PCR:
-
quantitative reverse transcription-polymerase chain reaction
- S100b:
-
S100 calcium-binding protein B
- shRNA:
-
short hairpin RNA
- siRNA:
-
short interfering RNA
- SOX10:
-
sex-determining region Y (SRY)-box 10
- TG:
-
transgenic
- TGF-β:
-
transforming growth factor β
- TYRP1:
-
tyrosinase-related protein 1
- TYR:
-
tyrosinase
- UV:
-
ultraviolet
- X-gal:
-
5-bromo-4-chloro-3-indolyl-β-D-galactopyranoside
- ZEB:
-
zinc finger E-box binding protein
References
Ernfors P . Cellular origin and developmental mechanisms during the formation of skin melanocytes. Exp Cell Res 2010; 316: 1397–1407.
Luciani F, Champeval D, Herbette A, Denat L, Aylaj B, Martinozzi S et al. Biological and mathematical modeling of melanocyte development. Development 2011; 138: 3943–3954.
Osawa M . Melanocyte stem cells (June 30, 2009), StemBook, ed. The Stem Cell Research Community, StemBook, doi:10.3824/stembook.1.46.1.
Pshenichnaya I, Schouwey K, Armaro M, Larue L, Knoepfler PS, Eisenman RN et al. Constitutive gray hair in mice induced by melanocyte-specific deletion of c-Myc. Pigment Cell Melanoma Res 2012; 25: 312–325.
Steingrimsson E, Copeland NG, Jenkins NA . Melanocyte stem cell maintenance and hair graying. Cell 2005; 121: 9–12.
White RM, Zon LI . Melanocytes in development, regeneration, and cancer. Cell Stem Cell 2008; 3: 242–252.
Kubic JD, Young KP, Plummer RS, Ludvik AE, Lang D . Pigmentation PAX-ways: the role of Pax3 in melanogenesis, melanocyte stem cell maintenance, and disease. Pigment Cell Melanoma Res 2008; 21: 627–645.
Hearing VJ . Determination of melanin synthetic pathways. J Invest Dermatol 2011; 131: E8–E11.
Schiaffino MV . Signaling pathways in melanosome biogenesis and pathology. Int J Biochem Cell Biol 2010; 42: 1094–1104.
Nishimura EK, Granter SR, Fisher DE . Mechanisms of hair graying: incomplete melanocyte stem cell maintenance in the niche. Science 2005; 307: 720–724.
Moriyama M, Osawa M, Mak SS, Ohtsuka T, Yamamoto N, Han H et al. Notch signaling via Hes1 transcription factor maintains survival of melanoblasts and melanocyte stem cells. J Cell Biol 2006; 173: 333–339.
Tsao H, Chin L, Garraway LA, Fisher DE . Melanoma: from mutations to medicine. Genes Dev 2012; 26: 1131–1155.
Gembarska A, Luciani F, Fedele C, Russell EA, Dewaele M, Villar S et al. MDM4 is a key therapeutic target in cutaneous melanoma. Nat Med 2012; 18: 1239–1249.
Ackermann J, Frutschi M, Kaloulis K, McKee T, Trumpp A, Beermann F . Metastasizing melanoma formation caused by expression of activated N-RasQ61K on an INK4a-deficient background. Cancer Res 2005; 65: 4005–4011.
Flaherty KT, Puzanov I, Kim KB, Ribas A, McArthur GA, Sosman JA et al. Inhibition of mutated, activated BRAF in metastatic melanoma. N Engl J Med 2010; 363: 809–819.
Nazarian RM, Prieto VG, Elder DE, Duncan LM . Melanoma biomarker expression in melanocytic tumor progression: a tissue microarray study. J Cutan Pathol 2010; 37 (Suppl 1): 41–47.
Flaherty KT, Infante JR, Daud A, Gonzalez R, Kefford RF, Sosman J et al. Combined BRAF and MEK inhibition in melanoma with BRAF V600 mutations. N Engl J Med 2012; 367: 1694–1703.
Colone M, Calcabrini A, Toccacieli L, Bozzuto G, Stringaro A, Gentile M et al. The multidrug transporter P-glycoprotein: a mediator of melanoma invasion? J Invest Dermatol 2008; 128: 957–971.
Sanchez-Tillo E, Liu Y, de Barrios O, Siles L, Fanlo L, Cuatrecasas M et al. EMT-activating transcription factors in cancer: beyond EMT and tumor invasiveness. Cell Mol Life Sci 2012; 69: 3429–3456.
Thiery JP, Acloque H, Huang RY, Nieto MA . Epithelial-mesenchymal transitions in development and disease. Cell 2009; 139: 871–890.
Peinado H, Olmeda D, Cano A . Snail, Zeb and bHLH factors in tumour progression: an alliance against the epithelial phenotype? Nat Rev Cancer 2007; 7: 415–428.
Sanchez-Martin M, Perez-Losada J, Rodriguez-Garcia A, Gonzalez-Sanchez B, Korf BR, Kuster W et al. Deletion of the SLUG (SNAI2) gene results in human piebaldism. Am J Med Genet A 2003; 122A: 125–132.
Perez-Losada J, Sanchez-Martin M, Rodriguez-Garcia A, Sanchez ML, Orfao A, Flores T et al. Zinc-finger transcription factor Slug contributes to the function of the stem cell factor c-kit signaling pathway. Blood 2002; 100: 1274–1286.
Jiang R, Lan Y, Norton CR, Sundberg JP, Gridley T . The Slug gene is not essential for mesoderm or neural crest development in mice. Dev Biol 1998; 198: 277–285.
Gupta PB, Kuperwasser C, Brunet JP, Ramaswamy S, Kuo WL, Gray JW et al. The melanocyte differentiation program predisposes to metastasis after neoplastic transformation. Nat Genet 2005; 37: 1047–1054.
Sanchez-Martin M, Rodriguez-Garcia A, Perez-Losada J, Sagrera A, Read AP, Sanchez-Garcia I . SLUG (SNAI2) deletions in patients with Waardenburg disease. Hum Mol Genet 2002; 11: 3231–3236.
Shirley SH, Greene VR, Duncan LM, Torres Cabala CA, Grimm EA, Kusewitt DF . Slug expression during melanoma progression. Am J Pathol 2012; 180: 2479–2489.
Caramel J, Papadogeorgakis E, Hill L, Browne GJ, Richard G, Wierinckx A et al. A switch in the expression of embryonic EMT-inducers drives the development of malignant melanoma. Cancer Cell 2013; 24: 466–480.
Vandewalle C, Van Roy F, Berx G . The role of the ZEB family of transcription factors in development and disease. Cell Mol Life Sci 2009; 66: 773–787.
Gheldof A, Hulpiau P, van Roy F, De Craene B, Berx G . Evolutionary functional analysis and molecular regulation of the ZEB transcription factors. Cell Mol Life Sci 2012; 69: 2527–2541.
Comijn J, Berx G, Vermassen P, Verschueren K, van Grunsven L, Bruyneel E et al. The two-handed E box binding zinc finger protein SIP1 downregulates E-cadherin and induces invasion. Mol Cell 2001; 7: 1267–1278.
De Craene BD, Berx G . Regulatory networks defining EMT during cancer initiation and progression. Nat Rev Cancer 2013; 13: 97–110.
Van de Putte T, Maruhashi M, Francis A, Nelles L, Kondoh H, Huylebroeck D et al. Mice lacking ZFHX1B, the gene that codes for Smad-interacting protein-1, reveal a role for multiple neural crest cell defects in the etiology of Hirschsprung disease-mental retardation syndrome. Am J Hum Genet 2003; 72: 465–470.
Van de Putte T, Francis A, Nelles L, van Grunsven LA, Huylebroeck D . Neural crest-specific removal of Zfhx1b in mouse leads to a wide range of neurocristopathies reminiscent of Mowat-Wilson syndrome. Hum Mol Genet 2007; 16: 1423–1436.
Colombo S, Champeval D, Rambow F, Larue L . Transcriptomic analysis of mouse embryonic skin cells reveals previously unreported genes expressed in melanoblasts. J Invest Dermatol 2012; 132: 170–178.
Delmas V, Martinozzi S, Bourgeois Y, Holzenberger M, Larue L . Cre-mediated recombination in the skin melanocyte lineage. Genesis 2003; 36: 73–80.
Mackenzie MA, Jordan SA, Budd PS, Jackson IJ . Activation of the receptor tyrosine kinase Kit is required for the proliferation of melanoblasts in the mouse embryo. Dev Biol 1997; 192: 99–107.
Thurber AE, Douglas G, Sturm EC, Zabierowski SE, Smit DJ, Ramakrishnan SN et al. Inverse expression states of the BRN2 and MITF transcription factors in melanoma spheres and tumour xenografts regulate the NOTCH pathway. Oncogene 2011; 30: 3036–3048.
Strub T, Giuliano S, Ye T, Bonet C, Keime C, Kobi D et al. Essential role of microphthalmia transcription factor for DNA replication, mitosis and genomic stability in melanoma. Oncogene 2011; 30: 2319–2332.
Chin L, Garraway LA, Fisher DE . Malignant melanoma: genetics and therapeutics in the genomic era. Genes Dev 2006; 20: 2149–2182.
Southall TD, Brand AH . Neural stem cell transcriptional networks highlight genes essential for nervous system development. EMBO J 2009; 28: 3799–3807.
Xu Y, Brenn T, Brown ER, Doherty V, Melton DW . Differential expression of microRNAs during melanoma progression: miR-200c, miR-205 and miR-211 are downregulated in melanoma and act as tumour suppressors. Br J Cancer 2012; 106: 553–561.
van Kempen LC, van den Hurk K, Lazar V, Michiels S, Winnepenninckx V, Stas M et al. Loss of microRNA-200a and c, and microRNA-203 expression at the invasive front of primary cutaneous melanoma is associated with increased thickness and disease progression. Virchows Arch 2012; 461: 441–448.
Higashi Y, Maruhashi M, Nelles L, Van de Putte T, Verschueren K, Miyoshi T et al. Generation of the floxed allele of the SIP1 (Smad-interacting protein 1) gene for Cre-mediated conditional knockout in the mouse. Genesis 2002; 32: 82–84.
Nyabi O, Naessens M, Haigh K, Gembarska A, Goossens S, Maetens M et al. Efficient mouse transgenesis using Gateway-compatible ROSA26 locus targeting vectors and F1 hybrid ES cells. Nucleic Acids Res 2009; 37: e55.
van den Berghe V, Stappers E, Vandesande B, Dimidschstein J, Kroes R, Francis A et al. Directed migration of cortical interneurons depends on the cell-autonomous action of Sip1. Neuron 2013; 77: 70–82.
Berlin I, Luciani F, Gallagher SJ, Rambow F, Conde-Perez A, Colombo S et al. General strategy to analyse coat colour phenotypes in mice. Pigment Cell Melanoma Res 2012; 25: 117–119.
Denecker G, Hoste E, Gilbert B, Hochepied T, Ovaere P, Lippens S et al. Caspase-14 protects against epidermal UVB photodamage and water loss. Nat Cell Biol 2007; 9: 666–674.
Gallagher SJ, Luciani F, Berlin I, Rambow F, Gros G, Champeval D et al. General strategy to analyse melanoma in mice. Pigment Cell Melanoma Res 2012; 24: 987–988.
Acknowledgements
This research was funded by grants from the FWO, the Geconcerteerde Onderzoeksacties of Ghent University, the Stichting tegen Kanker, the Association for International Cancer Research (Scotland), the EU-FP6 framework program BRECOSM LSHC-CT-2004-503224, the EU-FP7 framework programs TuMIC 2008-201662 and Target-Melanoma (www.targetmelanoma.com). Work in the lab of ID was supported by grants from the Ligue National Contre le Cancer, the INCa, the Université de Strasbourg and the ANR. The IGBMC high throughput sequencing facility is a member of the ‘France Génomique’ consortium (ANR10-INBS-09-08). We acknowledge Riet DeRycke for expert electron microscopic analysis, Dr. Amin Bredan for critical reading of the manuscript and the members of our research group for valuable discussions. We thank Professor D Darling for providing the ZEB1 antibody, Professor K Hearing for providing the TYRP1 antibody.
Author contributions
GD, NV, ÖA, DK, JT, KL, AG, BDC, BB, ID, JCM and GB performed the experiments and analyzed the interpreted data. MVG, LB, GMU, MR, WMG, GG, DH, JvdO, LL and JH provided valuable reagents/material. GD and GB conceived-designed the project, analyzed the interpreted data and wrote the paper with inputs particularly from NV, DH, LL, JH, JCM and all other authors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by G Melino
Supplementary Information accompanies this paper on Cell Death and Differentiation website
Rights and permissions
About this article
Cite this article
Denecker, G., Vandamme, N., Akay, Ö. et al. Identification of a ZEB2-MITF-ZEB1 transcriptional network that controls melanogenesis and melanoma progression. Cell Death Differ 21, 1250–1261 (2014). https://doi.org/10.1038/cdd.2014.44
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2014.44
This article is cited by
-
Sexual dimorphism in melanocyte stem cell behavior reveals combinational therapeutic strategies for cutaneous repigmentation
Nature Communications (2024)
-
Microenvironmental Snail1-induced immunosuppression promotes melanoma growth
Oncogene (2023)
-
The epithelial–mesenchymal plasticity landscape: principles of design and mechanisms of regulation
Nature Reviews Genetics (2023)
-
BCAT2 promotes melanoma progression by activating lipogenesis via the epigenetic regulation of FASN and ACLY expressions
Cellular and Molecular Life Sciences (2023)
-
Newly identified form of phenotypic plasticity of cancer: immunogenic mimicry
Cancer and Metastasis Reviews (2023)