Abstract
Transcription regulates axon outgrowth and regeneration. However, to date, no transcription complexes have been shown to control axon outgrowth and regeneration by regulating axon growth genes. Here, we report that the tumor suppressor p53 and its acetyltransferases CBP/p300 form a transcriptional complex that regulates the axonal growth-associated protein 43, a well-characterized pro-axon outgrowth and regeneration protein. Acetylated p53 at K372-3-82 drives axon outgrowth, GAP-43 expression, and binds specific elements on the neuronal GAP-43 promoter in a chromatin environment through CBP/p300 signaling. Importantly, in an axon regeneration model, both CBP and p53 K372-3-82 are induced following axotomy in facial motor neurons, where p53 K372-3-82 occupancy of GAP-43 promoter is enhanced as shown by in vivo chromatin immunoprecipitation. Finally, by comparing wild-type and p53 null mice, we demonstrate that the p53/GAP-43 transcriptional module is specifically switched on during axon regeneration in vivo. These data contribute to the understanding of gene regulation in axon outgrowth and may suggest new molecular targets for axon regeneration.
Similar content being viewed by others
Main
Axon outgrowth, sprouting and regeneration require an orchestrated sequence of events. These consist of the expression of extracellular and cellular signaling proteins, which remodel the cytoskeleton to modulate axon and growth cone plasticity. At the transcriptional level, they are regulated by a set of transcription factors that include c-jun, C/EBP, CREB, STAT3 and p53.1, 2 However, the fine-tuning of their expression is likely provided by the appropriate formation of transcriptional modules, which include transcription factors, co-activators and promoter sequences.
Thus far, neither specific transcription factors nor transcriptional complexes have been described to play a direct role in axon outgrowth by occupying promoters of pro-axon growth genes. To address this issue, we have chosen to investigate the transcriptional regulation of a prototypical axon growth factor, the growth-associated protein 43 (GAP-43), in a neurite/axon outgrowth and regeneration context. In fact, we believe that clarification of GAP-43 transcriptional environment may provide insights into the general transcription-dependent molecular mechanisms that regulate axon outgrowth, sprouting and regeneration in specific domains of the nervous system.
GAP-43 is a neurotrophin-dependent membrane bound phosphoprotein found in the axon and the growth cone of neurons.3, 4 It is highly expressed in several plastic regions of the nervous system during development,5, 6 following acute peripheral nerve and CNS injuries such as stroke, brain and spinal cord trauma and in neurons undergoing synaptic plasticity and axon sprouting.7, 8, 9, 10, 11 GAP-43 contributes to axon sprouting and regeneration in the peripheral nervous system, in the optic nerve, in the Purkinje cells of the cerebellum, in mossy fibers of the hippocampus and in the raphe-spinal descending spinal tracts.12, 13, 14, 15 However, GAP-43 alone does not significantly promote axon regeneration of the cortico-spinal tracts and the dorsal root ganglia ascending spinal fibers, but does so after dorsal column lesions when overexpressed together with CAP-23.16
Thus far, it has been shown that GAP-43 is regulated at both the transcriptional and post-transcriptional levels. Members of the ELAV-Hu family of RNA-binding proteins are essential post-transcriptional regulators of GAP-43 in PC-12 cells, developing neurons and certain mature neurons.17, 18, 19, 20 In fact, by binding to conserved motifs in the 3′-UTR of GAP-43 mRNA, they stabilize GAP-43 in the cytoplasm and the growth cone.21
Studies aimed at elucidating the transcriptional regulation of GAP-43 have highlighted the presence of a 1000 bp promoter element upstream of the protein-coding region, which is important for its neuronal expression.22, 23 Within this promoter region an AP-1 responsive element was identified,24 but neither the Jun nor Fos transcription factors, which bind to AP-1, have been shown to directly regulate GAP-43 expression. In addition, in vitro assays have demonstrated that the basic helix-loop-helix differentiation factor Nex1/MATH-2 binds to the promoter region of GAP-43 in proximity to the transcription start site.25
Recently, we have discovered that p53 is required for neurite outgrowth and axon regeneration following peripheral nerve injury, in part by modulating the expression of the cytoskeleton-associated proteins Coronin 1b and Rab13.26 Importantly, GAP-43 was observed to be co-expressed with Coronin 1b and Rab13 in cultured neurons during axon outgrowth and in vivo within sprouting axons after spinal cord injury.26, 27
As a result, we asked whether axon outgrowth and GAP-43 could be regulated by p53 within a specific transcriptional environment.
Results
p53 binds to specific GAP-43 promoter elements and drives their expression
To identify putative transcription factors for GAP-43 neuronal gene expression, we performed an in silico analysis using the MatInspector tool from Genomatix (www.genomatix.de) on the entire rat GAP-43 5′-promoter region. This included the 1000 bp upstream from the protein-coding sequence, which was a previously identified region required for GAP-43 neuronal gene expression.22, 23 Our in silico analysis led to the discovery of a p53 putative-binding site located about 600 bp upstream from GAP-43 protein-coding sequence. The p53-binding motif is identical in rat and mouse, and highly conserved in human, thus supporting its physiological relevance (Figure 1a).
p53 occupies the TFBS of the GAP-43 promoter and modulates its transcription in neuronal cells. (a) Alignment of the p53 TFBS from the GAP-43 promoter in rat, mouse and human. The core conserved binding domain is in bold within the box. These TFBS were identified in silico by using the MatInspector tool from Genomatix. (b) A ChIP assay shows in vivo interaction in a chromatin environment of wt p53 with the p53 TFBS derived from the rat GAP-43 promoter. PC-12 cells were left untreated or treated with NGF (100 ng/ml) for 30 h to induce differentiation. Thereafter, cells were crosslinked with formaldehyde and immunoprecipitated with a polyclonal antibody recognizing full length p53 (FL-393). The input samples were collected before precipitation and used as a normalization control. The WAF1/p21 promoter region was used as a positive control for p53 binding. 3′end regions of the GAP-43 gene were used as a control of the specificity of the occupancy by p53 on GAP-43 promoter. (c) An analogous ChIP assay was performed as in (b) in PC-12 cells stably overexpressing p53DN. In this case, p53 does not show binding to the GAP-43 or the p21/WAF1 promoter (right). (d) A ChIP assay, performed as in (b), shows that the GAP-43 promoter is occupied by endogenous p53 in cultured rat cortical neurons. (e–g) A wt rat GAP-43 promoter element containing the p53 TFBS (e and f) but not a scrambled sequence (g) is responsive to wt p53 overexpression in rat cortical neurons (e), and PC-12 cells (f and g). Following p53 overexpression by transfection of full-length p53, a dual luciferase assay was performed in rat cortical neurons (e), and in NGF-treated (100 ng/ml) or untreated PC12 cells (f and g). The activity expressed in fold changes of both the scrambled and the p53 TFBS containing luciferase construct were normalized to the pcDNA3 empty vector transfected cells. Asterisks indicate a significant difference analyzed by an unpaired two-tailed t-test with a P-value <0.01, triplicate samples. Below each bar graph are western blots that show the expression levels of p53 after transfection. β-actin was used as a loading control
To evaluate whether endogenous p53 can bind within a chromatin environment to the GAP-43 promoter region containing the in silico identified putative-binding site, chromatin immunoprecipitation assays (ChIP) were performed. First, neuronal-like PC-12 cells were used, as they express both p53 and GAP-43. Endogenous p53 was found to occupy the in silico predicted transcription factor-binding site (TFBS) of endogenous GAP-43 promoter in PC-12 cells (Figure 1b). This occupancy is not significantly increased by the addition of nerve growth factor (NGF) (Figure 1b). Occupancy prior to NGF stimulation suggests that p53 binding to the GAP-43 promoter also occurs independently of NGF signaling. Previously, we have shown that a dominant-negative p53 (p53DN R175H) inhibits p53-dependent transcription.28 Here, we specifically show that only wild-type endogenous p53 binds to the GAP-43 promoter elements. In fact, in a ChIP assay performed in PC-12 cells stably transfected with p53DN mutant, p53 does not occupy this GAP-43 promoter region (Figure 1c). Then, more importantly, we performed ChIP assays in primary cortical neurons, which also show occupancy by endogenous p53 of GAP-43 promoter elements (Figure 1d). A standard p53 target, represented by p21 promoter elements, was used here as a positive control for p53 occupancy (Figure 1b–d). In addition, 3′end regions of GAP-43 DNA were used as an internal control of the specificity of the binding by p53 on GAP-43 (Figure 1b–d).
To test the functionality of p53 interaction with the GAP-43 promoter elements, we cloned this p53-binding site region of the GAP-43 promoter into a luciferase expression vector. Transfection of wild-type p53 into both PC-12 and rat cortical neurons showed that p53 overexpression leads to a substantial increase in GAP-43 promoter-induced luciferase activity (Figure 1e and f). In PC-12 cells, administration of NGF amplifies this induction, but again is not required for it (Figure 1f). Specificity of p53 for this interaction was established by using a luciferase reporter containing a scrambled GAP-43 p53 core element TFBS, which does not result in an induction of luciferase activity following p53 overexpression (Figure 1g).
Taken together, these data show that endogenous p53 binds to in silico identified GAP-43 specific promoter elements in cell lines as well as primary neurons, to drive GAP-43 expression.
p53 is required for GAP-43 expression during neurite and axon outgrowth
The confirmation that p53 occupies and positively regulates GAP-43 promoter elements in neurons led to the hypothesis that p53 should regulate the expression levels of GAP-43 during neurite and axon outgrowth. First, we tested this hypothesis in differentiated PC-12 and P19 cells that express high levels of GAP-43 and p53 (see online Supplementary Information). In summary, those experiments support the conclusion that p53 is required for GAP-43 expression along with neurite-axon outgrowth in neuronal-like cells (Supplementary Figures 1–3).
Next, experiments were conducted in mouse brains and in primary neurons to investigate p53's regulation of GAP-43 expression in vivo and in neurons. We demonstrate by real-time RT-PCR that expression of GAP-43 mRNA is reduced by 60% in the mouse brain of p53 null versus wild-type (wt) mice, thereby showing that p53 is necessary for physiological levels of GAP-43 expression in the brain (Figure 2a). In addition, GAP-43, which is induced early on in cultured primary cortical neurons during axon development and outgrowth, is inhibited in neurons infected with a p53DN lentivirus (Figure 2b). By immunocytochemistry for GAP-43, we observed that p53DN expression also inhibits axon development and outgrowth as compared with neurons infected with GFP only (Figure 2c). In addition, a marked reduction in axon length, correlating to reduced GAP-43 expression is observed in cultured mouse cortical neurons from p53 null mice (Figure 2d).
p53 regulates GAP-43 expression and neurite/axon outgrowth in the brain and in cortical neurons. (a) Real-time RT-PCR experiments show a significant decrease in mRNA expression for GAP-43 in the brain of p53 null as compared with wt mice. Asterisk indicates a significant difference analyzed by unpaired two-tailed t-test with a P-value <0.01, triplicate experiments. (b) Immunoblotting shows inhibition of GAP-43, following infection with p53DN lentivirus at day 4 in rat cortical neurons. Anti-V5 antibodies were used to identify p53DN levels in infected cells. β-actin was used as a loading control. Bar graph shows densitometric data analysis: intensity of representative bands for GAP-43 was normalized to β-actin. (c) Immunofluorescence shows the inhibition of neurite outgrowth and branching by p53DN expressed in the nucleus versus eGFP in lentivirus infected rat cortical neurons. Neurite measurements show a significant difference in several parameters between eGFP- and p53DN-infected neurons (Asterisks: unpaired two-tailed t-test with a P-value<0.01, triplicate experiments). Scale bar: 50 μm. (d) E16 mouse cortical neurons (DIV3) from p53 null mice display diminished GAP-43 expression and neurite elongation when compared with wt neurons. Neurite measurements show a significant difference in neurite length (Asterisks: unpaired two-tailed t-test with a P-value<0.01, triplicate experiments). Scale bar: 50 μm
These results support the conclusion that p53 is required for adequate GAP-43 expression in the mouse brain along with axon outgrowth in primary neurons.
GAP-43 is preferentially regulated by a CBP/p300 dependent p53 acetylation complex
Cellular hyperacetylation has been found to have a neuroprotective effect in neurons both in vitro and in vivo.29 p53 can be acetylated at several Lysine residues in its C-terminal domain, including at Lysine 320 (K320), mediated by the acetyltransferase P/CAF,30 and at Lysines 372-3 and 382 (K373), mediated by CBP/p300.31, 32 Each of these two acetylation patterns re-directs p53 towards specific promoters.33 However, the role of the acetylation of p53, in driving neurite-axon outgrowth and pro-growth promoters has not been addressed in primary neurons.
Here, we examined whether GAP-43 expression is regulated by specific p53 acetylation pathways. First, we measured acetylated p53 protein levels. Immunoblotting for specific p53-acetylated forms shows similar neuronal protein expression levels for p53 acetyl K373 and p53 acetyl K320 (Figure 3a). Then, we show by immunofluorescence the nuclear expression of p53 acetyl K373, and P/CAF and the expression of p53 acetyl K320 and CBP in both nuclei and in neuronal processes (Supplementary Figure 4).
p53 acetyl K373 preferentially occupies the GAP-43 promoter through CBP/p300. (a) Immunoblotting for p53 full length (FL393), p53 acetyl K320 and acetyl K373 in cultured primary neurons at DIV 3. Expression levels of neuronal p53 K373 and K320 are similar. β-actin was used as a loading control. (b) ChIP assays show in vivo interaction in a chromatin environment of p53 K373 and p53 K320 with p53 TFBS derived from the rat GAP-43 promoter in cultured rat cortical neurons. Specific antibodies recognizing p53 acetyl K373 and p53 acetyl K320 were employed for immunoprecipitation. Results display preferential binding of p53 acetyl K373 versus acetyl p53 K320 to the GAP-43 promoter. TheWAF1/p21 promoter region was used as a positive control for p53 binding. PCR was also performed by using primers that amplify a 3′end site on GAP-43 gene as a control to assess the specificity of the occupancy of the promoter by p53 in all the ChIP assays (b–g). (c) ChIP assays performed as in (b), were followed by real-time quantitative PCR. The bar graph displays the fold increase in the binding affinity to the GAP-43 promoter elements of p53 acetyl K373 versus p53 acetyl K320 as analyzed following ChIP. TheWAF1/p21 promoter region was used as a positive control for p53 promoter occupancy. The input samples, collected before precipitation and the no antibody samples, were used as a normalization control. Asterisks indicate a significant difference analyzed by unpaired two-tailed t-test with a P-value <0.01, triplicate experiments. (d) ChIP assays were performed in PC-12 cells, which were left untreated or were treated with NGF (100 ng/ml) for 30 h to induce differentiation. p53 acetyl K373 binds GAP-43 promoter elements preferentially as compared with p53 acetyl K320. TheWAF1/p21 promoter region was used as a positive control for p53 promoter occupancy. (e) ChIP assays performed as in (d), were followed by real-time quantitative PCR. The bar graph shows the fold increase in specific promoter occupancy between p53 acetyl K373 and p53 acetyl K320 in untreated PC-12 cells as analyzed by real-time quantitative PCR following ChIP. These data are similar to the ones obtained in primary neurons (c). In addition, results show that NGF facilitates the occupancy to GAP-43 promoter for both p53 acetyl K373 and acetyl K320, but significantly more so for K373. The input samples, collected before precipitation and the no antibody samples, were used as a normalization control. Asterisks indicate a significant difference analyzed by unpaired two-tailed t-test with a P-value <0.01, triplicate experiments. (f) ChIP assays show GAP-43 promoter occupancy by the co-activators CBP/p300. The assay was performed as in (d), and specific antibodies for CBP and p300 were employed. (g) ChIP and real-time PCR assays following CBP/p300 overexpression or CBP/p300 gene silencing, respectively show a marked induction or inhibition of p53 acetyl K373 binding to the GAP-43 promoter elements. PC-12 cells were transfected with full-length CBP/p300 genes or control pcDNA, and RNAi for CBP/p300 or scrambled RNAs (scRNA). A specific anti-p53 acetyl K373 antibody was used for immunoprecipitation as in (d). The bar graph shows the fold change increase or decrease in p53 acetyl K373 binding to the GAP-43 promoter elements observed by a quantitative ChIP analysis after CBP/p300 overexpression or CBP/p300 silencing as compared with control pcDNA or scrambled RNAs, (scRNA) respectively. Asterisks indicate a significant difference analyzed by unpaired two-tailed t-test with a P-value below 0.01, triplicate experiments. (h) RT-PCR shows the expression levels of CBP and p300 after overexpression and RNAi experiments performed in (g)
To investigate the affinity of these two p53 acetylation forms for the GAP-43 promoter elements, we performed ChIP assays in both primary neurons and PC-12 cells. These experiments show occupancy by both acetylated p53 at K373 and K320 (Figure 3b–e). However, they show much higher promoter occupancy for acetylated p53 at K373 than p53 acetylated at K320 (Figure 3c and e). These effects were enhanced by NGF signaling in PC-12 cells (Figure 4e). ChIP experiments were also performed by immunoprecipitating GAP-43 promoter elements with antibodies against CBP/p300 acetyltransferases. Promoter occupancy was present for CBP and p300, demonstrating that CBP/p300 does form a transcriptional complex upon the GAP-43 promoter (Figure 3f). These data suggest that a specific CBP/p300-dependent p53 acetylation pattern (p53 K372-3-82) preferentially occupies GAP-43 promoter elements.
p53 K373Q specifically induces GAP-43 transcription elements and expression, and promotes the highest increase in neurite outgrowth in cortical neurons. (a) A dual luciferase assay was performed in PC-12 cells following overexpression of different p53 mutants that mimic acetylation (p53 K373Q, K320Q) or are unable to be acetylated (p53 K373R) at specific Lysine residues. The bar graphs show that only p53 K373Q drives the GAP-43 promoter elements. Neither p53 K320Q nor p53 K373R activate the GAP-43 promoter. The fold induction in relative luciferase activity for each construct was normalized against a pcDNA3 control vector. Asterisks indicate a significant difference between the pcDNA3 vector and the p53 mutants, analyzed by unpaired two-tailed t-test with a P-value <0.01, triplicate samples. Western blots for p53 show the expression levels after transfection (also in b and c). β-actin was used as a loading control. (b and c) Immunoblotting reveals an induction of GAP-43 protein expression following p53 K373Q overexpression in PC-12 p53DN, and P19 cells, but not with p53 K320Q overexpression. β-actin was used as a loading control. Bar graphs show densitometric data analysis: intensity of representative bands for GAP-43 was normalized to β-actin. (d) Cultured rat cortical neurons were transfected at day 3 for 72 h with eGFP, p53 K373Q-GFP, p53 K320Q-GFP or p53 K373R-GFP mutant plasmid DNAs. Representative images of transfected neurons demonstrate that overexpression of p53 K373Q as well as p53 K320Q enhances neurite outgrowth when compared with eGFP-transfected cells. However, p53 K373Q drives significantly enhanced neurite development than p53 K320Q. Transfection with p53 K373R does not increase neurite outgrowth as compared with eGFP-transfected neurons. The bar graph shows quantitation of neurite outgrowth measurements following transfections expressed as percentage values to control eGFP-transfected cells. Highest increase is observed following p53 K373Q transfection. Neurite measurements show a significant difference between eGFP and p53 K373Q, or K320Q transfected neurons. A significant difference is also found between p53 K373Q and K320Q. Asterisks: unpaired two-tailed t-test with a P-value<0.01, triplicate experiments. Scale bar: 200 μm
We therefore sought to investigate the role of CBP/p300 acetyltransferases on the modulation of the p53 binding to GAP-43 promoter elements.
ChIP assays were performed after silencing CBP/p300 by using specific RNA interference oligonucleotides and by immunoprecipitation of p53 with a specific antibody against p53 acetyl K373. Results show that silencing of CBP/p300 markedly reduces the ability of p53 K373 to occupy GAP-43 promoter elements (Figure 3g). In an analogous ChIP experiment, overexpression of full-length CBP/p300 increases the affinity of p53 acetyl K373 for the same promoter elements (Figure 3g). CBP/p300 overexpression or silencing led respectively to a strong increase or reduction in mRNA levels (Figure 3h).
Therefore, here we have demonstrated that CBP/p300 signaling increases the ability of p53 to occupy the GAP-43 promoter.
To address the question of whether specific K373 acetylation of p53 is also responsible and required for driving the expression of these GAP-43 promoter elements, we performed dual luciferase assays in PC-12 cells. In this experiment, we overexpressed p53 acetylation mimic mutants, which were obtained by a substitution of a positively charged Lysine with a neutral or uncharged Glutamine at K372-3-82 (K373Q) or K320 (K320Q). Results show that only p53 K373Q induces a significant increase in luciferase activity for the GAP-43 promoter elements (Figure 4a). We also studied another p53 mutant, which cannot be acetylated at Lysine 372-3-82 (K373R) due to a substitution of Lysine with Arginine. This mutant fails to drive luciferase activity supporting the specific activity of p53 K373Q on the GAP-43 promoter elements (Figure 4a).
Overexpression experiments by transfecting both PC-12 p53DN and P19 neuronal-like cells with p53 K373Q and K320Q show that only the 373 form drives protein expression of GAP-43 (Figure 4b and c). Importantly, p53 K373Q overexpression overcomes GAP-43 inhibition in PC-12 cells stably transfected with p53DN (Figure 5b). In comparison, transfection of a mutant incapable of acetylation, p53 K373R, does not increase GAP-43 expression (data not shown).
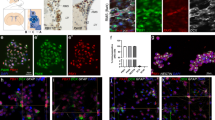
p53 acetyl K373 is induced in facial motor neurons following axonal transection. (a and b) Immunofluorescence displays the expression of p53 acetyl K373 (a) and CBP (b) in FMN (NeuN-positive cells) 48 h following facial nerve axotomy in mice. Increased expression levels are present in the transected (trans) side compared with the contralateral control (ctr) side. Arrows indicate the expression in specific FMNs. Scale bar: 40 μm. (c) Quantitation of pixel intensity shows a significant increase of p53 acetyl K373 and CBP expression in the transected versus the control side (average intensity of multiple sections of the facial nucleus per mice in three different mice). Asterisks: unpaired two-tailed t-test with a P-value<0.01. (d) Nuclear or (e) cytoplasmic extracts of dissected FMN were employed for immunoblotting for p53 acetyl K373 and CBP (d) and GAP-43 (e) 48 h after facial axotomy. β III tubulin and β-actin were used as loading control. Results show an increase in the expression of p53 acetyl K373 and CBP following transection (trans). Bar graphs are shown for quantitation (arbitrary units)
Taken together, these data demonstrate that modulation of GAP-43 transcription and expression is highly dependent upon p53-specific acetylation at Lysine 372, 3–82 through the CBP/p300 signaling.
Acetylated p53 promotes neuronal outgrowth in primary neurons
To test whether p53 acetylation at Lysine 372-3-82 does play a role in neuronal outgrowth in primary cortical neurons, we transfected rat primary cortical neurons with either one of the mutant p53 plasmid DNAs K373Q-GFP or K320Q-GFP for 72 h in comparison to the control eGFP vector. As expected, the p53 K320Q mutant was able to increase total process length (39.9%±9.24 S.D.), and the length of the longest process (30.1%±9.24) (Figure 4d) to a similar level as previously shown.26 However, here we show that the p53 K373Q mutant promotes the highest increase in process outgrowth compared with eGFP by driving an increase of 64.4%±11.3 in total process length and of 50.6%±5.9 in the length of the longest process (Figure 4d). In addition, p53 K373Q induces a significantly higher outgrowth in comparison to p53 K320Q (Figure 4d). The longest process corresponded almost invariably to axons as detected by GAP-43 and tau immunostaining (not shown). Importantly, we observed no increase in neuronal cell death following transfections with p53 acetylation mimics by using the MTT assay that accounts for cell viability in non-dividing cells such as neurons, and after cell counting and analysis of nuclear morphology (Supplementary Figure 5).
These data provide evidence that the C-terminal p53 acetylation promotes neurite and axon outgrowth, but does so preferentially via acetylation of p53 at position 372-3-82, which is also responsible for the activation of GAP-43 expression.
p53 acetyl K373 signaling shows enhanced expression and occupancy of GAP-43 promoter elements only in transected facial motor neurons following nerve cut
Facial nerve transection, a model of axon regeneration and target reinnervation34 allows for the study of the GAP-43-related molecular response to axonal injury. In fact, GAP-43 mRNA levels peak between 48 and 72 h post-axotomy in facial motor neurons (FMN) and contribute to axon regeneration.8, 35 Here, we have measured the expression of p53, p53 acetyl K320 and acetyl K373 as well as of CBP in FMN after facial axotomy. We have performed double immunofluorescence experiments for each of the above proteins and for the neuronal marker NeuN in cryosections throughout the whole facial nuclei pools, and quantified the expression levels by measurement of pixel intensity specifically in NeuN-positive motorneurons. Data analysis show that expression of p53 acetylated at K373 and CBP is significantly increased in the nucleus of FMN 48 h post-axotomy (Figure 5a–c). In addition, neurons expressing CBP and p53 acetyl K373 do not show apoptotic features (data not shown). Either no induction or weak nuclear localization was found for total p53 and p53 acetyl K320 in FMN following axotomy (Supplementary Figure 6). Immunoblotting was also performed in nuclear and cytoplasmic extracts of FMN 48 h post-axotomy, and showed increased expression of p53 acetyl K373, CBP and GAP-43 in the axotomized FMN as compared with the contralateral control side (Figure 5d and e).
Then, we performed ChIP assays in facial motor nuclei following nerve transection (see Materials and methods and Supplementary Figure 7 for visualization of the procedure), which showed that p53 acetyl K373 has a significantly enhanced occupancy of the GAP-43 promoter in axotomized FMN (Figure 6).
Facial axotomy promotes GAP-43 promoter occupancy by p53 acetyl K373. ChIP–real time PCR shows the in vivo GAP-43 promoter occupancy by p53 acetyl K373 from transected versus contralateral FMN. Briefly, in vivo crosslinked FMN were dissected 24 and 48 h post facial axotomy. Control contralateral and transected FMN were separately collected and processed for ChIP assay, which was followed by real-time PCR for quantitation of the promoter occupancy. Three mice per experiment were employed. On the left, ChIP results show the in vivo GAP-43 promoter occupancy by p53 acetyl K373 from transected versus contralateral FMN. Input samples, collected before precipitation and the no antibody samples, were used as a normalization control. Here, 3′end regions of the GAP-43 gene were used as a control of the specificity of the occupancy by p53 on GAP-43 promoter. On the right, the bar graph shows the fold increase (2.4 fold at 24 h and 2.8 fold at 48 h) in specific promoter occupancy by p53 acetyl K373 between axotomized and control contralateral FMN as analyzed by real-time quantitative PCR following ChIP. The input samples, collected before precipitation and the no antibody samples, were used as a normalization control. Asterisks indicate a significant difference analyzed by unpaired two-tailed t-test with a P-value <0.01, triplicate experiments
Taken together, these data suggest the presence of a direct transcriptional regulation of the pro-regeneration factor GAP-43 by CBP/p300-dependent acetylated p53 K373 in an in vivo model of nerve regeneration.
p53/GAP-43 transcriptional module is required during axon regeneration in vivo
To assess whether GAP-43 expression is dependent upon p53 during axon regeneration in vivo, we compared the levels of GAP-43 expression and the extent of nerve regeneration in p53 null versus wt mice after facial nerve axotomy. The extent of regeneration was assessed by counting FMN marked by a retrograde nerve tracer FluoroGold 28 days after injury, which corresponds to regenerating fibers. In wt mice the neuronal regeneration percentage was 29.7%±2.6 (S.E.M.), significant in comparison to 10.9%±2.9 (S.E.M.) in the p53 null mice (Figure 7a). Nissl-stained FMN counted at 28 days post-axotomy revealed fewer surviving motor neurons on the transected side compared with the control side in wt at 64.8%±7 (S.E.M.) versus p53 null mice at 76.2%±8 (S.E.M.) (Figure 7a). This further supports the role of p53 in the elimination of highly damaged cells as well as in axon regeneration of surviving neurons, as reported earlier.26
p53 is required for GAP-43 expression in axonal sprouts during axonal regeneration. (a) As a marker of regeneration, FluoroGold retrogradely positive FMN are shown 28 days after facial nerve transection. Shown is a significant reduction in the number of FluoroGold-positive neurons in p53−/− compared with p53 wt after nerve transection (trans). The first bar graph shows the quantitation of regenerating facial motorneurons 28 days after facial axotomy. Shown is the overall ratio of labeled neurons in the operated/unoperated side after sectioning of the entire facial nucleus (five mice per group, Asterisks: unpaired two-tailed t-test with a P-value<0.01). The second bar graph shows the quantitation of surviving neurons 28 days following facial transection in wt versus p53−/− mice. Shown is the overall ratio of surviving facial motorneurons in the operated/unoperated side (five mice per group). Scale bar: 100 μm. (b) Real-time RT-PCR experiments show a significant increase in mRNA expression for GAP-43 in the FMN of wt but not p53−/− mice in axotomized versus contralateral side 28 days post-axotomy. Asterisks indicate a significant difference analyzed by unpaired two-tailed t-test with a P-value <0.05, triplicate experiments. (c) Immunofluorescence displays the expression of GAP-43 in axonal sprouts 28 days following axotomy in wt and p53−/− mice. Typical GAP-43 immunolocalization in FMN in the cytoplasm (arrows) and in axonal sprouts (arrowheads). Quantitation of GAP-43 pixel intensity shows a significant increase in wt versus p53−/− mice (Asterisks: unpaired two-tailed t-test with a P-value<0.01). Scale bar: 50 μm. (d) A schematic diagram summarizes our novel findings. The transcriptional complex formed by p53 acetylated at K372, 3-82 and the acetyltransferases CBP/p300 is recruited to the GAP-43 promoter during development and following injury, and drives GAP-43 expression, thus leading to axon outgrowth
The analysis of GAP-43 mRNA expression by qRT-PCR shows a significant increase (48%) in FMN in the transected versus the contralateral side at 28 days post-axotomy, when regeneration occurs, only in the wt, but not in the p53 null mice (Figure 7b). Strikingly, immunofluorescence experiments in the brain stem 28 days post-axotomy also show an induction of GAP-43 expression in FMN post-axotomy, which is significantly impaired in p53 null mice (Figure 7c). GAP-43 expression levels are in fact not significantly different in the unlesioned side in wt and p53 null mice. In contrast, GAP-43 expression is induced in both the cell bodies and sprouting axons following axotomy during regeneration in the wt, but not in the p53 null mice (Figure 7c).
In summary, we show that direct p53-dependent transcriptional regulation and protein expression of GAP-43 are required and present only during FMN axon regeneration.
Discussion
This work identifies a novel transcriptional module for axon outgrowth and regeneration formed by CBP/p300 and acetylated p53 on the GAP-43 promoter (a schematic summarizes our findings, Figure 7d).
Our novel findings show that p53 drives the expression of neuronal GAP-43 and binds to specific promoter elements in neurons in a chromatin environment. The p53 binding site described here lies within the first 1000 bp (619–639) upstream from the protein-coding sequence, a promoter region considered important for GAP-43 neuronal gene expression.36 This GAP-43-p53-binding site is novel and its core binding element aCATGt and two flanking nucleotides (low case) contain a classical p53 consensus binding site,37 which is conserved in mouse, rat and human.
The diversity and specificity of p53 activity is regulated by post-translational modifications, that can direct p53 transcriptional activity toward specific promoters, thereby activating different transcriptional cascades.38 It is important to note that pro-apoptotic neuronal p53 is frequently phosphorylated through cascades involving MAPKs, JNKs and p38.39, 40, 41
In contrast, we have recently demonstrated that a mutant p53, which mimics acetylation at K320, is involved in the promotion of neurite outgrowth and in the regulation of the expression of the pro-outgrowth factors Coronin 1b and Rab13.26 These data along with the increasing evidence that an enriched acetylated environment protects neurons from cell death in vitro and in vivo,29 has prompted us to investigate the yet undiscovered role of endogenous acetylation of p53 in primary neurons on the activation of the pro-axon growth gene GAP-43.
Histone acetyltransferases (HATs) acetylate many proteins important for the initiation of transcription, such as nucleosomal histones and transcription factors (TFs). HATs modify core histone tails by post-translational acetylation of specific Lysine residues, inducing chromatin modification and enhanced DNA accessibility to TFs.42 Acetylation of p53 by CBP/p300 and P/CAF enhances its protein stability and its transactivation potential by facilitating interactions with specific promoters and other proteins of the transcriptional machinery.33, 43
Here, we show that GAP-43 promoter elements are preferentially occupied by acetylated p53 in cortical neurons and in vivo in facial motor neurons following axotomy through CBP/p300 signaling, which is responsible for maximal GAP-43 transcriptional activation.
Both mutant p53 that mimic acetylation by endogenous CBP/p300 and P/CAF respectively at K372-3, 82 and K320, drive neurite outgrowth in primary cortical neurons and do not induce cell death. However, CBP/p300-dependent p53 acetylation pattern induces the highest increase in neurite outgrowth in primary neurons.
A specific role for CBP/p300 acetylated p53 in axon outgrowth was also found following axotomy. In fact, we show that expression of p53 K373 and CBP is induced in FMN in the brain stem following facial nerve axotomy, a model of axon regeneration. In transected FMN, p53 K373 preferentially occupies the GAP-43 promoter as shown by in vivo ChIP, at a time that coincides with the well-known upregulation of GAP-43 transcript and protein,8, 35 whose expression we demonstrate to be dependent upon p53 following axotomy.
Importantly, our data demonstrate that C-terminal acetylated p53 induces pro-axon growth promoters and does not trigger cell death in primary neurons. In contrast, acetylation of p53 in cancer cell lines and models promotes cell cycle arrest and DNA repair as well as apoptosis.33, 44 This supports the concept that similar p53 post-translational patterns serve distinct functions in diverse cell types and signaling pathways, and shows for the first time that in neurons acetylation may be not only neuroprotective, but also pro-axon growth.
Post-translational modifications not only affect DNA–protein interactions but also influence protein–protein interaction events within transcriptional complexes. Therefore, the pro-axon growth properties of CBP–p53 signaling might cooperate with the CREB-dependent transcription pathway. In fact, CREB was recently shown to protect neurons from cell death and to promote neurite outgrowth by inducing arginase I expression.45
Therefore, future challenges include the identification of additional transcriptional machinery that participates at pro-axonal growth and regeneration promoters. In addition, it will be paramount to establish the effectiveness of different transcription factors on pro-growth promoters, as well as identifying a set of target genes that drive cytoskeleton remodeling and axon growth.
In conclusion, our work defines a novel role for CBP/p300-p53-GAP-43 transcriptional module in axon growth and regeneration that, ultimately, may offer a novel molecular target to foster axon re-growth in neurodegenerative conditions.
Materials and Methods
Cell cultures
See online Supplementary Materials.
In silico transcription factor-binding sites analysis
The entire promoter sequence of the selected gene GAP-43 was analyzed using MatInspector, a software tool provided by Genomatix Software that identifies TFBS in nucleotide sequences using a large library of weight matrices.
Chromatin immunoprecipitation assays
ChIP assays were performed according to the manufacturer's recommendations (Upstate). Experiments were done in RCN and PC-12 cells (at least 4 × 106 cells were used for each assay) and in facial motor neurons (FMN). PC-12 cells were grown in the absence or presence of NGF (100 ng/ml). Cultured cells were fixed in a 1% formaldehyde-PBS solution. Following cell lysis (0.5% SDS, 100 mM NaCl, 50 mM Tris-HCl, pH 8.0, 5 mM EDTA), extracts were sonicated to shear DNA to lengths of 200–400 bp. Also, in a set of experiments, transfection of full length p300 and CBP plasmid DNA expressed from pcDNA vector or siRNA for p300 and CBP were performed before fixing the cells. Chromatin solutions were incubated overnight with rotation using 5 μg of rabbit polyclonal anti-p53 antibody (FL-393, Santa Cruz, USA), rabbit polyclonal anti-p53 acetyl Lys320 (06–915, Upstate) or rabbit polyclonal anti p53 acetyl Lys373 (06–916, Upstate). The following day protein A agarose beads, that had been blocked with salmon sperm DNA, were added to each reaction to precipitate antibody complexes. The precipitated complexes were washed and then incubated for 4 h at 65 °C in parallel with input samples to reverse the crosslink. DNA was isolated by P:C:I extraction, which was followed by ethanol precipitation in the presence of sodium acetate.
Input, IP and no-Ab fractions were then analyzed by PCR and quantitative PCR (ABI 7000) analysis with appropriate primer pairs to amplify p53 responsive elements. The primers used were as follows: rat WAF1/p21 5′ site Forward 5′-TTGAGGGAAAGATCTCCAGAAAA-3′ rat WAF1/p21 5′ site Reverse 5′-CCAGATGGAGACCAGCTTA-3′; rat WAF1/p21 3′ site Forward 5′-GCCTTTGCAGAGAACATTCCT-3′ rat WAF1/p21 3′-site Reverse 5′-AGCCTAGGGCGTGGACTA-3′; rat GAP-43 5′ site Forward 5′-GCAGCTGTAACTTGTGTGCA-3′ rat GAP-43 5′ site Reverse 3′-GGTCCAGATTGGAGGTGTTTA-5′; rat GAP-43 3′ site Forward 5′-TTCCTTAGGCAATGTTTTGGAAAG-3′ rat GAP-43 3′ site Reverse 5′-TCAGGCATGTTCTTGGTCAG-3′; mouse GAP-43 3′ site Forward 5′-ACAATAGCTTGCTGGGGATG-3′ mouse GAP-43 3′ site Reverse 5′-CCATAGCAGAAAGCCTCCAG-3′.
For real-time quantitation of PCR products and fold change measurements after ChIP, we have first calculated the presence of enrichment in our p53 IP assays on GAP-promoter and the control 3′end regions as compared with input DNA by real-time PCR. All samples using primers on GAP-43 promoter regions presented enrichment as expected, whereas no signal was detected in the 3′end sequences. Then, each experimental sample was normalized to input and no-Ab fractions and presented as fold change between individual experimental conditions throughout our results.
Facial nerve transection-ChIP real-time PCR
Facial nerve transection was performed as described above in male B6. On day one and two following facial nerve transection, three mice per time point were deeply anesthetized and perfused with 0.9% saline solution followed by 4% paraformaldehyde in 0.1 M phosphate buffer. The brain stem was then removed and placed in 4% paraformaldehyde at 4 °C for 10 h. Here, 50-μm thick cross serial sections were cut using the vibratome and Nissl stain was performed to identify slices containing FMN. Subsequently, FMN were finely dissected from the axotomized and the contralateral side from serial sections of the brain stem spanning the whole facial nuclei. Dissected FMN from the contralateral and axotomized sides were collected in different vials and homogenized with an Ultra Turrax Disperser (IKA-Werke, Staufen, Germany) in lysis buffer (0.5% SDS/100 mM NaCl/50 mM Tris–HCl, pH 8/5 mM EDTA) containing protease inhibitors. Extracts were sonicated to shear DNA. Chromatin solutions were precipitated as described earlier in the chromatin immunoprecipitation assays section. An antibody against p53 acetyl K373 was employed. ChIP was followed by PCRs, which were performed as described above. ChIP reactions from contralateral and axotomized FMN were compared to evaluate the degree of GAP-43 promoter occupancy by using real-time PCR. Each experimental sample was normalized to input and no-Ab fractions. Fold change measurements after ChIP included three independent PCRs and three individual samples and were performed according to real-time PCR standard procedures.
Luciferase assays
For luciferases assays, the region of the GAP-43 promoter containing the putative p53-binding site identified by in silico analysis and the scrambled DNA control site, were generated. For further details, see online Supplementary Information.
Real-time RT-PCR analysis
RNA was extracted from wt, p53DN, p53-silenced, CBP/p300-silenced and CBP/p300-overexpressing PC-12 cells and from both wt and p53−/− mouse brain using TRIZOL reagent (Invitrogen) and cDNA was synthesized from 1 μg of RNA using random hexamers from the SuperScript II Reverse Transcriptase kit (Invitrogen). See online Supplementary Information.
Clones and transfection/infection experiments
See online Supplementary Information.
RNA interference
See online Supplementary Information.
Antibodies, immunoblotting and immunocytochemistry
See online Supplementary Information.
Neurite outgrowth assay
See online Supplementary Information.
p53 null mice and facial nerve transection
Homozygous p53−/− FVB.129S2 (B6)-Trp53tmltyj (Jackson Laboratories), and FVB.129S2 (B6) control mice were used for our experiments. The genotype of p53 null mice was confirmed by Southern blot analysis. Our experiments were performed starting at 6 weeks of age in male homozygous p53 null and wt mice (N=5). Facial nerve transection was performed as described earlier.26 Briefly, mice were anesthetized by an intramuscular injection of a combination of xylazine (13 mg/kg) and ketamine HCl (85 mg/kg). A skin incision was made behind the left ear and the facial nerve exposed, and transected about 2 mm posterior to the foramen stylomastoideum. The same incisions without transection were also made on the contralateral side for sham-operated intra-animal controls. The wounds were sutured, and the animals were allowed to recover in a darkened room on isothermal pads warmed at 37 °C for approximately 30 min.
Regeneration in facial nuclei in the brain stem following facial nerve transection was evaluated at day 28 by retrograde axonal tracing with FluoroGold. Three days before being killed, mice from each group were anesthetized and 1.5 μl of 3% FluoroGold was injected through a micropipette with a tip diameter of 25 μm into the whisker pads bilaterally. After 3 days, the mice were deeply anesthetized and perfused with 0.9% saline solution followed by 4% paraformaldehyde in 0.1 M phosphate buffer. The brain stem was removed and placed in 30% sucrose for 24 h. The entire facial nuclei were sectioned and reconstructed and all sections of both facial nuclei per each animal were evaluated (235 for the control and 225 for the transected side). Transverse serial frozen sections, 20-μm thick, were cut and mounted on slides and photographed under a fluorescence microscope. Only labeled neurons with visible nuclei were counted and taken to represent the number of regenerated motor neurons. For analysis, retrograde-labelled regenerating motor neurons were divided by total cells stained with cresyl violet, and this number was normalized to the uninjured (contralateral) side in each animal for intra-animal control. Thus, the overall ratio of FluoroGold-positive neurons was compared between p53 null and wt animals. The Abercrombie formula for cell counting was included in our analysis to correct for tissue volume.
Immunofluorescence (IF) for p53, p53 acetyl K320, p53 acetyl K373 and CBP was performed in 35 μm thick brain stem sections spanning the whole facial nucleus 48 h following axotomy in three wt mice. Evaluation of the intensity of the fluorescence was performed in multiple sections per mouse to measure the expression levels in FMN for each protein in the axotomized transected side versus the control contralateral side. Intensity of IF in FMN was calculated by using the AlphaEaseFC software and intensity values for each protein were normalized to the intensity of NeuN expression in FMN and compared between the control and transected side.
Abbreviations
- C/EBP:
-
CCAAT-enhancer-binding proteins
- CBP:
-
CREB-binding protein
- CREB:
-
cAMP-responsive element-binding protein
- JNKs:
-
c-Jun N-terminal kinases
- MAPK:
-
mitogen-activated protein kinases
- NFAT:
-
nuclear factor of activated T-cells
- P/CAF:
-
p300/CBP-associated factor
- STAT3:
-
signal transducer and activator of transcription 3
References
Makwana M, Raivich G . Molecular mechanisms in successful peripheral regeneration. FEBS J 2005; 272: 2628–2638.
Teng FY, Tang BL . Axonal regeneration in adult CNS neurons--signaling molecules and pathways. J Neurochem 2006; 96: 1501–1508.
Laux T, Fukami K, Thelen M, Golub T, Frey D, Caroni P . GAP43, MARCKS, and CAP23 modulate PI(4,5)P(2) at plasmalemmal rafts, and regulate cell cortex actin dynamics through a common mechanism. J Cell Biol 2000; 149: 1455–1472.
Perrone-Bizzozero NI, Weiner D, Hauser G, Benowitz LI . Extraction of major acidic Ca2+ dependent phosphoproteins from synaptic membranes. J Neurosci Res 1988; 20: 346–350.
De la Monte SM, Federoff HJ, Ng SC, Grabczyk E, Fishman MC . GAP-43 gene expression during development: persistence in a distinctive set of neurons in the mature central nervous system. Brain Res Dev Brain Res 1989; 46: 161–168.
Jacobson RD, Virag I, Skene JH . A protein associated with axon growth, GAP-43, is widely distributed and developmentally regulated in rat CNS. J Neurosci 1986; 6: 1843–1855.
Benowitz LI, Perrone-Bizzozero NI, Neve RL, Rodriguez W . GAP-43 as a marker for structural plasticity in the mature CNS. Prog Brain Res 1990; 86: 309–320.
Tetzlaff W, Alexander SW, Miller FD, Bisby MA . Response of facial and rubrospinal neurons to axotomy: changes in mRNA expression for cytoskeletal proteins and GAP-43. J Neurosci 1991; 11: 2528–2544.
Tetzlaff W, Zwiers H, Lederis K, Cassar L, Bisby MA . Axonal transport and localization of B-50/GAP-43-like immunoreactivity in regenerating sciatic and facial nerves of the rat. J Neurosci 1989; 9: 1303–1313.
Hulsebosch CE, DeWitt DS, Jenkins LW, Prough DS . Traumatic brain injury in rats results in increased expression of Gap-43 that correlates with behavioral recovery. Neurosci Lett 1998; 255: 83–86.
Carmichael ST . Plasticity of cortical projections after stroke. Neuroscientist 2003; 9: 64–75.
Gianola S, Rossi F . GAP-43 overexpression in adult mouse Purkinje cells overrides myelin-derived inhibition of neurite growth. Eur J Neurosci 2004; 19: 819–830.
Aigner L, Arber S, Kapfhammer JP, Laux T, Schneider C, Botteri F et al. Overexpression of the neural growth-associated protein GAP-43 induces nerve sprouting in the adult nervous system of transgenic mice. Cell 1995; 83: 269–278.
Aigner L, Caroni P . Depletion of 43-kD growth-associated protein in primary sensory neurons leads to diminished formation and spreading of growth cones. J Cell Biol 1993; 123: 417–429.
Alonso G, Ridet JL, Oestreicher AB, Gispen WH, Privat A . B-50 (GAP-43) immunoreactivity is rarely detected within intact catecholaminergic and serotonergic axons innervating the brain and spinal cord of the adult rat, but is associated with these axons following lesion. Exp neurol 1995; 134: 35–48.
Bomze HM, Bulsara KR, Iskandar BJ, Caroni P, Skene JH . Spinal axon regeneration evoked by replacing two growth cone proteins in adult neurons. Nat Neurosci 2001; 4: 38–43.
Beckel-Mitchener AC, Miera A, Keller R, Perrone-Bizzozero NI . Poly(A) tail length-dependent stabilization of GAP-43 mRNA by the RNA-binding protein HuD. J Biol Chem 2002; 277: 27996–28002.
Chung S, Eckrich M, Perrone-Bizzozero N, Kohn DT, Furneaux H . The Elav-like proteins bind to a conserved regulatory element in the 3′-untranslated region of GAP-43 mRNA. J Biol Chem 1997; 272: 6593–6598.
Bolognani F, Tanner DC, Merhege M, Deschenes-Furry J, Jasmin B, Perrone-Bizzozero NI . In vivo post-transcriptional regulation of GAP-43 mRNA by overexpression of the RNA-binding protein HuD. J Neurochem 2006; 96: 790–801.
Bolognani F, Tanner DC, Nixon S, Okano HJ, Okano H, Perrone-Bizzozero NI . Coordinated expression of HuD and GAP-43 in hippocampal dentate granule cells during developmental and adult plasticity. Neurochem Res 2007; 32: 2142–2151.
Smith CL, Afroz R, Bassell GJ, Furneaux HM, Perrone-Bizzozero NI, Burry RW . GAP-43 mRNA in growth cones is associated with HuD and ribosomes. J Neurobiol 2004; 61: 222–235.
Nedivi E, Basi GS, Akey IV, Skene JH . A neural-specific GAP-43 core promoter located between unusual DNA elements that interact to regulate its activity. J Neurosci 1992; 12: 691–704.
Starr RG, Lu B, Federoff HJ . Functional characterization of the rat GAP-43 promoter. Brain Res 1994; 638: 211–220.
Weber JR, Skene JH . The activity of a highly promiscuous AP-1 element can be confined to neurons by a tissue-selective repressive element. J Neurosci 1998; 18: 5264–5274.
Uittenbogaard M, Martinka DL, Chiaramello A . The basic helix-loop-helix differentiation factor Nex1/MATH-2 functions as a key activator of the GAP-43 gene. J Neurochem 2003; 84: 678–688.
Di Giovanni S, Knights CD, Rao M, Yakovlev A, Beers J, Catania J et al. The tumor suppressor protein p53 is required for neurite outgrowth and axon regeneration. EMBO J 2006; 25: 4084–4096.
Di Giovanni S, De Biase A, Yakovlev A, Finn T, Beers J, Hoffman EP et al. In vivo and in vitro characterization of novel neuronal plasticity factors identified following spinal cord injury. J Biol Chem 2005; 280: 2084–2091.
Yakovlev AG, Di Giovanni S, Wang G, Liu W, Stoica B, Faden AI . BOK and NOXA are essential mediators of p53-dependent apoptosis. J Biol Chem 2004; 279: 28367–28374.
Saha RN, Pahan K . HATs and HDACs in neurodegeneration: a tale of disconcerted acetylation homeostasis. Cell Death Differ 2006; 13: 539–550.
Liu L, Scolnick DM, Trievel RC, Zhang HB, Marmorstein R, Halazonetis TD et al. p53 sites acetylated in vitro by PCAF and p300 are acetylated in vivo in response to DNA damage. Mol Cell Biol 1999; 19: 1202–1209.
Avantaggiati ML, Ogryzko V, Gardner K, Giordano A, Levine AS, Kelly K . Recruitment of p300/CBP in p53-dependent signal pathways. Cell 1997; 89: 1175–1184.
Lill NL, Grossman SR, Ginsberg D, DeCaprio J, Livingston DM . Binding and modulation of p53 by p300/CBP coactivators. Nature 1997; 387: 823–827.
Knights CD, Catania J, Di Giovanni S, Muratoglu S, Perez R, Swartzbeck A et al. Distinct p53 acetylation cassettes differentially influence gene-expression patterns and cell fate. J Cell Biol 2006; 173: 533–544.
Raivich G, Bohatschek M, Da Costa C, Iwata O, Galiano M, Hristova M et al. The AP-1 transcription factor c-Jun is required for efficient axonal regeneration. Neuron 2004; 43: 57–67.
Namgung U, Routtenberg A . Transcriptional and post-transcriptional regulation of a brain growth protein: regional differentiation and regeneration induction of GAP-43. Eur J Neurosci 2000; 12: 3124–3136.
Takahashi M, Sato Y, Nakagami Y, Miyake K, Iijima S . Identification of cis-acting regions that contribute to neuron-specific expression of the GAP-43 gene. Biosci Biotechnol Biochem 2006; 70: 1492–1495.
Hoh J, Jin S, Parrado T, Edington J, Levine AJ, Ott J . The p53MH algorithm and its application in detecting p53-responsive genes. Proc Natl Acad Sci USA 2002; 99: 8467–8472.
Xu Y . Regulation of p53 responses by post-translational modifications. Cell Death Differ 2003; 10: 400–403.
Song J, Chao C, Xu Y . Ser18 and Ser23 phosphorylation plays synergistic roles in activating p53-dependent neuronal apoptosis. Cell Cycle 2007; 6: 1412–1414.
Zhu Y, Mao XO, Sun Y, Xia Z, Greenberg DA . p38 Mitogen-activated protein kinase mediates hypoxic regulation of Mdm2 and p53 in neurons. J Biol Chem 2002; 277: 22909–22914.
Chen RW, Qin ZH, Ren M, Kanai H, Chalecka-Franaszek E, Leeds P et al. Regulation of c-Jun N-terminal kinase, p38 kinase and AP-1 DNA binding in cultured brain neurons: roles in glutamate excitotoxicity and lithium neuroprotection. J Neurochem 2003; 84: 566–575.
Glozak MA, Sengupta N, Zhang X, Seto E . Acetylation and deacetylation of non-histone proteins. Gene 2005; 363: 15–23.
Luo J, Li M, Tang Y, Laszkowska M, Roeder RG, Gu W . Acetylation of p53 augments its site-specific DNA binding both in vitro and in vivo. Proc Natl Acad Sci USA 2004; 101: 2259–2264.
Gu W, Roeder RG . Activation of p53 sequence-specific DNA binding by acetylation of the p53 C-terminal domain. Cell 1997; 90: 595–606.
Gao Y, Deng K, Hou J, Bryson JB, Barco A, Nikulina E et al. Activated CREB is sufficient to overcome inhibitors in myelin and promote spinal axon regeneration in vivo. Neuron 2004; 44: 609–621.
Acknowledgements
We thank Jorge Garay for conducting part of the facial nerve transection experiments, Sergio Schinelli for critically reviewing the paper, and Sanam Vakil for her help editing the paper. In addition, we are grateful to Dr. Maria Laura Avantaggiati for providing the p53 acetylation mutant plasmid DNAs, to Dr. Ulrike Neumann for the p53 R273H and R248W mutant plasmid DNAs, and to Dr. Katsuhide Miyake for providing the CBP/p300 RNAi. This work was supported by the Hertie Foundation; the Fortune Grant, University of Tuebingen, the NIH R21 NS052640 and the DFG DI 1497/1-1 grants (all granted to Simone Di Giovanni).
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by D Kaplan
Supplementary Information accompanies the paper on Cell Death and Differentiation website (http://www.nature.com/cdd)
Rights and permissions
About this article
Cite this article
Tedeschi, A., Nguyen, T., Puttagunta, R. et al. A p53-CBP/p300 transcription module is required for GAP-43 expression, axon outgrowth, and regeneration. Cell Death Differ 16, 543–554 (2009). https://doi.org/10.1038/cdd.2008.175
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2008.175