Abstract
Background:
When tumour tissue is unavailable, cell-free DNA (cfDNA)can serve as a surrogate for genetic analyses. Because mutated alleles in cfDNA are usually below 1%, next-generation sequencing (NGS)must be narrowed to target only clinically relevant genes. In this proof-of-concept study, we developed a panel to use in ultra-deep sequencing to identify such mutations in cfDNA.
Methods:
Our panel (‘SiRe’) covers 568 mutations in six genes (EGFR, KRAS, NRAS, BRAF, cKIT and PDGFRα)involved in non-small-cell lung cancer (NSCLC), gastrointestinal stromal tumour, colorectal carcinoma and melanoma. We evaluated the panel performance in three steps. First, we analysed its analytical sensitivity on cell line DNA and by using an artificial reference standard with multiple mutations in different genes. Second, we analysed cfDNA from cancer patients at presentation (n=42), treatment response (n=12) and tumour progression (n=11); all patients had paired tumour tissue and cfDNA previously genotyped with a Taqman-derived assay (TDA). Third, we tested blood samples prospectively collected from NSCLC patients (n=79) to assess the performance of SiRe in clinical practice.
Results:
SiRe had a high analytical performance and a 0.01% lower limit of detection. In the retrospective series, SiRe detected 40 EGFR, 11 KRAS, 1 NRAS and 5 BRAF mutations (96.8% concordance with TDA). In the baseline samples, SiRe had 100% specificity and 79% sensitivity relative to tumour tissue. Finally, in the prospective series, SiRe detected 8.7% (4/46) of EGFR mutations at baseline and 42.9% (9/21) of EGFR p.T790M in patients at tumour progression.
Conclusions:
SiRe is a feasible NGS panel for cfDNA analysis in clinical practice.
Similar content being viewed by others
Main
Precision medicine, coupled with the tissue-based assessment of biomarkers predictive of treatment outcome, has transformed pathology practice (Papadopoulos et al, 2006). RAS and BRAF mutation testing in colorectal cancer (CRC; Di Nicolantonio et al, 2008; Lièvre et al, 2008), EGFR in non-small-cell lung cancer (NSCLC; Lynch et al, 2004) BRAF in melanoma (Chapman et al, 2011) and cKIT and PDGFRα in gastrointestinal stromal tumours (GIST; Antonescu, 2008) has added a genotypic element to the phenotypic diagnostics of solid tumours. However, tumour tissue is not always available or may be insufficient for molecular testing, especially when cancer is diagnosed at advanced stages on small biopsy specimens. On other occasions, due to tumour location or small size, tissue sampling can be challenging and risky, particularly in extensively treated patients. As an alternative to cancer tissue, predictive biomarkers can be non-invasively assessed in cell-free DNA (cfDNA; Schwarzenbach et al, 2011; Crowley et al, 2013).
Using a Taqman-derived assay (TDA) we previously identified EGFR mutations in NSCLC (Karachaliou et al, 2015) and BRAF mutations in melanoma patients (Gonzalez-Cao et al, 2015) with a specificity of 100% and with sensitivities of 69% and 78%, respectively. One of the factors contributing to this high sensitivity was the concomitant analysis, in each patient, of serum- and plasma-derived cfDNA (Karachaliou et al, 2015; Gonzalez-Cao et al, 2015). This performance may be further improved by next-generation sequencing (NGS), which can be multiplexed across several genes to cover less common and even novel variants (Malapelle et al, 2016a). Large gene panels or whole-exome approaches to screen for a large number of genomic regions may not be easily implemented in cfDNA analysis (Cancer Genome Atlas Research Network, 2014). Conversely, small NGS panels tailored to target a limited number of actionable genes can be an effective tool in daily clinical practice (Paweletz et al, 2016). This strategy, known as ‘ultra-deep sequencing’, can significantly increase sensitivity, which is essential considering that circulating tumour DNA represents only a small fraction (<0.5%) of the total cfDNA (Mead et al, 2011) in most patients with solid tumours. Since the low threshold levels of mutant alleles required to detect clinically relevant alterations may easily lead to false-positive results (van Dijk et al, 2014), implementation of the ultra-deep sequencing of cfDNA in the clinical setting must be validated in terms of blood collection, cfDNA extraction, automated library preparation, sequencing and variant calling (Gargis et al, 2012; Malapelle et al, 2016c).
In this proof-of-concept study, we report the development, performance evaluation and implementation in a clinical setting of a narrow gene panel that targets 568 clinically relevant mutations in six genes (EGFR, KRAS, NRAS, BRAF, cKIT and PDGFRα) involved in non Small cell lung cancer, gastroIntestinal stromal tumour, metastatic coloRectal carcinoma and mElanoma (whose acronym is SiRe). This panel has a high sensitivity and specificity and enables the detection and quantification of mutations in cfDNA purified from the plasma and serum of patients with different types of solid tumours.
Materials and methods
Design of the SiRe panel
The Ion AmpliSeq Designer suite v5.3.1 with hg19 was used as reference genome to develop a customised panel targeting six genes (EGFR, KRAS, NRAS, BRAF, cKIT and PDGFRα) that are associated with treatment outcome in NSCLC, GIST, CRC and metastatic melanoma (Lynch et al, 2004; Antonescu, 2008; Di Nicolantonio et al, 2008; Lièvre et al, 2008; Chapman et al, 2011). A single primer pool leading to the selection of 42 amplicons (ranging from 125 to 175 bp) enabled us to cover all COSMIC annotated mutations (n=568) in the selected exons of the target genes. The complete reference range of SiRe is reported in Supplementary Material (Supplementary Table S1). The amplicon design (available on request) covering 5.2 kb of genomic DNA was optimised for the simultaneous analysis of 16 samples with the 316v2 chip (Thermofisher, Foster City, CA, USA) on a Personal Genome Machine Torrent (Thermofisher).
Study design, patients and samples
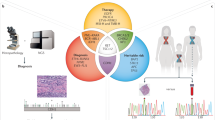
The panel performance was evaluated in three steps (Figure 1). First, the analytical sensitivity of the assay was assessed on DNA from two cell lines and by using an artificial reference standard with multiple mutations in different genes. Second, clinical sensitivity and specificity was determined using archival cfDNA from 63 cancer patients (Table 1) with paired tumour tissue, previously genotyped with a TDA. As exploratory analysis, to confirm that our NGS approach cover the mutations in cKit and PDGFRα genes, two GIST samples (bloods and tissues) were tested with SiRe and the relative data are reported only in Supplementary Material. Third, the performance of the panel in daily clinical practice was assessed using blood samples prospectively collected from patients with advanced NSCLC. Written informed consent was obtained from all patients and documented in accordance with the general authorisation to process personal data for scientific research purposes from ‘The Italian Data Protection Authority’ (http://www.garanteprivacy.it/web/guest/home/docweb/-/docwebdisplay/export/2485392). All information regarding human material was managed using anonymous numerical codes, and all samples were handled in compliance with the Helsinki Declaration (http://www.wma.net/en/30publications/10policies/b3/).
Study design.cfDNAs (A) extracted with the QIAsymphony virus/pathogen kit (B) from paired (P) plasma and (S) serum (C) samples were analysed by quantitative 5′-nuclease TaqMan PCR (D) and by the NGS SiRe panel (E). Any discordance between the two techniques was evaluated by dPCR (F). After preclinical validation, the SiRe panel was applied in clinical practice in cases in which tissues were not available to select patients for TKI treatment, at baseline (G), and to evaluate the selection of resistant clones after disease progression (H).
DNA purification
DNA from the two cell lines was isolated using the QIAamp Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. Circulating-free DNA was purified as follows: 15 ml blood was withdrawn from patients and collected in Vacutainer tubes (BD, Plymouth, UK). Plasma and serum were isolated by centrifugation twice at 2300 r.p.m. for 10 min. The supernatant (serum or plasma) was aliquoted and used immediately for cfDNA isolation or stored at −80 °C. Cell-free DNA was purified from serum and plasma for each patient (1.2 ml). In the rare instances that the volume of the serum and plasma sample obtained from a patient was between 1 and 1.2 ml, PBS up to 1.2 ml was added to the samples, which were then purified using the QIAsymphony robot (Qiagen) and the QIAsymphony DSPVirus/Pathogen Midi Kit, according to the manufacturer’s instructions, and cfDNA was eluted in a final volume of 30 μl. Since correct preanalytical handling of blood specimens is crucial to maintain the sample informative, the process was standardised (in terms of blood collection, sample centrifugation and cfDNA extraction) in the Department of Public Health of the University of Naples Federico II, and all procedures were performed in-house by a nurse belonging to the laboratory staff.
Sample sequencing
We analysed the serum and plasma cfDNAs of each patient enrolled in the study. Libraries were constructed and purified on the Ion Chef (Thermofisher), and eight samples (corresponding to 4 patients) were added per run. Library generation was as follows: 6 μl of cfDNA were dispensed on Ion Code plates and amplified using Ion AmpliSeq DL8 (Thermofisher). We used 22 cycles for cfDNA amplification and 6 cycles for library reamplification after barcoding, under the thermal conditions defined by the manufacturer. Purified libraries derived from eight cfDNA samples were diluted to 60 pM and combined with eight additional cfDNA-derived libraries to obtain a 16 Ion Code pooled library. The two-pooled libraries were re-loaded into the Ion Chef instrument, and templates were prepared using the Ion PGM Hi-Q IC Kit (Thermofisher). Finally, templates were loaded into the 316v2 chip and sequenced on PGM.
Data analysis
Signal processing and base calling were carried out using the default base-caller parameters on Torrent Suite [v.5.0.2] and coverage analysis was performed using SiRe designed bed files with coverage plug-in (v.5.0.2.0). BAM files were visually inspected with the Golden Helix Genome Browser v.2.0.7 (Bozeman,MT, USA). Variants were automatically annotated using variant caller plug-in (v.5.0.2.1) at specific optimised parameters of the SiRe panel (Supplementary Table S2). In particular, only variants with ⩾5X allele coverage and a quality score ⩾20, within an amplicon that covered at least 1000X alleles, were called, and the frequency of each mutant allele was recorded.
Preclinical assessment
Genomic DNA from the HCC827 (EGFR p.E746-A750del; KRAS wt) and A549 (EGFR wt; KRAS p.G12S) cell lines was used to assess analytical performance. Both cell lines were obtained from the National Research Council/Institute of Experimental Endocrinology and Oncology on courtesy of Dr Pierlorenzo Pallante (Naples, Italy). The analytical sensitivity of the assay for point mutation and indel detection was determined by diluting DNA from the appropriate mutated cell line (A549 for point mutations and HCC827 for indels) into increasing concentrations of DNA from the appropriate wt cell line (HCC827 for point mutations and A549 for indels). DNA dilutions ranged between 1 : 10 and 1 : 10 000, which correspond to allelic fractions from 1 : 20 to 1 : 20 000 of the mutated allele (both cell lines are heterozygous). Each dilution was analysed in duplicate to estimate inter-run assay reproducibility, and the library obtained from each dilution was sequenced twice to evaluate intra-run assay reproducibility. In addition, customised Horizon Diagnostics Multiplex gDNA reference standard, with mutation in EGFR (p.E746_A750del and p.G719S), KRAS (p.G12D), NRAS (p.Q61L) and BRAF (p.V600E), each of them at three different dilution points (1, 0.5 and 0.1%), were assessed to provide stronger evidence on SiRe analytical performance.
Clinical validation
We determined the specificity and sensitivity of our assay by analysing archival serum and plasma cfDNA from 40 cancer patients at presentation attending the Quiron Dexeus University Hospital (33 NSCLC, 2 CRC and 5 metastatic melanoma) with paired tumour tissue. In addition, we tested archival serum and plasma cfDNAs from 12 responder patients and 11 patients at the time of tumour progression after treatment (18 NSCLC, 2 CRC and 3 metastatic melanoma; Table 1). All of the 63 cfDNA samples and tumour tissues had previously been genotyped for EGFR, KRAS, NRAS and BRAF mutations using a TDA (Gonzalez-Cao et al, 2015; Karachaliou et al, 2015). In the case of tumour tissues, genotyping had been confirmed by standard PCR followed by Sanger sequencing. Cases showing discordance between the NGS SiRe panel and the TDA were further investigated by digital PCR (dPCR) on a QuantStudio 3D Digital PCR System platform (Thermofisher) as previously described (Malapelle et al, 2016b).
Performance of the SiRe panel in prospective clinical samples
To evaluate the performance of the SiRe panel in the clinical setting, we prospectively genotyped 79 advanced NSCLC patients (37 men and 42 women; mean age: 65 years) using blood samples collected at the Department of Public Health of the University of Naples Federico II. According to the European Medicines Agency guidelines, mutations related to EGFR disease were tested in patients when tissue was not available at presentation (n=46), or at tumour progression (n=33) in patients previously treated with erlotinib (n=14), gefitinib (n=14) or afatinib (n=5) in the attempt to detect the emergence of resistance secondary mutations. In 21 of the 33 cases with tumour progression, first-line TKI administration had been based on the demonstration of an EGFR mutation in tissue, whereas in the remaining 12/33 cases, TKI treatment had been administrated in second line without evidence of EGFR mutations.
Results
Panel design and preclinical performance evaluation
The SiRe panel was designed to cover 568 clinically relevant mutations in six genes (EGFR, KRAS, NRAS, BRAF, cKIT and PDGFRα) involved in NSCLC, GIST, CRC and metastatic melanoma (see Supplementary Table S1). The panel was intended for use in cfDNA purified from patients with advanced cancer. On cell line derived DNA, the SiRe panel detected the EGFR deletion p.E746_A750del and the KRAS point mutation p.G12S at a level as low as one copy of the mutated allele in a background of 20000 copies of wild-type alleles (0.005% mutated allele fraction), with 100% of intra- and inter-run reproducibility. In addition, regarding the results obtained on multiplex gDNA reference standard (Horizon Diagnostics), p.E746_A750del and p.G719S point mutation in EGFR, p.G12D mutation in KRAS exon 2, p.Q61L mutation in NRAS exon 3 and p.V600E mutation in BRAF exon 15 were correctly identified for each different dilution point.
This high analytical performance was achieved thanks to the use of optimised parameters set in variant caller plug-in (v.5.0.2.1) which detected low abundant mutated alleles with a specificity of 100% (see Supplementary Table S2).
Clinical sensitivity and specificity of the SiRe panel in cfDNA samples
The retrospective series of cfDNAs (Supplementary Table S3) was constituted by 126 paired serum and plasma samples from 63 patients. In each run, up to 16 paired serum and plasma samples from eight patients on 316v2 were processed. Run median output was 257Mbases, median read length was 124 bp, mean read depth was 2821 × and coverage uniformity was 97%. Technical performance data relative to each processed sample are reported in Supplementary Material (Supplementary Table S4). When the 63 samples were tested with the SiRe panel, the cfDNA of all eight patients with wild-type tumour tissue was negative (specificity 100%, CI 64.6-100%). In the remaining 55 patients with EGFR, KRAS, NRAS or BRAF mutations in tumour tissue, the SiRe panel detected the same mutation in the serum and/or plasma cfDNA in 46 cases (sensitivity 83.6%, CI 67.3–94.3%; Table 2).
Comparison of the SiRe panel with a TDA in cfDNA samples
We compared the performance of the SiRe panel for mutation analysis in cfDNA with that of a previously reported TDA (Karachaliou et al, 2015; Gonzalez-Cao et al, 2015) in 63 samples: (i) the 40 cfDNA samples obtained at presentation mentioned above; (ii) archival serum and plasma cfDNAs from 12 patients in response to different types of antitumour drugs; and 11 patients mutations in the cfDNA of 46 of 63 patients. The test was positive in both serum and plasma cfDNA in 35 patients (76.1%), positive in plasma but not in serum in 5 patients (10.9%), and positive in serum but not in plasma in 6 patients (13%). An EGFR sensitising mutation and the p.T790M resistance mutation were detected simultaneously in 10 patients at progression to EGFR TKIs.
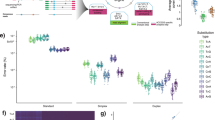
As reported in Table 2, there was a high concordance (Cohen’s Kappa 0.85) between the results obtained with the NGS SiRe panel and the TDA, although the performance of the SiRe was slightly better. All 42 patients with mutation-positive cfDNA at TDA were also positive with the SiRe panel, and the 17 negative samples with the panel were also negative at TDA. In addition, NGS detected mutations in the cfDNA of four patients, whereas TDA did not. The mutations in these four patients appeared also in paired tumour tissue. One was a p.L597R mutation in BRAF not covered by the TDA, and was confirmed by dPCR (Supplementary Figure S2). The remaining three mutations were a p.L861Q mutation in EGFR and two KRAS mutations, p.G12C and p.G12A. Both TDA and NGS using the SiRe panel enable quantification of the mutated alleles (Figure 2). There was a significant correlation in the levels of serum cfDNA between the two techniques (r=0.64). In contrast, correlation was lower in the case of plasma (r=0.35), but improved significantly when three outlier samples were removed (r=0.61).
Evaluation of the SiRe panel for prospective analysis of clinical samples
The performance of the SiRe panel in the clinical setting was evaluated by prospectively testing the serum and plasma cfDNA of patients with advanced NSCLC for whom no tissue was available in order to select them for TKI treatment. Seventy-nine patients were tested, 46 at presentation and 33 at the time of tumour progression after first-line TKI treatment (Table 1). The NGS procedure was adequate for variant calling in the 79 cfDNA paired serum and plasma samples. The run metrics parameters were not dissimilar from those of the retrospective samples. In fact, in prospective cfDNA samples, the median output was 210Mbases, the median read length 125.57 bp, the mean read depth 3385.45 and coverage uniformity 97.49%. Among the 46 patients analysed at baseline (Supplementary Table S5), we detected four EGFR mutations (8.7%), one point mutation in exon 18 (p.G719A), two deletions in exon 19 (both p.E746_A750delELREA) and one insertion in exon 20 (p.H773-V774insH). In all four patients, the mutant alleles were detected in both serum and plasma cfDNA and were confirmed by digital PCR (data not shown).
Regarding samples at progression (Supplementary Table S6), the SiRe panel did not detect mutations in 12 patients, whose tissues had been identified as EGFR wild type in biopsies at presentation. In contrast, among the 21 patients EGFR positive in baseline tissue, the SiRe panel confirmed the same mutation in cfDNA in 19 cases (Table 3). Thus, sensitivity and specificity in this cohort of patients at progression were within the range of those observed in the retrospective cohort. Interestingly, in 9 of those 19 cases (47%), we observed the emergence of the EGFR p.T790M mutation in addition to the original EGFR activating mutation. The appearance of EGFR p.T790M mutation in relation to TKIs treatment regimen was reported in Figure 3. Of the 28 mutations (sensitising+p.T790M) detected, 10 (35.70%) were present in both serum and plasma, 7 (25%) in plasma alone and 11 (39.3%) in serum alone. All mutations detected by the SiRe panel at progression were confirmed by dPCR.
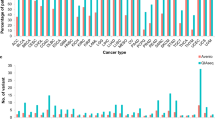
Frequency of the EGFR p.T790M mutation (green: T790M−; red T790M+) after progression to thyrosine kinase inhibitors (TKIs) in the serum and plasma cfDNA of EGFR-mutated patients evaluated with SiRe panel NGS.A full colour version of this figure is available at the British Journal of Cancer journal online.
Discussion
In this proof-of-concept study, we demonstrate that the performance of ultra-deep sequencing using a narrow NGS panel on Ion Torrent PGM is excellent, and that this procedure can be used for the routine testing of relevant tumour mutations in cfDNA. The high sensitivity (90.5%) and analytical specificity (100%) of this panel equal or even surpass those of such other procedures as real-time PCR-based methods. Unlike earlier NGS applications that cover large genomic regions (Cancer Genome Atlas Research Network, 2014), our small gene panel (5.2 kb) focuses on biomarkers that are currently used in the clinical setting.
The ultra-deep sequencing procedure reported herein has various advantages. In fact, using a single panel, we were able to detect up to 568 relevant mutations in six genes (EGFR, KRAS, NRAS, BRAF, cKIT and PDGFRα). These mutations included less common, but actionable variants such as the BRAF p.L597R mutation in melanoma (case #38 in Supplementary Table S3). Sequencing with the SiRe panel was more efficient than real-time PCR target techniques in detecting deletions (n=2) and point mutations (n=6) on cfDNA samples. In addition, NGS per se is a time-effective procedure for analysing large numbers of samples, thereby optimising the work flow in molecular pathology laboratories (Malapelle et al, 2016a). With our procedure, different types of samples (DNA from tumour tissues and cfDNAs from biological fluids) from patients affected by different types of diseases (e.g., NSCLC, GIST, CRC and melanoma) can be processed simultaneously. Consequently, sample batching is more effective and does not require a minimum number of a given tumour type. As a result, turnaround time (TAT) can be as short as three working days, as recommended by international guidelines (Lindeman et al, 2013). The recently developed Ion Chef automated library preparation station, which has a better procedure reproducibility and standardisation than manual procedures, also contributes to the short TAT (Malapelle et al, 2016a).
The Ion Torrent PGM protocols, panels and variant caller do not detect low abundant mutations diluted in a large amount of WT DNA. Therefore, we used several in-house strategies specifically tailored to cfDNA. Firstly, we reduced the number of genes and exons vs commercially available tests, and we modified the thresholds for variant calling, in particular all the variants with ⩾5X allele coverage and a quality score ⩾20, within an amplicon that covered at least 1000X alleles, were called (Supplementary Table S2).
We also adapted the Ion Chef template preparation protocol by pooling two 16-sample libraries in each run. Thus, using this well standardised procedure, we were able to sequence simultaneously up to 32 paired plasma/serum samples in less than 3 h on the PGM, with a consequent reduction in the total consumable cost. In a previous study (Malapelle et al, 2016a)we showed that by using a commercially available 22 gene panel(AmpliSeq Colon and Lung Cancer Panel)on the Ion Torrent PGM, the consumable cost was €196 per sample. Using the modified protocol that we developed in this current study the cost per sample was lowered to 98 euro for simultaneously analysis of six different genes. This is comparable with the cost of the most commercially available Real Time PCR based kits.
The simultaneous analysis of paired plasma/serum samples is a crucial feature of this new procedure since the sensitivity of somatic mutation analysis in cfDNA increases when serum and plasma are analysed together (Gonzalez-Cao et al, 2015; Karachaliou et al, 2015). Our results are in agreement with this finding. In fact, of the 89 patients found to carry mutations in cfDNA, 58 (65.17%)were positive in both serum and plasma, 15 (16.85%) in plasma alone and 16 (17.98%)in serum alone.
From the technical point of view, even when sequencing 16 samples simultaneously in a run, the SiRe panel had optimal run metrics in our daily clinical practice in terms of both mean depth reads and uniformity of coverage, which resulted in a high assay sensitivity in cfDNA vs tumour tissue (90.5%) and a specificity of 100%. This is a very high degree of concordance, particularly given the 91.7% concordance between paired surgical resection and cytological samples (Sun et al, 2013). Thanks to the high sensitivity of our assay, the EGFR mutational rate of 8.7% that we identified in NSCLC patients prospectively tested on cfDNA at baseline is in keeping with previous data on tissue samples (Malapelle et al, 2013). Similarly, the frequency of the EGFR p.T790M mutation, which was detected in the cfDNA of 9 of 19 (47.4%) patients progressing after TKI treatment (n=5 gefitinib, n=3 afatinib, n=1 erlotinib), is in line with data obtained on tissues samples collected after disease progression (Karachaliou et al, 2015).
The performance of our methodology compares favorably with that of NGS for mutational analysis in the blood of cancer patients. An Ion Torrent-derived sequencing of five genes in cfDNA purified from never smoking lung cancer patients achieved a modest 58% sensitivity and 87% specificity (Couraud et al, 2014). An analysis of 23 amplicons in five genes using cfDNA from breast cancer patients identified 10 mutations but missed 6 identified by droplet digital PCR (Guttery et al, 2015). When restricted to EGFR, deep sequencing achieved 61–80% sensitivity and 94–98% specificity in advanced NSCLC (Uchida et al, 2015). The 90.5% sensitivity of our assay also exceeds the 77% recently reported when NSCLC plasma-derived cfDNA was analysed on an Illumina NGS platform with a panel covering amplicons of 11 clinically relevant genes (Paweletz et al, 2016). Despite the variations inherent to the platforms used, such as the library preparation and the longer TAT (6 days), the Illumina-based NGS approach featured similar run metrics and analytical parameters as our assay, which supports the use of ultra-deep sequencing in the clinical setting (Paweletz et al, 2016). It is conceivable that the higher sensitivity achieved by our panel is due not only to technical differences but also to the simultaneous testing of serum and plasma in each patient.
Besides being an alternative to molecular diagnosis at presentation when tumour tissue is not available, liquid biopsy is also a noninvasive test with which to monitor response to targeted therapy and to detect the emergence of resistance mutations in genes such as EGFR (Sundaresan et al, 2016) and ESR1 (Chu et al, 2016). Monitoring would consist in quantifying the mutant allelic fractions in cfDNA over time, which can be reliably assessed by our NGS assay. The SiRe panel detects the appearance of resistance mutations such as EGFR p.T790M (Figure 3). Finally, the non-synonymous mutation burden correlates with a good response to immunotherapy in NSCLC (Rizvi et al, 2015) and other tumours, and NGS has been proposed as a tool with which to design customised immunotherapies that target common driver mutations (Nielsen et al, 2016). Our panel, which covers several exons in frequently mutated genes, can be useful also in this setting.
In conclusion, we have developed and translate in clinical setting an NGS assay based on a narrow gene panel. The assay detects relevant mutations in cfDNA purified from the serum and plasma of patients with the tumours most commonly tested for molecular alterations (such as NSCLC, CRC and metastatic melanoma). The SiRe panel has excellent sensitivity and specificity, and is hence suitable for testing blood samples in the clinical setting. Finally, it enables the application of NGS on a prospective basis in daily molecular predictive pathology practice, particularly when tumour tissue is not available, and is a tool with which to monitor disease course.
Change history
14 March 2017
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Antonescu CR (2008) Targeted therapies in gastrointestinal stromal tumors. Semin Diagn Pathol 25: 295–303.
Cancer Genome Atlas Research Network (2014) Comprehensive molecular profiling of lung adenocarcinoma. Nature 511: 543–550.
Chapman PB, Hauschild A, Robert C, Haanen JB, Ascierto P, Larkin J, Dummer R, Garbe C, Testori A, Maio M, Hogg D, Lorigan P, Lebbe C, Jouary T, Schadendorf D, Ribas A, O’Day SJ, Sosman JA, Kirkwood JM, Eggermont AM, Dreno B, Nolop K, Li J, Nelson B, Hou J, Lee RJ, Flaherty KT, McArthur GA BRIM-3 Study Group (2011) Improved survival with vemurafenib in melanoma with BRAF V600E mutation. N Engl J Med 364: 2507–2516.
Chu D, Paoletti C, Gersch C, VanDenBerg DA, Zabransky DJ, Cochran RL, Wong HY, Toro PV, Cidado J, Croessmann S, Erlanger B, Cravero K, Kyker-Snowman K, Button B, Parsons HA, Dalton WB, Gillani R, Medford A, Aung K, Tokudome N, Chinnaiyan AM, Schott A, Robinson D, Jacks KS, Lauring J, Hurley PJ, Hayes DF, Rae JM, Park BH (2016) ESR1 mutations in circulating plasma tumor DNA from metastatic breast cancer patients. Clin Cancer Res 22: 993–999.
Couraud S, Vaca-Paniagua F, Villar S, Oliver J, Schuster T, Blanché H, Girard N, Trédaniel J, Guilleminault L, Gervais R, Prim N, Vincent M, Margery J, Larivé S, Foucher P, Duvert B, Vallee M, Le Calvez-Kelm F, McKay J, Missy P, Morin F, Zalcman G, Olivier M, Souquet PJ BioCAST/IFCT-1002 Investigators (2014) Noninvasive diagnosis of actionable mutations by deep sequencing of circulating free DNA in lung cancer from never-smokers: a proof-of- conceptstudy from BioCAST/IFCT-1002. Clin Cancer Res 20: 4613–4624.
Crowley E, Di Nicolantonio F, Loupakis F, Bardelli A (2013) Liquid biopsy: monitoring cancer-genetics in the blood. Nat Rev Clin Oncol 10: 472–484.
Di Nicolantonio F, Martini M, Molinari F, Sartore - Bianchi A, Arena S, Saletti P, De Dosso S, Mazzucchelli L, Frattini M, Siena S, Bardelli A (2008) Wild-type BRAF is required for response to panitumumab or cetuximab in metastatic colorectal cancer. J Clin Oncol 26: 5705–5712.
Gargis AS, Kalman L, Berry MW, Bick DP, Dimmock DP, Hambuch T, Lu F, Lyon E, Voelkerding KV, Zehnbauer BA, Agarwala R, Bennett SF, Chen B, Chin EL, Compton JG, Das S, Farkas DH, Ferber MJ, Funke BH, Furtado MR, Ganova-Raeva LM, Geigenmüller U, Gunselman SJ, Hegde MR, Johnson PL, Kasarskis A, Kulkarni S, Lenk T, Liu CS, Manion M, Manolio TA, Mardis ER, Merker JD, Rajeevan MS, Reese MG, Rehm HL, Simen BB, Yeakley JM, Zook JM, Lubin IM (2012) Assuring the quality of next-generation sequencing in clinical laboratory practice. Nat Biotechnol 30: 1033–1036.
Gonzalez-Cao M, Mayo-de-Las-Casas C, Molina-Vila MA, De Mattos-Arruda L, Muñoz-Couselo E, Manzano JL, Cortes J, Berros JP, Drozdowskyj A, Sanmamed M, Gonzalez A, Alvarez C, Viteri S, Karachaliou N, Martin Algarra S, Bertran-Alamillo J, Jordana-Ariza N, Rosell R (2015) BRAF mutation analysis in circulating free tumor DNA of melanoma patients treated with BRAF inhibitors. Melanoma Res 25: 486–495.
Guttery DS, Page K, Hills A, Woodley L, Marchese SD, Rghebi B, Hastings RK, Luo J, Pringle JH, Stebbing J, Coombes RC, Ali S, Shaw JA (2015) Noninvasive detection of activating estrogen receptor 1 (ESR1) mutations in estrogen receptor positive metastatic breast cancer. Clin Chem. 61: 974–982.
Karachaliou N, Mayo-de las Casas C, Queralt C, de Aguirre I, Melloni B, Cardenal F, Garcia- Gomez R, Massuti B, Sánchez JM, Porta R, Ponce-Aix S, Moran T, Carcereny E, Felip E, Bover I, Insa A, Reguart N, Isla D, Vergnenegre A, de Marinis F, Gervais R, Corre R, Paz- Ares L, Morales-Espinosa D, Viteri S, Drozdowskyj A, Jordana-Ariza N, Ramirez-Serrano JL, Molina-Vila MA, Rosell R Spanish Lung Cancer Group (2015) Association of EGFR L858R mutation in circulating free DNA with survival in the EURTAC Trial. JAMA Oncol 1: 149–157.
Lièvre A, Bachet JB, Boige V, Cayre A, Le Corre D, Buc E, Ychou M, Bouché O, Landi B, Louvet C, André T, Bibeau F, Diebold MD, Rougier P, Ducreux M, Tomasic G, Emile JF, Penault-Llorca F, Laurent-Puig P (2008) KRAS mutations as an independent prognostic factor in patients with advanced colorectal cancer treated with cetuximab. J Clin Oncol 26: 374–379.
Lindeman NI, Cagle PT, Beasley MB, Chitale DA, Dacic S, Giaccone G, Jenkins RB, Kwiatkowski DJ, Saldivar JS, Squire J, Thunnissen E, Ladanyi M College of American Pathologists International Association for the Study of Lung Cancer and Association for Molecular Pathology (2013) Molecular testing guideline for selection of lung cancer patients for EGFR and ALK tyrosine kinase inhibitors: guideline from the College of American Pathologists, International Association for the Study of Lung Cancer, and Association for Molecular Pathology. J Mol Diagn. 15: 415–453.
Lynch TJ, Bell DW, Sordella R, Gurubhagavatula S, Okimoto RA, Brannigan BW, Harris PL, Haserlat SM, Supko JG, Haluska FG, Louis DN, Christiani DC, Settleman J, Haber DA (2004) Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med. 350: 2129–2139.
Malapelle U, Bellevicine C, De Luca C, Salatiello M, De Stefano A, Rocco D, de Rosa N, Vitiello F, Russo S, Pepe F, Iaccarino A, Micheli P, Illiano A, Carlomagno C, Piantedosi FV, Troncone G (2013) EGFR mutations detected on cytology samples by a centralized laboratory reliably predict response to gefitinib in non small cell lung cancer patients. Cancer Cytopathol 121: 552–560.
Malapelle U, Pisapia P, Sgariglia R, Vigliar E, Biglietto M, Carlomagno C, Giuffrè G, Bellevicine C, Troncone G (2016a) Less frequently mutated genes in colorectal cancer: evidences from next-generation sequencing of 653 routine cases. J Clin Pathol 69: 767–771.
Malapelle U, de Luca C, Vigliar E, Ambrosio F, Rocco D, Pisapia P, Bellevicine C, Troncone G (2016b) EGFR mutation detection on routine cytological smears of non-small cell lung cancer by digital PCR: a validation study. J Clin Pathol 69: 454–457.
Malapelle U, Pisapia P, Rocco D, Smeraglio R, di Spirito M, Bellevicine C, Troncone G (2016c) Next generation sequencing techniques in liquid biopsy: focus on non-small cell lung cancer patients. Transl Lung Cancer Res 5: 505–510,, Review.
Mead R, Duku M, Bhandari P, Cree IA (2011) Circulating tumour markers can define patients with normal colons, benign polyps, and cancers. Br J Cancer 105: 239–245.
Nielsen JS, Sedgwick CG, Shahid A, Zong Z, Brumme ZL, Yu S, Liu L, Kroeger DR, Treon SP, Connors JM, Gascoyne RD, Berry BR, Marra MA, Morin RD, Macpherson N, Nelson BH (2016) Toward personalized lymphoma immunotherapy: identification of common driver mutations recognized by patient CD8+ T cells. Clin Cancer Res 22: 2226–2236.
Papadopoulos N, Kinzler KW, Vogelstein B (2006) The role of companion diagnostics in the development and use of mutation-targeted cancer therapies. Nat Biotechnol 24: 985–995.
Paweletz CP, Sacher AG, Raymond CK, Alden RS, O’Connell A, Mach SL, Kuang Y, Gandhi L, Kirschmeier P, English JM, Lim LP, Jänne PA, Oxnard GR (2016) Bias-corrected targeted next-generation sequencing for rapid, multiplexed detection of actionable alterations in cell-free DNA from advanced lung cancer patients. Clin Cancer Res 22: 915–922.
Rizvi NA, Hellmann MD, Snyder A, Kvistborg P, Makarov V, Havel JJ, Lee W, Yuan J, Wong P, Ho TS, Miller ML, Rekhtman N, Moreira AL, Ibrahim F, Bruggeman C, Gasmi B, Zappasodi R, Maeda Y, Sander C, Garon EB, Merghoub T, Wolchok JD, Schumacher TN, Chan TA (2015) Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348: 124–128.
Schwarzenbach H, Hoon DS, Pantel K (2011) Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer 11: 426–437.
Sun PL, Jin Y, Kim H, Lee CT, Jheon S, Chung JH (2013) High concordance of EGFR mutation status between histologic and corresponding cytologic specimens of lung adenocarcinomas. Cancer Cytopathol 121: 311–319.
Sundaresan TK, Sequist LV, Heymach JV, Riely GJ, Jänne PA, Koch WH, Sullivan JP, Fox DB, Maher R, Muzikansky A, Webb A, Tran HT, Giri U, Fleisher M, Yu HA, Wei W, Johnson BE, Barber TA, Walsh JR, Engelman JA, Stott SL, Kapur R, Maheswaran S, Toner M, Haber DA (2016) Detection of T790M, the acquired resistance EGFR mutation, by tumor biopsy versus noninvasive blood-based analyses. Clin Cancer Res 22: 1103–1110.
Uchida J, Kato K, Kukita Y, Kumagai T, Nishino K, Daga H, Nagatomo I, Inoue T, Kimura M, Oba S, Ito Y, Takeda K, Imamura F (2015) Diagnostic accuracy of noninvasive genotyping of EGFR in lung cancer patients by deep sequencing of plasma cell-free DNA. Clin Chem 61: 1191–1196.
van Dijk EL, Auger H, Jaszczyszyn Y, Thermes C (2014) Ten years of next-generation sequencing technology. Trends Genet 30: 418–426.
Acknowledgements
We thank Jean Ann Gilder (Scientific Communication srl., Naples, Italy) for editing the manuscript. The project has been supported by Department of Public Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Malapelle, U., Mayo de-Las-Casas, C., Rocco, D. et al. Development of a gene panel for next-generation sequencing of clinically relevant mutations in cell-free DNA from cancer patients. Br J Cancer 116, 802–810 (2017). https://doi.org/10.1038/bjc.2017.8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2017.8
Keywords
This article is cited by
-
Non-Small Cell Lung Cancer Testing on Reference Specimens: An Italian Multicenter Experience
Oncology and Therapy (2024)
-
Development and externally validate MRI-based nomogram to assess EGFR and T790M mutations in patients with metastatic lung adenocarcinoma
European Radiology (2022)
-
Longitudinal analysis of cell-free mutated KRAS and CA 19–9 predicts survival following curative resection of pancreatic cancer
BMC Cancer (2021)
-
Overcoming therapy resistance in EGFR-mutant lung cancer
Nature Cancer (2021)
-
From single gene analysis to single cell profiling: a new era for precision medicine
Journal of Experimental & Clinical Cancer Research (2020)