Abstract
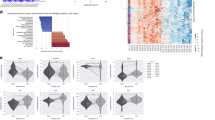
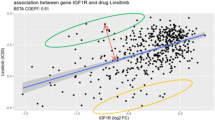
Cancer stem cells (CSCs) are thought to promote resistance to chemotherapeutic drugs in glioblastoma, the most common and aggressive primary brain tumor. However, the use of high-throughput drug screens to discover novel small-molecule inhibitors for CSC has been hampered by their instability in long-term cell culture. We asked whether predictive models of drug response could be developed from gene expression signatures of established cell lines and applied to predict drug response in glioblastoma stem cells. Predictions for active compounds were confirmed both for 185 compounds in seven established glioma cell lines and 21 compounds in three glioblastoma stem cells. The use of established cell lines as a surrogate for drug response in CSC lines may enable the large-scale virtual screening of drug candidates that would otherwise be difficult or impossible to test directly in CSCs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Riddick G, Fine HA . Integration and analysis of genome-scale data from gliomas. Nat Rev Neurol 2011; 7: 439–450.
Jiang Y, Uhrbom L . On the origin of glioma. Ups J Med Sci 2012; 117: 113–121.
Hemmati H D, Nakano I, Lazareff JA, Masterman-Smith M, Geschwind DH, Bronner-Fraser M et al. Cancerous stem cells can arise from pediatric brain tumors. Proc Natl Acad Sci 2003; 100: 15178–15183.
Lee J, Kotliarova S, Kotliarov Y, Li A, Su Q, Donin NM et al. Tumor stem cells derived from glioblastomas cultured in bFGF and EGF more closely mirror the phenotype and genotype of primary tumors than do serum-cultured cell lines. Cancer Cell 2006; 9: 391–403.
Godlewski J, Nowicki MO, Bronisz A, Williams S, Otsuki A, Nuovo G et al. Targeting of the Bmi-1 oncogene/stem cell renewal factor by microRNA-128 inhibits glioma proliferation and self-renewal. Cancer Res 2008; 68: 9125–9130.
Groszer M . Negative regulation of neural stem/progenitor cell proliferation by the Pten Tumor suppressor gene in vivo. Science 2001; 294: 2186–2189.
Häyry V, Tanner M, Blom T, Tynninen O, Roselli A, Ollikainen M et al. Copy number alterations of the polycomb gene BMI1 in gliomas. Acta Neuropathol 2008; 116: 97–102.
Holland EC, Celestino J, Dai C, Schaefer L, Sawaya RE, Fuller GN et al. Combined activation of Ras and Akt in neural progenitors induces glioblastoma formation in mice. Nature Genet 2000; 25: 55–57.
Kwon C-H, Zhao D, Chen J, Alcantara S, Li Y, Burns DK et al. Pten haploinsufficiency accelerates formation of high-grade astrocytomas. Cancer Res 2008; 68: 3286–3294.
Liu C, Tu Y, Sun X, Jiang J, Jin X, Bo X et al. Wnt/beta-Catenin pathway in human glioma: expression pattern and clinical/prognostic correlations. Clin Exp Med 2010; 11: 105–112.
Purow BW, Haque RM, Noel MW, Su Q, Burdick MJ, Lee J et al. Expression of Notch-1 and its ligands, Delta-like-1 and Jagged-1, is critical for glioma cell survival and proliferation. Cancer Res 2005; 65: 2353–2363.
Alcantara Llaguno S, Chen J, Kwon CH, Jackson ELLi Y, Burns DK et al. Malignant astrocytomas originate from neural stem/progenitor cells in a somatic tumor suppressor mouse model. Cancer Cell 2009; 15: 45–56.
Zhu Y, Guignard F, Zhao D, Liu L, Burns DK, Mason RP et al. Early inactivation of p53 tumor suppressor gene cooperating with NF1 loss induces malignant astrocytoma. Cancer Cell 2005; 8: 119–130.
Chen J, Li Y, Yu TS, McKay RM, Burns DK, Kernie SG et al. A restricted cell population propagates glioblastoma growth after chemotherapy. Nature 2012; 488: 522–526.
Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin AA, Kim S et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 2012; 483: 603–307.
Garnett MJ, Edelman EJ, Heidorn SJ, Greenman CD, Dastur A, Lau KW et al. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 2012; 483: 570–575.
Riddick G, Song H, Ahn S, Walling J, Borges-Rivera D, Zhang W et al. Predicting in vitro drug sensitivity using Random Forests. Bioinformatics 2010; 27: 220–224.
Lamb J . The connectivity map: using gene-expression signatures to connect small molecules, genes, and disease. Science 2006; 313: 1929–1935.
Shoemaker RH . The NCI60 human tumour cell line anticancer drug screen. Nat Rev Cancer 2006; 6: 813–823.
Cancer Genome Atlas Research Network, McLendon R, Friedman A, Bigner D, Van Meir EG, Brat DJ et al. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 2008; 455: 1061–1068.
Acknowledgements
We thank the Developmental Therapeutics Program (NCI) for providing drug sensitivity data for the NCI-60 cell lines and the Sanger Wellcome Trust for providing drug sensitivity and gene expression data for 485 cell lines.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the The Pharmacogenomics Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Riddick, G., Song, H., Holbeck, S. et al. An in silico screen links gene expression signatures to drug response in glioblastoma stem cells. Pharmacogenomics J 15, 347–353 (2015). https://doi.org/10.1038/tpj.2014.61
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tpj.2014.61