Abstract
Somatic cells can be reprogrammed to induced hepatocyte-like cells (iHeps) by overexpressing certain defined factors in direct reprogramming techniques. Of the various methods to deliver genes into cells, typically used genome-integrating viral vectors are associated with integration-related adverse events such as mutagenesis, whereas non-integrating viral vectors have low efficiency, making viral vectors unsuitable for clinical application. Therefore, we focused on developing a transposon system to establish a non-viral reprogramming method. Transposons are unique DNA elements that can be integrated into and removed from chromosomes. PiggyBac, a type of transposon, has high transduction efficiency and cargo capacity, and the integrated transgene can be precisely excised in the presence of transposase. This feature enables the piggyBac vector to achieve efficient transgene expression and a transgene-free state, thus making it a promising method for cell reprogramming. Here, we attempted to utilize the piggyBac transposon system to generate iHeps by integrating a transgene consisting of Hnf4a and Foxa3, and successfully obtained functional iHeps. We then demonstrated removal of the transgene to obtain transgene-free iHeps, which still maintained hepatocyte functions. This non-viral, transgene-free reprogramming method using the piggyBac vector may facilitate clinical applications of iHeps in upcoming cell therapy.
Similar content being viewed by others
Introduction
Therapy for lethal liver failure has mostly been limited to liver transplantation up to the present date, and the shortage of donor livers is a serious problem. Recently, cell transplantation is considered to be an alternative therapeutic choice. Although the leading cell-source candidate should be hepatocytes, it remains difficult to establish stable procedures for their isolation, preservation, and supply1,2,3,4. To overcome this hurdle, hepatocyte-like cells generated from the patient’s own somatic cells are expected to be an attractive alternative. Experimental procedures have been reported for differentiation of other cell types into hepatocyte-like cells, such as mesenchymal stem cells (MSCs)5,6,7, embryonic stem cells (ESCs)8,9, and induced pluripotent stem cells (iPSCs)10,11, under specific culture conditions. Since 2011, when Suzuki et al. and Huang et al. established a direct reprogramming technique to reprogram mouse fibroblasts to induced-hepatocyte like cells (iHeps) by overexpressing defined factors12,13, research in this field has proceeded rapidly.
Viral vectors have commonly been used as gene delivery tools because of their capability to transduce genes efficiently. However, although integrative vectors, such as retro- or lentivirus, have been utilized in many reprogramming trials for their strong ability to integrate and drive the expression of genes of interest, they present risks of genome mutagenesis and genotoxicity14,15 resulting from random genome integration. In contrast, non-integrative vectors, such as adenovirus, only provide transient gene expression and exhibit lower reprogramming effciency16,17. Therefore, given that viral vectors require special care in handling, non-viral vectors are preferable for generating clinically applicable iHeps.
Recently, non-viral methods to generate iHeps using synthetic mRNA18, microRNA19, or episomal vectors20 have been reported; however, these methods have low reprogramming efficiency and the procedures are complicated. In the present study, we focused on a piggyBac transposon system to establish a non-viral, transgene-free reprogramming method. Transposons are unique DNA segments that were discovered in animals and plants as reversibly “mobile elements” in the genome21. In the presence of transposase, transposons are excised from a region on the host genome and integrated into another region of the genome. PiggyBac, a transposon discovered in the cabbage looper moth Trichoplusia ni22, possesses a large cargo size and high transposition efficiency in mammalian cells23,24. Additionally, the excision of the transposon segment is precise and leaves no footprint on the host genome in which the transgene was integrated, thereby reducing the risk related to integration and making piggyBac a suitable vector for cell reprogramming25,26. In the present study, we attempted to generate iHeps from mouse MSCs by using piggyBac-based direct reprogramming to overexpress a transgene, and then removed the transgene to generate transgene-free iHeps. With this strategy, we successfully obtained functional transgene-free iHeps. This approach has the potential to advance iHep generation toward clinical application.
Results
Generating factor-integrated iHeps from mouse MSCs
First, we confirmed the maintenance of multipotency by differentiating the mouse MSCs into adipogenic and osteogenic lineages. The lipid droplets in adipocytes were visualized using Oil Red O staining and calcified osteocytes were visualized using Alizarin Red S staining (Fig. 1A). To construct the factor-expressing vector, we selected Hnf4a and Foxa3 as reprogramming factors; Suzuki et al.12 selected these factors as one of the appropriate combinations to generate iHeps, and linked them with a 2A sequence to transcribe both genes equally. Then Hnf4a-2A-Foxa3 was inserted into the multi cloning site (MCS) of a dual-promoter piggyBac plasmid vector (pPBd), resulting in pPBHF3 (Fig. 1B). In pPBHF3, the CMV promoter was expected to drive Hnf4a and Foxa3 and the EF1a promoter to drive mRuby-puro, each present within a region of the transposon segment. pPBHF3 and pPBase were co-transfected into MSCs and the cells underwent the following protocol for iHep generation; additionally, pPBd and pPBase were co-transfected into MSCs as mock-transfected samples (pPBd-TCs) and the cells underwent the same steps. Consequently, the transgene integrated into pPBHF3-transfected cells contained Hnf4a, Foxa3 and mRuby-puro, whereas the transgene of pPBd-TCs contained only mRuby-puro.
(A) The characteristics of MSCs-derived adipocytes and osteocytes were confirmed by Oil Red O staining and Arizarin Red S staining, respectively. (B) pPBd possesses transposon segment indicated between inverted terminal repeat sequences (ITR), carrying CMV promoter-MCS and EF1a promoter-mRuby-puro. A set of reprogramming factors, Hnf4a and Foxa3, were linked with T2A sequence and inserted into MCS to construct pPBHF3. (C) pPBHF3 and pPBase were co-transfected into MSCs. After antibiotics selection, all surviving cells expressed Hnf4a and Foxa3 simultaneously. Scale bar : 100 um.
From the day after transfection, antibiotic selection with puromycin at 5 μg/ml was performed until post-transfection day 7. When the surviving cells reached confluence, all cells showed red fluorescence, suggesting that they carried the integrated transgenes. Immunofluorescent staining detected the expression of Hnf4a and Foxa3 in the nuclei of surviving cells (Fig. 1C), indicating the successful activation of the transgenes in the pPBHF3-transfected cells.
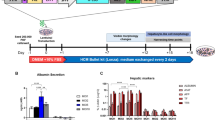
Cell differentiation was initiated on post-transfection day 7, beginning with step A culture in expansion medium for 12 days followed by step B culture with maturation medium (Fig. 2A). 5′-azacytidine was added for the first three days of differentiation step A. When expression of mRNA extracted every seven days was analyzed by quantitative PCR (qPCR), pPBd-TCs showed a slight but steady increase in albumin mRNA expression after post-transfection day 7, whereas pPBHF3-transfected cells showed a marked increase on day 7, with further increases thereafter (Fig. 2B). pPBHF3-transfected cells changed morphologically from a fibroblast-like shape to a polygonal shape and were densely proliferating (Fig. 2C). To evaluate reprogramming efficiency, immunofluorescent staining for E-cadherin was performed for pPBHF3-transfected cells on day 15 of the differentiation step; cells were then observed and counted using microscopy (Supplementary Fig. S1). Of the first seeded 1.34 × 103 cells per field, 336 ± 8.1 fluorescence-positive cells and colonies per field were detected; therefore, the reprogramming ratio was estimated to be 25.1%. To evaluate the mRNA expression pattern, PCR products of cultured pPBHF3-transfected cells and pPBd-TCs, along with those of undifferentiated MSCs and adult mouse hepatocytes, were compared. In pPBHF3-transfected cells, representative markers typically seen in hepatocytes, such as Cdh1 (E-cadherin), Alb, CK18, G6Pase, and Cyp3a11, were detected. Interestingly, Afp and CK19, which are hepatic progenitor markers and usually downregulated in hepatocytes, were also detected in pPBHF3-transfected cells (Fig. 2D). Thus, we considered that the pPBHF3-transfected cells had acquired hepatocyte characteristics and had been reprogrammed to iHeps.
(A) Antibiotics selection proceeded until post-transfection day 7, followed by initiation of differentiation steps. The differentiation protocol was consisted of 2 steps. Components in each medium are shown. (B) Albumin mRNA expression of pPBHF3-transfected cells and mock-transfected cells (pPBd-TCs) from the day of vector transfection. Differentiation step was initiated on post-transfection day7. Data were normalized to MSCs on day 0. (C) The morphological change in pPBHF3-transfected cells was observed on day 15 of differentiation step. All pPBHF3-transfected cells and pPBd-TCs expressed red fluorescence, indicating that transposon segments from pPBHF3 or pPBd were integrated, respectively. Scale bar: 100 um. (D) PCR bands of representative hepatic markers and hepatic progenitor markers are shown. Expression pattern of iHeps were similar to that of Hepatocytes but for Afp and CK19 expression. iHeps and pPBd-TCs expressed bands specific to the transgene.
Characterization of transgene-integrated iHeps
We next attempted to analyze the characteristics of these iHeps. To evaluate their proliferative capability, we passaged the cells at a 1:4 split ratio when they reached confluence. Although the time until confluence became gradually longer in iHeps than in pPBd-TCs and MSCs, iHeps still exhibited steady proliferation, even at passage 9 (Fig. 3A). When evaluating mRNA expression by qPCR, the levels of Cdh1, Alb, CK18, Ttr, Aat, G6P, Tat, Cyp2e1, and Cyp3a11 were elevated in iHeps as in hepatocytes, and were higher than those in pPBd-TCs. Again, marked elevation of Afp and CK19 was observed in iHeps. Among fibroblast markers the level of Acta2 mRNA decreased, suggesting that transdifferentiation from mesenchymal to epithelial cells was taking place (Fig. 3B). To evaluate protein production ELISA was performed, revealing that secreted albumin was present in the culture medium of iHeps (Fig. 3C). We also evaluated urea production by measuring urea every 12 hours until 48 hours, and revealed significant increase in iHeps compared with PBd-TCs (Fig. 3D). Furthermore, immunofluorescent staining revealed the production of E-cadherin, albumin, and Cyp1a2 in iHeps (Fig. 3E). To assess mature hepatocyte function, Oil Red O staining revealed accumulation of lipid droplets, periodic acid-Schiff (PAS) staining revealed storage of glycogen, and indocyanine-green (ICG) assay showed the capability for uptake of ICG in iHeps (Fig. 3F). These mature hepatocyte functions were not observed in pPBd-TCs or in undifferentiated MSCs.
(A) Each type of cells were passaged at 1:4 split when they reach confluence, until passage 9. (B) mRNA expression level of hepatic, hepatic progenitor and mesenchymal markers were analyzed by quantitative PCR. Data were normalized to MSCs. *p < 0.05. (C) Albumin secretion from each cell types were evaluated by ELISA. *p < 0.05. (D) Urea production of transgene-integrated iHeps and pPB-TCs were measured every 12 hours. Data were normalized to the value at 0 hour. *p < 0.05. (E) Immunofluorescent staining showed expression of albumin, E-cadherin and Cyp1a2 in iHeps. Scale bar: 100 um. (F) Storage of lipid droplets and glycogen inside the cells were evaluated by (a) Oil Red O staining and (b) PAS staining. (c) Uptake of ICG was evaluated. Scale bar: 100 um.
Removal of the transposon and generation of transgene-free iHeps
To remove the transposon segment including the transgene, the iHeps were reseeded onto dishes and transfected with pPBase alone. As a result, when the cells proliferated and reached confluence, scattered fluorescence-negative colonies were observed (Fig. 4A). Then, the cells were dispersed and analyzed using FCM. The intensity of red fluorescence was attenuated for the majority of the cells, compared to those that did not undergo pPBase transfection. An increase in the number of fluorescence-negative iHeps, for which the fluorescence intensity was under the threshold level, was observed (Fig. 4B), and these fluorescence-negative cells were sorted. pPBd-TCs underwent the same procedure. Sorted cells were then seeded onto a low-attachment 96-well plate to form spheroids (Fig. 4C). To confirm the removal of the transposon segment in these fluorescence-negative spheroidal iHeps and pPBd-TCs, PCR was performed using a primer specific to the transposon region of pPBd; a negative result indicated that transgene-free spheroidal iHeps (TFSiHeps) and pPBd-TCs (TFSpPBd-TCs) had been obtained, respectively (Fig. 4D).
(A) After the transfection of pPBase into iHeps, fluorescence-negative iHeps were observed (arrow). (B) Majority of the iHeps which underwent pPBase transfection showed a shift toward fluorescence attenuation. The population of cells under the threshold level (red line) increased, and they were sorted. (C) Fluorescent and bright field observation of iHeps and mock-transfected cells (pPBd-TCs) in the form of spheroid, fluorescence(+) and (–), respectively. Scale bar: 300 um. (D) Removal of the transgene in fluorescence(–) spheroidal iHeps and pPBd-TCs were ascertained by lack of PCR bands with the primer specific to the transgene.
We then evaluated the mRNA expression level of albumin and Cyp3a11. When TFSiHeps were compared with monolayer transgene-integrated iHeps, which were cultured on dishes and did not undergo the transgene-removal process, no significant difference was observed. However, when fluorescence-negative iHeps were seeded onto dishes, the monolayer transgene-free iHeps showed lower albumin and Cyp3a11 expression levels than both monolayer transgene-integrated iHeps and TFSiHeps (Fig. 5A). Therefore, we considered that the spheroidal form was better at maintaining the hepatocyte characteristics of transgene-free iHeps. To evaluate the mRNA expression pattern, PCR products of TFSiHeps and TFSpPBd-TCs, along with those of the spheroidal undifferentiated MSCs and adult mouse hepatocytes, were compared. TFSiHeps expressed the representative markers Cdh1, Alb, CK18, G6Pase, Cyp3a11, AFP, and CK19 in the same pattern observed for monolayer transgene-integrated iHeps (Fig. 5B). To evaluate protein production ELISA was performed, revealing that secreted albumin was present in the culture medium of TFSiHeps (Fig. 5C). We also evaluated urea production by measuring urea every 12 hours until 48 hours, and revealed significant increase compared with TFSpPBd-TCs (Fig. 5D). Immunofluorescent staining revealed the production of E-cadherin, albumin, and Cyp1a2 in TFSiHeps (Fig. 5E). Additionally, PAS staining revealed the storage of glycogen in TFSiHeps (Fig. 5F).
(A) Quantitative PCR analysis for mRNA expression of Alb and Cyp3a11. Monolayer-cultured transgene-integrated iHeps and transgene-free iHeps, and TFSiHeps were compared with each other. Data were normalized to monolayer-cultured pPBd-TCs. *p < 0.05. (B) PCR bands of representative markers are shown for MSCs, transgene-free pPBd-TCs, transgene-free iHeps (TFSiHeps) and hepatocytes, each in the form of spheroid. (C) Albumin secretion of each spheroidal cells were evaluated by ELISA. *p < 0.05. (D) Urea production of TFSiHeps and TFSpPB-TCs were measured every 12 hours. Data were normalized to the value at 0 hour. *p < 0.05. (E) Immunofluorescent staining showed expression of albumin, E-cadherin and Cyp1a2 in TFSiHeps. Scale bar: 50 um. (F) PAS staining showed glycogen storage in TFSiHeps. Scale bar: 100 um. (G) Quantitative PCR analysis for mRNA expression of Hnf4a, Foxa3 and other hepatocyte-specific genes. TFSiHeps which were cultured for 0, 6, 12 days after FCM were compared with monolayer-cultured transgene-integrated iHeps. Data were normalized to TFSpPBd-TCs. *p < 0.05. (H) Albumin secretion of TFSiHeps which were cultured for 0, 6, 12 days after FCM were evaluated by ELISA. *p < 0.05.
Finally, we evaluated how these hepatic characteristics change after the transgene removal at mRNA and protein level. We performed qPCR for Hnf4a, Foxa3 and other representative hepatic markers as Alb, Cdh1, Cyp3a11, G6Pase and AFP (Fig. 5G), and also performed ELISA for measuring albumin (Fig. 5H) from TFSiHeps which were cultured for 0, 6, 12 days after FCM, respectively. After 12 days of culture, all genes still maintained higher expression level compared with TFSpPBd-TCs cultured for 12 days. Although a significant decrease of expression was observed in Foxa3 compared with transgene-integrated iHeps, no significant change was observed during the culture period, indicating that expression profile was maintained after spheroid formation. No significant difference was also observed in albumin secretion between the three time points.
These results suggest that even after transgene removal, TFSiHeps maintained the same level of hepatocyte function as monolayer transgene-integrated iHeps.
Discussion
iHeps are expected to be an alternative to hepatocytes in many situations27, such as a serving as a cell source for transplantation in severe liver dysfunction, in vitro disease modeling, pharmacokinetic analyses, and differentiation or regeneration modeling in the hepatic lineage. In the initial reports on mouse iHeps, Suzuki et al.12 transduced Hnf4a along with Foxa1, Foxa2, or Foxa3 as reprogramming factors into mouse fibroblasts using a retrovirus, and Huang et al.13 transduced Gata4, Hnf1a, and Foxa3 using a lentivirus. Both research groups confirmed that the iHeps acquired hepatic function and exhibited supportive effects on transplantation in a Fah−/− mouse model. Thereafter, various research groups successfully generated human fibroblast-derived iHeps28,29,30,31, and progress also occurred in the development of culture protocols and the search for suitable reprogramming factors. However, most studies on direct reprogramming utilize integrative viral vectors. These vectors carry in vitro risks such as insertional mutagenesis and genotoxicity14,15 as well as in vivo risks such as unexpected reactivation of the transgene and virus-related toxicity32,33. Therefore, a safer reprogramming vector is essential if iHep-related cell therapy is to be applied in human medicine in the future. The piggyBac transposon vector system used in this study is a non-viral vector that provides efficient integration. It has several advantages such as constant gene expression as well as retention of the transgene in the transfected cells even over long culture periods or repeated passages, which should enable antibiotic selection. Owing to its suitability for reprogramming, piggyBac has also been applied to generate iPSCs33,34,35 and has recently attracted attention as an effective vector for genetic engineering36.
To our knowledge, this is the first report to demonstrate iHep generation using piggyBac-based direct reprogramming. In this study, we showed that forced expression of an integrated transgene containing Hnf4a and Foxa3 successfully reprogrammed MSCs into hepatocyte-like cells. The obtained iHeps exhibited hepatocyte-specific gene expression and protein production. The iHeps also exhibited hallmarks of mature hepatocyte function, such as lipid accumulation, glycogen storage, and ICG uptake. However, the maintenance of proliferative capability and expression of hepatic progenitor markers, such as AFP and CK19, suggest that the iHeps possess the characteristics of premature hepatocytes as well.
MSCs have been investigated as a possible source of cells that can acquire hepatic characteristics when cultured in specific conditions5; therefore, we speculated that they would be appropriate for generating iHeps by direct reprogramming. In the present study, MSCs without overexpression of reprogramming factors were treated as mock-transfected cells and exhibited limited expression of hepatocyte-specific genes; additionally, they did not exhibit any function of maturated hepatocytes in this differentiation protocol, even in the form of spheroids. In contrast, when reprogramming factors were overexpressed in MSCs, they manifested mature hepatocyte function, indicating that efficient reprogramming was achieved.
We also demonstrated that fluorescence-negative iHeps, which were confirmed to be transgene-free iHeps, still exhibited hepatocyte function in spheroidal form comparable to that of transgene-integrated iHeps. Moreover, we showed that hepatic profile of transgene-free iHeps was maintained in the form of spheroid. Although the expression of Foxa3 decreased as expected after the exogenous expression was lost, Hnf4a did not show significant decrease. The persistent endogenous expression of Hnf4a is considered to be one of the reasons for maintaining hepatocyte function, because Hnf4a plays a central role for the maintenance of hepatocyte differentiation37,38.
It is also suspected that the reprogrammed cells that underwent gene removal and FCM processes were susceptible to a decrease of viability and acquired hepatocyte function, whereas the cell-cell interactions in the spheroidal iHeps supported the maintenance of viability and promotion of differentiation into hepatocytes39,40,41. Thus, 3D culture environments including spheroids are expected to contribute to the maintenance and maturation of iHeps as reported for MSCs, iPSCs, and ESCs42,43,44, which could be an interesting avenue for future research. Although the present study utilized a simple model with 2 reprogramming factors, other factor combinations can also be used in the piggyBac system to achieve iHeps with higher levels of hepatocyte functions; as many as 6 factors have been reported4 (HNF1A, HNF4A, HNF6, ATF5, PROX1, and CEBPA).
Because risk-reduced transgene-free iHeps are expected to be useful for achieving therapeutic effects against liver disorder, studies to transplant these iHeps into the liver disorder models are desirable. In general, a large number of cells are required for cell transplantation, but cells do not proliferate in spheroidal form, which makes it difficult for the present model to supply abundant iHeps for cell transplantation. To overcome this problem, exploration of culture conditions that support the proliferation of transgene-free iHeps while maintaining hepatocyte functions is necessary. For example, the above mentioned 3D culture environment could serve as an alternative to spheroids, and progress in cell culture techniques with small compounds or cytokines may further support such efforts.
In conclusion, we suggest that the piggyBac-mediated method is a novel and promising approach to generate non-viral, transgene-free iHeps. Further advancement in achieving higher function and optimal culture conditions will facilitate the application of iHeps for upcoming cell therapies against severe liver dysfunction.
Methods
Preparing MSCs
C57BL/6 mouse bone marrow-derived mesenchymal stem cells (MSCs), passage 6, were purchased from Cyagen Biosciences. MSCs were seeded and expanded on collagen 1-coated dishes (AGC TECHNO GLASS) in DMEM containing 10% FBS (Invitrogen) and 1% penicillin/streptomycin (Invitrogen). MSCs of passage 8–9 were used for experiments.
MSCs underwent adipogenic and osteogenic differentiation steps using differentiation kits (Cyagen) and the protocol was carried out according to the manufacturer’s instructions. In brief, for adipogenesis, adipogenic differentiation medium was fed for three days, followed by MSC maintenance medium for a day, and repeat this process for three times. Then cells were cultured in MSC maintenance medium for seven days. Differentiated cells were evaluated by Oil Red O staining. For osteogenesis, cells were cultured in osteogenic differentiation medium for three weeks and observed after Arizarin Red S staining. MSCs were certified to exhibit a cell surface marker pattern positive for CD29, CD44, CD31 and Sca-1 and negative for CD117 by flow cytometry (FCM) by manufacturer.
Vector construction
PiggyBac dual promotor transposon vector (pPBd) along with piggyBac transposase expressing vector (pPBase) were purchased from System Biosciences. pPBd has multi-cloning site (MCS) down stream of CMV promotor, and mRuby and puro down stream of EF1a promotor.
Hnf4a and Foxa3 cDNA were obtained from mouse total mRNA by reverse transcription PCR. Then both of them were linked together via T2A sequence by recombinant PCR to construct Hnf4a-2A-Foxa3 segment, adding Xba1 and BamH1 sequence on each end. This segment was inserted into Xba1/BamH1 restriction site among MCS of pPBd by digestion and ligation, resulting in pPB-Hnf4a-2A-Foxa3-mRubyPuro (pPBHF3).
Cell culture/generation of transgene-integrated iHeps
pPBHF3 and pPBase were co-transfected into MSCs with Lipofectamine 3000 (Invitrogen). pPBd and pPBase were transfected into MSCs in the same manner as mock samples (pPBd-TCs). After medium was changed, puromycin was added for antibiotic selection and when cells reached confluent the differentiation step was initiated.
For the first twelve days, as step A, cells were cultured in expansion Medium consisted of DMEM/F-12 (Invitrogen) containing 10% fetal bovine serum (FBS) (Invitrogen), 1% penicillin/streptomycin (Wako), 10 mM nicotinamide (Sigma Aldrich), 100 μM 2-melcaptomethanol (Wako), 100 μM ascorbic acid (Wako), 1 μM Dexamethasone (Dex) (Wako) and ITS (BD Biosciences), supplemented with 20 ng/ml hepatocyte growth factor (HGF) (R&D Systems) and 20 ng/ml epidermal growth factor (EGF) (R&D systems). Then step B was initiated with maturation medium consisted of IMEM + GlutaMax (Invitrogen) containing 0.5% bovine serum albumin (BSA) (Wako), 1% penicillin/streptomycin, 10 mM nicotinamide, 100 μM 2-melcaptomethanol, 500 μM ascorbic acid, 1 μM Dex and ITS, supplemented with 20 ng/ml oncostatin M (OSM) (R&D Systems) and 20 ng/ml EGF. For the first three days of the differentiation step, 200 uM 5′-azacytidine (5′-aza) was added to expansion medium.
Generation of transgene-free iHeps
pPBase alone were transfected into iHeps with ViaFect (Promega) and cells were expanded until confluence for a few days. Then iHeps which were negative for red fluorescence were sorted by FCM, detected in PE-Texas Red filter. We prepared cells as described previously. The threshold level for red fluorescence positive was determined as the level under which less than 0.05% of the transgene-integrated cells, which did not undergo transgene-removal process, remained. Sorted cells were reseeded onto low attachment 96well V bottom plate (SUMILON) at 1 × 104 cells per well to form spheroids. After 1 week of culture, TFSiHeps along with TFSpPBd-TCs were collected for mRNA extraction, staining and ELISA, and culture medium for urea measurements. The image of whole protocol is depicted in Supplementary Fig. S2.
Primary hepatocyte isolation and culture
Adult C57BL/6 mice were subjected to two-step collagenase perfusion for isolation of primary hepatocytes as described previously4,45. Briefly, after laparotomy portal veins were cannulated with 25-gauge needles. Through these needles, the livers were preperfused with 10 ml Hanks-EGTA. Subsequently, the livers were perfused with 15 ml collagenase solution containing 0.3% dispase II (Sanko Junyaku) and 0.3% collagenase II (Gibco). Then the livers were minced with a scissor and filtered thorough 50 N polypropylene mesh, and the suspension was centrifuged two times at 50 g for 4 min. Cell pellets were resuspended and cultured with HBM (Lonza) onto collagen 1-coated dish for two days for RNA extraction or other experiments. For spheroid formation cells were seeded into 96well V bottom plate and cultured for four days until analyses.
PCR/Quantitative PCR
PCR and RT-PCR assays were performed as previously described46,47,48. Total RNA was extracted with PureLink RNA mini kit (Thermo Fischer Scientific) and then reverse-transcribed into cDNA with Revertra Ace (Toyobo). qPCR was performed with Power SYBR Green (Applied Biosystems). The quantification of the expression of genes were evaluated as the relative level normalized to endogenous Actin beta. Primer sequences for PCR analyses are listed in Supplementary Table S1.
Immunofluorescent staining
The cultured cells were fixed and stained as previously described49. Primary antibodies were as follows; mouse anti-human Hnf4a (PPMX), goat anti-human Foxa3 (Santa Cruz), goat anti-mouse albumin antibody (Bethyl Laboratories), mouse anti-mouse E-cadherin (Abcam), mouse anti-mouse cytchromeP450 1A2 (Abcam) at 1:200 respectively. The sections were incubated for two hours with secondary antibodies; Alexa 488-conjugated rabbit anti-goat IgG or goat anti-mouse IgG at 1:1000. After washing, the stained sections were covered with Vectashield mounting medium with DAPI (Vector Laboratories).
Spheroids were first embedded in iPGel (NIPPON Genetics) and fixed with 4% PFA overnight, then embedded in paraffin and cut in 5 μm thick. After deparaffinization and antigen retrieval using a Target Retrieval Solution (Dako), nonspecific binding was blocked with 1% bovine serum albumin (Sigma Aldrich) dissolved in 0.1% Polyoxyethylene Octylphenyl Ether (Wako) in PBS. The primary and secondary antibodies reactions were carried out in the same way for cultured cells.
Measurements of albumin and urea
Cells on 35 mm collagen1-coated dishes and spheroids in 96 well plate were incubated in newly fed maturation medium, and culture supernatants were collected. For albumin measurement, culture medium for 24 hour was collected. The quantity of albumin was calculated using mouse albumin ELISA kit (Bethyl Labolatories) and urea using QuantiChrom Urea Assay Kit (BioAssay Systems), each according to the manufacturer’s instructions.
Oil Red O staining, Periodic Acid-Schiff (PAS) staining, ICG uptake assay
For Oil Red O staining, linoleic acid (Wako) was added into the medium at 10 mg/ml and incubated for 30 minutes. After washed with PBS, cells were fixed with 4% PFA at room temperature for 30 minutes and washed with 60% isopropanol, followed by incubation for 30 minutes with Oil Red solution which Oil Red powder (Sigma Aldrich) was dissolved into isopropanol and diluted to 60% with distilled water. Then cells were washed with distilled water three times. For PAS staining, cultured cells were stained by periodic acid-Schiff (MUTO PURE CHEMICALS) following the manufacturer’s instructions. For paraffin-embedded spheroids, the same procedure was carried out after deparaffinization. For ICG uptake assay, indocyanine green (Wako) was added in the medium at 0.1 mg/ml and incubated for 1 hour, followed by washing with PBS three times.
Statistics
The results are expressed as mean ± SEM. Statistical significance was assessed using unpaired t-test; p-values < 0.05 were considered statistically significant. Data analysis was performed by GraphPad Prism. For all statistics, data from at least three independent samples were used.
Additional Information
How to cite this article: Katayama, H. et al. Generation of non-viral, transgene-free hepatocyte like cells with piggyBac transposon. Sci. Rep. 7, 44498; doi: 10.1038/srep44498 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Strom, S. C. et al. Hepatocyte transplantation as a bridge to orthotopic liver transplantation in terminal liver failure. Transplantation 63, 559–569 (1997).
Ohashi, K., Park, F. & Kay, M. A. Hepatocyte transplantation: clinical and experimental application. J. mol. med. 79, 617–630 (2001).
Puppi, J. et al. Improving the techniques for human hepatocyte transplantation: report from a consensus meeting in London. Cell Transplant. 21, 1–10 (2012).
Kita, S. et al. The Protective Effect of Transplanting Liver Cells Into the Mesentery on the Rescue of Acute Liver Failure After Massive Hepatectomy. Cell Transplant. 25, 1547–1559 (2016).
Lee, K. D. et al. In vitro hepatic differentiation of human mesenchymal stem cells. Hepatology 40, 1275–1284 (2004).
Shu, S. N. et al. Hepatic differentiation capability of rat bone marrow-derived mesenchymal stem cells and hematopoietic stem cells. World J. Gastroentero. 10, 2818–2822 (2004).
Banas, A. et al. Adipose tissue-derived mesenchymal stem cells as a source of human hepatocytes. Hepatology 46, 219–228 (2007).
Soto-Gutiérrez, A. et al. Reversal of mouse hepatic failure using an implanted liver-assist device containing ES cell–derived hepatocytes. Nat. Biotechnol. 24, 1412–1419 (2006).
Gouon-Evans, V. et al. BMP-4 is required for hepatic specification of mouse embryonic stem cell–derived definitive endoderm. Nat. Biotechnol. 24, 1402–1411 (2006).
Si-Tayeb, K. et al. Highly efficient generation of human hepatocyte-like cells from induced pluripotent stem cells. Hepatology 51, 297–305 (2010).
Sullivan, G. J. et al. Generation of functional human hepatic endoderm from human induced pluripotent stem cells. Hepatology 51, 329–335 (2010).
Sekiya, S. & Suzuki, A. Direct conversion of mouse fibroblasts to hepatocyte-like cells by defined factors. Nature 475, 390–393 (2011).
Huang, P. et al. Induction of functional hepatocyte-like cells from mouse fibroblasts by defined factors. Nature 475, 386–389 (2011).
Nair, V. Retrovirus-induced oncogenesis and safety of retroviral vectors. Curr. Opin. Mol. Ther. 10, 431–438 (2008).
Schambach, A. et al. Lentiviral vectors pseudotyped with murine ecotropic envelope: increased biosafety and convenience in preclinical research. Exp. Hematol. 34, 588–592 (2006)
Stadtfeld, M., Nagaya, M., Utikal, J., Weir, G. & Hochedlinger, K. Induced pluripotent stem cells generated without viral integration. Science 322, 945–949 (2008).
Okita, K., Nakagawa, M., Hyenjong, H., Ichisaka, T. & Yamanaka, S. Generation of mouse induced pluripotent stem cells without viral vectors. Science 322, 949–953 (2008).
Simeonov, K. P. & Uppal, H. Direct reprogramming of human fibroblasts to hepatocyte-like cells by synthetic modified mRNAs. PLOS ONE 9, e100134 (2014).
Davoodian, N., Lotfi, A. S., Soleimani, M. & Mowla, S. J. MicroRNA-122 Overexpression Promotes Hepatic Differentiation of Human Adipose Tissue-Derived Stem Cells. J. Cell. Biochem. 115, 1582–1593 (2014).
Kim, J. et al. Generation of integration-free induced hepatocyte-like cells from mouse fibroblasts. Sci. Rep. 5 (2015)
Ding, S. et al. Efficient transposition of the piggyBac (PB) transposon in mammalian cells and mice. Cell 122, 473–483 (2005).
Prak, E. T. L. & Kazazian, H. H. Mobile elements and the human genome. Nat. Rev. Genet. 1, 134–144 (2000).
Cary, L. C. et al. Transposon mutagenesis of baculoviruses: analysis of Trichoplusia ni transposon IFP2 insertions within the FP-locus of nuclear polyhedrosis viruses. Virology 172, 156–169 (1989).
Wu, S. C. et al. piggyBac is a flexible and highly active transposon as compared to Sleeping Beauty, Tol2, and Mos1 in mammalian cells. P. Nat. Acad. Sci. USA 103, 15008–15013 (2006).
Elick, T. A., Bauser, C. A. & Fraser, M. J. Excision of the piggyback transposable element in vitro is a precise event that is enhanced by the expression of its encoded transposase. Genetica 98, 33–41 (1996).
Ivics, Z. et al. Transposon-mediated genome manipulation in vertebrates. Nat. Methods 6, 415–422 (2009).
Ji, S., Zhang, L. & Hui, L. Cell fate conversion: Direct induction of hepatocyte‐like cells from fibroblasts. J. Cell. Biochem. 114, 256–265 (2013).
Huang, P. et al. Direct reprogramming of human fibroblasts to functional and expandable hepatocytes. Cell stem cell 14, 370–384 (2014).
Du, Y. et al. Human hepatocytes with drug metabolic function induced from fibroblasts by lineage reprogramming. Cell stem cell 14, 394–403 (2014)
Zhu, S. et al. Mouse liver repopulation with hepatocytes generated from human fibroblasts. Nature 508, 93–97 (2014).
Kogiso, T., Nagahara, H., Otsuka, M., Shiratori, K., Dowdy, S. F. et al. Transdifferentiation of human fibroblasts into hepatocyte-like cells by defined transcriptional factors. Hepatol. Int. 7, 937–944 (2013).
Thomas, C. E., Ehrhardt, A. & Kay, M. A. Progress and problems with the use of viral vectors for gene therapy. Nat. Rev. Genet. 4, 346–358 (2003).
Kaji, K. et al. Virus-free induction of pluripotency and subsequent excision of reprogramming factors. Nature 458, 771–775 (2009).
Woltjen, K. et al. piggyBac transposition reprograms fibroblasts to induced pluripotent stem cells. Nature 458, 766–770 (2009)
Yusa, K. et al. Generation of transgene-free induced pluripotent mouse stem cells by the piggyBac transposon. Nat. Methods. 6, 363–369 (2009).
Zhao, S. et al. PiggyBac transposon vectors: the tools of the human gene encoding. Transl. Lung Cancer Res. 5, 120–125 (2016).
Hayhurst, G. P., Lee, Y. H., Lambert, G., Ward, J. M. & Gonzalez, F. J. Hepatocyte nuclear factor 4α (nuclear receptor 2A1) is essential for maintenance of hepatic gene expression and lipid homeostasis. Mol. Cell. Biol. 21, 1393–1403 (2001).
Kyrmizi, I. et al. Plasticity and expanding complexity of the hepatic transcription factor network during liver development. Genes Dev. 20, 2293–2305 (2006).
Tong, J. Z., Sarrazin, S., Cassio, D., Gauthier, F., Alvarez, F. et al. Application of spheroid culture to human hepatocytes and maintenance of their differentiation. Biol. Cell 81, 77–81 (1994).
Subramanian, K., Owens, D. J. & O’Brien, T. D. Enhanced differentiation of adult bone marrow-derived stem cells to liver lineage in aggregate culture. Tissue Eng. Pt. A 17, 2331–2341 (2011).
Berger, D. R., Ware, B. R., Davidson, M. D., Allsup, S. R., Khetani, S. R. et al. Enhancing the functional maturity of induced pluripotent stem cell–derived human hepatocytes by controlled presentation of cell–cell interactions in vitro . Hepatology 61, 1370–1381 (2015).
Ji, R. et al. The differentiation of MSCs into functional hepatocyte-like cells in a liver biomatrix scaffold and their transplantation into liver-fibrotic mice. Biomaterials 33, 8995–9008 (2012).
Takayama, K. et al. 3D spheroid culture of hESC/hiPSC-derived hepatocyte-like cells for drug toxicity testing. Biomaterials 34, 1781–1789 (2013).
Gieseck, R. L. III et al. Maturation of induced pluripotent stem cell derived hepatocytes by 3D-culture. PloS ONE 9, e86372 (2014).
Ogiso, S. et al. Efficient recellularisation of decellularised whole-liver grafts using biliary tree and foetal hepatocytes. Sci. Rep. 6, 35887 (2016).
Kawai, T. et al. SOX9 is a novel cancer stem cell marker surrogated by osteopontin in human hepatocellular carcinoma. Sci. Rep. 6, 30489 (2016).
Kawai, T. et al. Identification of keratin19-positive cancer stem cells associating human hepatocellular carcinoma using 18F-fluorodeoxyglucose positron emission tomography. Clin. Cancer Res. doi: 10.1158/1078-0432.CCR-16-0871 (2016).
Kawai, T. et al. Keratin 19, a Cancer Stem Cell Marker in Human Hepatocellular Carcinoma. Clin. Cancer Res. 21, 3081–3091 (2015).
Ishii, T. et al. Transplantation of embryonic stem cell-derived endodermal cells into mice with induced lethal liver damage. Stem Cells 25, 3252–3260 (2007).
Acknowledgements
This work was supported by grants from the Scientific Research Fund of the Japan Science and Technology Agency (Research Project Number: 24591999).
Author information
Authors and Affiliations
Contributions
Conception and design; H.K., K.Y. and T.I. Acquisition of data; H.K. and K.Y. Analysis and interpretation of data; H.K., K.Y., N.S, K.F., J.K. and T.I. Writing, review, and/or revision of the manuscript; H.K., K.Y. and S.U. Statistical analysis; H.K., K.Y. and T.I. Administrative, technical, or material support; Y.M., H.K., R.Y., T.K., E.Y.Y., S.O., S.K. and K.Y. Study supervision; K.Y., N.S., K.F., J.K., T.I. and S.U.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Katayama, H., Yasuchika, K., Miyauchi, Y. et al. Generation of non-viral, transgene-free hepatocyte like cells with piggyBac transposon. Sci Rep 7, 44498 (2017). https://doi.org/10.1038/srep44498
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep44498
This article is cited by
-
Phenotype instability of hepatocyte-like cells produced by direct reprogramming of mesenchymal stromal cells
Stem Cell Research & Therapy (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.