Abstract
The pig is recognized as a valuable model in biomedical research in addition to its agricultural importance. Here we describe a means for generating skeletal muscle efficiently from porcine induced pluripotent stem cells (piPSC) in vitro thereby providing a versatile platform for applications ranging from regenerative biology to the ex vivo cultivation of meat. The GSK3B inhibitor, CHIR99021 was employed to suppress apoptosis, elicit WNT signaling events and drive naïve-type piPSC along the mesoderm lineage, and, in combination with the DNA methylation inhibitor 5-aza-cytidine, to activate an early skeletal muscle transcription program. Terminal differentiation was then induced by activation of an ectopically expressed MYOD1. Myotubes, characterized by myofibril development and both spontaneous and stimuli-elicited excitation-contraction coupling cycles appeared within 11 days. Efficient lineage-specific differentiation was confirmed by uniform NCAM1 and myosin heavy chain expression. These results provide an approach for generating skeletal muscle that is potentially applicable to other pluripotent cell lines and to generating other forms of muscle.
Similar content being viewed by others
Introduction
One of the main driving forces for pursuing studies on porcine pluripotent stem cells has been to create tissue precursors in vitro that might be used to explore engraftment procedures in an animal model with close physiological and morphological similarity to the human1. Unfortunately, true embryonic stem cells from agriculturally important large animals, including the pig, have proven difficult to generate, and piPSC, which are readily created, have only rarely been used for directed differentiation purposes2. As far as we are aware, for example, no studies to date have described differentiation of piPSC along the mesoderm lineage and overcoming the barrier to myogenesis, despite the widespread interest in using the pig in modeling infarction and cardiovascular disease, muscle regeneration and muscle biology3,4,5. In addition, skeletal muscle tissue is the major constituent of pork, and piPSC are a potential source of self-renewing, skeletal muscle progenitors for creating cultured meat and other livestock products6. Thus, the molecular control mechanisms directing porcine skeletal muscle development have considerable implications for medicine, agriculture and food technology.
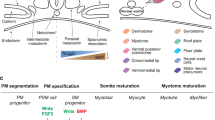
From the earliest stages of embryonic development through terminal differentiation, skeletal myogenesis is directed by CTNNB1 (β-catenin) and GSK3B (glycogen synthase kinase-3β). Phosphorylation at the targeted motif of CTNNB1 by GSK3B marks the protein for proteasomal degradation, while hypophosphorylated CTNNB1 can either stably associate with the cell membrane or form complexes with lymphoid enhancer binding factor (LEF1, sometimes known as TCF) to activate an expression suite of target genes within the nucleus7. Multiple signaling pathways converge at GSK3B to modulate CTNNB1 activity during embryonic development and myogenesis. WNT signaling, for example, can activate CTNNB1 by GSK3B inactivation8. During epiblast gastrulation, early mesodermal patterning is mediated by the transcription factor T (BRACHYURY)9. Like T, expression of a premyogenic transcription factor, PAX3, is directed by WNT signaling10,11,12,13 and is CTNNB1-activated14,15,16. PAX3 is expressed initially in the pre-segmented paraxial mesoderm, and later increases in the somite17 where, in combination with other factors, a NOTCH signaling checkpoint prevents precocious myotome maturation18. Subsequent WNT signaling19,20,21 directs sequential expression of the functionally overlapping22, but distinct myogenic regulatory factors (MRFs), MYF5 and MYOD1, signifying commitment to the skeletal muscle lineage. WNT and IGF signaling23,24,25,26,27, coupled with repression of activin receptor-like kinase (ALK) activity28,29, support terminal differentiation, marked by cell-cycle exit and cell fusion into multinucleated skeletal myofibers capable of generating mechanical force by contraction. To evaluate the potential of piPSC to give rise to mesodermal lineages and skeletal muscle, in particular, we activated developmental pathways characterized in these human and mouse studies to support lineage specification of piPSC to skeletal muscle in vitro, and assessed the capacity of the lineage-specified precursor cells to differentiate terminally into contractile myotubes.
Results
CHIR99021 attenuates apoptosis following ground-state withdrawal
A naïve piPSC line derived from the inner cell mass of a porcine blastocyst that expressed POU5F1 and KLF4 reprogramming genes from a doxycycline (DOX)-inducible promoter (Supplementary Fig. 1) was employed throughout30. As with naive mouse embryonic stem cells (ESC)31, naïve piPSC culture was supported by a medium (Renewal Medium, RM-1; Supplementary Table 1) containing CHIR99021/PD032591/PD173074 (inhibitors of GSK3B/MEK/FGF signaling) and LIF under a 5% O2, 5% CO2, 90% N2 atmosphere. To identify conditions that might permit differentiation from the ground state, piPSC colonies were dissociated to single cells and then sub-cultured on a poly-D-lysine, laminin and gelatin (PLG) substratum in RM-1 containing DOX in order to maintain expression of the two reprogramming genes, which are essential for self-renewal in this cell line. By three days, the cells had formed colonies (Fig. 1a, left panel). To support differentiation, the RM-1 renewal medium was replaced with a differentiation medium (DM-1; Table S1) formulated without DOX, LIF and CHIR99021/PD032591/PD173074. The cultures were transferred to a 20% O2, 5% CO2, 75% N2 atmosphere. Extensive cell death was observed over the two days following the transition (Fig. 1a, right panel). Increased cell-surface labeling by ANXA5 (Fig. 1b, Supplementary Fig. 2) and onset of CPP32 cleavage (Fig. 1c) indicated that cell death was linked to an intrinsic apoptosis pathway.
(a) Morphology of ground-state piPSC colonies cultured for three days from single cells in renewal conditions (RM-1, left panel) and following 48 hour (h) differentiation (RM-2, right panel). (b) ANXA5 (annexin V) labeling of apoptotic cells prior to, and following, 24 h differentiation. Error bars represent n = 3 experimental replicates ± SD. (c) Western blot detection of full-length CPP32 (~32 kD procaspase 3a) and the large cleaved fragment (~17 kD cleaved-caspase 3a) prior to (0 h) and following (12–48 h) differentiation. (d) Phase contrast images of adherent (upper panels) and detached colonies (lower panels), following 48 h differentiation + 1–9 μM CHIR99021, as indicated. (a,d) Scale bars, 500 μM. (e) Percent cell adhesion following 48 h differentiation without, as shown in (a), right panel, or with 1–9 μM CHIR99021, as shown in (d). *viable cells within detached colonies unaccounted. Error bars represent n = 3 experimental replicates ± SEM. (f) ANXA5 labeling of apoptotic cells following 24 h differentiation ± CHIR99021, as indicated. Error bars represent n = 3 experimental replicates ± SEM. *Significance (P < 0.05) determined by t-test. (g) Western blot detection of full-length and cleaved CPP32 following 42 h differentiation ± CHIR99021, as indicated. (c,g) TUBA detected as an internal protein loading control. Western blots were cropped for clarity. Examples of uncropped blots are found in Supplementary Figure 9.
Next, retention of CHIR99021 was tested as an approach to prevent the massive amount of cell death encountered in its absence. In these experiments, colonies were cultured in differentiation medium (DM-1) supplemented with CHIR99021. Improved cell survival and cell adhesion was observed over the full range (1–9 μM) of CHIR99021 concentrations tested. All concentrations between 3–9 μM supported formation of attached, rounded colonies, but, at the higher (6–9 μM) concentration cell aggregates break free from the colonies to form floating structures resembling embryoid bodies (Fig. 1d). The improved adhesion to the matrix observed between 3 to 6 μM CHIR99021 (Fig. 1e), plus the reduced presence of cell-surface ANXA5 (Fig. 1f, Supplementary Fig. 3) and intracellular, cleaved CPP32 (Fig. 1g), indicated that CHIR99021 had attenuated apoptosis.
CHIR99021 activates WNT signaling events during piPSC differentiation
To determine whether CHIR99021/DM-1 activated canonical WNT signaling events, relative phospho- and total CTNNB1 was monitored after transfer of piPSC to DM-1 culture medium with and without CHIR99021. As anticipated, CHIR99021 exposure (3–9 μM) resulted in a dose-dependent increase of CTNNB1. CHIR99021 exposure also increased CTNNB1 phosphorylation at the GSK3B-targeted motif, but there appeared to be less accumulation of phospho-CTNNB1 at concentrations above 3 μM (Fig. 2a).
(a) Upper Panel: Western blot detection of CTNNB1 and p-CTNNB1 (total and phospho-S33,37,T41 β-catenin, respectively) following one day differentiation ± CHIR99021, as indicated. Middle Panel: ratios of p-CTNNB1/CTNNB1 bands. Lower Panel: ratios of CTNNB1/TUBA bands. Middle and Lower Panels: Error bars represent n = 3 experimental replicates ± SEM. *Significance (P < 0.05) determined by t-test. (b) Adhesion and outgrowth morphology following transfer of day (d) 2 (Fig. 1d) detached colonies differentiating in 6 μM CHIR99021/DM-1 to PLM-coated culture dishes, for one (d3, left panel) or three (d5, right panel) additional days. Scale bar, 500 μM. (c) Western blot analysis of T, PAX3, POU5F1 and KLF4 expression in ground-state (d0) and differentiating cultures (d1–d5) during the regimen described in (b). (a,c) TUBA detected as an internal protein loading control. Western blots were cropped for clarity. Examples of uncropped blots are found in Supplementary Figure 9.
CTNNB1 stabilization and morphological changes observed during CHIR99021/DM-1 culture suggested that mesodermal determination might have been initiated. To investigate this possibility, embryoid body-like structures from day 2 of differentiation (Fig. 1d) were transferred to poly-D-lysine, laminin and Matrigel (PLM) substratum in CHIR99021/DM-1, where they adhered and formed outgrowths (Fig. 2b). Expression of ground-state pluripotency markers POU5F1 and KLF4, and mesodermal markers T (BRACHYURIA) and PAX3, were monitored over six days, from ground-state colony formation (day 0), to detachment of differentiating colonies (days 1–2), to adherence and outgrowth of detached colonies (days 3–5). Expression of POU5F1 and KLF4 were down-regulated to basal levels by day 3. An antibody against human T detected two ~49 kD bands within a migration range expected for the protein. The faster migrating isoform was detected at day 0, increased by day 2, and was maintained through day 5. Expression of a higher Mr isoform was observed on days 1 and 2, and then declined, the significance of which is unclear. Transient expression of PAX3 was observed on days 3–5 (Fig. 2c). Together, these features suggested that the cultures had attained a mesodermal state primed for myogenic induction.
Efficient myogenic induction of piPSC
Commitment to the skeletal muscle lineage is directed through transcription program activation by MYF5, MYOD1 or both. Ectopic MYOD1 expression or exposure to 5-aza-cytidine (5AC) has been reported to activate the endogenous myogenic program in certain cell lines32,33. To impart ectopic MYOD1 activity, the parental piPSC line was modified by integration of a selectable cassette expressing an estrogen receptor (ESR1) ligand-binding domain insertion within a MYOD1 open reading frame (MyoDER) (Fig. 3a). The expressed MyoDER protein can be activated conditionally by 17β-estradiol (E2) (Supplementary Fig. 1)34.The modified piPSC (piPSC-M) expressed a ~75 kD MyoDER fusion protein (Fig. 3a), and, in the absence of the E2 agonist, continued to form compact, refractive colonies characteristic of the parental colonies (Supplementary Fig. 4).
(a) piPSC modification by an integrated MyoDER expression cassette. Left panel: blasticidin (BLAST)-selectable MyoDER expression cassette. Arrows and boxes indicate promoter and gene sequences, respectively. Right panel: Western blot detection of MyoDER in the unmodified piPSC and the expression cassette-modified piPSC-M. MyoDER was detected with an antibody raised against a mouse MYOD1 peptide. (b) Morphology of differentiating piPSC-M cultures following expansion and differentiation regimens shown in Supplementary Figure 5a. Scale bar, 100 μM. (c) Western blot detection of MYOD1 antigens, MYF5 and MYOG following piPSC-M expansion and differentiation regimens shown in Supplementary Figure 5a. Double arrowheads: partially resolved MYOG isoforms. (d,e) Determinants of MYF5 activation: Western blot analyses. (d) Detection of MYF5 and CPP32 following piPSC differentiation regimens, as shown in Supplementary Figure 5b. (e) Detection of MYF5 following three days of piPSC-M culture in alternate (RM-1, RM-2) renewal or expansion conditions. (a,c,d,e) TUBA detected as an internal protein loading control. Western blots were cropped for clarity. Examples of uncropped blots are found in Supplementary Figure 9. (f) Immunofluorescent detection of nuclei (DAPI) and NCAM1 (AlexaFluor 568) in differentiating cultures prior to (d0, left panel) and following (d2, right panel) myogenic induction (RM-1→IM-1). Scale bar, 150 μM.
Dosages of 5AC and E2, alone and in combination, were then screened for toxicity and effects on cell morphology. Although 5AC exposure reduced colony growth across the dosage range tested (Supplementary Fig. 4), a 250 nM concentration was tolerated sufficiently well to provide viable cells for subsequent differentiation steps, and was selected for use in future procedures. Since a PLM substratum supported cell adhesion by differentiating piPSC and expression of PAX3 (Fig. 2b,c), we conjectured that it might also support skeletal muscle differentiation in CHIR99021/DM-1. Therefore, 3 μM CHIR99021, a concentration that supported cell adhesion, survival (Fig. 1d–g), and CTNNB1 phosphorylation (Fig. 2a) was added to the DM-1 to facilitate differentiation in the presence of 5AC and/or E2. As shown in Supplementary Figure 5a, piPSC-M were first expanded in RM-1 ± 5AC on a PLM substratum for three days, followed by culture in CHIR99021/DM-1 supplemented with combinations of 5AC and E2 for two additional days.
E2 (5–20 μM) induced a bipolar morphology characteristic of skeletal myocytes. 5AC enhanced the E2 effects, but, on its own, failed to induce this phenotype (Fig. 3b). Both 5AC and E2 increased MyoDER protein abundance in the cells (Fig. 3c). Expression of the endogenous MYOD1 protein (~45 kD) was not observed. MYF5 expression was activated by 5AC, independently of E2. In contrast, MYOG expression was activated by E2, independently of 5AC. However, relative MYOG abundance in the E2-induced cultures was reduced by increasing the E2 concentration. MYOG migrated as two molecular weight forms, with the faster migrating band favored by the combination of 5AC and E2. Together, these data suggested that, in combination with 5AC, induction of MYOD1 by10 μM E2 might be optimal for skeletal muscle lineage induction. Therefore, a medium containing 10 μM E2 and 3 μM CHIR99021 in DM (induction medium; IM Supplementary Table 1) was subsequently used for myogenic induction.
Our observation that CHIR99021/DM-1 failed to promote MYF5 expression in the absence of 5AC (Fig. 3c) suggested synergistic roles of 5AC and CHIR99021. To examine these roles, unmodified piPSC were expanded for three days on PLM substratum in renewal medium (RM-2, Supplementary Table 1) ±5AC and differentiated for two days in differentiation medium (DM-2, Supplementary Table 1) with combinations of CHIR99021 and 5AC, as shown in Supplementary Figure 5b. A sequential RM-2 expansion and DM-2 differentiation regimen supported detectable, but low MYF5 abundance. In the absence of CHIR99021, 5AC failed to increase MYF5. Rather, the highest MYF5 abundance occurred in medium supplemented with both 5AC and CHIR99021 (Fig. 3d). CPP32 cleavage associated with apoptotic cell death was not elicited under these conditions (compare to Fig. 1a,c).
We next examined whether MYF5 was expressed during the initial three-day expansion regimen. The piPSC were cultured for three days in renewal medium on PLG substratum (renewal conditions) or in renewal medium on PLM substratum plus 5AC (expansion conditions). For comparison, the procedure was conducted in two different renewal media, RM-1 and RM-2 (Supplementary Table 1). As expected, MYF5 was not detected in either renewal condition. However, it was detected in both expansion conditions (Fig. 3e), indicating early myogenic determination in the presence of DOX, LIF and CHIR99021/PD032591/PD173074, prior MyoDER induction by E2.
After a sequential three-day expansion and two-day induction course with 5AC exposure on PLM substratum, piPSC-M progeny exhibited myocyte-like morphology, enhanced ectopic MYOD1 expression, and elevated expression of endogenous MYF5 and MYOG. This regimen led to uniform onset of expression of a skeletal muscle cell-surface glycoprotein, NCAM1, during, but not before, the two-day induction course (Fig. 3f). Collectively, these results demonstrate efficient piPSC specification to the skeletal muscle lineage.
Terminal differentiation of lineage-specified piPSC-derived progeny
To assess the terminal differentiation potential of lineage-specified cells, cultures were maintained for up to six additional days in terminal differentiation medium (TDM) (Fig. 4a). The TDM was formulated without E2, which acted as a pleiotropic antagonist of MYOG expression (Fig. 3c), and with CHIR99021, IGF-1 and an ALK inhibitor, A83–01, to support terminal differentiation (Supplementary Table 1). Expression of DES (desmin), a muscle intermediate filament structural protein, was detected at low levels by day 0, and was up-regulated progressively throughout the induction and terminal differentiation steps (Fig. 4e). By day 4, multinucleated cells with elongated, refractive morphologies characteristic of myotubes were evident (Fig. 4b,d). Otherwise equivalent differentiation regimens excluding 5AC treatment during the expansion and induction steps failed to yield such characteristic morphologies (Fig. 4b, Supplementary Fig. 6). By day 6, myotubes exhibited regional anisotropic patterning (Fig. 4b), uniform expression of myosin heavy chain (MyHC) class proteins (Fig. 4c), and myofibrils with well-defined sarcomeric subunits (Fig. 4g). The average sarcomere length observed was 1.76 ± 0.23 μM SD, n = 21, which is within the resting range documented for metazoan skeletal muscle35. Following induction, MYOG expression was maintained through day 6 and was strongly up-regulated by day 8. Endogenous MYOD1 was not detected over the differentiation course (Fig. 4e). Spontaneous contractions by random myotube populations were observed beginning on day 4 during the terminal differentiation period, a process that led to partial myotube detachment from the substratum (Supplementary Video 1). By day 8, most cells had fused into polyploid myotubes (Fig. 4d, Supplementary Fig. 7), and most nuclei had stopped replicating DNA, indicating that cells had exited from the cell cycle and had committed to terminal differentiation (Fig. 4f, Supplementary Fig. 7).
(a) 11-day culture regimen, with terminal differentiation. (b) Myotube morphology and conformation during a RM-2→IM-2→TDM regimen. Scale bar, 125 μM. (c) Day 0 (d0) and 6 (d6) cultures were stained for nuclei (DAPI) and MyHC class proteins (AlexaFluor 568). (d) Myotube multinucleation. Left panel: terminal differentiation morphology, day 4 (RM-1→IM-1→TDM). Bracketed arrows mark multiple nuclei within single myotubes. Right panel: myonuclei distribution within day 8 (d8) di- and polyploid myotubes. Error bars represent n = 3 experimental replicates ± SD. (c,d) Scale bars, 100 μM. (e) Western blot analyses. MYOD1 antigen, MYOG and DES detection, days 0–8. Double arrowhead: partially resolved MYOG isoforms. TUBA detected as an internal protein loading control. (f) Cell-cycle exit and terminal differentiation. Western blots were cropped for clarity. Examples of uncropped blots are found in Supplementary Figure 9. (d,f) Populations labeled and enumerated as shown in Supplementary Figure 7. Error bars represent n = 3 experimental replicates ± SD. *d3 expansion is equivalent to day 0 (d0) induction, as shown in (a). (g) Transmission Electron Microscopy, day 6 myotubes. Myofibrils and sarcomeric subunits within the myotube sarcolemma. Left panels: Single myofibril within the sarcolemma (upper left panel) and an enlarged inset (red box, lower left panel) to visualize the sarcomere structure. Red bracketed arrows mark the sarcomere length, from Z-line to Z-line. Right panel: multiple, parallel myofibrils within the sarcolemma, marked by arrowheads. Scale bars: upper left panel, 2.0 μM; lower left panel, 0.4 μM; right panel, 0.5 μM.
Spontaneous and stimuli-responsive excitation-contraction coupling of piPSC-derived myotubes
In skeletal muscle, excitation-contraction coupling (ECC) occurs via voltage-dependent Ca2+ release into the cytosol, where dihydropyridine receptors sense depolarization at the plasma membrane and open adjacent ryanodine receptor Ca2+ release channels of the sarcoplasmic reticulum by allosteric interaction36. To characterize the contractile physiology of the differentiating myotubes, intracellular Ca2+ signaling was monitored during terminal differentiation. Asynchronous oscillations of intracellular Ca2+ transient cycles of varied frequency and amplitude were observed in spontaneously contracting populations (Fig. 5a, Supplementary Video 2). Aside from these populations, myotubes remained mechanically quiescent in the absence of external stimuli. Electrical field stimulation (1.0 Hz) elicited both a synchronized Ca2+ transient response corresponding to the stimulation frequency, and elevated cytosolic Ca2+ during field stimulation intervals. During stimulation, the average cyclic period (peak-to-peak) recorded was 1.0 ± 0.02 seconds SD, n = 15 (Fig. 5b, Supplementary Video 3). Electrically-induced Ca2+ transients were observed in the absence of extracellular Ca2+, indicating a voltage-dependent Ca2+ release as the underlying mechanism for the phenomenon (n = 2, data not shown), characteristic of skeletal muscle37. Caffeine, an agonist that triggers cytosolic Ca2+ release through ryanodine receptors, caused an initial high-amplitude, synchronous Ca2+ response followed by asynchronous Ca2+ oscillations (Fig. 5c, Supplementary Video 4). Application of acetylcholine, a native agonist of skeletal muscle contraction, also induced significant Ca2+ signaling within myotube cultures (Fig. 5d, Supplementary Video 5). Single-cell analysis revealed that acetylcholine exposure initiated both a gradual rise in cytosolic Ca2+ and cyclical Ca2+ oscillations (Fig. 5e). Individual acetylcholine responses varied in stimulation lag, amplitude, activity and magnitude (Supplementary Fig. 8), accounting for the high variance in the aggregate data (Fig. 5d).
Myotubes were loaded with the Ca2 + indicator dye Fluo-4 AM at T0 and imaged by high-speed confocal fluorescence microscopy. Dynamic Ca2+ signaling events within single cells or an entire field-of-view (FOV) were recorded and plotted. Ca2+ signal is shown as F1/F0; F1 represents the variable raw signal during the image sequence; F0 represents the constant baseline signal. For (a), F0 corresponds to the lowest signal during the respective trace. For (b–e), F0 corresponds to the respective signal recorded at 0.0 s (seconds). (a) Spontaneous Ca2+ transient activity in regionally active day 6 myotubes. Representative single-cell traces show simultaneous activities in four myotubes coinciding in the same FOV. (b) Ca2+ transient response by mechanically quiescent day 6 myotubes to 1.0 Hz field stimulation. (c) Ca2+ transient response by mechanically quiescent day 6 myotubes to 10 mM caffeine perfusion. (b,c) The response trace shows total signal from the respective FOVs over the duration indicated. (d) Mean 5 s Ca2+ transient interval response following perfusion of mechanically quiescent day 7 myotubes with 100 nM acetylcholine; n = 31 ± SD. *Significance (P < 0.05) determined by t-test. (e) Ca2+ transient response to acetylcholine perfusion by a single myotube represented in (d). The vertical dashed red line marks the threshold activation lag at F1/F0 = 3.0. The horizontal dashed red line marks the maximum signal amplitude (AmpMAX). The horizontal dotted green line marks the baseline signal (F1/F0 = 1).
Discussion
Our hybrid approach to induce skeletal muscle lineage specification by piPSC employs a coordinated application of CHIR99021, 5AC and an ectopically expressed MYOD1. Although each of these factors has been reported to enhance skeletal muscle derivation from pluripotent progenitors in vitro, a combinatorial application provides multiple advantages. A previous study that targeted GSK3B inhibition during mouse ESC differentiation by LiCl exposure included an intermediate cell sorting step to enrich myogenic progenitors prior to terminal differentiation38. More recent studies that used CHIR99021 or the GSK3B inhibitor (2′Z,3′E)-6-bromoindirubin-3′-oxime (BIO) to direct human iPSC differentiation reported sarcomeric development following 21 to 40-day differentiation regimens at varied efficiencies39,40,41,42. In our study, both enhanced MYOG expression and sarcomeres were observed within eleven days (Fig. 4e,g), demonstrating an accelerated, enriched development of NCAM+/MyHC+ cells without cell sorting.
One study has described an enrichment of skeletal muscle markers in differentiating human ESC populations by 5AC treatment43. However, this enrichment was incomplete. Unlike the results of our study (Figs 3c–f, and 4c,e), differentiating populations failed to express MYF5, DES, MyHC or NCAM1, in vitro. Results from multiple studies that have used an ectopic MYOD1 to derive skeletal muscle from pluripotent progenitors have varied. An early study demonstrated that, although ectopic MYOD1 was sufficient to activate MYOG expression in differentiating mouse ESC, an intermediate embryoid body step was required to support expression of MYF5 and MyHC. Moreover, the efficiency of skeletal muscle differentiation was low44. A later study reported myogenic differentiation directed by a Tet-Off MYOD1 cassette-modified mouse ESC line, but neither expression of endogenous MYOD1, nor endogenous MYF5, were confirmed45. Studies that have reported the conversion of human ESC and iPSC to skeletal muscle by ectopic MYOD1 expression in pluripotent cells or their derivatives differentiated in vitro46,47,48,49 have failed to demonstrate the activation of endogenous MYF5 expression.
In this study, we applied the self-renewal capacity of a piPSC line to allow indefinite expansion of myogenic progenitors30. This cell line exhibits certain characteristics of a recently described F-class mouse iPSC50, including a renewal dependence upon DOX-enforced reprogramming gene expression (not shown), and a low-threshold 5AC tolerance (Supplementary Fig. 4). In the absence of DOX, LIF, PD032591 and PD173074, cell survival and differentiation were maintained conditionally by CHIR99021 (Figs 1 and 2), thereby elucidating a fundamental role for GSK3B in cell death during differentiation by naïve piPSC. This study further revealed an unexpected role for 5AC in promoting expression of MYF5 rather than endogenous MYOD1, as previously reported during myogenic conversion of other cell types33. Expression of the MYF5 gene is regulated directly by CTNNB1/LEF1 binding and transcriptional co-activation of the MYF5 epaxial enhancer51. In our study, phosphorylation and stabilization of CTNNB1 during piPSC differentiation was directed by CHIR99021 (Fig. 2a). Moreover, expression of the MYF5 protein was either reduced or not detected (Fig. 3c–e) in the absence of 5AC or CHIR99021 exposure. We speculate that in combination, 5AC and CHIR99021 may activate MYF5 gene expression by stabilizing CTNNB1 interaction with a hypomethylated MYF5 epaxial enhancer. During the ontological progression from embryonic progenitors to skeletal muscle, MYF5 expression precedes MYOD1 activity52. In chronological agreement, MYF5 and its transcription target, DES53, were expressed during our differentiation course (Fig. 4a) by the third day of expansion culture (Figs 3e and 4e), and prior to the activation of ectopic MYOD1 by E2.
A faster-migrating MYOG isoform (Fig. 3c), possibly corresponding to the active, hypophosphorylated protein54, was enriched by 5AC exposure. Moreover, without 5AC exposure, myotube development was impaired (Fig. 4b, Supplementary Fig. 6), implicating a role for 5AC in directing a myogenic identity prior to terminal differentiation. Like myotubes from somatic myogenic progenitors55, piPSC-derived myotubes exhibited spontaneous contractile activities (Supplementary Videos 1, 2). Additionally, these myotubes responded to electrical and chemical stimuli in a manner consistent with the physiological signaling pathways actuating skeletal muscle excitation-contraction coupling, in vivo (Fig. 5b–e, Supplementary Videos 3,4,5). Together, these data provide molecular, cytological and physiological lines of evidence supporting the generation of skeletal muscle during our differentiation regimen.
The methods describe combinatorial approaches to optimize lineage specification of piPSC to skeletal muscle. They demonstrate production of contractile porcine myotubes over an expedited course with efficiency and fidelity in serum-free culture media. These results signify piPSC as a promising alternative to somatic myogenic progenitors, such as satellite cells, which have limited self-renewal properties for various applications that use porcine cells1, and furthermore, validate the use of piPSC for production of porcine skeletal muscle at a potentially unlimited scale. Further studies will be required to customize and extend this approach for additional species and cell lines.
Methods
Cell culture
Derivation of the DOX-inducible piPSC line (O2K) used in this study has been described previously28. For naïve piPSC renewal, cultures were maintained on poly-D-lysinea, mouse lamininb, and porcine gelatina (PLG)-coated culture surfaces under 5% atmospheric O2 in renewal medium (RM) consisting of serum replacement, neurobasal mediumc and DMEM-F12d combined at a 1:1 ratio, 1X non-essential amino acidsa, 0.5X GlutaMAXc, 0.1 mM β-mercaptoethanola, 0.1 mg/mL protease-free bovine serum albumine, 2 μg/mL doxacycline hyclate (DOX)f, 10 ng/mL human leukemia inhibitory factor (LIF)g, 3 μM CHIR99021b, 0.8 μM PD032591h and 0.1 μM PD173074a. Ground-state piPSC colonies were enzymatically dissociated and passaged every three days. For general differentiation, day 3 piPSC colonies were cultured under 20% atmospheric O2 in a basal differentiation medium (DM) formulated as RM, but with β-mercaptoethanol, DOX, LIF, CHIR99021, PD032591 and PD173074 omitted. For myogenic expansion, cultures were maintained as renewal cultures, except that piPSC were cultured on poly-D-lysine, laminin and Matrigeli (PLM)-coated culture surfaces, and RM was supplemented with 250 nM 5-aza-cytidine (5AC)a each day for three days prior to myogenic induction. For myogenic induction, the RM of day 3 myogenic expansion cultures was replaced with induction medium (IM), and cultures were differentiated for two additional days with 5AC supplementation. IM was prepared as DM, but additionally supplemented with 3 μM CHIR99021 and 10 μM 17β-estradiol (E2)a. For terminal differentiation of skeletal muscle lineage-specified cultures following induction, IM was replaced with terminal differentiation medium (TDM) and differentiating cells were cultured for up to six additional days. TDM was prepared as IM, except that E2 was excluded, and the medium was formulated with 100 ng/mL human IGF-1j and 4 μM A83–01k. 15% KnockOut Serum Replacement (KSR)c or a combination of 0.5X N-2 Supplementc + 0.5X B-27 Supplement without Vitamin Ac were alternately substituted as serum replacements. Culture media formulations are summarized in Supplementary Table 1.
piPSC modification
The MyoDER DNA sequence was PCR amplified from the pBabeMDER plasmidl and recombined into the pLentiCMVBlast lentiviral destination vectorl by using a pENTR/D-TOPO entry vectorc. Pseudovirus prepared from supernatants of 293FT cells co-transfected with the modified lentiviral vector, pMD2.G envelope plasmidl and the psPAX2 packaging plasmidl was used for transduction of the piPSC. Stably-transduced piPSC were selected with 10 μg/mL blasticidinm for four days and 15 μg/mL for two additional days. The modified piPSC were designated ‘piPSC-M’ and maintained as the unmodified piPSC, except to avoid unscheduled induction of the MyoDER protein, RM used to culture the piPSC-M was formulated with phenol red-free neurobasalc and DMEM-F12c basal media.
Immunodetection and flow cytometry
For Western blotting, cell lysates were normalized to 50 μg total protein/lane, resolved by 5–15% gradient SDS-PAGE and transferred to a PVDF membrane. Membranes were probed with antibodies against CPP32 (#9662)n, PAX3 (Pax3)°, POU5F1 (#sc-5279)p, KLF4 (#sc-20691)p, CTNNB1 (#8480)n, phospho-CTNNB1 (#9561)n, MYOD1 (#sc-304)p, MYF5 (#sc-302)p, MYOG (F5D)°, DES (#sc-14026)p and TUBA (#A01490)q. PVDF membranes were developed with SuperSignal West Dura Extended Duration Substrater and chemiluminescent images were captured with a Fujifilm LAS-300 CCD camera. To stabilize relative phospho-CTNNB1 to levels sufficient for detection, phosphatase and proteasome activities were inhibited by exposing the cultures to 50 nM calyculin An and 30 μM MG132n for three hours prior to cell lysate preparation. Densitometry data were obtained by using ImageJ image quantification software (version 1.40 g). The AlexaFluor 488 Annexin V/Dead Cell Apoptosis Kit3 was used for ANXA5 labeling of apoptotic cells. To label replicating DNA, live cells were exposed 10 mM 5-ethynyl-2′-deoxyuridine (EdU)c for two hours and then suspended in cold ethanol. Integrated EdU was fluorophore-conjugated by using the Click-iT Plus AlexaFluor 647 Flow Cytometry Assay Kitc. Total DNA was labeled overnight on ice with 20 μg/mL propidium iodidec solution. ANXA5, EdU and propidium iodide labeling data were collected by means of an Accuri C6 flow cytometer running CFlow software (version 1.0.202.1) and data were processed by FlowJo flow cytometry analysis software (version 10). For immunofluorescent imaging, cells were cultured on PLM-coated glass coverslips and bound primary antibodies were labeled with AlexaFluor 568-conjugated goat anti-mouse secondary antibodyc. For NCAM1 detection, coverslips were fixed in 4% paraformaldehyde and labeled with primary antibody 5.1H11°. For detection of myosin heavy chain (MyHC) class proteins, coverslips were fixed in −20 °C methanol and labeled with a pan-MyHC reactive primary antibody (MF 20)°. Immunolabeled coverslips were mounted on microscope slides in Vectasheild mounting medium with DAPIs and images were captured with an Olympus IX70 inverted microscope.
Ca2+ transient analyses
For live-cell Ca2+ transient imaging, piPSC were differentiated into myotubes on PLM-coated glass coverslips. The following steps were conducted at room temperature. Coverslips were washed with physiological saline solution (PSS; 1 mM CaCl2, 135 mM NaCl, 5 mM KCl, 1 mM MgCl2, 10 mM glucose and 10 mM HEPES pH 7.4) and loaded with 10 μM Fluo-4 AMc Ca2+ indicator for 10 min and washed for ~50 min with PSS. While submerged in PSS, coverslips were transferred to a custom stage and fluorescence images (XYt mode) were obtained by using the resonance scanhead of a Lecia SP5/MP confocal microscope at 27.56 frames/s. Field stimulation (1.0 Hz, 2 ms pulse duration, ~20 volts), and perfusion with caffeinea or acetylcholinea were introduced immediately following (~1 s) initiation of the image sequence capture course. Data were processed and quantified using ImageJ image quantification software.
aSigma-Aldrich, St. Louis, MO; bStemgent, Cambridge, MA; cLife Technologies, Grand Island, NY; dHyClone Laboratories, Logan, UT; eJackson ImmunoResearch Laboratories, Inc., West Grove, PA; fMP Biomedicals North America, Solon, OH; gEMD Millipore, Billerica, MA; hCayman Chemical Company, Ann Arbor, MI; iCorning Incorporated Life Sciences, Tewksbury, MA; jShenandoah Biotechnology, Inc., Warwick, PA; kTocris Bioscience, Bristol, UK; lAddgene, Cambridge, MA; mInvivoGen USA, San Diego, CA; nCell Signaling Technology, Inc., Danvers, MA; °Developmental Studies Hybridoma Bank, Iowa City, IA; pSanta Cruz Biotechnology, Inc., Dallas, TX; qGenScript USA Inc., Piscataway, NJ; rPierce Biotechnology, Inc., Rockford, IL; sVector Laboratories, Inc., Burlingame, CA. Unless otherwise noted, materials were used in accordance with the supplier’s provided specifications.
Additional Information
How to cite this article: Genovese, N. J. et al. Enhanced Development of Skeletal Myotubes from Porcine Induced Pluripotent Stem Cells. Sci. Rep. 7, 41833; doi: 10.1038/srep41833 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Roberts, R. M., Yuan, Y., Genovese, N. & Ezashi, T. Livestock models for exploiting the promise of pluripotent stem cells. ILAR J 56, 74–82, doi: 10.1093/ilar/ilv005 (2015).
Ezashi, T., Yuan, Y. & Roberts, R. M. Pluripotent Stem Cells from Domesticated Mammals. Ann Rev Animal Biosciences 4, 223–253, doi: 10.1146/annurev-animal-021815-111202 (2016).
Laumonier, T. et al. Myogenic precursor cell transplantation in pigs: a step towards a clinical use for muscle regeneration? Swiss Med. Wkly. 137 Suppl 155, 44S–48S (2007).
Vainzof, M. et al. Animal Models for Genetic Neuromuscular Diseases. J. Mol. Neurosci. 34, 241–248 (2008).
Dodson, M. V. et al. Skeletal muscle stem cells from animals I. Basic cell biology. Int J Biol Sci 6, 465–474 (2010).
Post, M. J. Cultured meat from stem cells: challenges and prospects. Meat Sci 92, 297–301 (2012).
Novak, A. & Dedhar, S. Signaling through β-catenin and Lef/Tcf. Cell. Mol. Life Sci. 56, 523–537 (1999).
Jin, T., George Fantus, I. & Sun, J. Wnt and beyond Wnt: Multiple mechanisms control the transcriptional property of β-catenin. Cell. Signal. 20, 1697–1704 (2008).
Martin, B. L. & Kimelman, D. Brachyury establishes the embryonic mesodermal progenitor niche. Genes Dev. 24, 2778–2783 (2010).
Maroto, M. et al. Ectopic Pax-3 Activates MyoD and Myf-5 Expression in Embryonic Mesoderm and Neural Tissue. Cell 89, 139–148 (1997).
Capdevila, J., Tabin, C. & Johnson, R. L. Control of Dorsoventral Somite Patterning by Wnt-1 and β-Catenin. Dev. Biol. 193, 182–194 (1998).
Fan, C.-M., Lee, C. S. & Tessier-Lavigne, M. A Role for WNT Proteins in Induction of Dermomyotome. Dev. Biol. 191, 160–165 (1997).
Yamaguchi, T. P., Takada, S., Yoshikawa, Y., Wu, N. & McMahon, A. P. T (Brachyury) is a direct target of Wnt3a during paraxial mesoderm specification. Genes Dev. 13, 3185–3190 (1999).
Wong, J. et al. beta-catenin is essential for efficient in vitro premyogenic mesoderm formation but can be partially compensated by retinoic acid signalling. PLoS ONE 8, e57501 (2013).
Arnold, S. J. et al. Brachyury is a target gene of the Wnt/β-catenin signaling pathway. Mech. Dev. 91, 249–258 (2000).
Haegel, H. et al. Lack of beta-catenin affects mouse development at gastrulation. Development 121, 3529–3537 (1995).
Williams, B. A. & Ordahl, C. P. Pax-3 expression in segmental mesoderm marks early stages in myogenic cell specification. Development 120, 785–796 (1994).
Schuster-Gossler, K., Cordes, R. & Gossler, A. Premature myogenic differentiation and depletion of progenitor cells cause severe muscle hypotrophy in Delta1 mutants. Proc. Natl. Acad. Sci. USA 104, 537–542 (2007).
Hoppler, S., Brown, J. D. & Moon, R. T. Expression of a dominant-negative Wnt blocks induction of MyoD in Xenopus embryos. Genes Dev. 10, 2805–2817 (1996).
Tajbakhsh, S. et al. Differential activation of Myf5 and MyoD by different Wnts in explants of mouse paraxial mesoderm and the later activation of myogenesis in the absence of Myf5. Development 125, 4155–4162 (1998).
Shi, D.-L., Bourdelas, A., Umbhauer, M. & Boucaut, J.-C. Zygotic Wnt/β-Catenin Signaling Preferentially Regulates the Expression of Myf5 Gene in the Mesoderm of Xenopus. Dev. Biol. 245, 124–135 (2002).
Rudnicki, M. A. et al. MyoD or Myf-5 is required for th e formation of skeletal muscle. Cell 75, 1351–1359 (1993).
Vyas, D. R., Spangenburg, E. E., Abraha, T. W., Childs, T. E. & Booth, F. W. GSK-3β negatively regulates skeletal myotube hypertrophy. Am. J. Physiol. Cell Physiol. 283, C545–C551 (2002).
Rochat, A. et al. Insulin and Wnt1 Pathways Cooperate to Induce Reserve Cell Activation in Differentiation and Myotube Hypertrophy. Mol. Biol. Cell 15, 4544–4555 (2004).
Tanaka, S., Terada, K. & Nohno, T. Canonical Wnt signaling is involved in switching from cell proliferation to myogenic differentiation of mouse myoblast cells. J. Mol. Signal. 6, 12 (2011).
Rommel, C. et al. Mediation of IGF-1-induced skeletal myotube hypertrophy by PI(3)K/Akt/mTOR and PI(3)K/Akt/GSK3 pathways. Nat. Cell Biol. 3, 1009–1013 (2001).
Pansters, N. A. M. et al. Segregation of myoblast fusion and muscle-specific gene expression by distinct ligand-dependent inactivation of GSK-3β. Cell. Mol. Life Sci. 68, 523–535 (2011).
Furutani, Y., Umemoto, T., Murakami, M., Matsui, T. & Funaba, M. Role of endogenous TGF-β family in myogenic differentiation of C2C12 cells. J. Cell. Biochem. 112, 614–624 (2010).
Trendelenburg, A. U. et al. Myostatin reduces Akt/TORC1/p70S6K signaling, inhibiting myoblast differentiation and myotube size. Am. J. Physiol. Cell Physiol. 296, C1258–C1270 (2009).
Telugu, B. P. et al. Leukemia inhibitory factor (LIF)-dependent, pluripotent stem cells established from inner cell mass of porcine embryos. J. Biol. Chem. 286, 28948–28953 (2011).
Ying, Q.-L. et al. The ground state of embryonic stem cell self-renewal. Nature 453, 519–523 (2008).
Chiu, C.-P. & Blau, H. M. 5-azacytidine permits gene activation in a previously noninducible cell type. Cell 40, 417–424 (1985).
Davis, R. L., Weintraub, H. & Lassar, A. B. Expression of a single transfected cDNA converts fibroblasts to myoblasts. Cell 51, 987–1000 (1987).
Hollenberg, S. M., Cheng, P. F. & Weintraub, H. Use of a conditional MyoD transcription factor in studies of MyoD trans-activation and muscle determination. Proc. Natl. Acad. Sci. USA 90, 8028–8032 (1993).
Hilber, K., Sun, Y. B. & Irving, M. Effects of sarcomere length and temperature on the rate of ATP utilisation by rabbit psoas muscle fibres. J. Physiol. 531, 771–780 (2001).
Rios, E. & Brum, G. Involvement of dihydropyridine receptors in excitation-contraction coupling in skeletal muscle. Nature 325, 717–720 (1987).
Armstrong, C. M., Bezanilla, F. M. & Horowicz, P. Twitches in the presence of ethylene glycol bis(-aminoethyl ether)-N,N′-tetracetic acid. Biochim. Biophys. Acta 267, 605–608 (1972).
Sakurai, H. et al. Bidirectional induction toward paraxial mesodermal derivatives from mouse ES cells in chemically defined medium. Stem Cell Res. 3, 157–169 (2009).
Xu, C. et al. A Zebrafish Embryo Culture System Defines Factors that Promote Vertebrate Myogenesis across Species. Cell 155, 909–921 (2013).
Choi, Y. et al. Concordant but Varied Phenotypes among Duchenne Muscular Dystrophy Patient-Specific Myoblasts Derived using a Human iPSC-Based Model. Cell Rep. 15, 1–2 (2016).
Chal, J. et al. Differentiation of pluripotent stem cells to muscle fiber to model Duchenne muscle dystrophy. Nat Biotechnol. 33 (2015).
Shelton, M. et al. Derivation and Expansion of PAX7-Positive Muscle Progenitors from Human and Mouse Embryonic Stem Cells. Stem Cell Reports. 3 (2014).
Zheng, J. K. et al. Skeletal myogenesis by human embryonic stem cells. Cell Res. 16, 713–722 (2006).
Dekel, I., Magal, Y., Pearson-White, S., Emerson, C. P. & Shani, M. Conditional conversion of ES cells to skeletal muscle by and exogenous MyoD1 gene. New Biol. 4, 217–224 (1992).
Ozasa, S. et al. Efficient conversion of ES cells into myogenic lineage using the gene-inducible system. Biochem. Biophys. Res. Commun. 357, 957–963 (2007).
Tedesco, F. S. et al. Transplantation of genetically corrected human iPSC-derived progenitors in mice with limb-girdle muscular dystrophy. Sci. Transl. Med. 4, 140ra189 (2012).
Rao, L. et al. Highly Efficient Derivation of Skeletal Myotubes from Human Embryonic Stem Cells. Stem Cell Rev. 8, 1109–1119 (2012).
Tanaka, A. et al. Efficient and reproducible myogenic differentiation from human iPS cells: prospects for modeling Miyoshi Myopathy in vitro . PLoS ONE 8, e61540 (2013).
Goudenege, S. et al. Myoblasts Derived From Normal hESCs and Dystrophic hiPSCs Efficiently Fuse With Existing Muscle Fibers Following Transplantation. Mol. Ther. 20, 2153–2167 (2012).
Tonge, P. D. et al. Divergent reprogramming routes lead to alternative stem-cell states. Nature 516, 192–197 (2014).
Borello, U. et al. The Wnt/β-catenin pathway regulates Gli-mediated Myf5 expression during somitogenesis. Development 133, 3723–3732 (2006).
Bentzinger, C. F., Wang, Y. X. & Rudnicki, M. A. Building Muscle: Molecular Regulation of Myogenesis. Cold Spring Harb. Perspect. Biol. 4 (2012).
Li, H. & Capetanaki, Y. Regulation of the mouse desmin gene: transactivation by MyoD, myogenin, MRF4 and Myf5. Nucleic Acids Res. 21, 335–343 (1993).
Li, L., Heller-Harrison, R., Czech, M. & Olson, E. N. Cyclic AMP-dependent protein kinase inhibits the activity of myogenic helix-loop-helix proteins. Mol. Cell. Biol. 12, 4478–4485 (1992).
Lorenzon, P. et al. Properties of Primary Mouse Myoblasts Expanded in Culture. Exp. Cell Res. 278, 84–91 (2002).
Acknowledgements
This study was made possible with support from People for the Ethical Treatment of Animals and NIH grant R01 HD069979. We would like to express our gratitude to Dr. Vladimir Mironov for his initial project guidance, Ms. Terri Lyddon for providing operational supervision during the flow cytometry procedures and to Dr. Dawn Cornelison for offering helpful suggestions. Transmission electron microscopy sample preparation and assisted image capture by Dr. Tommi White and Ms. Deana Grace Grant was performed at the University of Missouri Electron Microscopy Core. We would like to thank Dr. Alexander Jurkevich and Ms. Arpine Mikayelyan at the University of Missouri Molecular Cytology Core for their help with immunofluorescent image acquisition.
Author information
Authors and Affiliations
Contributions
N.J.G. conceived and initiated the project. B.P.V.L.T. established conditions for piPSC renewal. T.L.D. designed, performed and provided analytical support for Ca2+ signaling experiments. N.J.G. designed, performed and conducted all other experiments, and analyzed data. N.J.G. and R.M.R. arranged project funding support. R.M.R., T.L.D. and N.J.G. arranged laboratory resources. R.M.R. supervised all experiments and data analyses. N.J.G. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The work of N.J.G. was sponsored in part by People for the Ethical Treatment of Animals. N.J.G., R.M.R., and B.P.V.L.T. are listed as co-inventors on a Patent Cooperation Treaty (PCT) patent application filed by the Curators of the University of Missouri through the University of Missouri-Columbia Office of Technology Management and Industry Relations.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Genovese, N., Domeier, T., Telugu, B. et al. Enhanced Development of Skeletal Myotubes from Porcine Induced Pluripotent Stem Cells. Sci Rep 7, 41833 (2017). https://doi.org/10.1038/srep41833
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep41833
This article is cited by
-
Stem cell-based strategies and challenges for production of cultivated meat
Nature Food (2023)
-
Generation of three-dimensional meat-like tissue from stable pig epiblast stem cells
Nature Communications (2023)
-
Spontaneous immortalization of chicken fibroblasts generates stable, high-yield cell lines for serum-free production of cultured meat
Nature Food (2022)
-
Combination of cell signaling molecules can facilitate MYOD1-mediated myogenic transdifferentiation of pig fibroblasts
Journal of Animal Science and Biotechnology (2021)
-
The use of induced pluripotent stem cells in domestic animals: a narrative review
BMC Veterinary Research (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.