Abstract
Prediction and confirmation of the presence of disease-related miRNAs is beneficial to understand disease mechanisms at the miRNA level. However, the use of experimental verification to identify disease-related miRNAs is expensive and time-consuming. Effective computational approaches used to predict miRNA-disease associations are highly specific. In this study, we develop the Network Consistency Projection for miRNA-Disease Associations (NCPMDA) method to reveal the potential associations between miRNAs and diseases. NCPMDA is a non-parametric universal network-based method that can simultaneously predict miRNA-disease associations in all diseases but does not require negative samples. NCPMDA can also confirm the presence of miRNAs in isolated diseases (diseases without any known miRNA association). Leave-one-out cross validation and case studies have shown that the predictive performance of NCPMDA is superior over that of previous method.
Similar content being viewed by others
Introduction
MicroRNAs (miRNAs) are a class of small non-coding RNAs which are about 20 to 25 nucleotides long. miRNAs are important regulatory RNAs, which mainly function in repressing gene expression at the post-transcriptional level by binding to the 3′-UTR of target mRNAs through base pairing1,2. However, some researchers found out that miRNAs may also positively regulate target mRNAs3. Substantial evidence indicated that miRNA dysregulation is related to a number of human diseases, such as cancer4,5. miRNAs affect human diseases by the interaction with various factors, such as miRNA-mRNA interactions6, miRNA-protein interactions7, miRNA-lncRNA (long non-coding RNA) interactions8,9, miRNA-environmental factors interactions10,11, and so on. More miRNA-disease associations have been reported in the last few years. By collecting data from experiments supporting human miRNAs and disease associations from published studies, Li et al.12 and Jiang et al.13 constructed two comprehensive databases, namely, Human miRNA-associated Disease Database (HMDD) and miR2Disease, respectively. Yang et al.14 presented a database of differentially expressed miRNAs in human cancers (dbDEMC) to explore the expression of aberrant miRNAs in different cancer conditions. Therefore, identifying disease-related miRNAs (disease miRNAs) will not only help in the investigation of the pathogenesis of diseases at the molecular level15 but will also facilitate the diagnosis, treatment, and prevention of diseases.
In recent decades, the prediction and ranking of disease-related miRNAs have received much attention16,17,18,19. Experimental verification method for the identification of disease-associated miRNAs is expensive and time-consuming20,21. However, given the large number of studies available, immense biological data about miRNAs have been generated, providing a strong basis for the development of powerful computational methods to predict novel human miRNA-disease associations on a large scale22. Such computational methods mainly aim to predict interactions between diseases and miRNAs23. The key problem for miRNA-disease association inference is similarity computation, some studies developed computation methods to measure miRNA similarity and disease similarity, Zou et al.24 reviewed the main similarity computation methods and the future works of them. Among these computational methods, machine-learning-based methods and network-based methods are the main representatives.
Machine-learning-based models and algorithms have been used to solve the problem in practice, and it is beneficial to improve the classification accuracy and prediction performance25,26. Several studies have proposed machine-learning-based methods to predict novel miRNA-disease associations. Ala Qabaja et al.27 used the Lasso regression model of protein interaction to find associations between miRNAs and diseases. Jiang et al.28 proposed a NaïveBayes model to prioritize disease-related miRNAs through genomic data integration. To distinguish positive miRNA-disease associations from negative ones, Jiang et al.21 proposed a support vector machine (SVM) classification method. Xu et al.29 recommended a prediction method by conducting functional enhancement of miRNA-target dysregulated network. Zeng et al.30 proposed two multipath methods to predict disease related genes based on gene-disease heterogeneous network and then applied to predict miRNA-disease associations31, and achieved good results. Unfortunately, such machine-learning-based approaches face a common limitation; the negative training samples consisting of non-association between miRNAs and diseases do not demonstrate sufficient statistical confidence because an association was not observed in a biological experiment that cannot draw a conclusion indicating no association between them. Considering that the negative training samples are difficult to obtain, Chen et al.32 developed the Regularized Least Squares for miRNA-Disease Associations (RLSMDA) to prioritize the discovery of potential miRNA-disease associations without utilizing negative samples. RLSMDA is a semi-supervised classification algorithm that can predict associations for isolated diseases. To predict different types of miRNA-disease associations, Chen et al.33 developed the model of Restricted Boltzmann Machine for Multiple types of MiRNA-Disease Association prediction (RBMMMDA). RBMMMDA can effectively predict different types of miRNA-disease associations.
Recently, more researchers started using the network-based approaches to predict relationships between miRNAs and diseases. These methods sort the results of the prediction, and recommend some of the previous entries to biologists for further validation. As a result, these methods can be regarded as a recommender system, which has a wide range of applications in many other fields such as movies, news and social tags34. The network-based methods to predict miRNA-disease associations are based on the common assumption that miRNAs with similar functions are normally associated with phenotypically similar diseases and vice versa35,36. On the basis of this assumption, Jiang et al.20 constructed a miRNA functional network and predicted potential miRNA-disease associations by phenotype similarity through hyper-geometric distribution of miRNA-target genes. Given that miRNA-target genes demonstrate a high probability of yielding false-positive results, the same probability of bearing false-positive results is highly possible in predicting miRNA-disease associations. By using data on miRNA-disease associations and the directed acyclic graph (DAG) of disease annotation, Wang et al.37 presented a method to calculate the miRNA functional similarity (referred to as MISIM) and constructed a network of miRNA functional similarities. However, considering the incomplete description of diseases, the miRNA similarities calculated by target-disease annotation appear too biased. Considering miRNA functional similarity and the known miRNA-disease associations simultaneously, Chen et al.38 developed the model of within and between score for miRNA-disease association prediction (WBSMDA), WBSMDA could predicted associations with disease without any known related miRNA. Liu et al.39 measured disease similarity and miRNA similarity by integrating multiple data sources and constructed a heterogeneous network using the known miRNA- disease associations. They extended random walk with restart to predict miRNA-disease associations in the heterogeneous network. The cross validation and case studies show a good performance for predicting potential miRNA-disease associations. Chen et al.22 adopted a universal network similarity measure to construct a miRNA–miRNA functional network and then proposed the Random Walk with Restart for miRNA-Disease Association (RWRMDA) to predict potential miRNA-disease associations. Chen et al.16 implemented a random walk on the Online Mendelian Inheritance in Man (OMIM) database40 disease similarity network to infer potential associations between diseases and miRNAs. Unfortunately, when a random walk is applied to a particular disease, the miRNA-related information is disregarded. In addition, RWRMDA does not recognize the miRNA family or cluster information and is incapable of predicting novel miRNAs for diseases without any known related miRNAs (isolated diseases). Xuan et al.41 presented an algorithm (called HDMP) to predict miRNA-disease associations. HDMP measures miRNA–miRNA functional similarity based on the k of highly similar miRNAs and the distribution information of miRNA verified through experiments of these miRNA groups. Recently, Chen et al.42 presented the network-consistency-based inference (NetCBI) method, which was based on global network measure, to predict potential miRNA-disease associations. NetCBI integrated miRNA similarities, disease similarities, and known miRNA-disease associations to construct a globally associated network for predicting miRNA-disease associations. NetCBI can also predict miRNA-disease associations in isolated diseases, but the performance of cross validation is not as good as that of RWRMDA.
According to previous narratives, the existing computation methods for predicting miRNA-disease associations are restricted by the following limitations. First, some machine-learning-based methods require negative samples that are difficult to obtain. Second, some approaches are unable to predict isolated disease-related miRNAs. Third, some approaches do not recognize the positive influence of miRNA family or cluster information. Finally, although some methods such as NetCBI can predict isolated diseases, their cross-validation performance is poor.
To solve these complications, we propose a method called Network Consistency Projection for miRNA-Disease Associations (NCPMDA) in this paper. NCPMDA calculates the score of each miRNA-disease pair by integrating the miRNA functional similarity network, the disease semantic similarity network, the known miRNA-disease associations, and the miRNA family information to discover the potential associations. NCPMDA shows a clear advantage over other methods, which involve various features, such as leave-one-out cross validation, case studies, global prediction for all diseases, and prediction of novel miRNAs for isolated diseases.
The main contributions of the paper are summarized as follows.
-
1
NCPMDA is a simple and effective method to predict the associations between miRNAs and diseases by integrating various molecular data.
-
2
NCPMDA is a non-parametric method.
Results
Leave-one-out cross validation of NCPMDA
In this study, leave-one-out cross validation (LOOCV) was implemented on known and experimentally verified miRNA-disease associations to evaluate the predictive performance of NCPMDA. NCPMDA was tested using the benchmark dataset to assess its power and infer potential miRNA-disease associations. To deduce a miRNA-disease association, the known association was left out, and the remaining associations were used as a training set to recover predictive score of the association. Moreover, we used the area under the receiver operating characteristic (ROC) curve (AUC) to evaluate the performance of the methods. The closer AUC is to 1, the better the predictive performance is. ROC curve plots test sensitivity or true-positive rate (TPR) versus 1-specificity or false-positive rate (FPR) at different thresholds. The sensitivity refers to the percentage of the test miRNAs with ranking above a given threshold, specifically the ratio of the successfully predicted miRNA-disease associations to the total experimentally verified miRNA-disease associations. Specificity refers to the percentage of associations below the threshold. However, considering the limited number of known and experimentally verified miRNA-disease associations, using only AUC to evaluate the performance of predictive method was too arbitrary; thus, we also used precision-recall (PR) curve and the area under PR curve (AUPR) to complement the performance evaluation. Generally, if the ROC curve and the PR curve show similar variation and AUPR more close to 1, the prediction performance is better. The PR curve plots precision versus recall at different thresholds. In PR-curve plots, the precision refers to the ratio of correctly predicted associations to all associations with scores higher than the given threshold; by contrast, the recall refers to the ratio of correctly predicted associations to all known miRNA-disease associations.
On the basis of miRNA functional similarities and disease semantic similarities, NCPMDA integrates the similarity of known miRNA-disease association network (SN) and miRNA family information to construct a global miRNA-disease network for predicting miRNA-disease associations. We tested the predictive performance of NCPMDA considering the following aspects: (1) NCPMDA with all information (NCPMDA); (2) NCPMDA without miRNA family information; (3) NCPMDA without SN; (4) NCPMDA in miRNA space projection only; (5) NCPMDA in disease space projection only. The ROC curves of the above mentioned features are plotted in Fig. 1, and the PR curves are represented in Fig. 2.
ROC curves and AUC values of NCPMDA based on LOOCV in different situations.
(1) NCPMDA with all information (NCPMDA), (2) NCPMDA without miRNA family information, (3) NCPMDA without SN, (4) NCPMDA in miRNA space projection only, (5) NCPMDA in disease space projection only. SN is the similarity of the known miRNA-disease association network.
PR curves and AUC values of NCPMDA based on LOOCV in different situations.
(1) NCPMDA with all information (NCPMDA), (2) NCPMDA without miRNA family information, (3) NCPMDA without SN, (4) NCPMDA in miRNA space projection only, (5) NCPMDA in disease space projection only. SN is the similarity of the known miRNA-disease association network.
Obviously, NCPMDA exhibits a commendable predictive performance with an AUC value of 0.9173. The miRNA family information is advantageous in improving the predictive performance of NCPMDA. The SN increases the AUC value from 0.9112 to 0.9173 and obviously improves AUPR by increasing it from 0.4967 to 0.5358. In a single-space projection (miRNA space projection or disease space projection) test, NCPMDA also performs well with AUC values of 0.8878 in miRNA space projection and 0.7976 in disease space projection. If SN is removed from NCPMDA, the predictive performance is reduced. Therefore, the predictive performance of miRNA-disease associations is practical to improve by integrating the SN and miRNA family information in the method.
Comparison with other methods
To the best of our knowledge, HDMP41, RLSMDA32, NetCBI42, and the global network algorithm developed by Shi et al.43 are state-of-the-art computational methods for predicting miRNA-disease associations. However, HDMP does not work for diseases without known related miRNAs; thus, this algorithm cannot be used for comparisons. With regard to the method developed by Shi et al. in constructing a global network by integrating disease gene associations, miRNA-target interactions, and protein interactions to predict miRNA-disease associations, the datasets vary significantly from the ones used in our method. Furthermore, they did not use known miRNA-disease associations; thus, the performances of this method and NCPMDA cannot be fairly and reasonably compared. NCPMDA, RLSMDA, and NetCBI, which were developed using similar datasets, can be used to predict novel miRNA-disease associations for diseases with no known miRNAs. On the basis of the above analyses, we compared the performance of NCPMDA with those of RLSMDA and NetCBI.
NCPMDA, RLSMDA, and NetCBI were tested on the benchmark dataset to assess their performance in deducing the potential miRNA-disease associations. We implemented a LOOCV for each method. The optimal parameters are selected for RLSMDA and NetCBI as described in the literature. Given that RLSMDA and NetCBI do not employ the similarity of known miRNA-disease associations and miRNA family information, the three methods were assessed using only miRNA functional similarities and disease semantic similarities to predict miRNA-disease associations. Figure 3 shows the ROC curves and the AUC values to predict the miRNA-disease associations by using the three methods. Without considering the similarity of the known miRNA-disease association network and miRNA family information (Fig. 3), the AUC value of NCPMDA is 0.8958, and the AUC values of RLSMDA and NetCBI are 0.8059 and 0.8001, respectively. Evidently, NCPMDA performs better than that of RLSMDA and NDBM in LOOCV.
To avoid data dependence, the generalization abilities and strength of the algorithms were further verified on the predictive dataset based on LOOCV. The AUC value of NCPMDA disregarding the similarity of the known miRNA-disease association network and the miRNA family information is 0.9605, and the AUC values of RLSMDA and NetCBI are 0.9511 and 0.9560, respectively. The AUC values of the three methods on the predictive dataset are higher than those on the benchmark dataset because miRNA-disease associations differ significantly. In the benchmark dataset, each miRNA is associated with 2.27 disease phenotypes, and each disease phenotype is associated with 4.41 miRNAs on the average. However, in the predictive dataset, the values are 5.147 and 10.18, respectively. Given the increase in the number of associations that the known miRNA-disease associations make which are closer to the real network, the predictive performance is better. For the same reason, the predictive performance of NCPMDA disregarding the similarity of the known miRNA-disease association network consistency and miRNA family information was excellent. NCPMDA also performs slightly better than the two other methods on the predictive dataset. Therefore, our method demonstrates strong data generalization ability. Our algorithm also shows an obvious advantage when the known and experimentally verified miRNA-disease associations are very few.
Comprehensive prediction of unknown associations
The predictive performance of our method is thoroughly discussed in the first section of this paper. In this section, we utilized NCPMDA to predict unknown miRNA-disease associations, including all possible miRNA-disease pairs from the predictive dataset. The reason for not using the benchmark dataset to do the prediction is that the benchmark dataset contains fewer miRNA numbers, many of these miRNAs associated with disease has been confirmed by the updated databases, which will lead to a very high accuracy of prediction. First, the projection score of each miRNA-disease pair was calculated using all known and verified miRNA-disease associations, similarity information, and miRNA family information. Second, the unknown associations were ranked according to the projection scores. Finally, the top 40 associations were manually verified through three online databases: HMDD12, miR2Disease13, and dbDEMC14 (The database is being updated, and the experimentally validated miRNA-disease associations were gained from the author.). The predictive results and verified evidences are presented in Table 1. Among the top 40 predictive associations, only four have not been confirmed in the aforementioned three databases and the top 10 were all confirmed.
Case studies of Breast cancer and Hepatocellular cancer
Increasing evidence indicates that miRNAs play critical roles in the development of breast cancer and hepatocellular carcinoma (HCC). Case studies were analyzed to further evaluate the ability of NCPMDA to predict miRNA-disease associations. Similarly, all known associations were used as training set, and the unknown associations were assigned as the testing set. The predictive results were manually verified by three online databases. The top 40 potential breast cancer and HCC-related miRNAs predicted by NCPMDA are listed in (Supplementary Informations S1 and S2), respectively.
Breast cancer is one of the most common form of cancer among women. In the predictive dataset, 78 miRNAs are associated with breast cancer. The potential breast cancer-related miRNAs were predicted by NCPMDA based on the 78 known associations. Among the top 40 predicted miRNAs, 38 miRNAs were confirmed by the three aforementioned databases; and only hsa-mir-30e and hsa-mir-142 were not confirmed. However, Lin et al.44 demonstrated that has-miR-30e were down regulated in both plasma and breast cancer tissues, as described in the literature45, which provided information that has-mir-142 inhibited breast cancer cell invasiveness. Given that this evidence came in after the last updates for the three databases, they were not incorporated in the databases in time for the study. The evidence found in the literature further demonstrated the reliability of NCPMDA in predicting new disease-related miRNAs.
HCC is the most common type of liver cancer. HCC most commonly occurs in countries where hepatitis B infections are common. A total of 34 HCC-related miRNAs are found in the predictive dataset. Among the top 40 HCC-related miRNAs predicted by NCPMDA, 37 have been confirmed by HMDD, mir2disease, and dbDEMC. Further evidences are found to support our prediction results. Using quantitative methylation analysis and real-time PCR, Tang et al.46 defined has-miR-429 as a key inducer for HCC pathogenesis and metastasis with potential utility for tumor intervention. Based on gene knockdown experiment, Jung et al.47 summarized that Gα12gep oncogene inhibits FOXO1 in HCC is caused by miR-135b and miR-194 dysregulation. Using methylation-specific PCR, Xie et al.48 suggest that DNA methylation may be involved in the inactivation of miR-34 b in HCC.
Application of NCPMDA to predict isolated diseases
An isolated disease refers to a disease without any known associated miRNA. To demonstrate the predictive ability of NCPDMA on isolated diseases, we removed the known and verified miRNA-disease associations related to predictive diseases. This operation ensured that we only used similarity information and known miRNA-disease associations of the other diseases to predict disease-related miRNAs. We conducted case studies on breast cancer and HCC, and the results are presented in (Supplementary Informations S3 and S4), respectively. For breast cancer, we removed 78 known miRNA-breast cancer associations to predict the unknown associations by NCPMDA. Out of the top 40 predicted miRNAs, 37 have been confirmed based on the updated HMDD, mir2disease, and dbDEMC databases. For HCC, 34 known miRNA-HCC associations were removed, and 36 of the top 40 predicted miRNAs were confirmed. Moreover, the top 20 predictions for breast cancer and HCC were all confirmed. Therefore, NCPMDA performs well in the prediction of isolated disease.
Discussions
Accumulative evidence indicated that miRNAs play important roles in the occurrence and development of diseases. Identification of disease-related miRNAs helps in understanding the mechanism of diseases. Effective computational methods for identifying miRNA-disease associations can provide support for experimental studies on miRNAs.
In this study, we presented a network-based approach (NCPMDA) to predict miRNA-disease associations. On the basis of miRNA similarities and disease similarities, NCPMDA integrated the known miRNA-disease association network and miRNA family information to restructure the miRNA–miRNA and disease–disease similarity networks is to predict miRNA-disease associations. LOOCV and case studies have shown that integrated information on known association networks and miRNA families aids in the improvement of the predictive performance of NCPDMA. Compared with the current state-of-the-art computational methods for predicting miRNA-disease associations, NCPMDA does not require the use of negative samples. NCPDMA is also a universal method that can be used in simultaneously restructuring the potential associations for all diseases. Specifically, NCPMDA is a non-parametric method and demonstrates an obvious advantage when the known and experimentally verified miRNA-disease associations are very few. Furthermore, NCPMDA can predict related miRNAs of isolated diseases. Therefore, NCPMDA can be a useful resource for the prediction of miRNA-disease associations.
Despite the favorable results obtained using NCPMDA, this study presents some limitations. First, we simply used the similarity of known associations and miRNA family information as variables to restructure the miRNA–miRNA and disease–disease similarity networks; a more reasonable measurement and integration method can improve the performance of NCPMDA. Second, the final prediction score of NCPMDA was computed using the miRNA space projection and disease space projection; obtaining a single measurement or a more accurate integration of results from two different spaces should be prioritized in future studies. Finally, the increase in the number of miRNA-disease associations being experimentally verified advances the development of computational approaches to predict miRNA-disease associations.
Methods
Dataset and preprocessing
Two datasets are used in this study. For ease of description, the two datasets are called benchmark dataset and predictive dataset. Each dataset contains four kinds of data, namely, miRNA functional similarities, miRNA family information, disease semantic similarities, and known miRNA-disease associations.
The disease similarities were obtained from the literature32. A strict system for disease classification from the MeSH database (http://www.ncbi.nlm.nih.gov/) was obtained, and diseases were described in a DAG, in which the nodes represent the disease itself and its precursor diseases, whereas the link from a parent node to a child node represents the relationship between these nodes. The disease similarity was calculated using disease DAGs, with the assumption that two diseases sharing larger parts of DAGs are more similar. To represent the variables, we used the matrix DD to denote disease similarities, where DD(i, j) in row i and column j represents the similarity between diseases i and j.
The miRNA functional similarity scores were obtained from a reliable webpage (http://www.cuilab.cn/). On the basis of the assumption implying that miRNAs with similar functions tend to be associated with similar diseases, the similarity score of miRNA pairs were measured the similarity of their associated disease DAG (the literature37 provides a detailed description of the score calculation). We used a matrix MM which denotes the miRNA–miRNA functional similarity, where the variable MM(i, j) in row i and column j is the functional similarity between miRNA i and j.
The information on miRNA families was obtained from the miRBase49 Sequence Database, Release 21 (http://www.mirbase.org/). We used the matrix FAM to represent the information on miRNA families, where the variable FAM(i, j) in row i and column j is 1 if miRNA i and j belong to the same family; otherwise, the value is 0.
In the benchmark dataset, the known miRNA-disease associations were obtained from literature20. These associations were collected from the HMDD12 and mir2disease13 databases. The databases include 271 high-qualityand experimentally verified associations of miRNA deregulation with disease development. However, 19 miRNAs were not found in literature37, which left 242 high-quality miRNA-disease associations, including 99 miRNAs and 51 diseases. In the predictive dataset, the known miRNA-disease associations were also acquired37 to evaluate the prediction accuracy. The association data included 1616 distinct, high-quality, and experimentally verified human miRNA-disease associations, which were obtained from HMDD in September 2009. The names of different mature miRNAs and diseases were unified, and the records of different miRNA copies were consolidated. Finally, 1395 miRNA-disease associations, including 271 miRNAs and 137 diseases, were obtained. We used the information on the predictive dataset for the evaluation of prediction accuracy and the prediction of new miRNA-disease associations. We preferred to use the updated version of HMDD and other databases (mir2disease and dbDEMC) to verify our prediction. Hence, we did not include the latest association dataset in HMDD and other new associations in other databases. To represent the variables, we used the matrix AS signifies the adjacency matrix of miRNA-disease association, where AS(i, j) in row i and column j is 1 if miRNA i is associated with disease j; otherwise, the value is 0.
Constructing a miRNA–miRNA similarity network
To compute the relationship between miRNA pairs more efficiently, we integrated the miRNA functional similarities, miRNA family information, and miRNA similarities of known miRNA-disease associations to construct a miRNA–miRNA similarity network.
On the basis of the human miRNA-disease associations (matrix AS) considered and the assumption that two miRNAs are associated more with common diseases, and the function of the miRNA pair are more similar, the miRNA similarity of the known associations is calculated using the Jaccard similarity measure, as shown in Equation (1):

Matrix NCM represents the miRNA–miRNA functional similarity scores of known associations, where NCM(i, j) indicates the similarity between miRNA i and j. Where D11 is the total number of variables with a value of 1 in both miRNA i and j, that is, the total number of diseases simultaneously associated with miRNA i and j. Similarly, D01 is the total number of variables with a value of 0 in miRNA i and 1 in miRNA j, whereas D10 is the total number of variables with a value of 1 in miRNA i and 0 in miRNA j. This study focuses on known miRNA-disease associations; thus, the total number of variables with 0 value in both miRNA i and j is disregarded. For certain diseases, the number of related-miRNA is only one, and remove this association for LOOCV will lead to D11, D10 and D01 are 0. In Equation (1), ε represents a very small positive real number which does not affect the final score. Its function is to avoid having 0 as the denominator. In our experiments, ε is set to 10−30. For similar reasons, the use of ε in the back of the article uses the same settings.
By using the miRNA functional similarities (matrix MM), miRNA family information (matrix FAM), and miRNA similarities of known miRNA-disease associations (matrix NCM) as variables, we incorporate them into Equation (2) to construct a miRNA–miRNA similarity network:

where SM(i, j) is the final similarity score of miRNA i and j. Obviously, if two miRNAs in the known association network are more similar and belong to the same family, the computed score is higher.
Constructing disease–disease similarity network
Disease semantic similarities and disease similarities of known miRNA-disease associations were applied to construct a disease–disease similarity network.
Parallel to the miRNA similarities of the known miRNA-disease associations and based on the assumption that two diseases associated with more common miRNAs are more similar, we use the Jaccard similarity measure to calculate the disease similarities of known miRNA-disease associations, as shown in Equation (3):

where M11 is the total number of variables with a value of 1 in both diseases i and j, that is, M11 is the total number of miRNAs simultaneously associated with disease i and j. Similarly, M01 is the total number of variables with a value of 0 in disease i and 1 in disease j, whereas M10 is the total number of variables with a value of 1 in disease i and 0 in disease j. The significance of ε is the same as that in the previous equation.
Analogous to Equation (2), we use the values for the disease semantic similarities (matrix DD) and the disease similarities of known associations and incorporate them into Equation (4) to construct a disease–disease similarity network:

where SD(i, j) is the final similarity score of disease i and j. Evidently, if two diseases in the known association network are more similar, the computed score for them is higher.
Network Consistency Projection of miRNA-Disease Association (NCPMDA)
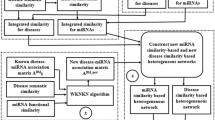
In accordance with the assumption that diseases associated with highly related miRNAs are more similar (and vice versa) and that miRNAs associated with highly related diseases are more similar (and vice versa), we developed the Network Consistency Projection for miRNA-Disease Associations (NCPMDA) method to predict potential miRNA-disease associations. The flowchart of NCPMDA method shows in Fig. 4.
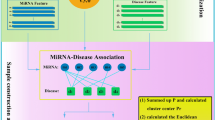
NCPMDA calculates the potential miRNA-disease association score consisting of two network consistency projection scores, miRNA space projection score and disease space projection score, separately. The network consistency mentioned here refers to the higher the spatial similarity of miRNA i associated miRNAs in miRNA-miRNA similarity network and disease j associated miRNAs in the known miRNA-disease network, the greater the association of miRNA i with disease j. We use vector space projection to represent it and named miRNA space projection. Similarly, disease space projection measure the association of miRNA i and disease j in disease space. Considering the miRNA-disease associations not verified by experiment cannot confirm that there is no association and to avoid having 0 as the denominator, we replace 0 in matrix AS to ε. In our experiments, ε is set to 10−30. Simultaneously, we use |C| to represent the length of vector C (the norm of vector C). The miRNA space projection score is calculated as

where SMi, is the ith row of matrix SM and a vector consisting of the similarities between miRNA i and all other miRNAs. Similarly, ASj is the jth column of matrix AS and the vector consisting of the associations of disease j and all miRNAs. Matrix ncp_m is the network consistency projection score of the miRNA similarity network SM on the known miRNA-disease association network, AS; the variable ncp_m(i, j) in row i and column j is the network consistency projection of SMi on ASj. Notably, the smaller angle between SMi and ASj, the more miRNAs associated with disease j, and the more similar miRNAs and miRNA i are, the greater the network consistency projection score ncp_m(i, j) is.
Similarly, the disease space projection score is calculated as follows:

where SDj is the jth column of matrix SD, the vector comprising the similarities of disease j and all other diseases. Similarly, ASi is the ith row of matrix AS, which consists of the associations of miRNA i and all diseases. Matrix ncp_d is the projection of the disease similarity network, SD, on the known miRNA-disease association network, AS; ncp_d(i, j) in row i and column j is the network consistency projection of SDj on ASi. Remarkably, the smaller angle between SDj and ASi, the more diseases are associated with miRNA i, and the more similar these diseases and disease j are, the greater the network consistency projection score ncp_d(i, j) is.
Finally, the miRNA space projection score and disease space projection score are combined and normalized as shown below:

where ncp(i, j) is the final score of network consistency projection of miRNA i and disease j; ncp_m(i, j) and ncp_d(i, j) are the miRNA space projection score and disease space projection score of miRNA i and disease j, respectively. The final score is used to predict miRNA-disease association. If only NCPMDA in miRNA space projection (remove ncp_d(i, j) and |SDj|) is considered, the final score of miRNA i and disease j is the cosine similarity of space vector SMi and ASj. Similarly, the cosine similarity of space vector SDj and ASi is the final score of miRNA i and disease j when considering NCPMDA in disease space projection only.
Additional Information
How to cite this article: Gu, C. et al. Network Consistency Projection for Human miRNA-Disease Associations Inference. Sci. Rep. 6, 36054; doi: 10.1038/srep36054 (2016).
References
Hammond, S. M. An overview of microRNAs. Advanced Drug Delivery Reviews 87, 3–14 (2015).
Meister, G. & Tuschl, T. Mechanisms of gene silencing by double-stranded RNA. Nature 431, 343–349 (2004).
Rajasekaran, S., Pattarayan, D., Rajaguru, P., Gandhi, P. S. S. & Thimmulappa, R. K. MicroRNA regulation of acute lung injury and acute respiratory distress syndrome. Journal of Cellular Physiology (2016).
Olson, E. N. MicroRNAs as therapeutic targets and biomarkers of cardiovascular disease. Science Translational Medicine 6, 53–74 (2014).
Yi, W. K., Ferland-Mccollough, D., Jackson, T. J. & Bushell, M. microRNAs in cancer management. Lancet Oncology 13, 249–258 (2012).
Li, Y., Liang, C., Wong, K. C., Luo, J. & Zhang, Z. Mirsynergy: detecting synergistic miRNA regulatory modules by overlapping neighbourhood expansion. Bioinformatics 30, 2627–2635 (2014).
Shi, H. et al. Integration of multiple genomic and phenotype data to infer novel miRNA-disease associations. Plos One 11 (2016).
Chen, X. Long non-coding RNAs and complex diseases: from experimental results to computational models. Briefings in Bioinformatics bbw060 (2016).
Chen, X., Huang, Y. A., Wang, X. S., You, Z. H. & Chan, K. C. FMLNCSIM: fuzzy measure-based lncRNA functional similarity calculation model. Oncotarget 7, 45948–45958 (2016).
Chen, X., Liu, M. X., Cui, Q. H. & Yan, G. Y. Prediction of disease-related interactions between microRNAs and environmental factors based on a semi-supervised classifier. Plos One 7, e43425–e43425 (2012).
Chen, X. MiREFRWR: a novel disease-related microRNA-environmental factor interactions prediction method. Molecular Biosystems 12, 624–633 (2015).
Li, Y. et al. HMDD v2.0: a database for experimentally supported human microRNA and disease associations. Nucleic Acids Research 42, 1070–1074 (2014).
Jiang, Q. et al. miR2Disease: a manually curated database for microRNA deregulation in human disease. Nucleic Acids Research 37, D98–104 (2009).
Yang, Z. et al. dbDEMC: a database of differentially expressed miRNAs in human cancers. BMC Genomics 11, 325–325 (2010).
Chen, R. W. et al. Truncation in ccnd1 mRNA alters mir-16-1 regulation in mantle cell lymphoma. Blood 112, 822–829 (2008).
Chen, H. & Zhang, Z. Prediction of associations between omim diseases and microRNAs by random walk on omim disease similarity network. Scientific World Journal 2013, 273–275 (2013).
Augustin, R. et al. Computational identification and experimental validation of microRNAs binding to the alzheimer-related gene adam10. BMC medical genetics 13, 1 (2012).
Madden, S. F. et al. Detecting microRNA activity from gene expression data. BMC Bioinformatics 11, 1–14 (2010).
Le, D. H. Network-based ranking methods for prediction of novel disease associated microRNAs. Computational Biology and Chemistry 58, 139–148 (2015).
Jiang, Q. et al. Prioritization of disease microRNAs through a human phenome-microRNAome network. BMC Systems Biology 4 Suppl 1, 1–9 (2010).
Jiang, Q., Wang, G., Jin, S., Li, Y. & Wang, Y. Predicting human microRNA-disease associations based on support vector machine. International journal of data mining and bioinformatics 8, 282–293 (2013).
Chen, X., Liu, M.-X. & Yan, G.-Y. RWRMDA: predicting novel human microRNA–disease associations. Molecular BioSystems 8, 2792–2798 (2012).
Zeng, X., Zhang, X. & Zou, Q. Integrative approaches for predicting microRNA function and prioritizing disease-related microRNA using biological interaction networks. Briefings in bioinformatics 17, 193–203 (2016).
Zou, Q., Li, J., Song, L., Zeng, X. & Wang, G. Similarity computation strategies in the microRNA-disease network: a survey. Briefings in functional genomics 15, 55–64 (2016).
Gu, B. et al. Incremental learning for n-support vector regression. Neural Networks 67, 140–150 (2015).
Wen, X., Shao, L., Xue, Y. & Fang, W. A rapid learning algorithm for vehicle classification. Information Sciences 295, 395–406 (2015).
Qabaja, A., Alshalalfa, M., Bismar, T. A. & Alhajj, R. Protein network-based lasso regression model for the construction of disease-miRNA functional interactions. EURASIP Journal on Bioinformatics and Systems Biology 2013, 1 (2013).
Jiang, Q., Wang, G. & Wang, Y. An approach for prioritizing disease-related microRNAs based on genomic data integration. In 2010 3rd International Conference on Biomedical Engineering and Informatics, vol. 6, 2270–2274 (2010).
Xu, J. et al. Prioritizing candidate disease miRNAs by topological features in the miRNA target–dysregulated network: Case study of prostate cancer. Molecular cancer therapeutics 10, 1857–1866 (2011).
Zeng, X., Liao, Y., Liu, Y. & Zou, Q. Prediction and validation of disease genes using HeteSim scores. IEEE/ACM Transactions on Computational Biology and Bioinformatics 1–1 (2016).
Zeng, X., Zhang, X., Liao, Y. & Pan, L. Prediction and validation of association between microRNAs and diseases by multipath methods. Biochimica et Biophysica Acta (BBA)-General Subjects (2016).
Chen, X. & Yan, G.-Y. Semi-supervised learning for potential human microRNA-disease associations inference. Scientific reports 4 (2014).
Chen, X. et al. RBMMMDA: predicting multiple types of disease-microRNA associations. Scientific reports 5 (2015).
Tinghuai, M. et al. Social network and tag sources based augmenting collaborative recommender system. IEICE transactions on Information and Systems 98, 902–910 (2015).
Lu, M. et al. An analysis of human microRNA and disease associations. PloS one 3, e3420 (2008).
Bandyopadhyay, S., Mitra, R., Maulik, U. & Zhang, M. Q. Development of the human cancer microRNA network. Silence 1, 1 (2010).
Wang, D., Wang, J., Lu, M., Song, F. & Cui, Q. Inferring the human microRNA functional similarity and functional network based on microRNA-associated diseases. Bioinformatics 26, 1644–1650 (2010).
Chen, X. et al. WBSMDA: within and between score for miRNA-disease association prediction. Scientific reports 6 (2016).
Liu, Y., Zeng, X., He, Z. & Quan, Z. Inferring microRNA-disease associations by random walk on a heterogeneous network with multiple data sources. IEEE/ACM Transactions on Computational Biology and Bioinformatics 1–1 (2016).
Hamosh, A., Scott, A. F., Amberger, J. S., Bocchini, C. A. & McKusick, V. A. Online mendelian inheritance in man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic acids research 33, D514–D517 (2005).
Xuan, P. et al. Prediction of microRNAs associated with human diseases based on weighted k most similar neighbors. PloS one 8, e70204 (2013).
Chen, H. & Zhang, Z. Similarity-based methods for potential human microRNA-disease association prediction. BMC medical genomics 6, 1 (2013).
Shi, H. et al. Walking the interactome to identify human miRNA-disease associations through the functional link between miRNA targets and disease genes. BMC systems biology 7, 101 (2013).
Lin, Z. et al. Abnormal miRNA-30e expression is associated with breast cancer progression. Clinical laboratory 62, 121–128 (2015).
Schwickert, A. et al. microRNA mir-142-3p inhibits breast cancer cell invasiveness by synchronous targeting of wasl, integrin alpha v, and additional cytoskeletal elements. PloS one 10, e0143993 (2015).
Tang, J. et al. Mir-429 increases the metastatic capability of hcc via regulating classic wnt pathway rather than epithelial–mesenchymal transition. Cancer letters 364, 33–43 (2015).
Jung, H. S. et al. Ga 12 gep oncogene inhibits foxo1 in hepatocellular carcinoma as a consequence of mir-135b and mir-194 dysregulation. Cellular signalling 26, 1456–1465 (2014).
Xie, K. et al. Methylation-associated silencing of microRNA-34b in hepatocellular carcinoma cancer. Gene 543, 101–107 (2014).
Kozomara, A. & Griffiths-Jones, S. MiRBase: integrating microRNA annotation and deep-sequencing data. Nucleic acids research gkq1027 (2010).
Acknowledgements
This study is supported by the Program for New Century Excellent Talents in university (Grant No. NCET-10-0365), National Nature Science Foundation of China (Grant Nos 60973082, 11171369, 61272395, and 61370171), National Nature Science Foundation of Hunan Province (Grant No. 12JJ2041), the Planned Science and Technology project of Hunan Province (Grant Nos 2009FJ3195 and 2012FJ1012), and the Fundamental Research Funds for the Central universities, Hunan university.
Author information
Authors and Affiliations
Contributions
C.G. conceived the project, developed the prediction method, designed and implemented the experiments, analyzed the result, and wrote the paper. B.L. analyzed the result, and wrote the paper. X.L. implemented the experiments, and analyzed the result. K.L. analyzed the result. All authors reviewed the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Gu, C., Liao, B., Li, X. et al. Network Consistency Projection for Human miRNA-Disease Associations Inference. Sci Rep 6, 36054 (2016). https://doi.org/10.1038/srep36054
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep36054
This article is cited by
-
DAE-CFR: detecting microRNA-disease associations using deep autoencoder and combined feature representation
BMC Bioinformatics (2024)
-
Identification of human microRNA-disease association via low-rank approximation-based link propagation and multiple kernel learning
Frontiers of Computer Science (2024)
-
BMPMDA: Prediction of MiRNA-Disease Associations Using a Space Projection Model Based on Block Matrix
Interdisciplinary Sciences: Computational Life Sciences (2022)
-
Predicting miRNA–disease associations using improved random walk with restart and integrating multiple similarities
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.