Abstract
Aged garlic extract (AGE) is widely used as a dietary supplement on account of its protective effects against oxidative stress and inflammation. But less is known about specific molecular targets of AGE and its bioactive components, including N-α-(1-deoxy-D-fructos-1-yl)-L-arginine (FruArg). Our recent study showed that both AGE and FruArg significantly attenuate lipopolysaccharide (LPS)-induced neuroinflammatory responses in BV-2 microglial cells. This study aims to unveil effects of AGE and FruArg on gene expression regulation in LPS stimulated BV-2 cells. Results showed that LPS treatment significantly altered mRNA levels from 2563 genes. AGE reversed 67% of the transcriptome alteration induced by LPS, whereas FruArg accounted for the protective effect by reversing expression levels of 55% of genes altered by LPS. Key pro-inflammatory canonical pathways induced by the LPS stimulation included toll-like receptor signaling, IL-6 signaling, and Nrf2-mediated oxidative stress pathway, along with elevated expression levels of genes, such as Il6, Cd14, Casp3, Nfkb1, Hmox1, and Tnf. These effects could be modulated by treatment with both AGE and FruArg. These findings suggests that AGE and FruArg are capable of alleviating oxidative stress and neuroinflammatory responses stimulated by LPS in BV-2 cells.
Similar content being viewed by others
Introduction
Garlic (Allium sativum L.) has been recognized to offer health benefits including cardiovascular protective effects and cancer preventive effects1,2,3. Aged garlic extract (AGE) is a nutritional supplement prepared by prolonged extraction (normally for 20 months) of fresh garlic with 15–20% aqueous ethanol at room temperature. This product is odorless and appears to be superior to normal garlic in its antioxidant properties. AGE can act as a superoxide radical scavenger4,5,6. AGE was further shown to promote a potent antioxidant protection in cells by enhancing activity of the cellular antioxidant enzymes superoxide dismutase, catalase and glutathione peroxidase, and by increasing glutathione7,8. As a dietary supplement, AGE was shown to reduce total serum cholesterol, low-density lipoprotein, and systolic pressure in hypercholesterolemic patients and to inhibit platelet aggregation in both animal and human subjects9,10,11.
N-α-(1-deoxy-D-fructos-1-yl)-L-arginine (FruArg) is a major carbohydrate derivative and a component of non-sulfur containing nutraceuticals from AGE12,13. Quantitative analysis revealed that AGE contained 2.1–2.4 mmol/L of FruArg, but none was detected in either raw or heated garlic juice14. FruArg belongs to a family of fructose-amino acids in AGE, and is an Amadori rearrangement product arising from the condensation reaction between free glucose and arginine during early stages of the Maillard reaction, which is responsible for color formation in foods and, to a large extent, antioxidant properties of AGE14,15,16. Studies indicated the anti-tumor effects of certain species of fructose-amino acids are due mainly to their ability to inhibit cancer cell proliferation and adhesion17,18. Meanwhile, FruArg was shown to exhibit antioxidant activity and hydrogen peroxide scavenging capacity that is comparable to the potent hydrogen peroxide scavenging compound ascorbic acid14,19. Our previous study demonstrated that AGE and FruArg could suppress NO production in a concentration-dependent manner without affecting the cell viability in lipopolysaccharide (LPS)-induced mouse BV-2 microglial cells, suggesting AGE and FruArg have beneficial effects in mitigating neuroinflammation20.
In the present study, RNA-Seq analysis was conducted to assess global gene expression affected by AGE and FruArg in LPS-induced BV-2 microglial cells. Our results show that AGE is capable of repressing levels of pro-inflammatory mRNAs in LPS-treated cells and that FruArg is an active functional component in AGE that accounts for the protective effects. Additionally, we have also annotated the canonical pathways and functional networks that may be responsible for the protective effects of these botanical compounds. The present study addresses the regulatory function of AGE and FruArg on LPS-stimulated cells with respect to global gene expression by providing quantitative information about the population of mRNA species. The data broaden our understanding of the effect of AGE and FruArg to the level of gene expression regulation and suggested their therapeutic potential for mitigating inflammatory abnormalities in the aging individuals.
Results
LPS induced transcriptome alteration in BV-2 microglial cells
Endotoxin LPS is one of the most potent stimuli for microglial cells. It is capable of activating multiple intracellular signaling pathways leading to cytokines secretion. The global gene expression profile alterations that were created by LPS stimulus was represented by the differentially expressed genes (DEGs) between BV-2 microglial cells treated with 100 ng/mL LPS and control cells. DEG analysis identified 2563 genes whose expression levels were altered significantly by LPS treatment (Fig. 1 and Supplementary Table S1). A total of 1172 genes were down-regulated and 1391 genes were up-regulated in response to the LPS. A number of genes associated with inflammation were significantly up-regulated, such as Cd14, Il1a, Il1b, Nfkb1, Nfkb2, Tlr1, Tlr2, Tlr3, Tlr6, and Tnf, which served as mRNA markers of LPS treatment in these cells. Their up-regulation proved the reliability of the LPS model used in the experiments21,22.
LPS induced gene changes in BV-2 microglial cells.
LPS treatment induced differential expression of 2563 genes. For a gene to be considered as differentially expressed gene, the “log2Fold Change” has to ≥1 or ≤−1, with a FDR adjusted p-value ≤ 0.05. Red region highlighted up-regulated genes (1391 genes), green region highlighted down-regulated genes (1172 genes). Vertical redline represented p-adj = 0.05, two horizontal lines indicate log2Fold Change of +1 and −1.
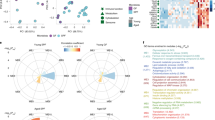
To evaluate which canonical signaling pathways and signaling networks could be activated upon LPS treatment, we performed a gene set enrichment analysis by Ingenuity Pathway Analysis (IPA). The most significant canonical pathways for the genes altered by LPS include death receptor signaling, toll-like receptor signaling, IL-10 signaling, TNFR2 signaling, role of pattern recognition receptors in recognition of bacteria and viruses, role of PKR in interferon induction and antiviral response, IL-6 signaling, role of macrophages, fibroblasts and endothelial cells in rheumatoid arthritis, TREM1 signaling, and activation of IRF by cytosolic pattern recognition receptors (Fig. 2A) (see Supplementary Table S4). Additionally, the disease and function networks were further analyzed. The top-ranked network is related to cell cycle, cellular development, and cellular growth and proliferation, with 34 focus genes, including Serp1, Ifrd1, and Nupr1 (Fig. 2B). Among these focus genes, 21 genes were up-regulated and 13 genes were down-regulated.
Effects of LPS in BV-2 microglial cells on top canonical pathways and disease/function networks.
(A) Top 10 canonical pathways predicted by the differentially expressed genes corresponding to LPS stimulation of BV-2 cells. The canonical pathways were ranked according to the –log (p-value). A ratio indicates the numbers of genes that were differentially expressed in each pathway over the total numbers of genes in that specific pathway. (B) The top disease and function network is associated with cell cycle, cellular development, and cellular growth and proliferation corresponding to the LPS stimulation of BV-2 cells. The identified genes involved in the networks were displayed in red (up-regulation) and green (down-regulation) color. The color intensity indicates the degree of regulation. Solid lines in the network imply direct interactions between genes, and dashed lines indicate indirect interactions. Geometric shapes represent different general functional families of gene regulation (diamond for enzyme, oval for transcription regulator, trapezoid for transporter, inverted triangle for kinase, double circle for complex/group, and circle for others).
AGE represses LPS-induced transcriptome alteration and FruArg partially contributes to the AGE’s LPS-repression activity
We co-treated the BV-2 cells with LPS and AGE in order to test whether AGE is capable of reversing the LPS-induced transcriptome alteration. Co-administration of AGE with LPS antagonized LPS-induced changes in the levels of 458 mRNAs (see Supplementary Figure S1). The expression levels of 215 LPS-stimulated mRNAs were reduced by AGE co-treatment relative to the levels in cells treated with LPS alone. Conversely, the expression levels of 92 LPS-repressed mRNAs were elevated by AGE co-treatment relative to the levels in cells treated with LPS alone (Fig. 3A,B, and Supplementary Figure S3A and Supplementary Table S2). A total of 307 genes accounted for 67% of the mRNAs whose expression was affected by AGE were also affected by LPS, in an opposing manner.
AGE and FruArg repress LPS-induced alteration of gene expression in BV-2 cells.
(A,B) Venn graph and heatmap showing AGE co-treatment with LPS stimulation was capable of repressing LPS-altered genes (LPS+/−). A large portion of AGE co-treatment down-regulated genes (AGE_LPS-) were up-regulated by LPS; similarly, a large portion of AGE co-treatment up-regulated genes (AGE_LPS+) were down-regulated by LPS treatment alone. Similar effect as AGE was also observed after FruArg treatment (FruArg_LPS+/−). In addition, there are 80 genes that were found to have the same LPS repressing activity after AGE or FruArg treatment. (C) Principle component analysis showed that these 80 genes effectively separated untreated control (UNT) from LPS stimulus, as well as co-treatment of LPS with either AGE or FruArg.
Because FruArg has been reported to be a bioactive component of AGE12,13, we tested its effect on LPS-altered gene expression in BV-2 cells, hypothesizing that some mRNAs would be altered by treatment with this compound in similar gene expression patterns treated by AGE. RNA-Seq analysis showed that the LPS-induced transcriptome alterations were antagonized by FruArg treatment. When compared to LPS alone, the co-treatment with FruArg altered 175 DEGs (see Supplementary Figure S2). The LPS-induced expression pattern of 96 mRNAs was reversed by FruArg co-treatment. Of these 96, the expression level of 77 LPS-stimulated mRNAs was reduced by FruArg co-treatment; the expression level of 19 LPS-repressed mRNAs was stimulated by FruArg co-treatment. In total, co-treatment with FruArg antagonized the alterations of 55% of the LPS-altered mRNA. Additionally, the expression levels of 80 out of the 96 mRNAs whose expression was altered by co-treatment with FruArg (83%) were also altered in the same way by co-treatment with AGE (Fig. 3A,B, and Supplementary Figure S3B and Supplementary Table S3).
To further establish the relationship between AGE, FruArg and LPS, we also performed Principal Component Analysis (PCA) on the normalized expression values of these 80 genes. The results demonstrated the consistency of these effects (Fig. 3C and Table 1). Along the first component, which accounts for 81% of variance, LPS-treated samples were clearly separated from the untreated controls. AGE co-treatments and FruArg co-treatments, although not clearly separated, were isolated from LPS treatments, supporting the idea that FruArg and AGE have similar activity on the expression of microglial mRNAs.
AGE and FruArg repressed LPS-altered canonical signaling pathways
To further understand the biological functions of AGE’s effects in LPS-stimulated BV-2 microglial cells, we mapped the mRNA identities to known signaling pathways and regulatory networks that were potentially affected by AGE treatment using IPA (Figs 4 and 5, and S4 and Supplementary Tables S5 and S6). The top 10 canonical pathways affected by the LPS stimulation and correlated genes, including Il1a, Il1b, Cd14, Casp3, Casp7, Nfkb1, Nfkb2, Tnf, Traf1, Traf2, Hmox1, Il6, and Tank, were repressed by AGE treatment (Fig. 4A) (see Supplementary Table S5). The most significant changes of canonical pathways modulated by AGE were TNFR2 signaling. Additionally, the disease/function networks that associated with AGE downregulated Casp1, Tlr3, Serp1, Irf1, Irf5, and Irf7 genes were involved in the antimicrobial response, inflammatory response, and infectious diseases (Fig. 4B). This gene interaction network was designated by IPA with the highest number of focus molecules (n = 27).
Effects of AGE on LPS-induced alterations of canonical pathways and disease/function networks in BV-2 microglial cells.
(A) Top 10 canonical pathways affected by LPS stimulation were repressed by AGE treatment. (B) The top disease and function network is associated with antimicrobial response, inflammatory response, and infectious diseases corresponding to the effects of AGE on LPS stimulation of BV-2 cells.
Effects of FruArg on LPS-induced alterations of canonical pathways and disease/function networks in BV-2 microglial cells.
(A) Top 10 canonical pathways predicted by the differentially expressed genes corresponding to the effects of FruArg on LPS induction of BV-2 cells. (B) The top disease and function network is associated with cellular movement, cell death and survival, and cell morphology corresponding to the effects of FruArg on LPS stimulation of BV-2 cells.
We also analyzed the canonical pathways related to disease/function networks to determine the regulation of FruArg on LPS-induced BV-2 cells. Similar to AGE, the top 10 canonical pathways and related genes, including Casp3, Casp7, Nfkb1, Nfkb2, Tnf, Traf2, Il1a, Il1b, Il6, Cd14, Il36g, Map2k3, Map2k6, and Hmox1, affected by the LPS stimulation were all repressed by FruArg treatment (Fig. 5A, Supplementary Table S6). The most significant change of canonical pathways modulated by AGE was TNFR2 signaling.
Subsequently, we compared the canonical pathways regulated by both AGE and FruArg (see Supplementary Figure S4). AGE and FruArg share major canonical pathways such as role of pattern recognition receptor in recognition of bacteria and viruses, interferon signaling, dendritic cell maturation, activation of IRF by cystolic pattern recognition receptors, p53 signaling, and retinoic acid mediated apoptosis signaling, endothelin-1 signaling, death receptor signaling, role of NFAT in regulation of the immune response, Nrf2-mediated oxidative stress response, and acute phase response signaling. The data indicated FruArg and AGE targeted common pathways that were modulated by LPS. Moreover, the top network is associated with cellular movement, cell death and survival, and cell morphology by regulating genes, such as Casp1, Cd81, Cdk5, Gapdh, Nupr1, and Serp1 (Fig. 5B). This gene interaction network was designated with the highest number of focus molecules (n = 16). Hence, the canonical pathways and cellular networks suggested that FruArg exhibits potent anti-inflammatory ability to reverse LPS-induced alterations.
Discussion
Microglia serve as resident immune cells in the central nervous system (CNS). Chronic neuroinflammatory responses mediated by activated microglial cells in the central nervous system have critical roles in the pathogenesis of numerous neurodegenerative diseases, including Alzheimer’s disease and Parkinson’s disease23,24,25,26. Various inflammatory mediators and cytotoxic molecules, such as IL-1α, IL-6, TNF-α, ROS, and NO, released by chronic activated microglial cells in response to infection, injury, or endotoxins (e.g. LPS) may trigger neuronal damage and even cell death27,28,29,30. NO serves as a lipid-soluble free signaling molecule that transports freely across cell membranes31,32,33. Nitrosative stress can be induced by excessive production of NO and other reactive nitrogen species (RNS), which contributes to the pathogenesis of traumatic brain injury, Parkinson’s disease, and Alzheimer’s disease34,35,36,37,38,39. LPS stimulation can significantly increase NO concentration in microglia. Antioxidants such as AGE and FruArg could alleviate microglial activation and suppress the nitrosative stress by reducing NO production, and thus have a potential to counteract the undesirable effects of chronic activation of microglial cells. In the present study, we showed the altered expression of LPS responsive genes, including Traf1, Nos2, Saa3, Lcn2, Il6, Irg1, Il1b, and Cxcl3. The up-regulation of these genes confirmed the activation of BV-2 microglial cells by LPS. Especially, NOS2 and its gene product, inducible NOS (iNOS), is involved in neuroinflammation by generating NO and plays key roles in the pathogenesis of certain neurodegenerative disorders40,41,42. Traf1 (tumor necrosis factor receptor-associated factor 1) gene is also reported to regulate the inflammatory response43,44. In addition, Il6 gene is related to not only proinflammatory, but also neurodevelopment functions45,46. A total of 2563 genes were shown to respond to LPS stimulation. These genes are associated with death receptor signaling, toll-like receptor signaling, IL-10 signaling, TNFR2 signaling, role of pattern recognition receptors in recognition of bacteria and viruses canonical pathways (Fig. 2A). Top networks modulated by LPS exposure are related to cell cycle, cellular development, cellular growth and proliferation, protein synthesis, and cellular function and maintenance (Fig. 2B). Overall, our gene expression profiling in LPS-treated BV-2 microglial cells provided a useful set of pathway markers.
Both AGE and its bioactive component FruArg demonstrated a potential in neuronal protection against cellular oxidative stress and inflammatory responses by directly scavenging superoxide free radicals4,5,6,18,20. The neuroprotective effects of AGE and FruArg were realized through anti-oxidant/anti-inflammatory activities and targeting certain intracellular proteins and molecular pathways, according to data obtained by quantitative proteomic approach in our previous study20. We have demonstrated the significant suppression of NO production by either AGE or FruArg treatment and the multi-modal regulation of protein expression in LPS-induced BV-2 microglial cells. It also suggested that AGE and FruArg are similar to a number of botanicals by modulating the toll-like receptor and nuclear translocation of transcription factors for gene regulation47,48,49,50,51. Here in this study, we analyzed the gene expression profiles of both AGE and FruArg treatment to further investigate their effects on LPS-induced BV-2 microglial cells. 67% of AGE altered genes and 55% of FruArg altered genes showed reversed activity compared to LPS stimulation. It is clear that the major activity of AGE and FruArg is to repress LPS’s activity in gene expression. Furthermore, the shared modulation of genes by co-treatment of LPS between AGE and FruArg show that FruArg serves as a functional compound in AGE, in consistence with our previous proteomic study20, that accounts for its protective effects. We conclude that FruArg contributed to the anti-inflammatory effect of AGE by affecting mRNA levels associated with stress and inflammation.
FruArg is an Amadori product from AGE generated by the amino-carbonyl (Maillard) reaction, which is a nonenzymatic browning reaction of amino acids with sugars16. At the protein level, it was shown by both our proteomic analysis and other reports that the Maillard reaction products may decrease pro-oxidant activity in cells by affecting enzymatic activity of peroxiredoxin-1, mitochondrial superoxide dismutase 2, and glutamate-cysteine ligase in LPS-induced BV-2 cells14,20. At gene expression levels, FruArg also exhibited ability to suppress the production of certain inflammatory mediators by inhibiting the gene expression of Il1a, Il1b, Nfkb1, Nfkb2, Traf1, Traf2, Casp1, Cd14, Il6, and Pi3kr5. Additionally, by reducing the levels of NOS2 gene, FruArg could inhibit the excessive production of NO induced by LPS. The data demonstrate that FruArg can modulate and reverse stimulation of microglia by LPS at gene level. The ability to repress the pro-inflammatory response and oxidative stress both at protein and gene levels suggests that FruArg may have a medicinal value. On the other hand, from the perspective of canonical pathway analysis, both our previous proteomic and current RNA-Seq study show that FruArg can alter the Nrf2-mediated oxidative stress response pathway, which plays a major role for antioxidant protection20,52,53,54. Genes as Gsta3, Gstm5, Gstp1, Hmox1, and Mgst2 related to Nrf2 mediated oxidative stress pathway were all up-regulated. The expression of these genes may enhance the subsequent activities of glutathione S-transferase and heme oxygenase 1 to catalyze the conjugation of reduced form of glutathione with excess oxidants, to detoxify and to catabolize free heme and produce carbon monoxide. Therefore, the neuroprotection mechanism of FruArg may involve both suppression of pro-inflammatory and promotion of antioxidant genes.
In conclusion, our results suggest that AGE and FruArg are capable of alleviating oxidative and neuroinflammatory responses stimulated by LPS in BV-2 cells by modulating gene expression. Inhibition of the expression of multiple immune- and inflammation-related genes suggested that AGE and FruArg could be drug candidates for the prevention of inflammation-mediated neurodegenerative diseases.
Materials and Methods
Cell culture and stimulation
The immortalized mouse microglial cells (BV-2) were originally obtained from Dr. R. Donato (University of Perugia, Italy) and were gifted from co-author Dr. Grace Y. Sun55,56. The cells, as previously described, were grown in Dulbecco’s modified Eagle’s medium (DMEM) (Gibco, Grand Island, NY, USA) supplemented with 5% (v/v) heat-inactivated fetal bovine serum (FBS) (Atlanta Biologicals, Inc., Lawrenceville, GA, USA), 25U/mL penicillin, and 25mg/mL streptomycin (Gibco, Grand Island, NY, USA) in a saturated humidity atmosphere containing 95% (v/v) air and 5% (v/v) CO2 at 37 °C57,58. Cells were cultured in DMEM without serum for 4 hours at 70–80% confluence and exposed to 100ng/mL LPS (rough strains from Escherichia coli F583 Rd mutant, Sigma-Aldrich, St. Louis, MO, USA) for 20 hours with or without aqueous AGE (0.5%, v/v) (Wakunaga of America, Mission Viejo, CA, USA) and FruArg (3 mM), which were added 1 hour prior to LPS exposure, respectively. FruArg was chemically synthesized and the purity and stability were measured as previously described20.
Total RNA extraction and RNA-Seq
Total RNA was extracted using TRIzol (Life Technologies) according to the manufacture’s protocol. Briefly, 0.2 mL of chloroform was added to the homogenized sample and then the tubes were shaken for 15 seconds and incubated for 2–3 minutes at room temperature. Next, the mixture was centrifuged at 12,000 × g for 10 minutes at 4 °C, and the aqueous phase of the sample was extracted and placed into new tubes. Then, 0.5 mL of 100% isopropanol was added to the aqueous phase and incubated at room temperature for 10 minutes. The lysis mixture was centrifuged 12,000 × g for 10 minutes at 4 °C. The supernatant from the tube was removed and washed with 1 mL of 75% ethanol. The RNA pellet was vortexed briefly, centrifuged at 7,500 × g for 5 minutes at 4 °C, and dried at room temperature for 5–10 minutes. Finally, the RNA pellet was resuspended in RNase-free water or 0.5% SDS solution and incubated in a water bath or heat block set at 55–60 °C for 10–15 minutes. Total RNA concentration was determined using a spectrophotometer (NanoDrop Technologies, Wilmington, DE, USA). RNA quality was verified on an Agilent Bioanalyzer (Agilent Technologies, Santa Clara, CA) to confirm that RIN scores were above 7 prior to sequencing. RNA sequencing was performed upon libraries constructed using the Illumina TrueSeq library preparation system that employs a poly-A tail selection. RNA libraries were sequenced as 100nt single-end fragments using Illumina Hi-Seq 2000 system (Illumina, Inc., San Diego, CA, USA). Adaptor sequences were clipped off first. Subsequently, sequencing reads were trimmed and filtered based on quality scores by using fastx_toolbox (http://hannonlab.cshl.edu/fastx_toolkit/) that removed low-scoring sections of each read and preserved the longest remaining sequencing reads.
Differential gene expression analysis
The trimmed and filtered reads were mapped to mouse genome (Mus_musculus.GRCm38) from Ensemble using Tophat259. Mapped reads were quantified as raw counts by the Subreads package function “featureCounts”. Differentially expressed genes were analyzed using R/Bioconductor package DESeq2. For a gene to be a Differentially Expressed Genes (DEGs), it has to alter at least 2 fold with False Discovery Rate adjusted p-value (p-adj) less than 0.05 (Log2FC ≥1 or Log2FC ≤−1, FDR <0.05). For data visualization, the normalized gene expression value in the format of Count per Million reads (cpm) was log2-transformed and plotted by R package gplots60.
Principal component analysis (PCA)
Raw gene counts were normalized to library sizes of each sample and further log2 transformed. PCA analysis was performed using R build-in package “prcomp”. The first three components, which accounted for 98% of the variance in dataset were plotted using R package “rgl”.
Functional annotation and canonical pathway analysis
The data were analyzed using QIAGEN’s Ingenuity® Pathway Analysis (IPA®, QIAGEN Redwood City, www.qiagen.com/ingenuity) for associated gene functions and pathways, as well as to predict gene interaction networks. The significance values for the canonical pathways and networks are calculated by right-tailed Fisher’s exact test. The cutoff p-value is 0.05. Taller bars equate to increased significance.
Accessibility of raw sequencing data and accession number
The raw sequencing reads files and process data files were submitted to NCBI GEO. The accession number is GSE87531.
Additional Information
How to cite this article: Song, H. et al. Effects of aged garlic extract and FruArg on gene expression and signaling pathways in lipopolysaccharide-activated microglial cells. Sci. Rep. 6, 35323; doi: 10.1038/srep35323 (2016).
References
Stabler, S. N., Tejani, A. M., Huynh, F. & Fowkes, C. Garlic for the prevention of cardiovascular morbidity and mortality in hypertensive patients. Cochrane Database Syst Rev 8, CD007653, 10.1002/14651858.CD007653.pub2 (2012).
Rahman, K. Effects of garlic on platelet biochemistry and physiology. Mol Nutr Food Res 51, 1335–1344, 10.1002/mnfr.200700058 (2007).
Amagase, H., Schaffer, E. M. & Milner, J. A. Dietary components modify the ability of garlic to suppress 7, 12-dimethylbenz (a) anthracene-induced mammary DNA adducts. The Journal of nutrition 126, 817 (1996).
Moriguchi, T., Saito, H. & Nishiyama, N. Anti‐ageing effect of aged garlic extract in the inbred brain atrophy mouse model. Clinical and experimental pharmacology and physiology 24, 235–242 (1997).
Kyo, E., Uda, N., Kasuga, S. & Itakura, Y. Immunomodulatory effects of aged garlic extract. J Nutr 131, 1075S–1079S (2001).
Morihara, N., Hayama, M. & Fujii, H. Aged garlic extract scavenges superoxide radicals. Plant Foods Hum Nutr 66, 17–21, 10.1007/s11130-011-0216-6 (2011).
Imai, J. et al. Antioxidant and radical scavenging effects of aged garlic extract and its constituents. Planta medica 60, 417–420 (1994).
Borek, C. Antioxidant health effects of aged garlic extract. The Journal of nutrition 131, 1010S–1015S (2001).
Steiner, M., Khan, A. H., Holbert, D. & Lin, R. A double-blind crossover study in moderately hypercholesterolemic men that compared the effect of aged garlic extract and placebo administration on blood lipids. The American journal of clinical nutrition 64, 866–870 (1996).
Rahman, K. & Billington, D. Dietary supplementation with aged garlic extract inhibits ADP-induced platelet aggregation in humans. The Journal of nutrition 130, 2662–2665 (2000).
Steiner, M. & Li, W. Aged garlic extract, a modulator of cardiovascular risk factors: a dose-finding study on the effects of AGE on platelet functions. The Journal of nutrition 131, 980S–984S (2001).
Amagase, H. Clarifying the real bioactive constituents of garlic. J Nutr 136, 716S–725S (2006).
Mossine, V. V. & Mawhinney, T. P. 1-Amino-1-deoxy-D-fructose (“fructosamine”) and its derivatives. Adv Carbohydr Chem Biochem 64, 291–402, 10.1016/S0065-2318(10)64006-1 (2010).
Ryu, K., Ide, N., Matsuura, H. & Itakura, Y. Nα-(1-deoxy-D-fructos-1-yl)-L-arginine, an antioxidant compound identified in aged garlic extract. The Journal of nutrition 131, 972S–976S (2001).
Ledl, F. Chemical pathways of the Maillard reaction. The Maillard reaction in food processing, human nutrition and physiology 19–42 (1990).
Martins, S. I., Jongen, W. M. & Van Boekel, M. A. A review of Maillard reaction in food and implications to kinetic modelling. Trends in Food Science & Technology 11, 364–373 (2000).
Mossine, V. V., Glinsky, V. V. & Mawhinney, T. P. Antitumor effects of the early Maillard reaction products, in The Maillard Reaction: Interface between Aging, Nutrition and Metabolism, M.C. Thomas and J. Forbes, Editors. Royal Society of Chemistry 170–179 (2010).
Mossine, V. V., Chopra, P. & Mawhinney, T. P. Interaction of tomato lycopene and ketosamine against rat prostate tumorigenesis. Cancer Research 68, 4384–4391 (2008).
Ide, N., Lau, B. H., Ryu, K., Matsuura, H. & Itakura, Y. Antioxidant effects of fructosyl arginine, a Maillard reaction product in aged garlic extract. The Journal of nutritional biochemistry 10, 372–376 (1999).
Zhou, H. et al. Proteomic analysis of the effects of aged garlic extract and its FruArg component on lipopolysaccharide-induced neuroinflammatory response in microglial cells. PLoS One 9, e113531, 10.1371/journal.pone.0113531 (2014).
Das, A. et al. Dual RNA sequencing reveals the expression of unique transcriptomic signatures in lipopolysaccharide-induced BV-2 microglial cells. PloS one 10, e0121117 (2015).
Schneiderman, A. I., Braver, E. R. & Kang, H. K. Understanding sequelae of injury mechanisms and mild traumatic brain injury incurred during the conflicts in Iraq and Afghanistan: persistent postconcussive symptoms and posttraumatic stress disorder. American journal of epidemiology 167, 1446–1452 (2008).
Streit, W. J. Microglial activation and neuroinflammation in Alzheimer’s disease: a critical examination of recent history. Frontiers in aging neuroscience 2 (2010).
Rogers, J., Mastroeni, D., Leonard, B., Joyce, J. & Grover, A. Neuroinflammation in Alzheimer’s disease and Parkinson’s disease: are microglia pathogenic in either disorder? International review of neurobiology 82, 235–246 (2007).
Qian, L. & Flood, P. M. Microglial cells and Parkinson’s disease. Immunologic research 41, 155–164 (2008).
Town, T., Nikolic, V. & Tan, J. The microglial. Journal of neuroinflammation 2, 24 (2005).
Wood, P. Neuroinflammation: mechanisms and management. (Springer Science & Business Media, 2002).
Block, M. L., Zecca, L. & Hong, J.-S. Microglia-mediated neurotoxicity: uncovering the molecular mechanisms. Nature Reviews Neuroscience 8, 57–69 (2007).
Graeber, M. B. & Streit, W. J. Microglia: biology and pathology. Acta neuropathologica 119, 89–105 (2010).
Di Filippo, M. et al. Persistent activation of microglia and NADPH drive hippocampal dysfunction in experimental multiple sclerosis. Scientific reports 6 (2016).
Pacher, P., Beckman, J. S. & Liaudet, L. Nitric oxide and peroxynitrite in health and disease. Physiological reviews 87, 315–424 (2007).
Bal-Price, A. & Brown, G. C. Inflammatory neurodegeneration mediated by nitric oxide from activated glia-inhibiting neuronal respiration, causing glutamate release and excitotoxicity. The Journal of Neuroscience 21, 6480–6491 (2001).
Gibbons, H. M. & Dragunow, M. Microglia induce neural cell death via a proximity-dependent mechanism involving nitric oxide. Brain research 1084, 1–15 (2006).
Tsang, A. H. & Chung, K. K. Oxidative and nitrosative stress in Parkinson’s disease. Biochimica et Biophysica Acta (BBA)-Molecular Basis of Disease 1792, 643–650 (2009).
Danielson, S. R. & Andersen, J. K. Oxidative and nitrative protein modifications in Parkinson’s disease. Free Radical Biology and Medicine 44, 1787–1794 (2008).
Gu, Z., Nakamura, T., Yao, D., Shi, Z. & Lipton, S. Nitrosative and oxidative stress links dysfunctional ubiquitination to Parkinson’s disease. Cell death and differentiation 12, 1202–1204 (2005).
Calabrese, V. et al. Nitrosative stress, cellular stress response, and thiol homeostasis in patients with Alzheimer’s disease. Antioxidants & redox signaling 8, 1975–1986 (2006).
Di Pietro, V. et al. Neuroglobin expression and oxidant/antioxidant balance after graded traumatic brain injury in the rat. Free Radical Biology and Medicine 69, 258–264 (2014).
Buchwalow, I. et al. L-arginine-NO-cGMP signalling pathway in pancreatitis. Scientific reports 3, 1899 (2013).
Colton, C. A. et al. The effects of NOS2 gene deletion on mice expressing mutated human AβPP. Journal of Alzheimer’s disease: JAD 15, 571 (2008).
Dawson, V. L. & Dawson, T. M. Nitric oxide in neurodegeneration. Progress in brain research 118, 215–229 (1998).
Uttara, B., Singh, A. V., Zamboni, P. & Mahajan, R. Oxidative stress and neurodegenerative diseases: a review of upstream and downstream antioxidant therapeutic options. Current neuropharmacology 7, 65 (2009).
Bryce, P. J., Oyoshi, M. K., Kawamoto, S., Oettgen, H. C. & Tsitsikov, E. N. TRAF1 regulates Th2 differentiation, allergic inflammation and nuclear localization of the Th2 transcription factor, NIP45. International immunology 18, 101–111 (2006).
Cerhan, J. R. et al. Genetic variation in 1253 immune and inflammation genes and risk of non-Hodgkin lymphoma. Blood 110, 4455–4463 (2007).
Scheller, J., Chalaris, A., Schmidt-Arras, D. & Rose-John, S. The pro-and anti-inflammatory properties of the cytokine interleukin-6. Biochimica et Biophysica Acta (BBA)-Molecular Cell Research 1813, 878–888 (2011).
Baune, B. T. et al. Interleukin-6 gene (IL-6): a possible role in brain morphology in the healthy adult brain. Journal of neuroinflammation 9, 125 (2012).
Yoon, Y. D. et al. Toll-like receptor 4-dependent activation of macrophages by polysaccharide isolated from the radix of Platycodon grandiflorum. International immunopharmacology 3, 1873–1882 (2003).
Chino, A. et al. Juzentaihoto, a Kampo medicine, enhances IL-12 production by modulating Toll-like receptor 4 signaling pathways in murine peritoneal exudate macrophages. International immunopharmacology 5, 871–882 (2005).
Patwardhan, B. & Gautam, M. Botanical immunodrugs: scope and opportunities. Drug discovery today 10, 495–502 (2005).
Pugh, N. D. et al. Melanin: dietary mucosal immune modulator from Echinacea and other botanical supplements. International immunopharmacology 5, 637–647 (2005).
Zhao, L., Lee, J. Y. & Hwang, D. H. Inhibition of pattern recognition receptor-mediated inflammation by bioactive phytochemicals. Nutrition reviews 69, 310–320 (2011).
Zhang, M. et al. Emerging roles of Nrf2 and phase II antioxidant enzymes in neuroprotection. Progress in neurobiology 100, 30–47 (2013).
Sethy, N. K., Singh, M., Kumar, R., Ilavazhagan, G. & Bhargava, K. Upregulation of transcription factor NRF2-mediated oxidative stress response pathway in rat brain under short-term chronic hypobaric hypoxia. Functional & integrative genomics 11, 119–137 (2011).
Jung, J.-S. et al. Anti-inflammatory mechanism of exogenous C2 ceramide in lipopolysaccharide-stimulated microglia. Biochimica et Biophysica Acta (BBA)-Molecular and Cell Biology of Lipids 1831, 1016–1026 (2013).
Blasi, E., Barluzzi, R., Bocchini, V., Mazzolla, R. & Bistoni, F. Immortalization of murine microglial cells by a v-raf/v-myc carrying retrovirus. J Neuroimmunol 27, 229–237 (1990).
Bocchini, V. et al. An immortalized cell line expresses properties of activated microglial cells. J Neurosci Res 31, 616–621, 10.1002/jnr.490310405 (1992).
Shen, S. et al. Distinct signaling pathways for induction of type II NOS by IFNgamma and LPS in BV-2 microglial cells. Neurochem Int 47, 298–307, 10.1016/j.neuint.2005.03.007 (2005).
Jiang, J. et al. Sutherlandia frutescens ethanol extracts inhibit oxidative stress and inflammatory responses in neurons and microglial cells. PLoS One 9, e89748, 10.1371/journal.pone.0089748 (2014).
Li, J. et al. From Gigabyte to Kilobyte: A Bioinformatics Protocol for Mining Large RNA-Seq Transcriptomics Data. PLoS One 10, e0125000, 10.1371/journal.pone.0125000 (2015).
Lu, Y., Li, J., Cheng, J. & Lubahn, D. B. Genes targeted by the Hedgehog-signaling pathway can be regulated by Estrogen related receptor β. BMC molecular biology 16, 19 (2015).
Acknowledgements
This publication was made possible by funding of the Department of Pathology and Anatomical Sciences research fund at the University of Missouri School of Medicine (to Z.G.), as well as by Grant Number P50AT006273 from the National Center for Complementary and Integrative Health (NCCIH), the Office of Dietary Supplements (ODS), and the National Cancer Institute (NCI). Its contents are solely the responsibility of the authors and do not necessarily represent the official views of the NIEHS, NCCIH, ODS, NCI, or the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: Z.G. Performed the experiments: H.S., Y.L., Z.Q., V.V.M., M.B.M. and B.A.P. Analyzed the data: H.S., Y.L., J.H., J.C., F.J.S. and Z.G. Contributed reagents/materials/analysis tools: V.V.M., T.P.M., J.C., C.M.G., D.B.L., R.B.W. and G.Y.S. Wrote the paper: H.S., Y.L. and Z.G. All authors reviewed and approved the final version of the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Song, H., Lu, Y., Qu, Z. et al. Effects of aged garlic extract and FruArg on gene expression and signaling pathways in lipopolysaccharide-activated microglial cells. Sci Rep 6, 35323 (2016). https://doi.org/10.1038/srep35323
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep35323
This article is cited by
-
Acute Diallyl Disulfide Administration Prevents and Reveres Lipopolysaccharide-Induced Depression-Like Behaviors in Mice via Regulating Neuroinflammation and Oxido-Nitrosative Stress
Inflammation (2021)
-
Harpagophytum procumbens Extract Ameliorates Allodynia and Modulates Oxidative and Antioxidant Stress Pathways in a Rat Model of Spinal Cord Injury
NeuroMolecular Medicine (2020)
-
From Analysis of Ischemic Mouse Brain Proteome to Identification of Human Serum Clusterin as a Potential Biomarker for Severity of Acute Ischemic Stroke
Translational Stroke Research (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.