Abstract
Human pluripotent stem cells (hPSCs) may significantly improve drug development pipeline, serving as an in vitro system for the identification of novel leads, and for testing drug toxicity. Furthermore, these cells may be used to address the issue of differential drug response, a phenomenon greatly influenced by genetic factors. This application depends on the availability of hPSC lines from populations with diverse ancestries. So far, it has been reported that most lines of hPSCs derived worldwide are of European or East Asian ancestries. We have established 23 lines of hPSCs from Brazilian individuals, and we report the analysis of their genomic ancestry. We show that embryo-derived PSCs are mostly of European descent, while induced PSCs derived from participants of a national-wide Brazilian cohort study present high levels of admixed European, African and Native American genomic ancestry. Additionally, we use high density SNP data and estimate local ancestries, particularly those of CYP genes loci. Such information will be of key importance when interpreting variation among cell lines with respect to cellular phenotypes of interest. The availability of genetically admixed lines of hPSCs will be of relevance when setting up future in vitro studies of drug response.
Similar content being viewed by others
Introduction
Human pluripotent stem cells (hPSCs) are an ideal cell source for the development of cell based assays for drug response. In addition to their extensive proliferation and genetic stability in culture, these human cells can give rise to primary cell types relevant for drug response, including cardiomyocytes, hepatocytes and neurons1. Individual differences in drug response can result from the effects of age, sex, disease, ancestry, or drug interactions, but genetic factors play a major role in influencing adverse drug reactions and ineffective therapy2. Thus, a collection of genetically diverse lines of hPSCs is required for a broader study of differential drug response in vitro3.
To date, most lines of hPSCs available are of European or Eastern Asian ancestry4, although one hiPSC line from a Native American, and one from an African (Yoruba) have been reported5. More recently, one hiPSC line of African American and of Hispanic Latino ancestry each have been described, although the authors do not show the genetic evidence of those ethnicities6.
The Brazilian population results from 500 years of admixture among the original Native Americans, Europeans (mostly Portuguese), and sub-Saharan Africans, most of which were brought to the country as slaves7. Different analyses of genomic ancestry in Brazil have shown that, on average, the urban population has 60% contribution from European, 25% from African, and 15% from Native American populations, although these proportions vary according to the Brazilian geographic region analyzed7,8,9,10,11. Therefore, from the genetic point of view the Brazilian population is significantly distinct from the ancestral populations, containing novel genotypes and haplotypes that may impact various phenotypes, including drug response.
We have previously established five lines of hPSCs from surplus human embryos generated for reproduction purpose and stored in private human reproduction clinics in Brazil12,13,14. Here we report the derivation of 18 lines of hPSCs from peripheral blood of participants of The Brazilian Longitudinal Study of Adult Health (ELSA-Brasil), a large multicenter cohort study of 15,105 Brazilians, focused on assessing incidence and risk factors for diabetes and cardiovascular diseases in the country15,16. We analyze the genomic ancestry of the different hPSC lines, and show that while the lines derived from human embryos (human embryonic stem cells, hESCs) are mostly of European genomic descent, those derived from the participants of the ELSA-Brasil present a broader range of ancestries, providing a better reflection of the genetic admixture that characterizes the Brazilian population. The hiPSCs described here significantly increase the spectrum of genetic admixture of the available lines, and can be an important resource for the study of the molecular basis of differential drug response.
Results and Discussion
Genomic ancestry of hPSCs
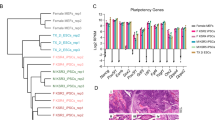
Mononuclear cells from peripheral blood of 1,872 participants of the ELSA study were isolated and cryopreserved. Eighteen of those were randomly selected for hiPSC derivation. Reprogrammed colonies were pooled and expanded for pluripotency evaluation. Cell lines were shown to have normal karyotype (data not shown), homogeneously express the pluripotency markers OCT4, NANOG and SSEA-4, and to differentiate in vitro into tissues from the three embryonic germ layers (Supplemental Fig. 1). In addition, PCR analysis showed no integration of the reprogramming vectors, and thus the hiPSC lines are footprint-free (Supplemental Fig. 1).
Genomic ancestries of five lines of hESCs derived from embryos generated in Brazil for reproductive purpose and of the eighteen hiPSCs were determined by the analysis of 192 ancestry informative markers and comparison with reference populations (Fig. 1). The data showed that Brazilian hESC lines are mostly of European ancestry, with the European genomic component ranging from 92.7% to 98.6%. In contrast, in the 18 randomly chosen lines of hiPSCs from the ELSA-Brasil, the European genomic contribution ranged from 14.2% to 95%, while African ancestry ranged from 1.6% to 55.1%, and Native American ancestry ranged from 7% to 56% (Fig. 1a,b). Principal component analysis (PCA) of hiPSCs showed that most of them are on the African - European variation axis (Fig. 1c).
Genomic ancestry of hPSCs.
(A) Ternary plot of African, European, and Native American ancestry in hPSC lines. Each yellow point represents hiPSC lines, purple points represents hESC lines. Each point is positioned within the triangle reflecting the amount of ancestry estimated from each cell line; (B) Percentage of the contributions of different ancestral genomes of each hPSCS; (C) Principal component (PC) analysis of the hiPSC lines. PC1 and PC2 are plotted in the x and y axes, respectively. Comparison populations from the HGDP and HapMap are shown in green (European), red (Africans) and purple (Native American). hiPSCs are represented by black circles.
Local ancestry of hPSCs around CYP genes
We randomly chose two hESCs and four hiPSCs to perform a local ancestry analysis, i.e., to determine the ancestry for each physical location in the genome. As expected under a scenario of admixture that started several generations ago, the chromosomes of the hPSC lines analyzed (hESCS: BR-4 and BR-5, and hiPSCS IPS2, IPS3, IPS4 and IPS5) are mosaics comprised of segments of the three ancestries (Fig. 2). We investigated the local ancestry of specific genes belonging to the Cytochrome P450 (CYP) family (CYP1A2, CYP2C8, CYP3A4, CYP2C9, CYP2C19, CYP2D6, CYP2B6, CYP2A6), known for its role in drug metabolism. We found a high level of heterogeneity of ancestry among the hPSC lines, with all CYP genes investigated having cell lines with representatives from African and European ancestries, and all loci except CYP2C8/CYP2C9/CYP2C19 having at least one instance in which it lies in a region of Native American ancestry (Fig. 3).
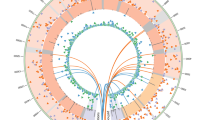
Local ancestry estimative in two hESC (BR-4 and BR-5) and four hiPSC lines (IPS2, IPS3, IPS4, IPS5).
Each row is a chromosome sample and every two consecutive rows is a pair of individual chromosomes. The column represents the SNPs and the colors represent the ancestry: Blue – African, Green –European, Red – Native American.
Local ancestry of hPSCs around CYP genes.
Local ancestry estimative for CYP family genes in two hESC (BR-4 and BR-5) and four hiPSC lines (IPS2, IPS3, IPS4, IPS5). Each row is a chromosome sample and every two consecutive rows is a pair of individual chromosomes. The column represents the SNPs and the colors represent the ancestry: Blue – African, Green – European, Red – Native American. The horizontal dashed lines indicate the location of the CYP gene indicated.
Human PSCs are a valuable model system in which to study Mendelian conditions, where a single gene mutation triggers the clinical phenotype17. The use of these cells to model complex disorders, where the phenotype results from the interaction of multiple mutations and environmental factors, is far more challenging18. In particular, differential drug response has been shown to be greatly influenced by genetic factors. Thus, collections of hPSCs with different genetic backgrounds must be used to dissect the molecular basis and to develop cell based assays of differential drug toxicity and efficacy. Nevertheless, three studies of genomic ancestry have shown limited ethnic diversity of the available lines of hPSCs, the great majority being of European and Asian ancestry4,5,19. Importantly, none of those lines derived from populations with recent African ancestry19.
Analysis of the genomic ancestry of all lines of hESCs derived in Brazil confirmed our initial hypothesis that the embryos available for research do not represent the ethnic diversity of the Brazilian population12. While the European contribution to the Brazilian population ranges from 37% to 82%20, the five Brazilian lines of hESCs were 92% to 98% of European genomic ancestry. In contrast, eighteen hiPSCs analyzed here presented a broad range of genomic ancestries, with the European contribution ranging from 14.2% to 95.0%. In addition, the PCA shows that the ancestry variability of the hiPSCs is mainly between the European and African components. Thus, we were able to find high genetic admixture in hiPSCs derived from participants of the ELSA-Brasil cohort study.
Moreover, we also analyzed the ancestry of 2 hESC and 4 hiPSC in the region containing eight genes from the CYP family, responsible for the metabolism of the majority of drugs and xenobiotics. We looked specifically at CYP family members that have polymorphisms described in different frequency between the ethnic groups, and different levels of metabolic response reported21. We showed the analyzed hPSC have great heterogeneity of ancestry in the regions of CYP genes. We also showed that although there is a predominance of European ancestry genomewide, different ancestries are often present in the genomic region of interest, including the occurrence of Native American ancestry for CYP genes in otherwise predominantly European genomes. This genetic heterogeneity is of great interest, since possible genetic variants of specific ancestry may be present and influence the functionality of gene products related to differential drug response and other complex phenotypes.
The ELSA-Brasil cohort study represents a unique opportunity for the generation of collections of hiPSCs for different research purposes. The study collects data from 15,000 35–74 year old participants from three regions of Brazil (Northeast, Southeast and South) every 3–4 years in the form of interviews, electrocardiogram, blood sample, blood pressure, as well as other clinical, biochemical and genetic tests15. Base-line findings identified several interesting clinical groups, including diabetes (19.7%), common mental disorders (26.7%) and coronary heart disease (4.7%)16. Thus, while a collection of genetically admixed hiPSCs can be generated from these participants for future in vitro clinical trials and other studies of drug response, the availability of extensive clinical data of each participant permits the generation of different collections of cells based on phenotypes of interest. For example, a group of 4,116 participants being treated for hypertension was identified, 11% of those being resistant to pharmacological intervention22 – we are currently generating a collection of hiPSCs from responsive and resistant hypertensive individuals to understand and to predict resistant hypertension.
Conclusions
We derived lines of hiPSCs with novel combinations of genomic ancestry, including Native American and African. In addition, we have established a collection of primary cells from a well clinically characterized population ready to be reprogrammed into hiPSCs for different research purposes. The new hiPSC lines described here significantly increase the ethnic diversity and genetic admixture of the currently available hPSCs.
Methods
Subjects
Participants of the ELSA-Brasil in São Paulo were invited to participate in this research. Exclusion criteria were current or recent (<4 months prior to the first interview) pregnancy, intention to quit working at the institution in the near future, severe cognitive or communication impairment, and, if retired, residence outside of a study center’s corresponding metropolitan area. The first examination was carried out from 2008 through 2010. Annual telephone surveillance for outcomes is now in its seventh year, and the first follow-up examination was conducted form 2012 through 2014. From the 5061 participants, 1872 were randomly chosen to be included in the present analysis. Participants signed informed consents. The project was carried out in accordance with the guidelines for research on human subjects, and approved by the Ethics Committee of the University Hospital, University of São Paulo.
Human PSC lines
Human embryonic stem cell lines BR-1, BR-2, BR-4, BR-5 and BR-6 were cultured as described12,13,14. Human iPSC lines were derived from erythroblasts with episomal vectors using a protocol described23,24 with modifications. Briefly, mononuclear cells were isolated from 10 ml of peripheral blood by Ficoll gradient. Erythroblasts were cultured in a serum-free mononuclear cell (MNC) medium containing the following cytokines diluted in Stem Span Serum Free Expansion Medium (Stem Cell Technologies, Cat No. 09650): insulin-like growth factor 1 (IGF-1): 40 ng/ml; Stem Cell Factor (SCF): 100 ng/ml; Interleukin 3 (IL-3):10 ng/ml; erythropoietin (EPO): 2 U/ml. Two million cells were transfected with plasmids pEB-C5 and pEB-Tg (Addgene), containing reprogramming factors Oct4, Sox2, Klf4, cMyc, Lin28 and SV40-T, using the Human CD34+ nucleofector kit and the Nucleofector II device, both by Lonza (Basel, Switzerland) following manufacturer’s instructions. Reprogrammed erythroblasts were incubated in MEF-coated plates in MEF medium and FBS ES-Cell Qualified (ESQ, Invitrogen) with basic fibroblast growth factor (bFGF; 20 ng/ml) overnight. Then, they were transferred into embryonic stem cell (ESC) medium containing Knockout DMEM (Life Technologies), Knockout Serum replacement (Life Technologies, Catalog No. 10828–028), Antibiotic-antimycotic (Life Technologies), Glutamax 200 mM (Life Technologies), MEM non‐essential amino acid solution (Life Technologies), 2‐mercaptoethanol (Life Technologies) supplemented with bFGF (20 ng/ml) and Sodium butyrate (0.25 mM). hiPSC colonies were passaged from a 6 well MEF-coated plate into Matrigel (BD)-coated plates with E8 medium (Invitrogen), using Gentle Cell Dissociation Reagent (Stemcell Technologies) and 10 μM ROCK inhibitor Y-27632 (Stemgent).
Flow Cytometry
Cells were fixed and labeled using Human Pluripotent Stem Cell Transcription Factor Analysis Kit from BD according to the manufacturer’s instructions. Cells were incubated with human monoclonal antibodies specific to Nanog-PE, Oct3/4-PerCP-Cy5.5 and Alexa 647-Sox2 and corresponding isotype controls for 30 minutes and washed twice with PBS. Fifty thousand events were acquired in BD Accuri C6 flow cytometer using the kit template. Cells were gated on light scatter properties and analyzed for expression of key pluripotency transcription factors using BD Accuri C6 software.
Immunocytochemistry
Cells from passages 6 and 29 were fixed and immunostained following standard protocols23. Primary antibodies used were: monoclonal anti-mouse stage-specific embryonic antigen-1 (SSEA-1) and -4 (SSEA-4), TRA-1-60, TRA-1-81 (dilution 1:50; Chemicon), and anti-mouse OCT-4 (dilution 1:100; Santa Cruz Biotechnology, Inc.). Human foreskin fibroblasts (HS27 cell line, ATCC) were used as negative controls. Images were analyzed in an Axiophot 2 fluorescent microscope (Carl Zeiss) and captured by CCD camera using the ISIS software (MetaSystem).
In Vitro Differentiation
To demonstrate spontaneous in vitro differentiation, hiPSC and hESC were grown to confluency and harvested by dispase. Cells were washed and re-suspended twice in E8 medium and transferred to non-tissue culture treated 6 well plates coated with 1% agarose. Day 5 embryoid bodies (EBs) were transferred to a 6 well plate coated with 0.1% gelatin, and cultured for a further 15 days in differentiation medium [DMEM, 20% FBS (ESQ), 2 mM L-glutamax, MEM-NEAA 2 mM, 0.1 mM β-mercaptoethanol, 1% HEPES, 1% Sodium piruvate. 1% penicillin-streptomicin (Life Tech)]. Differentiated cells were harvested with Trizol and RNA extraction was performed usingRNeasy kit (Qiagen). One mg of RNA was used for subsequent reverse transcriptase reactions using High Capacity cDNA kit (Life Technologies). Pluripotency and trilineage differentiation potential was assessed using the TaqMan® hPSC Scorecard™ kit 384w following manufacturer’s instructions and run on a StepOne Plus System (Life Technologies), as described25. Data analysis was performed using the cloud based TaqMan® hPSC Scorecard™ analysis software (Life Technologies).
Vector integration analysis
Absence of vector DNA integration into hiPSC lines was confirmed by PCR analysis as described16. Briefly, primers for the Epstein–Barr nuclear antigen 1 (EBNA1) gene region of the pCEP4 vector were used to amplify 150 ng of hiPSC DNA and 3 pg of vector DNA (positive control) for 30 cycles.
Genomic ancestry
Analysis of genomic ancestry was conducted using the Admixture program26. Admixture is a software tool for maximum likelihood estimation of individual ancestries from multilocus SNP genotype datasets. Specifically, Admixture uses a block relaxation approach to alternately update allele frequency and ancestry fraction, with parameter standard errors using bootstrapping. We used a supervised approach for ancestry determination27, with 200 bootstrap replicates (default) and k = 3 (number of parental populations assumed for the analysis). The analysis was done using 192 Ancestry Informative Markers developed by our group28, genotyped by OpenArray® Real-Time PCR (Applied Biosystems). We used as reference ancestral populations those from the Human Genome Diversity Project (HGDP) (Center SHG. Human Genome Diversity Project): Pima, Maya as Native Americans; and from HapMap project (International HapMap Project): Africans - YRI (Yoruba in Ibadan, Nigeria), LWK (Luhya in Webuye, Kenya), ASW (Americans of African Ancestry in SW, USA); European - CEU (Utah Residents (CEPH) with Northern and Western European ancestry) and TSI (Tuscan in Italia).
Local ancestry analysis was performed by genotyping samples with the Axiom Human Origins Array (approximately 600,000 SNPs) according to manufacturer’s instructions (Affymetrix). As parental populations we used a sample of 30 unrelated African and 30 unrelated Europeans from 1000 Genomes Project (1000 g) and the 30 unrelated Native Americans from the Centre d’Étude du Polymorphisme Humain and the Human Genetic Diversity Panel (CEPH-HGDP). The SNPs from these three datasets (1000 g+HGDP and present study) were filtered and paired according to Nunes et al.29 Local ancestry analyses were performed using the RFMix software30 via a discriminative modeling approach based on a Random Forest algorithm. We used an admixture model that assumed 8 generations since the start of admixture and windows of 0.2 cM. The genotypes were phased using the SHAPEIT software31.
Additional Information
How to cite this article: Tofoli, F. A. et al. Increasing The Genetic Admixture of Available Lines of Human Pluripotent Stem Cells. Sci. Rep. 6, 34699; doi: 10.1038/srep34699 (2016).
References
Laustriat, D., Gide, J. & Pechanski, M. Human pluripotent stem cells in drug discovery and predictive toxicology. Biochem. Soc. Trans. 38, 1051–1057 (2010).
Weinshilboum, R. Inheritance and drug response. New Eng. J. of Med. 348, 529–537 (2003).
Inoue, H., Nagata, N., Kurokawa, H. et al. iPS cells: a game changer for future medicine. EMBO J. 5, 409–417 (2014).
Amps, K., Andrews, P. W., Anyfantis, G. et al. Screening ethnically diverse human embryonic stem cells identifies a chromosome 20 minimal amplicon conferring growth advantage. Nat Biotech. 12, 1132–1144 (2011).
Laurent, L. C., Nievergelt, C. M., Lynch, C. et al. Restricted ethnic diversity in human embryonic stem cell lines. Nat. Meth. 1, 6–7 (2010).
Chang, E. A., Tomov, M. L., Suhr, S. T. et al. Derivation of Ethnically Diverse Human Induced Pluripotent Stem Cell Lines. Sci. Rep. 5, 15234 (2015).
Manta, F. S., Pereira, R., Caiafa, A. et al. Analysis of genetic ancestry in the admixed Brazilian population from Rio de Janeiro using 46 autosomal ancestry-informative indel markers. Ann. Hum. Biol. 1, 94–98 (2013).
Callegari-Jacques, S. M., Grattapaglia, D., Salzano, F. M. et al. Historical genetics: spatiotemporal analysis of the formation of the Brazilian population. Am. J. Hum. Biol. 6, 824–834 (2003).
Lins, T. C., Vieira, R. G., Abreu, B. S. et al. Genetic composition of Brazilian population samples based on a set of twenty-eight ancestry informative SNPs. Am. J. Hum. Biol. 2, 187–192 (2010).
Pena, S. D., Di Pietro, G., Fuchshuber-Moraes, M. et al.The genomic ancestry of individuals from different geographical regions of Brazil is more uniform than expected. PLoS One 2, e17063 (2011).
Giolo, S. R., Soler, J. M., Greenway, S. C. et al. Brazilian urban population genetic structure reveals a high degree of admixture. Eur. J. Hum. Genet. 1, 111–116 (2012).
Fraga, A. M., Sukoyan, M., Rajan, P. et al. Establishment of a Brazilian line of human embryonic stem cells in defined medium: implications for cell therapy in an ethnically diverse population. Cell Trans. 3, 431–440 (2011).
Georges de Oliveira, J. A., Vergani, N., Fonseca, S. A. et al. Aberrant Patterns of X Chromosome Inactivation in a New Line of Human Embryonic Stem Cells Established in Physiological Oxygen Concentrations. Stem Cell Rev. 4, 472–479 (2014).
Fonseca, S. A., Costas, R. M. & Pereira, L. V. Searching for naïve human pluripotent stem cells. World J. Stem Cells 3, 649–656 (2015).
Aquino, E. M., Barreto, S. M., Bensenor, I. M. et al. Brazilian Longitudinal Study of Adult Health (ELSA-Brasil): objectives and design. Am. J. Epid. 4, 315–324 (2012).
Schmidt, M. I., Duncan, B. B., Mill, J. G. et al. Cohort Profile: Longitudinal Study of Adult Health (ELSA-Brasil). Int J Epidemiol. 1, 68–75 (2015).
Ko, H. C. & Gelb, B. D. Concise review: drug discovery in the age of the induced pluripotent stem cell. Stem Cells Trans. Med. 3, 500–509 (2014).
Cherry, A. B. & Daley, G. Q. Reprogramming cellular identity for regenerative medicine. Cell 6, 1110–1122 (2012).
Mosher, J. T., Pemberton, T. J., Harter, K. et al. Lack of population diversity in commonly used human embryonic stem-cell lines. New Eng. J. Med. 362, 183–185 (2010).
Pena, S. D., Bastos-Rodrigues, L., Pimenta, J. R. et al. DNA tests probe the genomic ancestry of Brazilians. Braz. J. Med. Biol. Res. 42, 870–876 (2009).
Zanger, U. M. & Schwab, M. Cytochrome P450 enzymes in drug metabolism: regulation of gene expression, enzyme activities, and impact of genetic variation. Pharm. Ther. 138, 103–141 (2013).
Lotufo, P. A., Pereira, A. C., Vasconcellos, P. S. et al. Resistant hypertension: risk factors, subclinical atherosclerosis, and comorbidities among adults-the Brazilian Longitudinal Study of Adult Health (ELSA-Brasil). J. Clin. Hypertens. (Greenwich) 1, 74–80 (2015).
Chou, B. K., Mali, P., Huang, X. et al. Efficient human iPS cell derivation by a non-integrating plasmid from blood cells with unique epigenetic and gene expression signatures. Cell Res. 21, 518–529 (2011).
Dowey, S. N., Huang, X., Chou, B. K. et al. Generation of integration-free human induced pluripotent stem cells from postnatal blood mononuclear cells by plasmid vector transfection. Nat. Prot. 7, 2012–2021 (2012).
Tsankov, A. M., Akopian, V., Pop, R. et al. A qPCRScoreCard quantifies the differentiation potential of human pluripotent stem cells. Nature Biotech. 33, 1182–1192 (2015).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Gen. Res. 19, 1655–1664 (2009).
Cardena, M. M., Ribeiro-dos-Santos, A., Santos, S. et al. Amerindian genetic ancestry is associated with higher survival rates compared to African and European ancestry in Brazilian patients with heart failure. Int. J. Cardiol. 176, 527–528 (2014).
Santos, H. C., Horimoto, A. V. R., Tarazona-Santos, E. et al. A minimum set of ancestry informative markers for determining admixture proportions in a mixed American population: the Brazilian set. Eur. J. of Hum. Gen. 24, 725–731 (2015).
Nunes, K., Zheng, X., Torres, M. et al. HLA imputation in an admixed population: An assessment of the 1000 Genomes data as a training set. Hum Immunol. 77, 307–312 (2015).
Maples, B. K., Gravel, S., Kenny, E. E. et al. RFMix: a discriminative modeling approach for rapid and robust local-ancestry inference. Am. J. Hum. Gen. 2, 278–288 (2013).
Delaneau, O. & Marchini, J. 1000 Genomes Project Consortium et al. Integrating sequence and array data to create an improved 1000 Genomes Project haplotype reference panel. Nat Commun. 5, 1–5 (2014).
Acknowledgements
This work was supported by grants from Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP - CEPID 13/08135-2) Conselho Nacional de Desenvolvimento Científico e Tecnológico/Departamento de Ciência e Tecnologia do Ministério da Saúde (CNPq/MS/DECIT), Banco Nacional de Desenvolvimento Econômico e Social (BNDES) and Financiadora de Estudos e Projetos (FINEP).
Author information
Authors and Affiliations
Contributions
F.A.T., collection and assembly of data, final approval of manuscript; M.D., conception and design, collection and assembly of data, data analysis and interpretation, manuscript writing, final approval of manuscript; M.M.-M., collection and assembly of data, manuscript writing, final approval of manuscript; K.N., collection and assembly of data, data analysis and interpretation, manuscript writing, final approval of manuscript; L.A.P., collection and assembly of data, final approval of manuscript; G.S.d.S., collection and assembly of data, final approval of manuscript; S.F., collection and assembly of data, final approval of manuscript; R.M., collection and assembly of data, final approval of manuscript; H.C.S., collection and assembly of data, data analysis and interpretation, final approval of manuscript; A.d.C.P., collection and assembly of data, data analysis and interpretation, final approval of manuscript; P.A.L., collection and assembly of data, data analysis and interpretation, final approval of manuscript; I.M.B., collection and assembly of data, final approval of manuscript; D.M., collection and assembly of data, data analysis and interpretation, manuscript writing, final approval of manuscript; L.V.P., collection and assembly of data, financial support, data analysis and interpretation, manuscript writing, final approval of manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Tofoli, F., Dasso, M., Morato-Marques, M. et al. Increasing The Genetic Admixture of Available Lines of Human Pluripotent Stem Cells. Sci Rep 6, 34699 (2016). https://doi.org/10.1038/srep34699
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep34699
This article is cited by
-
Defective hematopoietic differentiation of immune aplastic anemia patient-derived iPSCs
Cell Death & Disease (2022)
-
Greater genetic diversity is needed in human pluripotent stem cell models
Nature Communications (2022)
-
Automated image analysis system for studying cardiotoxicity in human pluripotent stem cell-Derived cardiomyocytes
BMC Bioinformatics (2020)
-
PSCRIdb: A database of regulatory interactions and networks of pluripotent stem cell lines
Journal of Biosciences (2020)
-
Origin and age of the causative mutations in KLC2, IMPA1, MED25 and WNT7A unravelled through Brazilian admixed populations
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.