Abstract
Brettanomyces bruxellensis has been described as the main contaminant yeast in wine production, due to its ability to convert the hydroxycinnamic acids naturally present in the grape phenolic derivatives, into volatile phenols. Currently, there are no studies in B. bruxellensis which explains the resistance mechanisms to hydroxycinnamic acids, and in particular to p-coumaric acid which is directly involved in alterations to wine. In this work, we performed a transcriptome analysis of B. bruxellensis LAMAP248rown in the presence and absence of p-coumaric acid during lag phase. Because of reported genetic variability among B. bruxellensis strains, to complement de novo assembly of the transcripts, we used the high-quality genome of B. bruxellensis AWRI1499, as well as the draft genomes of strains CBS2499 and0 g LAMAP2480. The results from the transcriptome analysis allowed us to propose a model in which the entrance of p-coumaric acid to the cell generates a generalized stress condition, in which the expression of proton pump and efflux of toxic compounds are induced. In addition, these mechanisms could be involved in the outflux of nitrogen compounds, such as amino acids, decreasing the overall concentration and triggering the expression of nitrogen metabolism genes.
Similar content being viewed by others
Introduction
Brettanomyces bruxellensis is one of the main contaminant yeasts in wines, with the ability to metabolize hydroxycinnamic acids, that are naturally present in grapes, into volatile phenols1,2,3,4. Hydroxycinnamic acids are weak acids with lipophilic character, such as p-coumaric acid, ferulic acid, and caffeic acid, all of which are able to adversely affect the organoleptic qualities of the wine giving undesirable odors2. Inhibitory effects of ferulic acid has been described on Pichia anomala5 and Debaromyces hansenni6, where the inhibitory effect is related to the high polarity of the acids. In wine, a large number of hydroxycinnamic acids have a synergistic effect which could enhance the inhibitory power. However, Brettanomyces species can overcome this toxicity by converting these acids into volatile phenols. The formation of these compounds in B. bruxellensis is a result of the enzymatic processing of hydroxycinnamic acids by the action of two specific enzymes, cinnamate decarboxylase (CD) and vinylphenol reductase (VR)1,7,8.
The presence of hydroxycinnamic acids in the culture medium can inhibit the growth of a wide range of microorganisms. However, this inhibition depends on the acid concentration5,9,10,11,12 and the type of molecule, with p-coumaric and ferulic acids showing the greater degree of inhibition5,6,9,13,14. The concentrations that can inhibit B. bruxellensis growth have been previously studied6,15, as well as the mechanisms involved16,17,18. These are based primarily on the chemical nature of the side chain carboxylic group (R-COOH) and its antimicrobial activity is based on the effects of the form of undissociated acid, which depends on the pH of the medium and the pKa of the weak acid. The undissociated acid is able to go through the cell membrane by simple diffusion and activates several ATP demanding mechanisms that jeopardise the cell survivance19,20,21. In Saccharomyces cerevisiae, adaptation to weak acids has been has been related to the expression of different genes, mainly PMA1 and PDR12, together with VMA1, PDR5, MSN2, HSP26 and MSN422,23,24. The response to weak acids has been extensively characterized in S. cerevisiae, Candida albicans and Zygosaccharomyces bailii19,25,26,27. In contrast, for B. bruxellensis no studies have evaluated the genes that are associated with the resistance to weak acids, and how they are taking part in the metabolism of hydroxycinnamic acids.
Previous studies of our group have shown that B. bruxellensis LAMAP2480 has significant differences in its growth curve, as well as in the production levels of volatile phenols, when grown in the presence of 100 mg/L of p-coumaric. Growth experiments in synthetic and natural wine28, showed that LAMAP2480 has a shorter duration of the lag phase and increased production of 4-ethylphenol compared to another strain (LAMAP1359)28, suggesting that LAMAP2480 has a rapid adaptation response to medium with p-coumaric acid, with a mechanism that allows for efficient resistance to this compound. Similar observations have been made for B. bruxellensis strains grown in the presence of different hydroxycinnamic acids13. We have also shown that the presence of p-coumaric acid at a concentration of 100 mg/L has a positive effect on the growth rate for some isolates of Brettanomyces spp28. The exposed antecedents suggests that the effect of this acid on the growth of this organism would be strain-dependent. This evidence suggests that the mechanism of weak acid resistance by B. bruxellensis, particularly to p-coumaric acid, is strain-dependent phenomenon that occurs during the lag phase of the growth curve.
B. bruxellensis has been poorly studied at the genetic level, and due to its genetic variability it has been a problem to develop models that describe its behaviour29,30,31,32,33,34,35. In some B. bruxellensis strains, the number of chromosomes can vary between 4 to 936, with chromosome sizes in the range of 1 to 6 Mb, and a total genome size between 20 to 30 Mb33. B. bruxellensis karyotypic variation suggests speciation due to genome rearrangements. To this date, six B. bruxellensis strains from different sources (beer and wine) and geographical areas have been sequenced: AWRI149934, CBS249935, AWRI1608 and AWRI161337, ST05.12/2231 and LAMAP248031. Comparative genomic analysis of B. bruxellensis strains, showed that while CBS2499 is diploid, AWRI1499 and AWRI1608 are triploid, with two closely related alleles and a more divergent third one37. These findings suggested that some B. bruxellensis strains originate from three haplotypes. Preliminary evidence suggests that LAMAP2480 is triploid, and with the current draft state of its genome, this could introduce biases if only this genome is used as a reference for transcriptomic analysis. To complement this, and evaluate the effects of weak acids during lag phase in B. bruxellensis, we used two complementary approaches to study transcription profiles: de novo and genome guided transcriptome assembly.
Results and Discussion
Physiological evaluation of LAMAP2480 strain growth in p-coumaric acid
The presence of 100 mg/L p-coumaric acid in the culture medium caused an increase in the duration of the lag phase in LAMAP2480 (Fig. 1) from 14 hours (control) to 28 hours. Furthermore, we observed a reduction in the specific growth rate (μ) (Table 1) in presence of p-coumaric acid, which agrees with previous results16,38.
Measurements of extracellular pH during the lag phase showed a strong decrease in extracellular pH when LAMAP2480 was grown in the presence of p-coumaric acid, compared to the control condition. However, it has been reported that production of organic acids, such acetic acid, during yeast growth may also contribute to the process of extracellular acidification. Nevertheless, we previously demonstrated using High Performance Liquid Chromatography (HPLC) that acetic acid production starts during the exponential phase39,40. This suggests that the decrease of the culture pH was due to the action of the proton pump ATP-ase Pma141.
Pma1p activity was determined using protein extracts from cultures grown in Synthetic Dextose medium supplemented with p-coumaric acid during lag phase, and a higher activity was found in media containing p-coumaric acid (Fig. 2). This is similar to what has been previously reported for B. bruxellensis LAMAP1359 when grown in the presence of p-coumaric acid14. In the presence of weak acids, the role of Pma1p is vital to restore homeostasis20,42,43, which exerts a high energy demand, consuming until 60% of the total cellular ATP20, causing ATP levels to reduce to a point where growth rate declines20,44,45. Thus, the increase in the specific activity of Pma1p during lag phase in the medium supplemented with p-coumaric acid suggests that this proton pump is an important component of an adaptive response to hydroxycinnamic acids.
Transcriptome analysis of B. bruxellensis LAMAP2480
Triplicates of two pools of mRNA samples were used to build libraries for RNA sequencing, generating approximately 197.7 millions of high quality paired-end reads, with an average length of 90 bp and encompassing 17,792 million nucleotides. The clean reads are available at the Sequence Read Archive (SRA) under accession number SRP077865.
Due to genomic differences between B. bruxellensis strains31,32,33,34,35,36,46, we performed transcriptional analysis using two approaches: de novo and reference-guided. To obtain a de novo transcriptome in which we could compare the abundance of transcripts against the genome-guided approach, all high quality reads were assembled into contigs using Trinity47. The assembly resulted in 26,761 transcripts with an average length of 1,570 base pairs, which were used to predict open reading frames (ORFs) and generated 23,650 non-redundant coding sequences, denominated Unigenes (Table 2).
In order to obtain the most complete annotated set of transcripts, the total of Unigenes were compared against the NCBI non-redundant database (NR), using Blastx. Approximately 93.3% (22,065 unigenes) were annotated with high homology, with an e-value ≤10−5 (Fig. 3A). Out of this total, a 70% of the genes were annotated with a sequence similarity greater than 80% (Fig. 3B). These results provide transcript information for genes expressed during lag phase of LAMAP2480 in SD media with presence of p-coumaric acid that were used for transcriptional downstream analysis.
As part of the annotation process, we evaluated the taxonomic distribution of the Blastx results for the B. bruxellensis de novo transcripts. As expected, the most represented yeast species was B. bruxellensis AWRI1499 (Fig. 3C) with 13,834 assignations, followed by Ogataea parapolymorpha, Pichia kudriavzevii and others. Although B. bruxellensis AWRI1499 is considered to be the best annotated strain, the number of contigs exhibiting homology with AWRI1499 only represents 62.7% of annotations (51.9% of the total of unigenes). These results strongly suggest that de novo assembly, complemented with de novo annotation of transcripts, can capture LAMAP2480 genes that are not well-annotated in the reference strain, as it was observed previously in Schizosaccharomyces pombe47.
The total number of de novo hits (22,065 unigenes) is an improvement compared to the existing gene annotations reported for AWRI1499 (4,861 genes), CBS2499 (5,650 genes) and LAMAP2480 (9,008 genes) (Fig. 3D), because we cover a higher number of transcripts that were not included in previous annotations. However, a single unigene represents the collection of expressed sequences that match a common transcript in a determined locus. These results must be interpreted carefully, because if two contigs match the same gene, it will produce an overestimation of annotations. When the data is filtered by removing multiple unigenes matching common genes, only 6,314 are unique annotations. This shows that de novo and genome-guided are complementary approaches that can be used to overcome the lack of annotation information for the B. bruxellensis genomes.
None of B. bruxellensis genome annotations had Gene Ontology (GO) terms assigned, and GO terms assigned to S. cerevisiae did not covered the complete universe of annotated genes. To overcome this, we assigned GO annotation to all B. bruxellensis genomes and unigenes. Based on sequence homology of B. bruxellesis annotations, we annotated GO term to 3,432 AWRI1499 sequences, 3,779 CBS2499 sequences and 8,723 LAMAP2480 sequences. In the case of LAMAP2480 de novo assembled transcripts, a total of 16,054 sequences (72.76%) were annotated with GO terms. Sequence annotations against the GO database were performed using Blast2GO basic tool, and the data generated was stored in genome reference sessions known by the software as “work environments”, which file format has b2g extension. They can be used for enrichment analysis, KEGG pathways and are available for download (https://figshare.com/s/103665d0b83485b7e193). The distribution of functional categories for the three annotated genomes and the de novo transcriptome, was visualized looking at the higher hierarchical GO terms (Fig. 4). As expected, de novo assembly of LAMAP2480 transcripts generates a functional profile that is more similar to the genome annotation of this organism than to the other two strains (ARWI1499 and CBS2499). However, in the de novo transcripts we did not found categories such as biological process cell killing (GO:0001906) and rhythmic process (GO:0048511), and the molecular function auxiliary transport protein activity (GO:0015475), which were present in the genome annotation.
Expression profiling during p-coumaric stress
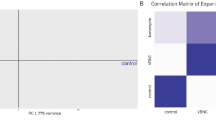
To compare de novo assembly results with reference genomes, reads were aligned against the available genomes of B. bruxellensis strains using TopHat248. As expected, the strain with a higher coverage was LAMAP2480 (91.78%), compared to strains AWRI1499 (62.22%) and CBS2499 (75.02%). Changes in gene expression produced by induction of p-coumaric stress were evaluated by normalizing B. bruxellensis gene expression levels using the RPKM method (Reads per kilobase transcriptome per million mapped reads). Principal component analysis (PCA) was used to plot the data of the control condition against the p-coumaric acid condition (Supplementary Fig. S1). The dispersion of the replicates indicates a high reproducibility per condition. Nevertheless, we observed that mapping against the CBS2499 genome did not show the same behavior on the replicates. Due to the low dispersion observed on the analysis with the other strains (AWRI1499 and LAMAP2480), we rule out that this distortion is due to technical problems. A possible source of variation for CBS2499 could be due to allelic variation, as CBS2499 has a diploid genome, while AWRI1499 is triploid, with a high degree of sequence divergence in one of the three alleles37. This evidence, combined with the low annotation frequency of de novo transcripts for CBS2499, may explain the distortion observed in the PCA plots, as part of the reads are not mapped to the genome.
Differentially expressed genes between the control and p-coumaric acid treatments were clustered based on the log2 fold-change and plotted against the p-value to observe overall gene expression patterns (Supplementary Fig. S2). Nevertheless, considering high genetic variability that exists between the genomes of B. bruxellensis strains, first we compared the expression of selected genes for validation related to the response to p-coumaric acid with each genome using log2 fold change expressed in RPKM (Fig. 5). The results, as expected, indicate that the expression patterns are different with each genome, however, pattern gene expression on de novo analysis and the strain LAMAP2480 are similar, and also in annotation and expression levels. Considering this, we used RNA-seq data generated at the draft genome LAMAP2480 as reference for validation. The results of gene expression in the figure revealed high expression differences between strains, which lead to validation of genes related to p-coumaric response.
Validation of specific gene expression by RT-qPCR
B. bruxellensis transcriptional response to stressing agents has been poorly studied49,50. In this study we evaluate B. bruxellensis LAMAP2480 response to p-coumaric acid stress during lag phase of growth curve. In this stage we observed differences of kinetic behaviour and we also found that in this stage p-coumaric acid is decarboxylated to produce 4-vinylphenol. Genes representative of functional categories or pathways that show a differential regulation by p-coumaric acid were selected for validation (Supplementary Table S1). Then, to validate the p-coumaric acid-induced transcriptional changes, qPCR was performed on three biological replicates per growth condition and relative expression was calculated (Fig. 6).
Data (means ± SD) were calculated by the 2(-Delta Delta C(T)) method104 on three independent experimental replicates.
PAD1 gene is overexpressed in the presence of p-coumaric acid. This gene codifies for a phenylacrylic acid decarboxylase51,52, and confers resistance to hydroxycinnamic acid by its decarboxylation to vinyl derivatives51,53,54. This gene was previously identified in LAMAP2480, and it was demonstrated that it plays a role in the decarboxylation of p-coumaric acid to 4-vinylphenol52. HPLC measurements shows that p-coumaric acid is decarboxylated at the start of the lag phase in the growth curve (Supplementary Fig. S3), as an early response mechanism to stress. PAD1 is also present in CBS2499, but it has not been described for AWRI1499. To verify the presence of PAD1 in AWRI1499, we aligned the PAD1 known nucleotide sequence52 against its reference genome and we had a match in a non-annotated region (AHIQ01000324:33,529-34,042). This result allow us to complement the current reference annotation on AWRI1499 and may support future studies of decarboxylation in this strain.
p-coumaric acid induces the activation of transporters involved in efflux of toxic compounds and drug resistance (Supplementary Table S2). These transporters have been associated with the efflux of anions produced by the dissociation of weak acids24. Although it has been demonstrated that p-coumaric acid can be metabolized to less toxic compounds, such as 4-ethylphenol, the presence of transporters for p-coumaric acid or other derivatives has not been proved1,24. One possibility is that these transporters are responsible for the efflux of cumarate and/or its derivates. To evaluate this, the genes related to detoxification of toxic compounds and transports of drugs were studied. We observed overexpression of the genes PDR13, PDR15, YLL056C, AMF1, YOR1, and PDR10. These genes have been previously described to respond to the presence of propionic acid55, sorbic acid23,55, 2,4-D56,57 and Neocarzinostatin58. The resistance mechanism to this type of molecules involves ATP powered pumps, whose action can contribute to decrease the ATP levels in the cell. In the case of the gene YLL056C, it encodes for a putative protein of unknown function, and its transcription has been previously associated to response to the transcription factors Yrm1p and Yrr1p, as well as genes involved in pleiotropic drug resistance (PDR)59. In addition, it has been observed that YLL056C expression is induced in cells treated with different drugs60, such as the mycotoxin patulin61.
The treatment with p-coumaric acid also induced the expression of genes related to sulfate (SUL1) and iron (FTR1, SEF1) uptake. SUL1 encodes for a sulphate transporter, and its overexpression has been previously reported as a resistance mechanism to neomycin in S. cerevisiae62. FTR1 encodes for an iron permease and it is up-regulated by weak acids, being associated with an overall decrease in intracellular iron concentrations in C. albicans60. SEF1 encodes for a putative transcription factor and has been related to be involved in controlling the expression of iron acquisition genes in C. albicans59. The overexpression of these genes suggest that the availability of cations is relevant to the resistance mechanism to weak acids, which is supported by previous evidence that the uptake of cations such as potassium, calcium and zinc are regulated in presence of propionic, acetic and lactic acids55,63,64.
In our experiment, genes that encode for monocarboxylate permeases (MCH2 and JEN1) were overexpressed. These transporters are involved in the movement of monocarboxylic acids such as lactate, pyruvate, and acetate across the plasma membrane65,66. For MCH2, it has been previously reported that it is overexpressed in response to vanillin, a major phenolic compound generated due to lignin breakdown67, which suggest that it is involved in the transport of this compound65. It has also been observed that JEN1 gene is induced by lactic and pyruvic acids68, which indicates that p-coumaric response is the induction of genes related to weak acids.
We also found that a gene-complex that is related to oxidative stress was regulated by the action p-coumaric acid. This complex is encoded by the genes PST2 (flavodoxin-like protein), PRX1 (mitochondrial peroxiredoxin) and SOD1 (cytosolic copper-zinc superoxide dismutase). PST2 is induced by oxidative stress in a Yap1p dependent manner and PRX1 is induced during respiratory growth and oxidative stress69. It has been observed that PST2 and PRX1 genes are induced by oxidative stress in S. cerevisiae67,70 and C. albicans66. Correspondingly, it has been reported that SOD1 is overexpressed in response to sorbic acid in S. cerevisiae23,71, suggesting that it plays a role in the resistance mechanism to ascorbic acid. Weak acids, such as sorbic, octanoic and decanoic acids has also been associated with the induction of multiple genes that take part in the response to oxidative stress. Because p-coumaric acid is also a weak acid, we can suggest that it induces a similar response and that the induction of PST2 and PRX1 are part of the resistance mechanisms of B. bruxellensis.
Under stress conditions mentioned above, the permeability of the cell wall is modified in order to reduce the rate of entry of weak acids to the cell72. Accordingly, we found genes involved in the synthesis of cell wall components that are DE in response to p-coumaric acid (Supplementary Table S2), such as chitin synthesis and cell wall assembly. We evaluated the expression of WSC4 and HSP150 by RT-qPCR. WSC4 is involved in the translocation of soluble secretory proteins and the insertion of membrane proteins into the endoplasmatic reticulum membrane, while HSP150 encodes for a O-mannosylated heat shock protein, that has been reported to be induced by heat shock, oxidative stress, and nitrogen limitation73. Our results indicate that WSC4 is repressed in the presence of p-coumaric acid. While it has been reported that this gene is up-regulated after exposure to acetic acid by two-fold in S. cerevisiae74, other studies indicate that is repressed in response to the accumulation of misfolded proteins in the endoplasmic reticulum75. In contrast, HSP150, also annotated as PIR2, is overexpressed in the presence of p-coumaric acid. Previous reports have shown that this gene is overexpressed under stress in response to lactic and acetic acids76. Overall, the action of proteins encoded by both identified genes help restrict access of the p-coumaric molecules to the cell membrane.
In our analysis, a large number of genes associated with the use of nitrogen sources are highly represented (Supplementary Table S2). A first group is constituted of genes encoding for allantoin transporters and amino acid transporters, while a second group are genes that codify for enzymes involved in the metabolism of these compounds. VBA2, PUT2, and DAL4 are overexpressed in the presence of p-coumaric acid, and were validated by qPCR. VBA2 encodes for a protein involved in vacuolar uptake of basic amino acids77,78. DAL4 encodes for a protein that takes part in the in the uptake of allantoin, a nitrogen-rich molecule generated from purine catabolism, which can be used as a nitrogen source by yeast cells79. PUT2 encodes for a delta-1-pyrroline-5-carboxylate dehydrogenase80. p-coumaric acid also induced MCH5, which encodes for riboflavin transporter81, which has been reported to be highly expressed on proline medium82 and overexpressed during the lag phase of cells treated with 5-hydroxymethylfurfural (HMF)83. While the relationship between the toxicity of p-coumaric acid and an increase in the expression of these genes does not appear clear, it has been proposed in S. cerevisiae that an efflux pump that confers resistance to weak acids, can also carry amino acids outside the cell84. Furthermore, it has been previously reported that under stress conditions S. cerevisiae and Arabidopsis thaliana accumulate proline80, which is defense mechanism to create a nitrogen storage required for protein biosynthesis that in consequence acts as a protective agent for cells under osmotic stress83. Also, the upregulation of genes related to uptake and biosynthesis of amino acids has been observed in cells cultivated in the presence of acetic acid85,86, propionic acid55 and 2,4-D57. Supporting this, we found that the the gene SSU1, which encodes for a sulfite/nitrite membrane pump, was overexpressed in the presence of p-coumaric acid, and validated by qPCR. This pump exports excess sulfite produced during sulfate assimilation and amino acid biosynthesis87,88, suggesting that in the presence of p-coumaric acid, MFS transporters related to cellular detoxification also allow the output of important sources of nitrogen, leading to the depletion of intracellular free nitrogen, and inducing an increase in the expression of genes involved with the uptake and biosynthesis of amino acid, helping to maintain the balance the amino acid pool within cells. In addition, it has been reported that basic amino acid have a buffering capacity, which improves yeast growth by maintaining the pH close to neutral89. These amino acids can be stored in vacuoles using a cation-dependent transport mechanism, which could be related to the overexpression of cation permeases90.
Two heat shock proteins, HSP12 and HSP26, were induced in the presence of p-coumaric acid. It has been previously reported that HSP12 is overexpressed in yeast cells exposed to heat shock, as well as to osmotic and oxidative stress conditions89,90,91,92. In the case of HSP26, it has been associated with the ability to counteract HMF stress damage to proteins83. In yeast cells treated with sorbic acid, HSP12 and HSP26 were overexpressed23 conferring resistance to weak acid stress, which suggests this acid could be due to intracellular protein denaturation.
Conclusion
We analyzed transcriptome data for B. bruxellensis LAMAP2480, for which prior genomic information was limited. The sequence information generated in this study allows to improve our understanding of the mechanisms that B. bruxellensis uses to respond to weak acid stress. In addition, this information restates the genetic differences between B. bruxellensis strains. Interpretations and accuracy of RNA-seq analysis results are highly dependent on the quality of a well-annotated reference genome. In this study we compared two draft genomes (LAMAP2480 and CBS2499), a complete reference genome (AWRI1499) and de novo assembly data. The results shows that when there is evidence of genetic variability among strains, de novo analysis complemented with a draft genome provides more accurate results than using a reference genome from a closely related strain.
The results presented allow us to propose a model of early response to stress by p-coumaric acid. Its inflow into the cell causes a generalized stress condition, in which the expression of proton pumps and mechanisms involved in the efflux of toxic compounds are induced (Fig. 7). The latter might be involved in the outflow of nitrogen sources, such as as amino acids or allantoin, decreasing the intracellular concentration of nitrogen and triggering the expression of genes related to nitrogen metabolism. p-coumaric acid also induces oxidative stress, activating the expression of genes as SOD1, PST2 and PRX1. Finally, the presence of this acid seems to cause protein denaturation, which is observed in the expression of chaperone proteins. At last this acid triggers a change of the permeability of the cell wall in order to reduce the rate of entry of weak acid into the cell.
Materials and Methods
Strain and culture media
Brettanomyces bruxellensis LAMAP2480 was originally isolated from Cabernet Sauvignon wine, and is part of the collection at the Laboratorio de Biotecnología y Microbiología Aplicada (LAMAP), Universidad de Santiago de Chile.
P-coumaric effect on B. bruxellensis LAMAP2480 growth was evaluated according to previous protocols developed in our laboratory with slight modifications14. The kinetic parameters evaluated were: specific growth rate (μ)93, Lag phase duration94, and efficiency was defined as area under curve (AUC) and expressed as a percentage using as 100% the control condition. Finally, growth inhibition (GI) was calculated as the percentage of the maximal specific growth rate of non-stressed control cultures95.
Evaluation of physiological parameters
In order to describe the response of B. bruxellensis LAMAP2480 to p-coumaric stress, we evaluated seven physiological parameters: extracelullar pH, p-coumaric acid, 4-vinylphenol, 4-ethylphenol, acetic acid and glucose levels, and Pma1 enzyme activity. Extracellular pH was evaluated using previously described methods22, with adaptations to our local conditions and culture media14. p-coumaric acid, 4-vinylphenol, 4-ethylphenol, acetic acid and glucose were quantified by high performance liquid chromatography (HPLC) (Shimadzu Scientific Instruments, Colombia, MD, USA) using a Shimazdu Shim-Pack VP-ODS column96 for p-coumaric acid, 4-vinylphenol and 4-ethylphenol, and a Bio-Rad HPX87H column97 for acetic acid and glucose. Finally, plasma membrane ATPase Pma1p activity was estimated by calculating the rate of phosphate production after ATP hydrolysis98, as previously described14.
RNA isolation, library preparation and Illumina sequencing
Cells were harvested after consumption of 10% of the glucose present in the medium, centrifuged at 2,850 g for 10 min and then resuspended in 200 μL RNA buffer (50 mM Tris-HCl pH 7.4; 100 mM NaCl; 10 mM EDTA) and 400 μL acidic phenol (pH 4.3). Cells were broken using acid-washed glass beads for 3 min with incubation on ice every 1 min. RNA buffer (200 μL) and 10% SDS (40 μL) were added and the mix was incubated for 6 min on 65 °C followed by centrifugation at 16,060 g for 15 min. Acidic phenol (400 μL) and 3 M NaOAc (40 μL) were added to the water phase99. After centrifugation at 16,060 g for 15 min, 1 mL of 96% ethanol was added and the mix was incubated 2 h. at −80 °C. The precipitated RNA was spun down at 4 °C for 10 min and was purified using the RNA Clean & Concentrator Column (Zymo Research). Total RNA was treated with DNase I (Promega, Madison, WI, USA). RNA quality was analyzed by CE-LIF (AATI Fragment Analyzer, Advanced Analytical Technologies Inc) to evaluate the RQN (RNA Quality Number.) All RNA samples had RNA Quality Numbers (RQNs) greater than 7.5.
RNA-seq library preparation and sequencing was carried out by the Beijing Genomics Institute (BGI) (Hong-Kong, China). Libraries were sequenced using Illumina HiSeqTM 2000 (Illumina Inc, San Diego, CA, USA) in paired-end mode with a read length of 100 bp. The raw reads were cleaned by removing adaptor sequences and low quality sequences (q < 30).
De novo and genome-guided RNA-seq analysis
With the aim of comparing gene expression of B. bruxellensis LAMAP2480 among strains, and evaluate differences between existing reference genomes, two different strategies were carried out; 1) De novo assembly and posterior annotation with Blast, 2) genome-guided mapping, aligning the raw reads to three B. bruxellensis strains: AWRI149934, CBS249933, and LAMAP248031.
In undertaking the first analysis, reads were assembled into transcripts using Trinity, as previously described for de novo transcriptome assembly without a reference genome47. Transdecoder, an external Trinity plugin, was used to identify predicted open reading frames (ORFs) within isoforms or contigs and clustered into non-redundant sequences with a minimum length of 200 base pairs, which are known as unigenes.
Annotation was performed using NCBI BLAST 2.2.28+ with a minimum e-value cutoff of 1e-5 for the NCBI non-redundant protein (NR) database. Blast2GO100 software was used for gene ontology (GO) annotation in genomes from both approaches. Posteriorly, WEGO101 was used to perform GO functional classification at the second GO hierarchical level.
For gene expression analysis, reads mapped were normalized to RPKM (Reads per Kilobase per Million mapped reads) and edgeR102 was used for differential of expression. Genes differentially expressed were cut-off at significant level of 0.001 (p-values were adjusted with Benjamini & Hochberg method).
For the second approach, three B. bruxellensis strains genome and gene information were downloaded from NCBI. Alignment of reads to AWRI149934, CBS 249933 and LAMAP248031 was performed using TopHat248. To improve accuracy of differential expression analysis, samples were merged before analysis with Cufflinks, improving with this the insufficient depth of coverage for genes with low expression values, as described in Tuxedo protocol103. For the differential expression analysis a gene was considered to have significant changes with FDR < 0.05.
Validation of differentially expressed genes by real-time PCR
Quantitative RT-PCR analysis was used to validate the expression of the candidate genes. RT-qPCR was conducted on Applied Biosystems StepOnePlus Real-Time PCR System using StepOne Software (v2.0) (Applied Biosystems, Foster City, CA, USA) according to the manufacturer’s instructions. All qRT PCR primers were designed using qRT primer design tools available online (idtdna.com), and designed to amplify fragments between 150 and 250 base pairs. All qRT-PCR reactions were run in 20 ul reactions using using 5x HOT FIREPol® EvaGreen® qPCR Mix Plus (ROX) (Solis BioDyne, Tartu, Estonia) according to the SYBR Green method. Each reaction contained 4 μL 5x EvaGreen® qPCR Mix Plus (ROX), 1 μL primer mix (250 nM each), 1 μL cDNA (1 ng), and 13 μL DNase/RNase free water. Amplification was carried out with the following cycling parameters: heating for 15 min at 95 °C, 40 cycles of denaturation at 95 °C for 15 s, annealing at 58 °C for 20 s and extension at 72 °C for 20 s. Each sample was analyzed in triplicates and the expression values were normalized against β−actin. The molecular weight of the products was confirmed via diagnostic agarose gel and the melting curves were analyzed. Analysis of the relative gene expression data was conducted using the 2−ΔΔCT method104.
Additional Information
How to cite this article: Godoy, L. et al. Comparative transcriptome assembly and genome-guided profiling for Brettanomyces bruxellensis LAMAP2480 during p-coumaric acid stress. Sci. Rep. 6, 34304; doi: 10.1038/srep34304 (2016).
References
Chatonnet, P., Pascal, C., Denis, D., Jean-noël, B. & Monique, P. The origin of ethylphenols in wines. J. Sci. Food Agric. 60, 165–178 (1992).
Suárez, R., Suárez-Lepe, J. A., Morata, A. & Calderón, F. The production of ethylphenols in wine by yeasts of the genera Brettanomyces and Dekkera: A review. Food Chem. 102, 10–21 (2007).
Fleet, G. H. Yeast interactions and wine flavour. Int. J. Food Microbiol. 86, 11–22 (2003).
Wedral, D., Danielle, W., Robert, S. & Joseph, F. The challenge of Brettanomyces in wine. LWT - Food Science and Technology 43, 1474–1479 (2010).
Stead, D. The effect of hydroxycinnamic acids and potassium sorbate on the growth of 11 strains of spoilage yeasts. J. Appl. Bacteriol. 78, 82–87 (1995).
Baranowski, J. D., Davidson, P. M., Nagel, C. W. & Branen, A. L. Inhibition of Saccharomyces cerevisiae by naturally occurring hydroxycinnamates. J. Food Sci. 45, 592–594 (1980).
Edlin, D. A. N., Arjan, N., Gasson, M. J. J., Richard, D. & David, L. Purification and characterization of hydroxycinnamate decarboxylase from Brettanomyces anomalus. Enzyme Microb. Technol. 22, 232–239 (1998).
Godoy, L., Martínez, C., Carrasco, N. & Ganga, M. A. Purification and characterization of a p-coumarate decarboxylase and a vinylphenol reductase from Brettanomyces bruxellensis. Int. J. Food Microbiol. 127, 6–11 (2008).
Edlin, D. A. N., Arjan, N., Richard Dickinson, J. & David, L. The biotransformation of simple phenolic compounds by Brettanomyces anomalus. FEMS Microbiol. Lett. 125, 311–315 (1995).
Herald, P. J. & Davidson, P. M. Antibacterial Activity of Selected Hydroxycinnamic Acids. J. Food Sci. 48, 1378–1379 (1983).
Stead, D. The effect of hydroxycinnamic acids on the growth of wine-spoilage lactic acid bacteria. J. Appl. Bacteriol. 75, 135–141 (1993).
Campos, F. M., Couto, J. A. & Hogg, T. A. Influence of phenolic acids on growth and inactivation of Oenococcus oeni and Lactobacillus hilgardii. J. Appl. Microbiol. 94, 167–174 (2003).
Harris, V., Ford, C. M., Jiranek, V. & Grbin, P. R. Dekkera and Brettanomyces growth and utilisation of hydroxycinnamic acids in synthetic media. Appl. Microbiol. Biotechnol. 78, 997–1006 (2008).
Godoy, L., Varela, J., Martínez, C. & Ganga, M. A. The effect of hydroxycinnamic acids on growth and H- ATPase activity of the wine spoilage yeast, Dekkera bruxellensis. African Journal of Microbiology Research doi: 10.5897/AJMR2013.6350 (2013).
Harris, V., Jiranek, V., Ford, C. M. & Grbin, P. R. Inhibitory effect of hydroxycinnamic acids on Dekkera spp. Appl. Microbiol. Biotechnol. 86, 721–729 (2010).
Fernandes, a. R., Mira, N. P., Vargas, R. C., Canelhas, I. & Sá-Correia, I. Saccharomyces cerevisiae adaptation to weak acids involves the transcription factor Haa1p and Haa1p-regulated genes. Biochem. Biophys. Res. Commun. 337, 95–103 (2005).
Piper, P. et al. The Pdr12 ABC transporter is required for the development of weak organic acid resistance in yeast. The EMBO Journal 17, 4257–4265 (1998).
Stratford, M. & Anslow, P. a. Evidence that sorbic acid does not inhibit yeast as a classic ‘weak acid preservative’. Lett. Appl. Microbiol. 27, 203–206 (1998).
Piper, P., Calderon, C. O. & Hatzixanthis, K. REVIEW ARTICLE Weak acid adaptation: the stress response that confers yeasts with resistance to organic acid food preservatives. Microbiology 2635–2642 (2001).
Holyoak, C. D. et al. Activity of the plasma membrane H(+)-ATPase and optimal glycolytic flux are required for rapid adaptation and growth of Saccharomyces cerevisiae in the presence of the weak-acid preservative sorbic acid. Appl. Environ. Microbiol. 62, 3158–3164 (1996).
Ullah, A., Chandrasekaran, G., Brul, S. & Smits, G. J. Yeast adaptation to weak acids prevents futile energy expenditure. Front. Microbiol. 4, 142 (2013).
Chambel, A., Viegas, C. A. & Sa-correia, I. Effect of cinnamic acid on the growth and on plasma membrane H 1 – ATPase activity of Saccharomyces cerevisiae. International Journal of Food Microbiology 50, 173–179 (1999).
de Nobel, H. et al. Parallel and comparative analysis of the proteome and transcriptome of sorbic acid-stressed Saccharomyces cerevisiae. Yeast 18, 1413–1428 (2001).
Holyoak, C. D., Bracey, D., Piper, P. W. & Kuchler, K. The Saccharomyces cerevisiae Weak-Acid-Inducible ABC Transporter Pdr12 Transports Fluorescein and Preservative Anions from the Cytosol by an Energy-Dependent Mechanism. Journal of Bacteriology 181, 4644–4652 (1999).
Zhao, J., Yang, Z., Wang, M., Lu, Y. & Yang, Z. Electrochemical evaluation of the inhibitory effects of weak acids on Zygosaccharomyces bailii. J. Agric. Food Chem. 52, 7246–7250 (2004).
Guerreiro, J. F., Mira, N. P. & Sá-Correia, I. Adaptive response to acetic acid in the highly resistant yeast species Zygosaccharomyces bailii revealed by quantitative proteomics. Proteomics 12, 2303–2318 (2012).
Mira, N. P., Teixeira, M. C. & Sá-Correia, I. Adaptive response and tolerance to weak acids in Saccharomyces cerevisiae: a genome-wide view. OMICS 14, 525–540 (2010).
Coronado, P. et al. Comparison of the behaviour of Brettanomyces bruxellensis strain LAMAP L2480 growing in authentic and synthetic wines. Antonie Van Leeuwenhoek 107, 1217–1223 (2015).
de Barros Lopes, M., Rainieri, S., Henschke, P. A. & Langridge, P. AFLP fingerprinting for analysis of yeast genetic variation. Int. J. Syst. Bacteriol. 49, 915–924 (1999).
Conterno, L., Joseph, C. M. L., Arvik, T. J., Henick-Kling, T. & Bisson, L. F. Genetic and Physiological Characterization of Brettanomyces bruxellensis Strains Isolated from Wines. Am. J. Enol. Vitic. 57, 139–147 (2006).
Crauwels, S. et al. Assessing genetic diversity among Brettanomyces yeasts by DNA fingerprinting and whole-genome sequencing. Appl. Environ. Microbiol. 80, 4398–4413 (2014).
Vigentini, I. et al. Intraspecific variations of Dekkera/Brettanomyces bruxellensis genome studied by capillary electrophoresis separation of the intron splice site profiles. Int. J. Food Microbiol. 157, 6–15 (2012).
Woolfit, M., Rozpedowska, E., Piskur, J. & Wolfe, K. H. Genome survey sequencing of the wine spoilage yeast Dekkera (Brettanomyces) bruxellensis. Eukaryot. Cell 6, 721–733 (2007).
Curtin, C. D., Borneman, A. R., Chambers, P. J. & Pretorius, I. S. De-novo assembly and analysis of the heterozygous triploid genome of the wine spoilage yeast Dekkera bruxellensis AWRI1499. PLoS One 7, e33840 (2012).
Piškur, J. et al. The genome of wine yeast Dekkera bruxellensis provides a tool to explore its food-related properties. Int. J. Food Microbiol. 157, 202–209 (2012).
Hellborg, L. & Piskur, J. Complex nature of the genome in a wine spoilage yeast, Dekkera bruxellensis. Eukaryot. Cell 8, 1739–1749 (2009).
Borneman, A. R., Zeppel, R., Chambers, P. J. & Curtin, C. D. Insights into the Dekkera bruxellensis Genomic Landscape: Comparative Genomics Reveals Variations in Ploidy and Nutrient Utilisation Potential amongst Wine Isolates. PLoS Genet. 10, (2014).
Valdes, J. et al. Draft genome sequence and transcriptome analysis of the wine spoilage yeast Dekkera bruxellensis LAMAP2480 provides insights into genetic diversity, metabolism and survival. FEMS Microbiol. Lett. 361, 104–106 (2014).
Godoy, L. et al. Study of the coumarate decarboxylase and vinylphenol reductase activities of Dekkera bruxellensis (anamorph Brettanomyces bruxellensis) isolates. Lett. Appl. Microbiol. 48, 452–457 (2009).
Gerós, H., Azevedo, M.-M. & Cássio, F. Biochemical Studies on the Production of Acetic Acid by the Yeast Dekkera anomala. Food Technology and Biotechnology 38, 59–62 (2000).
Leite, F. C. B. et al. Quantitative aerobic physiology of the yeast Dekkera bruxellensis, a major contaminant in bioethanol production plants. FEMS Yeast Res. doi: 10.1111/j.1567-1364.2012.12007.x (2012).
Sigler, K. & Höfer, M. Mechanisms of acid extrusion in yeast. Biochim. Biophys. Acta 1071, 375–391 (1991).
Stratford, M. et al. Weak-acid preservatives: pH and proton movements in the yeast Saccharomyces cerevisiae. Int. J. Food Microbiol. 161, 164–171 (2013).
Piper, P., Peter, P., Mehdi, M., Calderon, C. O. & Kostas, H. Weak acid adaptation: the stress response that confers yeasts with resistance to organic acid food preservatives. Microbiology 147, 2635–2642 (2001).
Viegas, C. A. & Sá-Correia, I. Activation of plasma membrane ATPase of Saccharomyces cerevisiae by octanoic acid. J. Gen. Microbiol. 137, 645–651 (1991).
Piper, P. W., Ortiz-Calderon, C., Holyoak, C., Coote, P. & Cole, M. Hsp30, the integral plasma membrane heat shock protein of Saccharomyces cerevisiae, is a stress-inducible regulator of plasma membrane H(+)-ATPase. Cell Stress Chaperones 2, 12–24 (1997).
Curtin, C. D. & Pretorius, I. S. Genomic insights into the evolution of industrial yeast species Brettanomyces bruxellensis. FEMS Yeast Res. 14, 997–1005 (2014).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Nardi, T., Remize, F. & Alexandre, H. Adaptation of yeasts Saccharomyces cerevisiae and Brettanomyces bruxellensis to winemaking conditions: a comparative study of stress genes expression. Appl. Microbiol. Biotechnol. 88, 925–937 (2010).
Tiukova, I. a. et al. Transcriptome of the alternative ethanol production strain Dekkera bruxellensis CBS 11270 in sugar limited, low oxygen cultivation. PLoS One 8, e58455 (2013).
Clausen, M., Lamb, C. J., Megnet, R. & Doerner, P. W. PAD1 encodes phenylacrylic acid decarboxylase which confers resistance to cinnamic acid in Saccharomyces cerevisiae. Gene 142, 107–112 (1994).
Godoy, L., García, V., Peña, R., Martínez, C. & Ganga, M. A. Identification of the Dekkera bruxellensis phenolic acid decarboxylase (PAD) gene responsible for wine spoilage. Food Control 45, 81–86 (2014).
Larsson, S., Nilvebrant, N. O. & Jönsson, L. J. Effect of overexpression of Saccharomyces cerevisiae Pad1p on the resistance to phenylacrylic acids and lignocellulose hydrolysates under aerobic and oxygen-limited conditions. Appl. Microbiol. Biotechnol. 57, 167–174 (2001).
Plumridge, A., Stratford, M., Lowe, K. C. & Archer, D. B. The weak-acid preservative sorbic acid is decarboxylated and detoxified by a phenylacrylic acid decarboxylase, PadA1, in the spoilage mold Aspergillus niger. Appl. Environ. Microbiol. 74, 550–552 (2008).
Mira, N. P., Lourenço, A. B., Fernandes, A. R., Becker, J. D. & Isabel, S.-C. The RIM101 pathway has a role in Saccharomyces cerevisiae adaptive response and resistance to propionic acid and other weak acids. FEMS Yeast Res. 9, 202–216 (2009).
Teixeira, M. C., Santos, P. M., Fernandes, A. R. & Sá-Correia, I. A proteome analysis of the yeast response to the herbicide 2,4-dichlorophenoxyacetic acid. Proteomics 5, 1889–1901 (2005).
Teixeira, M. C., Fernandes, A. R., Mira, N. P., Becker, J. D. & Sá-Correia, I. Early transcriptional response of Saccharomyces cerevisiae to stress imposed by the herbicide 2,4-dichlorophenoxyacetic acid. FEMS Yeast Res. 6, 230–248 (2006).
Schaus, S. E., Cavalieri, D. & Myers, A. G. Gene transcription analysis of Saccharomyces cerevisiae exposed to neocarzinostatin protein-chromophore complex reveals evidence of DNA damage, a potential mechanism of resistance, and consequences of prolonged exposure. Proc. Natl. Acad. Sci. USA 98, 11075–11080 (2001).
Linde, J., Wilson, D., Hube, B. & Guthke, R. Regulatory network modelling of iron acquisition by a fungal pathogen in contact with epithelial cells. BMC Syst. Biol. 4, 148 (2010).
Cottier, F. et al. The transcriptional stress response of Candida albicans to weak organic acids. G3 5, 497–505 (2015).
Iwahashi, Y. et al. Mechanisms of patulin toxicity under conditions that inhibit yeast growth. J. Agric. Food Chem. 54, 1936–1942 (2006).
Castro, I. M., Cabral, D. B., Trópia, M. J. M., Almeida, F. M. & Brandão, R. L. Yeast genes YOL002C and SUL1 are involved in neomycin resistance. World J. Microbiol. Biotechnol. 17, 399–402 (2001).
Abbott, D. A., Suir, E., van Maris, A. J. A. & Pronk, J. T. Physiological and transcriptional responses to high concentrations of lactic acid in anaerobic chemostat cultures of Saccharomyces cerevisiae. Appl. Environ. Microbiol. 74, 5759–5768 (2008).
Hazelwood, L. A., Walsh, M. C., Pronk, J. T. & Daran, J.-M. Involvement of vacuolar sequestration and active transport in tolerance of Saccharomyces cerevisiae to hop iso-alpha-acids. Appl. Environ. Microbiol. 76, 318–328 (2010).
Makuc, J. et al. The putative monocarboxylate permeases of the yeast Saccharomyces cerevisiae do not transport monocarboxylic acids across the plasma membrane. Yeast 18, 1131–1143 (2001).
Li, L., Naseem, S., Sharma, S. & Konopka, J. B. Flavodoxin-Like Proteins Protect Candida albicans from Oxidative Stress and Promote Virulence. PLoS Pathog. 11, e1005147 (2015).
Grant, C. M. Role of the glutathione/glutaredoxin and thioredoxin systems in yeast growth and response to stress conditions. Mol. Microbiol. 39, 533–541 (2001).
Casal, M., Paiva, S., Andrade, R. P., Gancedo, C. & Leão, C. The lactate-proton symport of Saccharomyces cerevisiae is encoded by JEN1. J. Bacteriol. 181, 2620–2623 (1999).
Morano, K. A., Grant, C. M. & Moye-Rowley, W. S. The response to heat shock and oxidative stress in Saccharomyces cerevisiae. Genetics 190, 1157–1195 (2012).
North, M. et al. Genome-wide functional profiling reveals genes required for tolerance to benzene metabolites in yeast. PLoS One 6, e24205 (2011).
Piper, P. W. Yeast superoxide dismutase mutants reveal a pro-oxidant action of weak organic acid food preservatives. Free Radic. Biol. Med. 27, 1219–1227 (1999).
Kapteyn, J. C. et al. Low external pH induces HOG1-dependent changes in the organization of the Saccharomyces cerevisiae cell wall. Mol. Microbiol. 39, 469–479 (2001).
Brennan, T. C. R., Kromer, J. O. & Nielsen, L. K. Physiological and Transcriptional Responses of Saccharomyces cerevisiae to d-Limonene Show Changes to the Cell Wall but Not to the Plasma Membrane. Appl. Environ. Microbiol. 79, 3590–3600 (2013).
Mira, N. P., Becker, J. D. & Sá-Correia, I. Genomic expression program involving the Haa1p-regulon in Saccharomyces cerevisiae response to acetic acid. OMICS 14, 587–601 (2010).
Kimata, Y., Ishiwata-Kimata, Y., Yamada, S. & Kohno, K. Yeast unfolded protein response pathway regulates expression of genes for anti-oxidative stress and for cell surface proteins. Genes Cells 11, 59–69 (2006).
Kawahata, M., Miho, K., Kazuo, M., Tsutomu, F. & Haruyuki, I. Yeast genes involved in response to lactic acid and acetic acid: acidic conditions caused by the organic acids in Saccharomyces cerevisiae cultures induce expression of intracellular metal metabolism genes regulated by Aft1p. FEMS Yeast Res. 6, 924–936 (2006).
Dias, P., Paulo, D. & Isabel, S.-C. The drug:H antiporters of family 2 (DHA2), siderophore transporters (ARN) and glutathione:H antiporters (GEX) have a common evolutionary origin in hemiascomycete yeasts. BMC Genomics 14, 901 (2013).
Shimazu, M., Sekito, T., Akiyama, K., Ohsumi, Y. & Kakinuma, Y. A family of basic amino acid transporters of the vacuolar membrane from Saccharomyces cerevisiae. J. Biol. Chem. 280, 4851–4857 (2005).
Gabriel, F. et al. Deletion of the uracil permease gene confers cross-resistance to 5-fluorouracil and azoles in Candida lusitaniae and highlights antagonistic interaction between fluorinated nucleotides and fluconazole. Antimicrob. Agents Chemother. 58, 4476–4485 (2014).
Morita, Y., Nakamori, S. & Takagi, H. Effect of proline and arginine metabolism on freezing stress of Saccharomyces cerevisiae. J. Biosci. Bioeng. 94, 390–394 (2002).
Reihl, P. & Stolz, J. The monocarboxylate transporter homolog Mch5p catalyzes riboflavin (vitamin B2) uptake in Saccharomyces cerevisiae. J. Biol. Chem. 280, 39809–39817 (2005).
Godard, P. et al. Effect of 21 different nitrogen sources on global gene expression in the yeast Saccharomyces cerevisiae. Mol. Cell. Biol. 27, 3065–3086 (2007).
Deuschle, K. et al. The role of [Delta]1-pyrroline-5-carboxylate dehydrogenase in proline degradation. Plant Cell 16, 3413–3425 (2004).
Hazelwood, L. A. et al. A new physiological role for Pdr12p in Saccharomyces cerevisiae: export of aromatic and branched-chain organic acids produced in amino acid catabolism. FEMS Yeast Res. 6, 937–945 (2006).
Almeida, B. et al. Yeast protein expression profile during acetic acid-induced apoptosis indicates causal involvement of the TOR pathway. Proteomics 9, 720–732 (2009).
Mira, N. P., Palma, M., Guerreiro, J. F. & Sá-Correia, I. Genome-wide identification of Saccharomyces cerevisiae genes required for tolerance to acetic acid. Microb. Cell Fact. 9, 79 (2010).
Clowers, K. J., Heilberger, J., Piotrowski, J. S., Will, J. L. & Gasch, A. P. Ecological and Genetic Barriers Differentiate Natural Populations of Saccharomyces cerevisiae. Mol. Biol. Evol. 32, 2317–2327 (2015).
Donalies, U. E. B. & Stahl, U. Increasing sulphite formation in Saccharomyces cerevisiae by overexpression of MET14 and SSU1. Yeast 19, 475–484 (2002).
Siderius, M., Rots, E. & Mager, W. H. High-osmolarity signalling in Saccharomyces cerevisiae is modulated in a carbon-source-dependent fashion. Microbiology 143, 3241–3250 (1997).
Rep, M. et al. Osmotic stress-induced gene expression in Saccharomyces cerevisiae requires Msn1p and the novel nuclear factor Hot1p. Mol. Cell. Biol. 19, 5474–5485 (1999).
Sales, K., Kurt, S., Wolf, B., Elaine, R. & George, L. The LEA-like protein HSP 12 in Saccharomyces cerevisiae has a plasma membrane location and protects membranes against desiccation and ethanol-induced stress. Biochimica et Biophysica Acta (BBA) - Biomembranes 1463, 267–278 (2000).
Varela, J. C., Praekelt, U. M., Meacock, P. A., Planta, R. J. & Mager, W. H. The Saccharomyces cerevisiae HSP12 gene is activated by the high-osmolarity glycerol pathway and negatively regulated by protein kinase A. Mol. Cell. Biol. 15, 6232–6245 (1995).
Barata, A. et al. The effect of sugar concentration and temperature on growth and volatile phenol production by Dekkera bruxellensis in wine. FEMS Yeast Res. 8, 1097–1102 (2008).
Buchanan, R. L. & Cygnarowicz, M. L. A mathematical approach toward defining and calculating the duration of the lag phase. Food Microbiol. 7, 237–240 (1990).
Ullah, A., Orij, R., Brul, S. & Smits, G. J. Quantitative analysis of the modes of growth inhibition by weak organic acids in Saccharomyces cerevisiae. Appl. Environ. Microbiol. 78, 8377–8387 (2012).
Ross, K. A., Beta, T. & Arntfield, S. D. A comparative study on the phenolic acids identified and quantified in dry beans using HPLC as affected by different extraction and hydrolysis methods. Food Chem. 113, 336–344 (2009).
Nissen, T. L., Schulze, U., Nielsen, J. & Villadsen, J. Flux distributions in anaerobic, glucose-limited continuous cultures of Saccharomyces cerevisiae. Microbiology 143 (Pt 1), 203–218 (1997).
Baykov, A. A., Kasho, V. N. & Avaeva, S. M. Inorganic pyrophosphatase as a label in heterogeneous enzyme immunoassay. Anal. Biochem. 171, 271–276 (1988).
Rozpędowska, E. et al. Parallel evolution of the make–accumulate–consume strategy in Saccharomyces and Dekkera yeasts. Nat. Commun. 2, 302 (2011).
Conesa, A. & Götz, S. Blast2GO: A comprehensive suite for functional analysis in plant genomics. Int. J. Plant Genomics 2008, 619832 (2008).
Ye, J. et al. WEGO: a web tool for plotting GO annotations. Nucleic Acids Res. 34, W293–W297 (2006).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Livak, K. J. & Schmittgen, T. D. Analysis of Relative Gene Expression Data Using Real- Time Quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 408, 402–408 (2001).
Acknowledgements
This work was supported by the Comisión Nacional de Investigación Científica y Tecnológica CONICYT Postdoctorado/FONDECYT 3140083 (L.G.), FONDECYT Iniciación 11140666 (J.A.U.), Fondecyt 1150522 (C.M.), Millennium Nucleus for Fungal Integrative and Synthetic Biology NC120043 (M.A.G.), and USA1555-USACH grant (M.A.G.).
Author information
Authors and Affiliations
Contributions
L.G. conceived, designed the project and prepared the experiment samples. P.V.-W. and J.A.U. performed the bioinformatics analysis. L.G., P.V.-W. and J.A.U. wrote the paper. C.M. and M.A.G. supervised the work. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Godoy, L., Vera-Wolf, P., Martinez, C. et al. Comparative transcriptome assembly and genome-guided profiling for Brettanomyces bruxellensis LAMAP2480 during p-coumaric acid stress. Sci Rep 6, 34304 (2016). https://doi.org/10.1038/srep34304
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep34304
This article is cited by
-
Understanding the Streptomyces albulus response to low-pH stress at the interface of physiology and transcriptomics
Applied Microbiology and Biotechnology (2023)
-
Novel antimicrobial peptides produced by Candida intermedia LAMAP1790 active against the wine-spoilage yeast Brettanomyces bruxellensis
Antonie van Leeuwenhoek (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.