Abstract
Cell senescence can limit proliferative potential and prevent tumorigenesis. Bmi1 is a key regulator in cell senescence by suppressing the Ink4a/Arf locus. However, how to regulate Bmi1 activity in cell senescence is largely unknown. Here, we show that SENP1 plays an important role in cell senescence by regulating Bmi1 SUMOylation. Senp1−/− primary MEF cells show resistance to cell senescence induced by passaging or other senescence inducing signals. SENP1 deficiency also reduces oncogene H-RasV12-induced senescence, and enhances H-RasV12-induced cell transformation. We further show that in Senp1−/− MEFs the expression of p19Arf, an important regulator in p53/p21-mediated cell senescence, is markedly reduced. Meanwhile, we demonstrate that SENP1 can specifically de-SUMOylate Bmi1 and thereby decreases the occupancy of Bmi1 on p19Arf promoter leading to decrease of H2AK119 mono-ubiquitination and up-expression of p19Arf. These data reveal a crucial role of SENP1 in regulation of cell senescence as well as cell transformation.
Similar content being viewed by others
Introduction
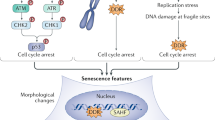
Cell senescence is an irreversible growth-arrested status that is characterized by flat cell morphology, senescence-associated-β-galactosidase (SA-β-gal) activity, and up-regulation of cell cycle inhibitors1. A variety of stresses, including telomere uncapping, DNA damage, oxidative stress, and oncogene can trigger cell senescence2. Senescence triggered by cell-intrinsic mechanisms, such as DNA damage and oxidative stress, are referred to as replicative senescence, whereas senescence induced by ectopic expression of activated onco-proteins, such as Ras and Raf, as oncogene-induced senescence (OIS)3,4,5. Cell senescence plays an important role in tumor suppression and organ aging2,6,7,8. Different stress signals induce cell senescence through activation of p16Ink4a/Rb or p19Arf/p53 pathway2,9,10,11.
PcG complexes are a family of transcriptional repressors and suppress the expression of lots of genes including p16Ink4a or p19Arf via modulation of histone modification. PcG has two multi-protein complexes, polycomb repressive complex 1 (PRC1) and 2 (PRC2). PRC1 includes Bmi1, CBXs, PHC1-3, RNF1-2, and SCML1-2 proteins12. PRC1 mono-ubiquitinates histone H2A at lysine 119 (H2AK119ub)13,14,15, which is associated with gene silencing16,17,18. PRC2 contains EZH2, EED, and SUZ12, which trimethylates histone H3 on Lys 27 (H3K27me3)19,20. It has been reported that PRC1 components Bmi1, CBX7, and CBX8 can delay the onset of senescence in mouse and human embryonic fibroblasts by repressing the expression of p16Ink4a or p19Arf 21,22,23.
SUMOylation has emerged as an important protein modification in regulation of many cellular processes24. SUMOylation is a dynamic process, which is sequentially catalyzed by SUMO-specific E1, E2 and E3 ligase and can be reversed by SUMO-specific proteases (SENPs)25,26. Recently, it was reported that DNA damage induces SUMOylation of Bmi127. SUMOylation can regulate Bmi1 to be recruited to DNA breaks. However, whether SUMOylation modulates Bmi1 activity in cell senescence is largely unknown. In this study, we show that Senp1−/− primary MEF cells are resistant to cell senescence triggered by passaging or other senescence inducing signals. Moreover, Senp1−/− MEF cells are also resistant to oncogene H-RasV12-induced senescence and show enhanced H-RasV12-induced cell transformation. Mechanistically, SENP1 de-SUMOylates Bmi1, and thus decreases the occupancy of Bmi1 on p19Arf promoter locus leading to the expression of p19Arf/p53, and subsequently promotes cell senescence.
Results
SENP1 deficiency reduces the senescence of MEF cells
Our previous study has demonstrated that Senp1−/− MEF cells were easily immortalized by using a 3T3 protocol compared to wild-type (WT) cells, indicating that SENP1 deficiency might keep MEFs away from cell senescence. To test it, we freshly isolated MEF cells from Senp1+/+ and Senp1−/− embryos at E12.5 and cultured them by using a 3T3 protocol. As shown in Fig. 1a, Senp1+/+ MEF cells entered growth plateau phase at 6 to 8 passages, whereas Senp1−/− MEF cells proliferated steadily up to 20 passages. Consistent with this observation, Senp1−/− MEF cells exhibited a higher BrdU incorporation rate than Senp1+/+ MEF cells did (Fig. 1b), indicating higher proliferation rate in Senp1−/− MEF cells. We further stained SA-β-gal and found that SA-β-gal positive cells were markedly less in Senp1−/− but not in Senp2−/− MEF cells when compared with control WT MEF cells (Fig. 1c). Furthermore, we determined whether SENP1 deficiency could affect the senescence induced by serum deprivation or oxidative stress. Senp1+/+ and Senp1−/− MEF cells at passage 5 were cultured in medium containing 50 μM H2O2 or 3% serum. Both conditions reduced the proliferation of Senp1+/+ MEFs but not Senp1−/− cells (Fig. 1d). We also confirmed that SENP1 deficiency could decrease the senescence in both conditions (Fig. 1e).
(a) Population doublings (PD) over serial passaging according to a 3T3 protocol were measured in Senp1+/+ and Senp1−/− cell lines. (b) Proliferation rate of primary MEF cells at passage 5 (P5) and passage 12 (P12) were measured by using BrdU labelling assay. (c) Senp1+/+, Senp1−/−, Senp2+/+ and Senp2−/− MEF cells at passage 12 (P12) were stained for SA-β-Gal activity. Representative images are shown in left panel. SA-β-Gal positive cells were counted at least in five fields. Data represent the mean ± S.E.M. of three independent experiments (right panel) (**P < 0.01, Student’s t-test). (d) Cell proliferation rates of Senp1+/+ and Senp1−/− MEF cells were measured under the conditions of low serum (3%) (upper) or sub-lethal concentration of H2O2 (50 μM) (lower). (e) Senp1+/+ and Senp1−/− MEF cells in low serum or 50 μM H2O2 were stained for SA-β-Gal activity. Data represent the mean ± S.E.M. of three independent experiments (**P < 0.01, Student’s t-test).
SENP1 deficiency decreases H-RasV12-induced senescence
Oncogene-induced senescence acts as a barrier to prevent oncogenic cell transformation and tumorigenesis5,28. To determine the role of SENP1 deficiency in oncogene-induced senescence, we stably transduced H-RasV12 into Senp1+/+ or Senp1−/− MEF cells at early-passages (passage 2 or 3). The cell proliferation assay showed that H-RasV12 inhibited less cell growth in Senp1−/− MEF cells than that in Senp1+/+ cells (Fig. 2a). The number of SA-β-gal positive cells was much less in H-RasV12-transduced Senp1−/− MEF cells than those in Senp1+/+ MEF cells (Fig. 2b). We also stained phosphorylated histone H2AX (γH2AX) and observed less γH2AX foci in H-RasV12-transduced Senp1−/− MEF cells when compared to Senp1+/+ MEF cells (Fig. 2c). Additionally, the percentages of γH2AX and macroH2A positive cells were also lower in H-RasV12-transduced Senp1−/− MEF cells than that in Senp1+/+ cells (Figure S1a–d). We further analysed the colony forming ability in H-RasV12-transduced MEF cells. As shown in Fig. 2d, more colonies were observed in H-RasV12-transduced Senp1−/− MEF cells than in control cells. These data suggest that SENP1 deficiency can decrease oncogene-induced cell senescence.
(a) Cell numbers were measured in primary Senp1+/+ and Senp1−/− MEF cells (passage 2 or 3) infected with mock or H-RasV12 lentiviral vectors. The cell numbers were counted for 6 days. Data represent the mean ± S.E.M. of three independent experiments. A p value of <0.05 (*) indicates a significant difference between H-RasV12 lentiviral vectors-infected Senp1+/+ and Senp1−/− MEFs. (b) Mock lentiviral vectors or H-RasV12 lentiviral vectors-infected cells were stained for SA-β-gal (left panel); the percentage of SA-β-gal positive cells was counted at least in five fields (right panel) (**P < 0.01, Student’s t-test). (c) Senp1+/+ and Senp1−/− MEF cells were stained for γH2AX (left panel); Percentage of cells with γH2AX foci per nucleus is shown in right panel. (d) Growth of Senp1+/+ and Senp1−/− MEF cells at early-passage (passage 2 or 3) transduced with H-RasV12 were measured by colony forming assay. Data represent the mean ±S.E.M. of three independent experiments (*P < 0.05, Student’s t-test).
Downregulation of p19Arf in Senp1−/− MEF cells
To understand the mechanism underlying SENP1 regulating senescence, we analysed the activity of the two major senescence-related signal pathways p19Arf/p53/p21 and p16Ink4a/Rb in Senp1+/+ and Senp1−/− MEF cells. As shown in Fig. 3a, the expressions of p19Arf, p53, and p21 proteins were markedly decreased in Senp1−/− MEF cells when compared to Senp1+/+ cells. However, p16Ink4a and Rb proteins showed only mild reduction in Senp1−/− MEF cells in comparison to Senp1+/+ cells. We also detected the expressions of p19Arf and p16Ink4a in H-RasV12-transduced Senp1+/+ or Senp1−/− MEF cells and found that H-RasV12 significantly induced p19Arf but not p16Ink4a expression in both cells (Fig. 3b). The expression of p19Arf was less increased in H-RasV12-transduced Senp1−/− MEFs when compared to Senp1+/+ cells. We further transfected SENP1 wild-type (SENP1 WT) or SENP1 catalytic mutant (SENP1m) in Senp1−/− MEF cells and found that re-expression of SENP1 WT but not SENP1m could rescue p19Arf expression in Senp1−/− MEF cells (Fig. 3c). To further confirm the role of SENP1 in regulating p19Arf expression, we constructed p19Arf promoter-driven luciferase reporter gene and showed that SENP1 WT but not SENP1m could induce p19Arf transcription (Fig. 3d). Taken together, these data reveal that SENP1 promotes p19Arf expression.
(a) Senp1+/+ or Senp1−/− MEF cells at passage 5 (P5) were analysed for expression of p19Arf, p53, p21, p16Ink4a, p-RB, and cyclin D1. The MEFs were cultured following a 3T3 protocol. Shown are the cropped blot images representing indicated proteins. Full-length blots are presented in the Supplementary Figure S3. (b) Expression of p19Arf and p16Ink4a was analysed in cell extracts from Senp1+/+ or Senp1−/− MEF cells infected with H-RasV12 lentiviral vectors. Cropped blot images shows the representing indicated proteins. Full-length blots are presented in the Supplementary Figure S4. (c) Primary Senp1−/− MEF cells at passage 5 transfected with RGS-SENP1, RGS-SENP1m or Vector were analysed the expression of p19Arf by western blotting. Shown are the cropped blot images representing indicated proteins. Full-length blots are presented in the Supplementary Figure S4. (d) P19Arf transcription was analyzed in MEF cells transfected with p19Arf luciferase plasmid and SENP1 or SENP1m plasmids. Data represent the mean ± S.E.M. of three independent experiments (*P < 0.05, Student’s t-test).
SENP1 promotes p19Arf expression via de-SUMOylation of Bmi1
Bmi1 is a critical negative regulator for p19Arf and p16Ink4a expression23,29,30. It has also been reported as a SUMOylated protein27. Therefore, we postulated that SENP1 might regulate p19Arf expression via de-SUMOylation of Bmi1. To test whether SENP1 directly de-SUMOylates Bmi1, Flag-Bmi1 and HA-SUMO1 were co-expressed with RGS-tagged SENP1 or SENP1m in 293T cell lines. As a result, the co-expression of SENP1, not SENP1m, could de-conjugate SUMOylated Bmi1, while the SENP1m increase the SUMOylated Bmi1 (Fig. 4a,b) through a dominant negative activity to block the de-SUMOylation activity of endogenous SENP131. More importantly, we observed that the SUMOylated Bmi1 proteins were accumulated in Senp1−/− MEF cells (Fig. 4c). To exclude the possibility that Bmi1 expression was affected by SENP1, we checked the expression of Bmi1, as well as the expressions of other PRC1 components CBX4 and Ring1B, and found no changes in Senp1−/− MEF cells (Figure S2A). These results indicate that SENP1 is a specific protease to de-SUMOylate Bmi1.
(a) Bmi1 was immunoprecipiated from cell lysates of 293T cells transfected with the indicated plasmids, and blotted for SUMO1 or Bmi1. Whole cell lysates were blotted with anti-HA, -Flag, or -RGS antibodies. Shown are the cropped blot images representing indicated proteins. Full-length blots are presented in the Supplementary Figure S5. (b) 293T cells were transfected with indicated plasmids. SUMO1-conjugated proteins were pulled down by talon beads from cell lysate. Bound proteins were blotted with anti-His, -Flag, or -RGS antibodies. Shown are the cropped blot images representing indicated proteins. Full-length blots are presented in the Supplementary Figure S5. (c) Senp1+/+ and Senp1−/− MEFs or Bmi1 silencing (shBmi1) MEFs were lysed. SUMO-conjugated proteins were immunoprecipitated with anti-SUMO1 and bound proteins were blotted with anti-SUMO1 or -Bmi1 antibodies. Whole cell lysates were blotted with anti-SUMO1 or -Bmi1 antibodies. Shown are the cropped blot images representing indicated proteins. Full-length blots are presented in the Supplementary Figure S5. (d) H2AK119ub was blotted in Senp1+/+ and Senp1−/− MEFs. Shown are the cropped blot images representing indicated proteins. Full-length blots are presented in the Supplementary Figure S5. (e) P19Arf transcription was analyzed in MEFs transfected with p19Arf luciferase plasmid, Flag-Bmi1 WT or K88R plasmids. Data represent the mean ± S.E.M. of three independent experiments (*P < 0.05, Student’s t-test). (f) P19Arf transcription was analyzed in MEFs transfected with p19Arf luciferase plasmid, Flag-Bmi1 WT or K88R, plus SENP1 or SENP1m plasmid. Data represent the mean ± S.E.M. of three independent experiments (*P < 0.05, Student’s t-test; n.s., non-significant). (g) Senp1−/− MEFs were transfected with Flag-Bmi1 WT, Flag-Bmi1 K88R, or vector as indicated. Chromatins prepared from these cells were subjected to IP with anti-Flag beads followed by Real-time PCR using primers for p19Arf promoter. Data represent the mean ± S.E.M. of three independent experiments (*P < 0.05, Student’s t-test). (h) Chromatins from Senp1+/+ and Senp1−/− MEFs were subjected to IP with anti-Bmi1, -H2AK119ub or IgG followed by Real-time PCR using primers for p19Arf promoter. Data represents the mean ± S.E.M. of three independent experiments (*P < 0.05, Student’s t-test).
We further assessed whether SENP1 could affect Bmi1 ubiquitin ligase activity in mono-ubiquitination of histone 2A at K119. As shown in Fig. 4d, mono-ubiquitination of H2AK119 was increased in Senp1−/− MEFs when compared to Senp1+/+ cells, suggesting that SENP1 mediated de-SUMOylation decreases Bmi1 ubiquitin ligase activity. We also measured the suppressive activity of Bmi1 WT and Bmi1 SUMOylation mutant (K88R) using p19Arf promoter-driven luciferase assay. Bmi1 WT significantly reduced p19Arf transcription, whereas Bmi1 K88R mutant showed a weaker repressive activity than Bmi1 WT (Fig. 4e). Meanwhile we found that SENP1, but not SENP1m could counteract the suppressive activity of Bmi1 WT on p19Arf transcription (Fig. 4f).
We reasoned that SUMOylation might promote Bmi1 binding to p19Arf promoter. To test it, we used chromatin immunoprecipitation (ChIP) assay to demonstrate that the occupancy of Bmi1 WT on p19Arf promoter was much higher than that of Bmi1 K88R mutant (Fig. 4g). We also illustrated that Bmi1 occupancy on p19Arf promoter was more profound in Senp1−/− MEFs than that in WT cells, and that the ubiquitination of H2A at K119 of p19Arf promoter was also increased in Senp1−/− MEFs when compared to WT cells (Fig. 4h). Taken together, these data suggest that SENP1 promotes p19Arf expression via de-SUMOylation of Bmi1.
SUMOylation is essential for Bmi1 repression of senescence
To determine whether SUMOylation affect Bmi1 repression of cell senescence, we transduced H-RasV12-Senp1−/− MEF cells either with Bmi1 WT or K88R mutant. SA-β-gal staining showed that both Bmi1 WT and Bmi1 K88R expressions decreased the percentage of SA-β-gal positive cells in H-RasV12-Senp1−/− MEF cells compared to vector control (Fig. 5a). However, more SA-β-gal positive cells were observed in Bmi1 K88R-transduced cells than that in Bmi1 WT control. We further found that both Bmi1 WT and Bmi1 K88R expressions increased colony formation in H-RasV12-Senp1−/− MEF cells compared to vector control. Less colony were found in Bmi1 K88R-transduced H-RasV12-Senp1−/− MEF cells in comparison to Bmi-1 WT control (Fig. 5b). These data suggest that SUMOylation is essential for Bmi1 repression of cell senescence.
(a) H-RasV12 transduced Bmi1 (WT or K88R)-Senp1−/− MEF cells were stained for SA-β-Gal activity. Primary MEF cells at passage 2 or 3 were sequentially transduced with Bmi1 and H-RasV12. The assay was performed at around passage 8. Representative images are shown in left panel. SA-β-Gal positive cells were counted at least in five fields (right panel). Data represent the mean ± S.E.M. of three independent experiments (*P < 0.05, Student’s t-test). (b) Growth of Bmi1 (WT or K88R)-Senp1−/− MEF cells transduced with H-RasV12 was measured by colony forming assay. Data represent the mean ± S.E.M. of three independent experiments (*P < 0.05, Student’s t-test).
SENP1 expression is associated with cell senescence in prostate PIN lesion
Previously, we have shown that SENP1 was overexpressed in prostrate intraepithelial neoplasia (PIN) lesion32 and tumor cells33,34. Overexpression of SENP1 in mouse prostate induced PIN lesions, but not tumor formation33. We speculated that overexpression of SENP1 could promote the growth of prostate epithelial cell as well as cell senescence in PIN lesion. We thus collected 44 human prostate PIN samples and analysed the expression of SENP1, senescence markers heterochromatin proteins 1 γ (HP1γ)35,36, p16INK4a, p14ARF and Bmi1 by immunohistochemistry staining in these samples. A score indicating SENP1, HP1γ, p16INK4a, p14ARF or Bmi1 was given ranging from 0 to 3 based on the percentage of the stained area and immunostaining intensity in these samples. As shown, SENP1 expression was positively related to HP1γ (rs = 0.553, P < 0.0001), p14ARF (rs = 0.6578, P < 0.0001), and p16INK4a (rs = 0.6755, P < 0.0001) expressions in human PIN samples (Fig. 6a,b) (Figure S3a,b). However, it was not related to Bmi1 expression (rs = −1501, P = 3309) (Figure S3a,b). These data indicate that SENP1 expression is associated with cell senescence in prostate PIN lesion.
(a) Human PIN samples stained for SENP1, p14ARF, HP1γ and Bmi1. (b) Immunohistochemistry was performed to evaluate SENP1, p14ARF and HP1γ expression on PIN TMA slides from patients. SENP1, p14ARF and HP1γ protein level in these samples was scored from 0 to 3 based on the stained area percentage and immunostaining intensity. Spearman correlation coefficient was performed to evaluate the association of SENP1 with p14ARF or HP1γ. (c) Working model shows the mechanism of SENP1 in regulating cell senescence.
Discussion
In this study, we demonstrate a role of SENP1 in cell senescence. SENP1 de-SUMOylates Bmi1 and reduces the occupancy of Bmi1 on p19Arf promoter, leading to p19Arf expression and cell senescence (Fig. 6c). SUMOylated Bmi1 was accumulated in Senp1−/− MEFs (Fig. 4c) and SUMOylation promoted Bmi1 occupancy on p19Arf promoter to suppress p19Arf expression (Fig. 4f–h). Interestingly, Bmi1−/− cells showed increased Ink4a/Arf expression, especially p16Ink4a expression23, whereas p16Ink4a expression is only mildly decreased in Senp1−/− MEFs compared to p19Arf. These results suggest that SUMOylation might selectively modulate Bmi1 effect on p19Arf by promoting its binding to the promoter locus, although currently it is unknown how SUMOylation modulates Bmi1 selection.
We showed the deficiency of SENP1 delays MEF cell senescence, which seems to contradict with the previously report on SENP1 represses cell senescence in human fibroblasts37. Yates et al. showed that shSENP1 in human fibroblasts induced senescence. They further showed that shSENP1 increased p53 expression via unknown mechanism. In the present study, we find that SENP1 deficiency reduces p19Arf/p16Ink4a/p53 expression via Bmi1. As mouse cells have very long telomeres (40–60 kb) when compared to human cells (5–15 kb)38, we currently don’t know whether this difference would contribute to their different responses to senescence. We also note another difference in the approaches that we have utilized for SENP1 deficiency from Yates et al.37. We deleted Senp1 gene in genome of Senp1−/− MEF cells, whereas SENP1 was silenced at mRNA level by shSENP1in HFF cells, which might cause off target effect.
The SUMOylated Bmi1 was first reported to accumulate at the DNA damage sites27. Our data showed the SUMOylation increased Bmi1 binding to DNA. It is possible the increased Bmi1 may recruit some DNA repairing proteins to repair oncogene induced DNA damage and to reduce the γH2Ax foci. Bmi1 is proposed to bind and increase Ring1B’s E3 ligase activity39. Bmi1/Ring1b, an autoinhibited RING E3 ubiquitin ligase, was also reported to promote SENP1 ubiquitination and degradation40, suggesting that this regulation might be a negative feedback mechanism to control SENP1 action on Bmi1, which would promote cell to be transformed during tumorigenesis. However, we don’t know whether SUMOylated Bmi1 further increases the E3 ligase activity of Ring1B and thereby affects ubiquitination of SENP1.
It is well-known that ROS-induced DNA damage can increase p53 activation, which in turn leads to cellular senescence38,41,42,43. Previously, we have shown that ROS production decreases in Senp1−/− MEF cells because of the defect in mitochondrial biogenesis44. Thus, the lower ROS production might partially contribute to the reduction of senescence in Senp1−/− MEF cells. It is evident that we observe a decrease in DNA damage marker γH2AX foci in Senp1−/− MEF cells.
We previously found SENP1 was overexpressed in PIN lesion32 and tumor cells33,34. We also showed that overexpression of SENP1 in mouse prostate induced PIN lesions, but no tumor formation in the transgenic model33. We show herein that SENP1 expression is associated with the senescence in PIN lesion. We expect to see if bypassing senescence would promote PIN to cancer in the SENP1 prostate transgenic model. This is supported by our recent observations that tumor suppressor PTEN deficiency promote cancer development in SENP1 prostate transgenic mice partially through the decrease of p53 expression and cell senescence (unpublished data), suggesting the role of SENP1-mediated senescence in tumorigenesis, which need to be further explored in future.
Material and Methods
Mouse embryo fibroblasts isolation and cell culture
All animal procedures were approved by the Institutional Animal Care and Use Committee of Shanghai Jiao Tong University School of Medicine, which were carried out in accordance with the guidelines (IACUC No. 2013027). The generation of Senp1+/+ or Senp1−/− MEF cells was previously described45. MEF cells were cultured in completed DMEM containing 10% FBS at 37 °C under 5% CO2. For replicative senescence assay, cells were passaged following a 3T3 protocol46,47. Cells were split every 3 days and replanted at a density of 1 × 106 cells per 10 cm dish for 20 passages. For colony forming assay, MEF cells were seeded at a density of 5 × 104 cells per 10 cm dish and cultured for 2 weeks. Then cells were washed with phosphate-buffered saline, fixed in 70% ethanol for 15 min and stained with crystal violet. Colonies were observed under microscope. The cell mass with more than 50 cells was counted as a colony.
Plasmids and antibodies
Flag-Bmi1 (WT or K88R) and Flag-GFP-Bmi1 (WT or K88R) were generated using standard cloning procedures. K88R mutation was introduced into Bmi1 by site directed mutagenesis using QuikChange Site-Directed Mutagenesis Kit (Agilent California, USA). HA-SUMO1, HA-UBC9, RGS-SENP1 and RGS-SENP1m were previously described45,48. Anti-FLAG (mouse; #F1804) and anti-HA(mouse; #H3663) antibodies were from Sigma; anti-RGS-his (mouse; #34610) antibody from QIAGEN; anti-H-Ras (mouse; #sc-53959) and p16Ink4a (mouse; #sc-1661) antibodies from Santa Cruz; anti-SUMO1 (rabbit; #ab5316), H3K27me3 (mouse; #ab6002) and p19Arf (rabbit; #ab80) antibodies from Abcam; anti-Bmi1 (mouse; #05–637), γH2AX (mouse; #05–636) and uH2AK119 (mouse; #05–678) antibodies from Millipore.
Senescence associated β-galactosidase staining
MEF cells were plated at a density of 1 × 105 cells in 35 mm dishes. Cells were fixed and stained according to the instructions supplied by the senescence β-galactosidase staining kit (Beyotime biotechnology, China). Briefly, cells were washed with PBS and fixed in SA-β-Gal fixing solution for 15 min at room temperature. Cells then were washed 3 times with PBS and stained with working solution (10 μl buffer A, 10 μl buffer B, 930 μl buffer C, and 50 μl X-Gal solution) overnight at 37 °C. Cells were counterstained with nuclear fast red. The population of SA-β-Gal positive cells was determined by counting 400 cells in at least 5 fields per dish and images were taken using a phase-contrast microscope at 400× magnification (Olympus, Japan). The proportions of cells positive for SA-β-Gal activity are shown as the percentage of the total number of cells counted in each dish.
BrdU incorporation assay
Sub-confluent Senp1+/+ and Senp1−/− MEF cells at passage 5 (P5) and passage 12 (P12) were labelled for 4 h with 10 μM 5-bromo-2′-deoxyuridine (BrdU; Amersham). Labelled MEF cells were planted on cover slips and fixed in 5% acetic acid and 95% ethanol for 15 min at −20 °C. After fixation, cells were washed with PBS and treated with 0.1 mg/ml RNase A for 30 min at 37 °C. After washed with PBS, cells were washed with 2 N HCl, 0.5% Triton X-100 for 30 min at room temperature. Thereafter the cells were washed with PBS and incubated with BrdU antibody and FITC conjugated secondary antibody for 1 h. After washing with PBS, cells were counterstained with DAPI and analyzed under fluorescence microscope.
Immunoprecipitation and immunoblotting
Cells were washed once and lysed in the presence of 10 mM N-ethylmaleimide (NEM) using ice-cold RIPA buffer (50 mM Tris-HCl (pH 7.4), 400 mM NaCl, 1% Triton X-100, 0.1% SDS, 1 mM PMSF, and protease inhibitors) for 30 min on ice. Cell lysates were centrifugated at 14,000 × g for 10 min at 4 °C. The supernatants were collected and incubated with appropriate antibodies overnight at 4 °C, followed by incubation with protein A/G-Sepharose beads (Amersham Biosciences) for another 2 h at 4 °C. After washing three times with RIPA buffer, the immunoprecipitates were eluted with Laemmli sample buffer, elute boiled, and subjected to western blot analysis.
Immunofluorescent staining
Paraformaldehyde-fixed cells were treated with 0.3% Triton X-100 in PBS for 15 min. After washing with PBS, cells were blocked with 1% BSA in PBS for 1 h, and then incubated with anti-γH2AX(1:200) antibody overnight at 4 °C, followed by incubation with APC conjugated secondary antibody(1:200) for 1 h. Cells were then counterstained with DAPI. Samples were visualized with an Olympus fluorescence microscopy.
Chromatin immunoprecipitation (ChIP) assays
Formaldehyde-crosslinked chromatin was prepared from MEF cells and immunoprecipitations were performed using the ChIP Assay Kit according to the manufacturer’s recommended protocol (Upstate Biotechnology). Antibodies used for ChIP assays were anti-Flag-M2, anti-Bmi1, anti-H2AK119ub or IgG. A pair of real-time PCR primers was used for amplification of the promoter segments of the mouse p19Arf gene as previously49: forward, 5′-AAAACCCTCTCTTGGAGTGGG-3′; reverse, 5′-GCAGGTTCTTGGTCACTGTGAG-3′.
Luciferase assays
P19Arf promoter containing 1202 nucleotides upstream the ATG start codon was cloned into the pGL3-basic luciferase reporter vector (Promega) using standard cloning procedures. For reporter assays, MEF cells were transfected with X-tremeGENE HP DNA Transfection Reagent (Roche Applied Science). 36 h after transfection, the luciferase was assayed using the Dual-Luciferase reporter assay system (Promega). Luciferase activity of the reporter construct was normalized on the basis of the Renilla luciferase activity.
Statistical analyses
All data are represented by the mean ± S.E.M. of at least three independent experiments. GraphPad Prim 5 (GraphPad Software, La Jolla, CA) was used for statistical analysis. Data was analysed using two-tailed student’s t-test. A p value of <0.05 was considered significant.
Additional Information
How to cite this article: Xia, N. et al. SENP1 Is a Crucial Regulator for Cell Senescence through DeSUMOylation of Bmi1. Sci. Rep. 6, 34099; doi: 10.1038/srep34099 (2016).
References
Kuilman, T., Michaloglou, C., Mooi, W. J. & Peeper, D. S. The essence of senescence. Genes Dev 24, 2463–2479 (2010).
Collado, M., Blasco, M. A. & Serrano, M. Cellular senescence in cancer and aging. Cell 130, 223–233 (2007).
Campisi, J. & di Fagagna, F. D. Cellular senescence: when bad things happen to good cells. Nat Rev Mol Cell Biol 8, 729–740 (2007).
Courtois-Cox, S., Jones, S. L. & Cichowski, K. Many roads lead to oncogene-induced senescence. Oncogene 27, 2801–2809 (2008).
Braig, M. & Schmitt, C. A. Oncogene-induced senescence: Putting the brakes on tumor development. Cancer Res 66, 2881–2884 (2006).
Krizhanovsky, V. et al. Implications of Cellular Senescence in Tissue Damage Response, Tumor Suppression, and Stem Cell Biology. Cold Spring Harb Symp Quant Biol 73, 513–522 (2008).
Roninson, I. B. Tumor cell senescence in cancer treatment. Cancer Res 63, 2705–2715 (2003).
Collado, M. et al. Tumour biology - Senescence in premalignant tumours. Nature 436, 642–642 (2005).
Sharpless, N. E. Ink4a/Arf links senescence and aging. Exp Gerontol 39, 1751–1759 (2004).
del Arroyo, A. G. & Peters, G. The Ink4a/Arf network–cell cycle checkpoint or emergency brake? Adv Exp Med Biol 570, 227–247 (2005).
Sharpless, N. E. INK4a/ARF: a multifunctional tumor suppressor locus. Mutat Res 576, 22–38 (2005).
Levine, S. S., King, I. F. & Kingston, R. E. Division of labor in polycomb group repression. Trends Biochem Sci 29, 478–485 (2004).
Otte, A. P. & Kwaks, T. H. Gene repression by Polycomb group protein complexes: a distinct complex for every occasion? Curr Opin Genet Dev 13, 448–454 (2003).
Margueron, R. & Reinberg, D. The Polycomb complex PRC2 and its mark in life. Nature 469, 343–349 (2011).
Gil, J. & O’Loghlen, A. PRC1 complex diversity: where is it taking us? Trends Cell Biol 24, 632–641 (2014).
Cao, R., Tsukada, Y. & Zhang, Y. Role of Bmi-1 and Ring1A in H2A ubiquitylation and Hox gene silencing. Mol Cell 20, 845–854 (2005).
Wang, H. et al. Role of histone H2A ubiquitination in Polycomb silencing. Nature 431, 873–878 (2004).
Buchwald, G. et al. Structure and E3-ligase activity of the Ring-Ring complex of polycomb proteins Bmi1 and Ring1b. Embo J 25, 2465–2474 (2006).
Cao, R. & Zhang, Y. The functions of E(Z)/EZH2-mediated methylation of lysine 27 in histone H3. Curr Opin Genet Dev 14, 155–164 (2004).
Pasini, D., Bracken, A. P., Jensen, M. R., Denchi, E. L. & Helin, K. Suz12 is essential for mouse development and for EZH2 histone methyltransferase activity. Embo J 23, 4061–4071 (2004).
Dietrich, N. et al. Bypass of senescence by the polycomb group protein CBX8 through direct binding to the INK4A-ARF locus. Embo J 26, 1637–1648 (2007).
Gil, J., Bernard, D., Martinez, D. & Beach, D. Polycomb CBX7 has a unifying role in cellular lifespan. Nat Cell Biol 6, 67–72 (2004).
Jacobs, J. J., Kieboom, K., Marino, S., DePinho, R. A. & van Lohuizen, M. The oncogene and Polycomb-group gene bmi-1 regulates cell proliferation and senescence through the ink4a locus. Nature 397, 164–168 (1999).
Geiss-Friedlander, R. & Melchior, F. Concepts in sumoylation: a decade on. Nat Rev Mol Cell Biol 8, 947–956 (2007).
Hay, R. T. SUMO: a history of modification. Mol Cell 18, 1–12 (2005).
Yeh, E. T. H. SUMOylation and De-SUMOylation: Wrestling with Life’s Processes. J Biol Chem 284, 8223–8227 (2009).
Ismail, I. H. et al. CBX4-mediated SUMO modification regulates BMI1 recruitment at sites of DNA damage. Nucleic Acids Res 40, 5497–5510 (2012).
Larsson, L. G. Oncogene- and tumor suppressor gene-mediated suppression of cellular senescence. Semin Cancer Biol 21, 367–376 (2011).
Liu, Y. et al. Akt phosphorylates the transcriptional repressor bmi1 to block its effects on the tumor-suppressing ink4a-arf locus. Sci Signal 5, ra77 (2012).
Bruggeman, S. W. et al. Ink4a and Arf differentially affect cell proliferation and neural stem cell self-renewal in Bmi1-deficient mice. Genes Dev 19, 1438–1443 (2005).
Bailey, D. & O’Hare, P. Characterization of the localization and proteolytic activity of the SUMO-specific protease, SENP1. J Biol Chem 279, 692–703 (2004).
Cheng, J. K., Bawa, T., Lee, P., Gong, L. M. & Yeh, E. T. H. Role of desumoylation in the development of prostate cancer. Neoplasia 8, 667–676 (2006).
Bawa-Khalfe, T., Cheng, J., Lin, S. H., Ittmann, M. M. & Yeh, E. T. H. SENP1 Induces Prostatic Intraepithelial Neoplasia through Multiple Mechanisms. J Biol Chem 285, 25859–25866 (2010).
Wang, Q. et al. SUMO-specific protease 1 promotes prostate cancer progression and metastasis. Oncogene 32, 2493–2498 (2013).
Majumder, P. K. et al. A prostatic intraepithelial neoplasia-dependent p27 Kip1 checkpoint induces senescence and inhibits cell proliferation and cancer progression. Cancer Cell 14, 146–155 (2008).
Bartkova, J. et al. Oncogene-induced senescence is part of the tumorigenesis barrier imposed by DNA damage checkpoints. Nature 444, 633–637 (2006).
Yates, K. E., Korbel, G. A., Shtutman, M., Roninson, I. B. & DiMaio, D. Repression of the SUMO-specific protease Senp1 induces p53-dependent premature senescence in normal human fibroblasts. Aging Cell 7, 609–621 (2008).
Cristofalo, V. J., Lorenzini, A., Allen, R. G., Torres, C. & Tresini, M. Replicative senescence: a critical review. Mech Ageing Dev 125, 827–848 (2004).
Li, Z. et al. Structure of a Bmi-1-Ring1B polycomb group ubiquitin ligase complex. J Biol Chem 281, 20643–20649 (2006).
Lee, J. Y. et al. MEL-18 loss mediates estrogen receptor-alpha downregulation and hormone independence. J Clin Invest 125, 1801–1814 (2015).
di Fagagna, F. D. Living on a break: cellular senescence as a DNA-damage response. Nat Rev Cancer 8, 512–522 (2008).
Parrinello, S. et al. Oxygen sensitivity severely limits the replicative lifespan of murine fibroblasts. Nat Cell Biol 5, 741–747 (2003).
Weyemi, U. et al. ROS-generating NADPH oxidase NOX4 is a critical mediator in oncogenic H-Ras-induced DNA damage and subsequent senescence. Oncogene 31, 1117–1129 (2012).
Cai, R. et al. SUMO-specific protease 1 regulates mitochondrial biogenesis through PGC-1alpha. J Biol Chem 287, 44464–44470 (2012).
Cheng, J., Kang, X. L., Zhang, S. & Yeh, E. T. H. SUMO-Specific protease 1 is essential for stabilization of HIF1 alpha during hypoxia. Cell 131, 584–595 (2007).
Chua, K. F. et al. Mammalian SIRT1 limits replicative life span in response to chronic genotoxic stress. Cell Metab 2, 67–76 (2005).
Todaro, G. J. & Green, H. Quantitative studies of the growth of mouse embryo cells in culture and their development into established lines. J Cell Biol 17, 299–313 (1963).
Gong, L., Millas, S., Maul, G. G. & Yeh, E. T. Differential regulation of sentrinized proteins by a novel sentrin-specific protease. J Biol Chem 275, 3355–3359 (2000).
Bracken, A. P. et al. The Polycomb group proteins bind throughout the INK4A-ARF locus and are disassociated in senescent cells. Genes Dev 21, 525–530 (2007).
Acknowledgements
This work was supported by National Natural Science Foundation of China (30900740 to Y.Z.),National Basic Research Program of China (973 Program) (No. 2013CB910902 to J.Cheng), National Natural Science Foundation of China (91229202 and 81430069 to J.Cheng).We thank Dr Ismail for technical suggestion to detect the SUMOylation of Bmi1.
Author information
Authors and Affiliations
Contributions
N.X., J.Cai and F.W. performed the experiments, analyzed the data and prepared the figures. N.X. wrote the main manuscript text. N.X., J.Cai and F.W. made equal contribution to this work. B.D., S.L. and F.C. helped to perform the experiments. B.D. provided the prostatic intraepithelial neoplasm tissue samples. J.Cheng and Y.Z. designed the experiments, helped with data analysis and interpretation, editing and final approval of manuscript. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Xia, N., Cai, J., Wang, F. et al. SENP1 Is a Crucial Regulator for Cell Senescence through DeSUMOylation of Bmi1. Sci Rep 6, 34099 (2016). https://doi.org/10.1038/srep34099
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep34099
This article is cited by
-
SUMO promotes longevity and maintains mitochondrial homeostasis during ageing in Caenorhabditis elegans
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.