Abstract
Whereas studies have extensively examined the ability of bacteria to influence Plasmodium infection in the mosquito, the tripartite interactions between non-entomopathogenic fungi, mosquitoes and Plasmodium parasites remain largely uncharacterized. Here we report the isolation of a common mosquito-associated ascomycete fungus, Penicillium chrysogenum, from the midgut of field-caught Anopheles mosquitoes. Although the presence of Pe. chrysogenum in the Anopheles gambiae midgut does not affect mosquito survival, it renders the mosquito significantly more susceptible to Plasmodium infection through a secreted heat-stable factor. We further provide evidence that the mechanism of the fungus-mediated modulation of mosquito susceptibility to Plasmodium involves an upregulation of the insect’s ornithine decarboxylase gene, which sequesters arginine for polyamine biosynthesis. Arginine plays an important role in the mosquito’s anti-Plasmodium defense as a substrate of nitric oxide production and its availability therefore has a direct impact on the mosquito’s susceptibility to the parasite. While this type of immunomodulatory mechanism has already been demonstrated in other host-pathogen interaction systems, this is the first report of a mosquito-associated fungus that can suppress the mosquito’s innate immune system in a way that would favor Plasmodium infection and possibly malaria transmission.
Similar content being viewed by others
Introduction
Several studies have demonstrated a profound influence of the Anopheles gambiae midgut bacterial microbiota on Plasmodium infection of the mosquito. This influence has been attributed to a bacteria-mediated stimulation of anti-Plasmodium immune defenses and the production of anti-Plasmodium metabolites1,2,3. In comparison to the bacterial microbiota, much less is known about the mosquito-associated fungi, or mycobiome and how these organisms may influence infection with and transmission of, disease-causing pathogens such as Plasmodium. Most studies of mosquito-associated fungi have focused on the entomopathogenic fungi because of their potential for mosquito control4,5. Filamentous fungi and yeasts have been isolated from mosquito midguts and other tissues, but their influence on mosquito-Plasmodium interaction has not been extensively addressed6,7,8,9. A recent study showed that a Anopheles-associated yeast strain produced an antiparasitic toxin10. Fungi produce an extensive range of proteins and secondary metabolites, making them attractive sources for the discovery of novel bioactive molecules with desirable properties11,12. Here we report the isolation of a commonly mosquito-associated ascomycete fungus, Penicillium chrysogenum, from the midgut of field-collected Anopheles mosquitoes. We show that the presence of Pe. chrysogenum in the mosquito midgut does not influence mosquito longevity but instead its susceptibility to the malaria parasite Plasmodium. We further show that the molecular basis of this phenomenon likely involves fungus-mediated upregulation of ornithine decarboxylase (ODC), sequestering L-arginine for polyamine biosynthesis and thereby resulting in diminished production of anti-parasitic nitric oxide (NO).
Results
A Penicillium chrysogenum fungus was isolated from the midgut of Anopheles mosquitoes collected in the Maunabo region of Puerto Rico. Visual inspection of conidial structure by light microscopy, in combination with DNA sequencing of the ribosomal internal transcribed spacer (ITS) region, identified this fungus as a strain of Penicillium chrysogenum13. Pe. chrysogenum is found globally in temperate and subtropical regions and it has been isolated from various mosquito species, including malaria vectors8,9. In fact, over 50% of fungi infecting field-caught Anopheles have been identified as Penicillium8. These results suggest that Pe. chrysogenum may be an opportunistic mosquito-associated fungus. Pe. chrysogenum is generally considered non-pathogenic, although some of its strains are capable of producing harmful mycotoxins14. Since the fungus was isolated from the mosquito’s midgut tissue, it is likely that mosquitoes are acquiring Pe. chrysogenum in nature by feeding on contaminated sugar and water sources. Thus, in the present study we investigated the influence of orally administered Pe. chrysogenum on A. gambiae in the context of mosquito longevity as well as susceptibility to Plasmodium infection and the midgut microbiota.
Pe. chrysogenum does not affect A. gambiae survival
To gain baseline information as to whether the presence of Pe. chrysogenum in the mosquito midgut could influence mosquito physiology in either an adverse or beneficial manner, we investigated its impact on mosquito longevity as a standard fitness parameter. We allowed adult female A. gambiae to sugar-feed on sucrose containing Pe. chrysogenum conidia (2.5 × 108 conidia/mL) for 48 h and then monitored mosquito mortality over a 14-day period. Mosquitoes fed on Pe. chrysogenum-laced sucrose experienced the same mortality level as did the naïve sucrose-fed controls (Fig. 1A). In general, Penicillium fungi are considered non-pathogenic and this result indicates that Pe. chrysogenum is not exerting a major detrimental effect on A. gambiae, corroborating a previous report showing low overall mosquito mortality after exposure to conidia at a concentration four times higher than the one we used15. To investigate the midgut colonization capacity of the fungus, we assayed fungal colony forming units recovered from A. gambiae midguts over the course of the 7 days immediately following feeding on a fungus-laced sucrose solution (Fig. 1B). Pe. chrysogenum reached its highest infection intensity by day 5 and dropped to a significantly lower level at day 7 (P = 0.0001 when compared to day 1).
Impact of Pe. chrysogenum on A. gambiae longevity and fungus colonization efficiency of the mosquito midgut.
(A) Survival of adult female A. gambiae, over 14 days, after feeding on a Pe. chrysogenum-laced sucrose solution for 48 h. Data represent 4 biological replicates (N = 79, P = 0.180), Kaplan-Meier statistical analysis. (B) Pe. chrysogenum colony-forming units (CFU) per midgut at 1, 2, 5 and 7 days after ingestion of fungus via a sucrose solution for 48 h (N = 40 at day 1, 5, 7 and N = 39 at day 2). CFU for Pe. chrysogenum were significantly lower (P = 0.0001) on day 7 than on day 1. Three biological replicates were performed; bars represent standard error of the mean (SEM). ****P ≤ 0.0001; two-tailed Mann-Whitney test.
Midgut colonization of Pe. chrysogenum enhances mosquito susceptibility to Plasmodium infection
To investigate whether the presence of Pe. chrysogenum in the mosquito midgut could influence Plasmodium infection, we fed adult female A. gambiae on a conidia-laced sucrose solution (2.5 × 108 conidia/mL) for 48 h before providing the mosquitoes with an infectious blood meal containing P. falciparum gametocytes. The presence of Pe. chrysogenum in the mosquito midgut resulted in an enhanced (P = 0.0001) P. falciparum infection when compared to naïve sugar-fed controls (Fig. 2A). To investigate whether the Pe. chrysogenum-mediated influence on parasite infection was Plasmodium species-specific, we performed a similar assay with the rodent malaria parasite P. berghei. The presence of Pe. chrysogenum in the mosquito midgut again resulted in a higher (P = 0.0161) P. berghei infection (Fig. 2B) when compared to controls, suggesting that the infection enhancement is parasite species-independent16. In nature, mosquitoes will consume sugar and water from a variety of sources, providing opportunities for fungi and other microorganisms to enter the midgut lumen17,18. It is impossible to exactly reproduce this fashion of fungus acquisition in the laboratory, since the concentrations of Pe. chrysogenum in these different natural habitats and media are unknown. However, to investigate whether the Pe. chrysogenum–dependent modulation of parasite infection is dose-dependent, we performed assays providing various concentrations of the fungus in the sucrose solution prior to P. falciparum infection. We observed a dose-dependent increase in P. falciparum infection using live fungi (Fig. 2C). These results suggest that both the dose of fungi and variance in individual mosquito susceptibility to fungal colonization play a role in determining whether Pe. chrysogenum will modulate Plasmodium susceptibility.
Influence of Pe. chrysogenum on Plasmodium infection.
(A) Influence of Pe. chrysogenum on P. falciparum infection of A. gambiae, as a measured by oocyst numbers 7 days after feeding on a P. falciparum gametocyte culture (infection intensity). The mosquito cohort (N = 58) that had been exposed to a Pe. chrysogenum-laced sucrose solution for 48 h prior to parasite infection had a significantly higher P. falciparum infection (P = 0.0001) than did the non-fungus-exposed control cohort (N = 83). (B) Influence of Pe. chrysogenum on P. berghei infection intensity in A. gambiae. The mosquito cohort (N = 89) that had been exposed to a Pe. chrysogenum-laced sucrose solution for 48 h prior to parasite infection had a significantly higher infection (P = 0.0161) than did the non-fungus-exposed control cohort (N = 104). (C) Influence of various doses of Pe. chrysogenum on P. falciparum infection intensity. Mosquito cohorts that had been exposed to Pe. chrysogenum at 2.5 × 108 conidia/mL (N = 55, P = 0.0137); 2.5 × 106 conidia/mL (N = 33, P = 0.1449), or 2.5 × 104 conidia/mL (N = 40, P = 0.1866) for 48 h prior to parasite infection had a significantly higher level of P. falciparum infection than did the non-fungus-exposed control cohort (N = 45). Graphs show three independent biological replicates. Each dot represents a single midgut and bars represent the median. *P < 0.05, ****P ≤ 0.0001; two-tailed Mann-Whitney test.
Pe. chrysogenum modulates Plasmodium infection through a secreted heat-stable factor
Fungi typically produce a variety of proteins and secondary metabolites17,18,19. To investigate whether Pe. chrysogenum’s influence on Plasmodium infection could be attributed to a secreted factor, we fed mosquitoes on a sucrose solution containing a Pe. chrysogenum culture filtrate (devoid of fungi) for 48 h prior to P. falciparum infection. Feeding on the fungus filtrate resulted in a significant increase in P. falciparum infection intensity (P = 0.0067) when compared to sterile sucrose-fed controls (Fig. 3A). This result suggests that Pe. chrysogenum influences parasite infection of the mosquito midgut through a secreted factor. We next investigated whether this secreted factor was heat-stable. We heat-treated the fungus filtrate at 95 oC for 2 h before providing it to mosquitoes and infecting them with P. falciparum. Surprisingly, we again observed a significant increase in the infection intensity (P = 0.0018) of P. falciparum infection when compared to the sterile sucrose-fed controls (Fig. 3A). These findings suggest that Pe. chrysogenum produces a heat-stable, secreted secondary metabolite, or peptide, that enhances Plasmodium infection in the mosquito.
Influence of Pe. chrysogenum culture filtrate on Plasmodium infection.
Measured by oocyst numbers at 7 days after feeding on a P. falciparum gametocyte culture, the mosquito cohort (N = 109) that had been exposed to a fungus culture filtrate-laced sucrose solution and the cohort (N = 120) that had been exposed to a heat-treated fungus culture filtrate-laced sucrose solution for 48 h prior to parasite infection had significantly higher infection levels (P = 0.0067 for filtrate and P = 0.0018 for heat-treated filtrate) when compared to the non-fungus-exposed control cohort (N = 143). Graph represent three independent biological replicates, each dot represents a single midgut and bars represent the median. **P < 0.01; two-tailed Mann-Whitney test.
Pe. chrysogenum does not affect Plasmodium development in a mosquito-independent fashion
Next, we investigated whether the effect of Pe. chrysogenum on Plasmodium infection could be attributed to a direct interaction between the fungus-secreted molecule(s) and the parasite, operating in a mosquito-independent fashion. We exposed an in vitro ookinete culture of a luciferase–expressing P. berghei parasite to the Pe. chrysogenum conidia (P = 0.5521) or filtrate (P = 0.8321) and monitored ookinete-stage parasite development and viability. The presence of the Pe. chrysogenum or fungus-secreted molecule(s) did not influence parasite development or viability when compared to the non–exposed control parasite culture (Fig. 4A,B). These results suggest that the effect of Pe. chrysogenum on Plasmodium infection is likely mediated through a modulation of mosquito susceptibility rather than parasite development.
Influence of Pe. chrysogenum on in vitro Plasmodium development and ookinete viability.
(A) No difference in in vitro P. berghei ookinete development or viability was observed between Pe. chrysogenum conidia-supplemented ookinete cultures (N = 18, P = 0.5521) and non-Pe. chrysogenum-exposed control cultures (N = 9). (B) No difference in in vitro P. berghei ookinete development or viability was observed between Pe. chrysogenum culture filtrate-supplemented cultures (N = 15, P = 0.8321) and non-supplemented control cultures (N = 12). Values are arbitrary fluorescence; three biological replicates were performed, bars represent standard error of the mean (SEM), two-tailed unpaired t-test.
The mosquito microbiota is not influencing the fungus-mediated modulation of mosquito susceptibility to Plasmodium infection
Penicillium fungi are known to produce antibiotics and our previous studies have shown that antibiotic-mediated suppression of the midgut microbiota enhances Plasmodium infection in Anopheles1,22. To investigate whether the mosquito-associated bacteria could in some way be involved in the fungus-mediated modulation of Plasmodium infection, we suppressed the A. gambiae midgut microbiota by maintaining the mosquitoes on a sucrose solution containing a broad-spectrum antibiotic cocktail, prior to ingestion of the fungus filtrate and subsequent infection with P. falciparum. Ingestion of the Pe. chrysogenum conidia (P = 0.0001) or filtrate (P = 0.0009) still caused a significant increase in P. falciparum infection of antibiotic-treated mosquitoes when compared to the sterile sucrose-fed controls (Fig. 5A,B). These results suggest a bacteria-independent mechanism underlying the observed increase in P. falciparum infection after Pe. chrysogenum exposure.
The influence of Pe. chrysogenum on Plasmodium infection is microbiota-independent.
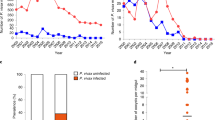
P. falciparum infection intensity, as measured by oocyst numbers at 7 days after feeding on a gametocyte culture, was significantly higher in (A) Pe. chrysogenum conidia-exposed mosquitoes (N = 72, P = 0.0001) and (B) fungus culture filtrate-exposed (N = 75, P = 0.0009) mosquitoes than in their respective bacteria-void, antibiotic-treated non-Pe. chrysogenum-exposed controls (N = 87, N = 59, respectively). Graphs show three independent biological replicates; each dot represents a single midgut and horizontal bars represent the median infection intensity. ***P < 0.001, ****P < 0.0001; two-tailed Mann-Whitney test. (C) Midgut bacterial load, as measured by colony forming units (CFU), was significantly higher in Pe. chrysogenum-exposed mosquitoes (P = 0.0001) than in non-exposed controls (N = 25 for each). Error bars represent the standard error of the mean. ****P ≤ 0.0001; two-tailed Mann-Whitney test. (D) Pe. chrysogenum does not produce anti-bacterial products. Image shows four mosquito-isolated bacteria plates: (a) S. marcences, (b) E. hormaechei, (c) B. subtilis and (d) S. capprae grown with a disk soaked in a fungus culture filtrate-laced sucrose solution (left disc) or a fungus culture filtrate-laced water solution (top disc), as compared to an antibiotic-soaked disc (right disc).
We also investigated the impact of Pe. chrysogenum exposure on the midgut bacterial load by feeding A. gambiae on conidia-laced sucrose solution for 2 days and subsequently enumerating the bacteria using colony forming unit (CFU) assays. Interestingly, the presence of Pe. chrysogenum in the mosquito midgut resulted in a significant increase (P = 0.0001) in the midgut bacterial load (Fig. 5C). We also found that Pe. chrysogenum did not produce any antibiotic compound with antibacterial activity (Fig. 5D). Pe. chrysogenum’s impact on both parasitic and bacterial infection suggests that it involves the suppression of a physiological system of the mosquito that can act against both bacteria and the parasite, such as the innate immune system1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23. Alternatively, the fungus-produced factor could enhance a mosquito physiological system that favors both parasitic and bacterial infection.
A Pe. chrysogenum filtrate induces A. gambiae genes that influence Plasmodium infection
The influence of Pe. chrysogenum on both Plasmodium infection and the midgut microbiota as well as the lack of influence on the in vitro development and viability of ookinete-stage parasites strongly implied the involvement of a physiological system in the mosquito. To gain insight into how the fungus might influence mosquito physiology at the molecular level, we compared the transcriptomes of mosquitoes that had been provided with heat-inactivated fungus filtrate through a sucrose solution for 48 h to a control cohort maintained on a sterile sucrose solution. The experiment showed an overall regulation of a relatively small number of genes that included 32 induced genes belonging to the following functional groups: metabolism (16%), immunity (16%), redox/stress (13%), digestion (13%), cytoskeletal and structural (3%), transport (3%), diverse functions (20%) and unknown function (16%) (Table 1). The gene showing the greatest increase (2.94-fold) in mRNA abundance upon exposure to Pe. chrysogenum filtrate encoded an ornithine decarboxylase (ODC). ODC is an enzyme that mediates polyamine synthesis, using L-arginine (L-Arg) as a substrate.
Because OCD has been shown to modulate infection by pathogens, including Plasmodium infection of mosquitoes24,25, we hypothesized that it was likely to be responsible for mediating the effect of Pe. chrysogenum on Plasmodium infection. Chemical inactivation of OCD by feeding mosquitoes on the inhibitor a-difluoromethylornithine (DFMO) via sucrose solution was previously shown to result in a greater resistance to P. berghei infection26. Another study has shown that Chlamydia trachomatis modulates the immune competence of macrophages by upregulating OCD, which in turn results in L-Arg sequestration, thereby diminishing the production of bacteriocidal nitric oxide25. Furthermore, several studies have shown that feeding mosquitoes on L-Arg after Plasmodium infection results in a greater resistance to the parasite by enhancing nitric oxide-mediated parasite killing24,27,28.
To determine whether ODC influences Plasmodium infection, we silenced the gene using dsRNA-mediated silencing after P. falciparuminfection (Fig. 6A,B). A reduction in ODC transcript abundance by 40% (Fig. 6A) resulted in a significantly greater resistance to Plasmodium infection (P = 0.0419) in the mosquito midgut when compared to GFP dsRNA-treated controls (Fig. 6B). This result indicates an antagonistic effect of OCD on Plasmodium infection, which, in turn, is consistent with the use of L-Arg as its substrate. We hypothesize that an ODC over-activation results in sequestration of L-Arg, which in turn enhances parasite infection by compromising the protective response against P. falciparum. In order to test this hypothesis, we fed mosquitoes with the heat-inactivated fungus filtrate along with L-Arg via a sucrose solution for 48 h prior to Plasmodium infection (Fig. 6C). Consistent with previous studies, mosquitoes treated with L-Arg alone showed a greater resistance to parasite infection (P = 0.0378) than did non-L-Arg-treated controls. L-Arg supplementation negated the agonistic effect of the Pe. chrysogenum filtrate on parasite infection when compared mosquitoes treated with filtrate alone (P = 0.0001). Interestingly, co-feeding mosquitoes on both L-Arg and the fungus filtrate resulted in a greater resistance to infection (P = 0.0225) than did feeding on L-Arg alone. This result may reflect a synergistic effect of greater OCD activity resulting from an unlimited supply of L-Arg, perhaps leading to the production of anti-parasitic metabolites. Our results, taken together with those of previous studies, strongly suggest that Pe. chrysogenum influences Plasmodium infection through a mechanism that involves increased OCD activity and sequestration of L-Arg, which is necessary for NO-mediated Plasmodium killing (Fig. 6D).
Ornithine decarboxylase (ODC) influences susceptibility to P. falciparum infection.
(A) ODC gene silencing efficiency, as measured by qRT-PCR of mRNA abundance in whole mosquitoes; bars represent the standard error of the mean (SEM). (B) P. falciparum infection intensity of OCD gene-silenced mosquitoes (N = 33, P = 0.419) compared to GFP dsRNA-treated control mosquitoes (N = 36). (C) P. falciparum infection intensity of sugar control groups (N = 50), fungus filtrate-exposed mosquitoes (N = 47), L-Arg-fed mosquitoes (N = 50) and fungus filtrate plus L-Arg–exposed mosquitoes (N = 35). Each dot represents a single midgut and bars represent the median infection intensity of three independent experiments, *P < 0.05, ***P < 0.001, ****P ≤ 0.0001; two-tailed Mann-Whitney test. (D) L-Arg is a substrate for polyamine synthesis and nitric oxide (NO) production. Pe. chrysogenum-secreted factors shift the balance toward polyamine synthesis, resulting in decreased parasite killing.
Discussion
Complex tripartite interactions between the Anopheles mosquito’s endogenous microbes, various mosquito physiological systems and Plasmodium dictate the success of parasite infection and malaria transmission1,29. Here, we show that the presence of the natural mosquito-associated fungus, Pe. chrysogenum, in the A. gambiae midgut results in an enhanced Plasmodium infection of the midgut tissue. This fungus-mediated activity is exerted by a secreted heat-stable molecule(s). The fungus-secreted factor does not directly influence parasite development, independently of the mosquito and its impact on infection is independent of the mosquito’s bacterial microbiota. The presence of Pe. chrysogenum in the mosquito gut also results in an increased bacterial load. The influence of the fungus-secreted molecule(s) on both parasite infection and the proliferation of the mosquito midgut bacteria suggest that it likely compromises a mosquito defense mechanism that can act against both Plasmodium and bacteria.
We show that ingestion of the Pe. chrysogenum filtrate results in the upregulation of the ODC gene, which encodes a rate-limiting enzyme of the polyamine biosynthesis pathway. Polyamine induction is known to interfere with NO production by NOS because of L-Arg sequestration30,31. The conversion of L-Arg into L-ornithine by ODC limits the availability of L-Arg for NO production by NOS32. Several studies have linked an increased L-Arg availability to greater mosquito resistance to Plasmodium through the action of NO and NO is also a potent antibacterial factor; furthermore, the NOS gene is strongly induced by both plasmodial and bacterial infection of A. gambiae24,33,34. Chemical inhibition or RNAi-mediated depletion of OCD resulted in a greater resistance to Plasmodium, which is consistent with a greater availability of L-Arg. Accordingly, the effect of the Pe. chrysogenum filtrate on mosquito susceptibility to Plasmodium could be reversed by provision of L-Arg. Our data suggest that a heat-stable Pe. chrysogenum-secreted molecule induces the expression of OCD, which in turn causes diminished NO production by sequestering L-Arg, leading to a greater susceptibility to the Plasmodium parasite and midgut bacteria. Interestingly, an immunomodulatory effect of a Chlamydia bacterium on macrophages has been shown to be mediated by a similar mechanism involving OCD upregulation that, in turn, diminishes NO production and increases bacterial survival35.
These findings may have significant implications for the epidemiology and transmission of Plasmodium by Anopheles mosquitoes in the field. Like many species of Penicillium, Pe. chrysogenum is non-pathogenic to mosquitoes and may therefore not exert strong selective pressure for the development of mosquito resistance. In fact, Penicillium fungi have been found to be abundantly associated with field mosquitoes; also, our work suggests that mosquitoes can acquire the fungi through sugar-feeding and that the fungi will persist in the midgut tissue for up to 5 days7,8,9. Whether the intensity of Pe. chrysogenum exposure in the field is sufficient to shift the mosquito’s susceptibility to Plasmodium is currently unknown and also difficult to assess. Plasmodium infection of mosquitoes in the field is usually much lower than that achieved in the laboratory, averaging between one and three oocysts per midgut. Hence, even a relatively small increase in mosquito susceptibility to the parasite could allow mosquitoes that would not have become infected in the absence of the fungus to become infected and Plasmodium transmission-capable. It will be interesting to survey the midguts of field-caught mosquitoes in malaria-endemic areas for this type of fungi and assess any possible correlation with Plasmodium infection and epidemiological parameters.
Methods
Mosquito Rearing
A. gambiae Keele strain and wild-type A. stephensi Liston strain were maintained on a 10% sucrose solution with 12-h light/dark cycles at 27 °C and 80% humidity.
Genomic DNA Extraction, PCR and DNA Sequencing
Pe. chrysogenum conidia were collected and resuspended in 150 μL of sterile 1x PBS, homogenized (Bullet Blender, Next Advance) with 0.5-mm glass beads for 2 min and the resulting homogenate incubated at 70 °C for 15 min to inactivate residual enzymes and prevent sample degradation. A phenol:chloroform extraction of nucleic acids was performed on this homogenate and DNA yields were quantified following RNase (Qiagen) treatment. Universal fungal primers were used to amplify a region of intertranscribed spacer region of ribosomal 18 s DNA and the PCR products were separated by electrophoresis in 1% agarose gel. The target amplicon was column-purified (Qiagen) and sent to the JHMI Genomics Core for Sanger sequencing and the returned nucleotide sequence was searched against a non-human nucleotide database using BlastN™14. Table S2 lists the primers used.
Culturing of Pe. chrysogenum and Collection of Conidia
Pe. chrysogenum was initially isolated on Yeast Peptone Dextrose Agar (YPDA) (Sigma) and subsequent cultures were maintained on Sabouraud Glucose Agar 4% (SGA) (Sigma). In brief, 4 mL of Sabouraud Dextrose Broth (SDB) was inoculated with Pe. chrysogenum and shaken at 30 °C for up to 72 h. These Pe. chrysogenum cultures were subsequently kept at 4 °C and used to inoculate SGA plates for up to 4 weeks. Using sterile technique, 100 μL of broth was added to SGA plates and Pe. chrysogenum was cultured at 27 °C for 1–2 weeks or until the surface became confluent with conidia. Conidia were collected by flushing plates with 1x PBS containing 0.1% Tween 80 and by gently scraping the conidia into the solution. Glass wool filtration of this solution removed residual agar and mycelium and the filtered conidia were centrifuged at 2,000 rpm for 10 min. The supernatant was discarded and the conidia pellet was washed with 50 mL of sterile 1x PBS. After a second centrifugation, the pellet was resuspended in 1 mL of sterile 1x PBS and serially diluted and a hemocytometer (Neubauer) was used to enumerate conidia by light microscopy (40x). Concentrated conidia were added to 25% glycerol for long-term storage at −80 °C in order to prepare fresh liquid starter cultures.
Preparation of Pe. chrysogenum Filtrate
Four Petri dishes confluent with conidia were used for each preparation of filtrate. After washing and centrifugation as described, the PBS supernatant was discarded and the conidia pellet was resuspended in 1 mL of sterile 10% sucrose solution. This suspension was vortexed briefly for 20 sec and then the conidia were pelleted by centrifugation at 2,000 rpm for 10 min. After centrifugation, the supernatant was collected in a 1-mL syringe and passed through a 0.2-micron filter (Millipore). For heat inactivation, filtrates were incubated at 95 °C for 2 h. A small aliquot of filtrate was cultured on SGA plates to confirm the absence of live fungi.
Sugar-feeding of A. gambiae on Conidia and Filtrate
Conidia were added to sterile 10% sucrose solution to obtain a final volume of 4 mL and the solution was absorbed on cotton and placed in a location accessible to the mosquitoes. For filtrate preparations, approximately 1 mL of either non-treated or heat-inactivated filtrate in a sterile 10% sucrose solution was absorbed on cotton. Mosquitoes were allowed to feed on conidia or filtrate for 48 h. Conidia were administered once, whereas fresh filtrate solutions were administered on days 1 and 2. Forty-eight hours after administering conidia or filtrate, the cotton pads were removed and replaced with sterile 10% sucrose.
A. gambiae Survival Assay
Three- to four-day-old adult female A. gambiae were allowed to feed on Pe. chrysogenum conidia as described above. At day 0, mosquitoes were given sterile 10% sucrose solution that was changed every 2 days. Survival of mosquitoes was monitored for 14 days and dead mosquitoes were removed.
Quantification of Microbiota and Pe. chrysogenum in the Anopheles Midgut
Colony forming units (CFU) from mosquito midguts were quantified in control untreated, fungi-fed and filtrate-fed mosquitoes as described1,36. Three- to four-day-old female A. gambiae were fed fungi or filtrate and then at 48 h post-feeding, the female mosquitoes were collected, surface-sterilized in ethanol and washed with 1x PBS and their midguts were dissected in sterile 1x PBS. Collected midguts were homogenized and serial dilutions of homogenate were added to LB agar plates for bacterial enumeration. After incubation for 2–3 days at 27 °C under aerobic conditions, the CFUs were counted. Pe. chrysogenum CFU from midgut samples were similarly quantified, except that midgut samples were cultured on SGA plates containing 75 μg/mL gentamicin sulfate (Quality Biological) and 100 units/μg per mL of penicillin-streptomycin (Invitrogen). Plates were incubated for up to 4 days at 27 °C and were inspected every day for hyphal nuclei, which were enumerated to determine the fungal CFU/midgut. For the Pe. chrysogenum antibiotic production test, four species of mosquito-isolated bacteria (Serratia marcences, Enterobacter hormaechei, Bacillus subtilis and Staphylococcus sp.) were grown at 30 °C on LB plates containing a disk soaked in a fungus culture filtrate solution and compared to an antibiotic-soaked disc containing 100 units/μg per mL of penicillin-streptomycin.
Antibiotic Treatment
Adult female mosquitoes were given a sterile 10% sucrose solution containing 75 μg/mL gentamicin sulfate and 100 units/μg per mL of penicillin-streptomycin for 3 days. To validate the efficiency of the antibiotic treatment, midguts from control untreated and antibiotic-treated mosquitoes were subjected to CFU assays22. Where indicated, conidia or filtrate solutions were supplemented with antibiotics. Antibiotic treated cohorts were maintained on antibiotic-treated sucrose following a blood meal.
Plasmodium Infection and Oocyst Enumeration
P. falciparum and P. berghei infectiuons were performed following a standard protocol16. For P. falciparum infection: At 48 h post-feeding on fungi or filtrate, mosquitoes were fed on an NF54W strain gametocyte culture mixed with human blood, through a membrane feeder at 37 °C. Engorged mosquitoes were maintained at 27 °C for up to 8 days. For P. berghei infection: At 2 days post-feeding on fungi or filtrate, mosquitoes were fed on Swiss Webster mice infected with the WT Anka 2.34 parasite strain. Engorged mosquitoes were maintained at 19 °C for 10 days. P. falciparum- and P. berghei-infected mosquito midguts were dissected and stained with 0.1% mercurochrome and oocyst numbers were determined using a light microscope.
In vitro P. berghei ookinete culture assays
In vitro P. berghei ookinete culture assays were performed as previously described using parasite infected blood from female Swiss Webster mice (6–8 weeks old) and a genetically modified P. berghei expressing Renilla luciferase under the control of an ookinete-specific promoter37. To assay the influence of Pe. chrysogenum on the ookinete culture, conidia or filtrate was added to each ookinete culture well in triplicate along with experiment-matched controls and incubated at 19 °C for 26–28 h prior to luciferase assay he percentage inhibition of ookinete development was calculated by subtracting the blanks and expressing the luciferase units as a percentage of the control values.
Genome-wide Microarray Analysis
Assays were conducted and analyzed as reported previously using a custom-designed full-genome 8 × 60 K A Agilent-based microarray platform38. In brief, midgut RNA from 20 mosquitoes, which had either been treated with Pe. chrysogenum heat-treated filtrate or left untreated, was purified using the RNeasy kit (Qiagen). Samples were labeled with the Low Input Quick Amp Labeling kit (Agilent Technologies). The array was scanned with an Agilent SureScan microarray scanner. The gene expression data were processed and analyzed as described previously38. Four independent biological replicate assays were performed. Self-self hybridizations have previously been used to determine the cutoff value for the significance of gene regulation to 0.75 in log2 scale16. Given the low number of regulated genes in these assays, we present genes up to a cutoff value of 0.50 in log2 scale, which corresponds to 1.40-fold regulation. Numeric gene expression data are presented in Table 1.
dsRNA-mediated Gene Silencing
The Odc gene was depleted from adult female mosquitoes using established RNAi methodology16. RNAi assays were repeated three times, using GFP dsRNA as a control. Gene silencing was verified at 3 days post-injection using RNA extracted from five whole mosquitoes per biological replicate and two technical replicates by real-time RT-PCR with Sybr Green PCR Master Mix (Applied Biosystems) using the ABI StepOnePlus Real-Time PCR System and ABI StepOne Software. The ribosomal protein S7 gene was used to standardize and verify ODC silencing. The primers used to produce PCR Amplicons for dsRNA synthesis and qRT-PCR are given in Table S2.
L-Arginine Treatment
Mosquitoes were treated for 48 h prior to P. falciparum infection with either Pe. chrysogenum heat-inactivated filtrate, 0.2% L-arginine (L-Arg), or both, all in 10% sucrose solution. After P. falciparum infection, the mosquitoes were maintained with sugar throughout the 8 days of infection incubation, or with 0.2% L-Arg-supplemented sugar in the case of the groups treated previously with L-Arg.
Statistical Analysis
The Graphpad Prism 6 (Graphpad Prism®) software package was used to perform statistical analyses. The particular test used is indicated in the legend of each respective figure. See Table S1 for a summary of the statistics.
Additional Information
How to cite this article: Angleró-Rodríguez, Y. I. et al. A natural Anopheles-associated Penicillium chrysogenum enhances mosquito susceptibility to Plasmodium infection. Sci. Rep. 6, 34084; doi: 10.1038/srep34084 (2016).
References
Dong, Y., Manfredini, F. & Dimopoulos, G. Implication of the mosquito midgut microbiota in the defense against malaria parasites. PLoS Pathog. 5 (2009).
Dennison, N. J., Jupatanakul, N. & Dimopoulos, G. The mosquito microbiota influences vector competence for human pathogens. Curr. Opin. Insect Sci. 3, 6–13 (2014).
Saraiva, R. G., Kang, S., Simões, M. L., Angleró-Rodríguez, Y. I. & Dimopoulos, G. Mosquito gut antiparasitic and antiviral immunity. Dev. Comp. Immunol. 64, 53–64 (2015).
Fang, W. et al. Development of transgenic fungi that kill human malaria parasites in mosquitoes. Science 331, 1074–1077 (2011).
Bukhari, T., Takken, W. & Koenraadt, C. J. M. Development of Metarhizium anisopliae and Beauveria bassiana formulations for control of malaria mosquito larvae. Parasit. Vectors 4, 23 (2011).
Frants, T. G. & Mertvetsova, O. A. [Yeast associations with mosquitoes of the genus Aedes Mg. (Diptera, Culicidae) in the Tom-Ob river region]. Nauchnye Doki. Vyss. Shkoly. Biol. Nauki 94–98 (1986).
Ricci, I. et al. The yeast Wickerhamomyces anomalus (Pichia anomala) inhabits the midgut and reproductive system of the Asian malaria vector Anopheles stephensi. Environ. Microbiol. 13, 911–921 (2011).
Da Costa, G. L. & De Oliveira, P. C. Penicillium species in mosquitoes from two Brazilian regions. J. Basic Microbiol. 38, 343–347 (1998).
da S Pereira, E., de M Sarquis, M. I., Ferreira-Keppler, R. L., Hamada, N. & Alencar, Y. B. Filamentous fungi associated with mosquito larvae (Diptera: Culicidae) in municipalities of the Brazilian Amazon. Neotrop. Entomol. 38, 352–359 (2009).
Valzano, M. et al. A yeast strain associated to Anopheles mosquitoes produces a toxin able to kill malaria parasites. Malar. J. 15, 21 (2016).
Liu, B.-H. et al. The fungal metabolite, citrinin, inhibits lipopolysaccharide/interferon-γ-induced nitric oxide production in glomerular mesangial cells. Int. Immunopharmacol. 10, 1608–1615 (2010).
Schueffler, A. & Anke, T. Fungal natural products in research and development. Nat. Prod. Rep. 31, 1425–1448 (2014).
Schoch, C. L. et al. Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc. Natl. Acad. Sci. USA. 109, 6241–6246 (2012).
Hidalgo, P. I. et al. Molecular characterization of the PR-toxin gene cluster in Penicillium roqueforti and Penicillium chrysogenum: cross talk of secondary metabolite pathways. Fungal Genet. Biol. 62, 11–24 (2014).
Maketon, M., Amnuaykanjanasin, A. & Kaysorngup, A. A rapid knockdown effect of Penicillium citrinum for control of the mosquito Culex quinquefasciatus in Thailand. World J. Microbiol. Biotechnol. 30, 727–736 (2014).
Dong, Y. et al. Anopheles gambiae immune responses to human and rodent Plasmodium parasite species. PLoS Pathog. 2, 0513–0525 (2006).
Müller, G. C., Xue, R.-D. & Schlein, Y. Differential attraction of Aedes albopictus in the field to flowers, fruits and honeydew. Acta Trop. 118, 45–49 (2011).
Impoinvil, D. E. et al. Feeding and survival of the malaria vector Anopheles gambiae on plants growing in Kenya. Med. Vet. Entomol. 18, 108–115 (2004).
Fitzpatrick, L. R., Wang, J. & Le, T. Gliotoxin, an inhibitor of nuclear factor-kappa B, attenuates peptidoglycan-polysaccharide-induced colitis in rats. Inflamm. Bowel Dis. 8, 159–167 (2002).
Wätjen, W., Debbab, A., Hohlfeld, A., Chovolou, Y. & Proksch, P. The mycotoxin beauvericin induces apoptotic cell death in H4IIE hepatoma cells accompanied by an inhibition of NF-κB-activity and modulation of MAP-kinases. Toxicol. Lett. 231, 9–16 (2014).
Wong, V. L., Ellison, C. E., Eisen, M. B., Pachter, L. & Brem, R. B. Structural variation among wild and industrial strains of Penicillium chrysogenum. PLoS One 9, e96784 (2014).
Blumberg, B. J., Trop, S., Das, S. & Dimopoulos, G. Bacteria- and IMD pathway-independent immune defenses against Plasmodium falciparum in Anopheles gambiae. PLoS One 8, e72130 (2013).
Bahia, A. C. et al. Exploring Anopheles gut bacteria for Plasmodium blocking activity. Environ. Microbiol. 16, 2980–2994 (2014).
Luckhart, S., Vodovotz, Y., Cui, L. & Rosenberg, R. The mosquito Anopheles stephensi limits malaria parasite development with inducible synthesis of nitric oxide. Proc. Natl. Acad. Sci. USA. 95, 5700–5705 (1998).
Abu-Lubad, M., Meyer, T. F. & Al-Zeer, M. a. Chlamydia trachomatis inhibits inducible NO synthase in human mesenchymal stem cells by stimulating polyamine synthesis. J. Immunol. 193, 2941–2951 (2014).
Gillet, J. M., Charlier, J., Boné, G. & Mulamba, P. L. Plasmodium berghei: Inhibition of the sporogonous cycle by α-difluoromethylornithine. Exp. Parasitol. 56, 190–193 (1983).
Zheng, L., Pan, Y., Feng, Y., Cui, L. & Cao, Y. L-Arginine supplementation in mice enhances NO production in spleen cells and inhibits Plasmodium yoelii transmission in mosquitoes. Parasit. Vectors 8, 326 (2015).
Motard, a. et al. The role of reactive nitrogen intermediates in modulation of gametocyte infectivity of rodent malaria parasites. Parasite Immunol. 15, 21–26 (1993).
Cirimotich, C. M., Dong, Y., Garver, L. S., Sim, S. & Dimopoulos, G. Mosquito immune defenses against Plasmodium infection. Dev. Comp. Immunol. 34, 387–395 (2010).
Szabó, C., Southan, G. J., Thiemermann, C. & Vane, J. R. The mechanism of the inhibitory effect of polyamines on the induction of nitric oxide synthase: role of aldehyde metabolites. Br. J. Pharmacol. 113, 757–766 (1994).
Chaturvedi, R. et al. Polyamines impair immunity to helicobacter pylori by inhibiting L-Arginine uptake required for nitric oxide production. Gastroenterology 139, 1686–1698 (2010).
Li, H. et al. Regulatory role of arginase I and II in nitric oxide, polyamine and proline syntheses in endothelial cells. Am. J. Physiol. Endocrinol. Metab. 280, E75–E82 (2001).
Hillyer, J. F. & Estevez-Lao, T. Y. Nitric oxide is an essential component of the hemocyte-mediated mosquito immune response against bacteria. Dev. Comp. Immunol. 34, 141–149 (2010).
Gupta, L. et al. The STAT Pathway Mediates Late-Phase Immunity against Plasmodium in the Mosquito Anopheles gambiae. Cell Host Microbe 5, 498–507 (2009).
Abu-Lubad, M., Meyer, T. F. & Al-Zeer, M. a. Chlamydia trachomatis inhibits inducible NO synthase in human mesenchymal stem cells by stimulating polyamine synthesis. J. Immunol. 193, 2941–2951 (2014).
Dong, Y., Taylor, H. E. & Dimopoulos, G. AgDscam, a hypervariable immunoglobulin domain-containing receptor of the Anopheles gambiae innate immune system. PLoS Biol. 4, e229 (2006).
Ramirez, J. L. et al. Chromobacterium Csp_P Reduces Malaria and Dengue Infection in Vector Mosquitoes and Has Entomopathogenic and In Vitro Anti-pathogen Activities. PLoS Pathog. 10, e1004398 (2014).
Sim, S. et al. Transcriptomic profiling of diverse Aedes aegypti strains reveals increased basal-level immune activation in dengue virus-refractory populations and identifies novel virus-vector molecular interactions. PLoS Negl. Trop. Dis. 7, e2295 (2013).
Acknowledgements
This work was supported by the National Institutes of Health/National Institute of Allergy and Infectious Disease R01AI061576, R01AI101431, R01AI081877 and a JHMRI pre-doctoral fellowship (to A.P.) and a UNCF-Merck Science Initiative Graduate Research Fellowship (to AMC). We would like to thank the Johns Hopkins Malaria Research Institute Insectary and Parasitology core facilities. We also thank Dr. Deborah McClellan for editorial assistance.
Author information
Authors and Affiliations
Contributions
Y.I.A.R., B.J.B. and G.D. conceived experiments. Y.I.A.R., B.J.B., Y.D., S.L.S., A.P. and A.M.C. performed experiments. Y.I.A.R., B.J.B., Y.D. and G.D. analyzed data obtained. Y.I.A.R., B.J.B. and G.D. wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Angleró-Rodríguez, Y., Blumberg, B., Dong, Y. et al. A natural Anopheles-associated Penicillium chrysogenum enhances mosquito susceptibility to Plasmodium infection. Sci Rep 6, 34084 (2016). https://doi.org/10.1038/srep34084
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep34084
This article is cited by
-
Interkingdom interactions shape the fungal microbiome of mosquitoes
Animal Microbiome (2024)
-
Holobiont perspectives on tripartite interactions among microbiota, mosquitoes, and pathogens
The ISME Journal (2023)
-
Mutualism promotes insect fitness by fungal nutrient compensation and facilitates fungus propagation by mediating insect oviposition preference
The ISME Journal (2022)
-
An Overview on the Impact of Microbiota on Malaria Transmission and Severity: Plasmodium–Vector–Host Axis
Acta Parasitologica (2022)
-
Effect of seasonal malaria chemoprevention plus azithromycin on Plasmodium falciparum transmission: gametocyte infectivity and mosquito fitness
Malaria Journal (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.