Abstract
Basmati rice is preferred internationally because of its appealing taste, mouth feel and aroma. Pusa Basmati 1121 (PB1121) is a widely grown variety known for its excellent grain and cooking quality in the international and domestic market. It contributes approximately USD 3 billion to India’s forex earning annually by being the most traded variety. However, PB1121 is highly susceptible to bacterial blight (BB) disease. A novel BB resistance gene Xa38 was incorporated in PB1121 from donor parent PR114-Xa38 using a modified marker-assisted backcross breeding (MABB) scheme. Phenotypic selection prior to background selection was instrumental in identifying the novel recombinants with maximum recovery of recurrent parent phenome. The strategy was effective in delimiting the linkage drag to <0.5 mb upstream and <1.9 mb downstream of Xa38 with recurrent parent genome recovery upto 96.9% in the developed NILs. The NILs of PB1121 carrying Xa38 were compared with PB1121 NILs carrying xa13 + Xa21 (developed earlier in our lab) for their resistance to BB. Both NILs showed resistance against the Xoo races 1, 2, 3 and 6. Additionally, Xa38 also resisted Xoo race 5 to which xa13 + Xa21 was susceptible. The PB1121 NILs carrying Xa38 gene will provide effective control of BB in the Basmati growing region.
Similar content being viewed by others
Introduction
Harmonious combination of cooking quality characteristics and pleasant aroma is the uniqueness of Basmati rice. Intensive program on genetic improvement of Basmati rice has led to the development of a landmark rice variety, Pusa Basmati 1121 (PB1121), at ICAR-Indian Agricultural Research Institute (IARI), New Delhi. This is a medium duration high-yielding Basmati rice variety characterized by extra-long slender grains with very high kernel length after cooking (22–25 mm), high cooked kernel elongation ratio (2.5), intermediate amylose content, high volume expansion after cooking (>4 times) and strong aroma1. These unique properties of PB1121 have attracted consumers worldwide. PB1121 currently occupies 1.35 million hectares, which is nearly 70% of the area under Basmati cultivation in India and contributes approximately 4 million tons to Basmati rice production annually. With the recognition of PB1121 as Basmati during 2008, the forex earning of the country has risen from USD 0.67 million to USD 4.5 billion in 2014–2015 2, in which the contribution of PB1121 is about 65%. This variety has not only revolutionized the international Basmati rice trade but also improved the livelihood of millions of Basmati farmers. However, major weakness of PB1121 is its susceptibility to a number of diseases such as to bacterial blight (BB)3, blast3 and bakanae. The BB disease is caused by a gram-negative bacteria, Xanthomonas oryzae pv. oryzae (Xoo), which severely constrains rice productivity and quality. Xoo breaches the plant epidermis and multiplies in the xylem vessels which lead to disease symptoms. The pathogen produces various virulence factors including EPS, extracellular enzyme, iron chelating siderophores and the type III-secretion dependent effectors, which are collectively essential for virulence4,5,6,7,8 and to counterattack plant’s immune response termed as PAMP-triggered immunity9. Xoo populations are highly diversed in terms of mode of action through virulence factors10,11,12,13,14,15. Mondal et al.15 characterized 780 Xoo isolates collected from thirteen states of India and grouped them into six races based on their reaction to 12 near isogenic lines (NILs). To date, 30 races of Xoo have been documented globally16,17,18.
Utilizing the host plant resistance is an effective means to manage this pathogen in rice. As of now, 41 BB resistance genes have been reported among which 8 have been cloned and characterized3,19,20. However, four genes namely, Xa4, Xa8, xa13 and Xa21 have been reported to be effective against the Xoo isolates prevalent in Basmati growing regions of India21. Furthermore, the combination of genes xa13 + Xa21, xa5 + xa13 + Xa21 and Xa4 + xa5 + xa13 + Xa21 were reported to be equally effective and has provided greater degree of resistance as compared to monogenic lines. Therefore, xa13 + Xa21 have been widely used in the Basmati rice improvement program which resulted in development of several BB resistant genotypes such as Improved Pusa Basmati 1 22,23, Pusa 1718 and Pusa 1728 3. Extensive utilization of these genes over a long period of time, may lead to resurgence of pathogen with virulence genes that can overcome this combination. Therefore, deployment of diverse genes is essential to prolong the durability of resistance genes and to counter the development of virulent races of the pathogen.
Recently, a novel BB resistance gene Xa38 was identified in Oryza nivara and mapped on to the long arm of chromosome 4 24. It was further fine mapped to a region of 38.4 kb and an InDel marker based on the putative candidate gene LOC_Os04gg53050-1 was developed, which co-segregated with the BB resistance25. It is expected that deployment of Xa38 in rice varieties may help in checking the breakdown of resistance to Xoo. Therefore, the present study was undertaken to transfer Xa38 into the elite Basmati rice variety PB1121 through marker assisted backcross breeding (MABB) and to study its resistance spectrum in comparison to the combination of genes xa13 + Xa21 in the genetic background of PB1121.
Materials and Methods
Marker assisted breeding scheme
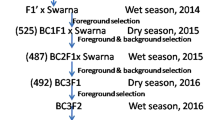
PR114-Xa38, an introgression line generated from a cross PR114/Oryza nivara carrying the BB resistance gene Xa38 24 was used as the donor parent and PB1121 was used as the recurrent parent in a MABB (Fig. 1). Individual F1 plant generated from the cross PB1121/PR114-Xa38 (designated as Pusa 1927), confirmed for hybridity using the Xa38 linked marker-Os04g53050-1 was backcrossed with the recurrent parent PB1121 to generate BC1F1 seeds. Foreground selection followed by phenotypic and background selection was carried out to identify the plants with maximum recovery for recurrent parent phenome (RPP) and genome (RPG). The best BC1F1 plant was further backcrossed to generate BC2F1 seeds. The BC2F1 plants were also subjected to foreground, phenotypic and background selection to identify the plant with maximum recovery for RPG and RPP. The superior plant was advanced to BC2F2 generation and foreground selection was repeated in order to identify the plants homozygous for Xa38. Further, these plants were advanced via pedigree based phenotypic selection to obtain Xa38 introgressed near isogenic lines (NILs) of PB1121.
Molecular Marker Analysis
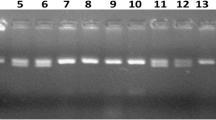
DNA from each of the leaf sample was extracted using the modified protocol of Murray and Thompson26. PCR reaction was carried out using 20 ng of template DNA, 1x PCR buffer [10 mM Tris–HCl (pH 8.4); 50 mMKCl], 5 pmol of each primer, 0.05 mM dNTPs, 1.8 mM MgCl2 and 1U of Taq DNA polymerase (Invitrogen, Life Technologies, Brazil). The PCR was carried out in a G-Storm thermal cycler with the following program: 1) initial denaturation at 94 °C for 5 min; 2) 35 cycles for denaturation for 30 s at 94 °C, annealing for 30 s at 56 °C, extension for 1 min at 72 °C; and 3) final extension at 72 °C for 7 min. The amplified product was resolved in a gel electrophoresis using 3.5% MetaphorTM Agarose gel and visualized on GelDocTM XR (Bio-Rad Laboratories Inc., USA).
Foreground selection was carried out using the Xa38 linked molecular marker-Os04g53050-1 25. For background selection, 628 genome-wide SSR markers were used in parental polymorphism survey between the parental lines PB1121 and PR114-Xa38, and a total of 70 polymorphic markers spanning uniformly across the genome were selected. Recombinant selection was carried out using the markers RM5511 and RM17523 flanking the BB resistance gene Xa38 on chromosome 4.
Screening for resistance to BB
The NILs, along with the checks PB1121 and PR114-Xa38, were grown in field conditions till maximum tillering stage. Different races of Xoo maintained as single spore culture was used to generate the BB suspension with a density of 109 cells/mL. Top five leaves from each of the entry were inoculated with Xoo isolate through the clip inoculation method27. After 21 days of inoculation, the BB lesion length was measured using the disease assessment scheme as adopted by Mondal et al.14, wherein, lesion length of <5 cm was considered resistant, 5–10 cm was considered moderately resistant, 10–15 cm was considered moderately susceptible and more than 15 cm was considered highly susceptible reaction. Additionally, the resistance spectrum of Pusa1927 (PB1121 + Xa38) was compared with Pusa1718 (PB1121 + xa13 + Xa21) using a set of six races of Xoo.
Agronomic and Cooking Quality Evaluation
The NILs, along with the recurrent and donor parents, were evaluated for agronomic traits in randomized complete block design (RCBD) with three replications. Standard agronomic management practices were followed for raising the rice crop at ICAR-IARI, New Delhi. The data was recorded on five plants from each of the entries for the characters namely: days to 50% flowering (DFF), plant height (PH), panicle length (PL), filled grains per panicle (FGP), spikelet fertility (SF), thousand grain weight (TGW) and grain yield (GY). Further, the lines were also analyzed for grain and cooking quality parameters such as kernel length before cooking (KLBC), kernel breadth before cooking (KBBC), length/breadth ratio (L/B), kernel length after cooking (KLAC), kernel breadth after cooking (KBAC) and aroma as described by Khanna et al.28.
Results
Development of PB1121-NILs carrying BB resistance gene Xa38
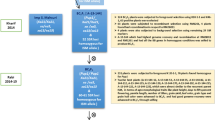
The F1s generated from the cross PB1121/PR114-Xa38 were tested for hybridity using the marker-Os04g53050-1. A single plant confirmed for hybridity was backcrossed to recurrent parent and a total of 468 BC1F1 seeds were produced, out of which 223 plants were found to be heterozygous for Xa38. All the heterozygous plants were subjected to phenotypic evaluation for agro-morphological traits. Ten superior plants, possessing relatively superior recovery for RPP were selected. The phenotypic selection for key Basmati traits such as KLAC and KBAC resulted in positive selection differential of 0.32 mm and 1.56 mm, respectively. Further, the background selection resulted in RPG recovery ranging from 70.6% to 79.29%. The best BC1F1 plant with RPG recovery of 79.29% was selected for further backcrossing with the recurrent parent PB1121 and a total of 198 BC2F1 seeds were produced. Foreground selection revealed that a total of 100 plants were heterozygous for Xa38, which were subjected to phenotypic selection for agronomic, grain and cooking quality traits. The selection differential was positive with a gain of 0.12 mm for KLAC and 0.36 mm for KBAC in the selected top ten superior BC2F1 plants. These plants when subjected to background selection using 29 markers which were heterozygous in the BC1F1 generation; the RPG recovery ranged from 83.6% to 88.6%.
The superior BC2F1 plant was selfed to produce 243 BC2F2 plants where 65 plants were homozygous for Xa38. Based on grain and cooking quality parameters, the selection differential was 0.29 mm in KLAC and 0.15 mm in KLBC. Furthermore, background selection with 16 markers remained heterozygous in BC2F1 generation, and the RPG recovery increased to the range of 88.57% to 92.86%. Finally, best plant was selected and advanced to BC2F3 generation. In ensuing BC2F3 generation, after phenotypic selection for the key Basmati traits KLAC and KLBC, the selection differential was 0.37 mm and 0.02 mm, respectively. The corresponding RPG recovery in this generation ranged from 93.18 to 94.70%. Repeating the same process in the next generation, BC2F4, the selection differential observed due to selection was 0.45 mm for KLAC and no gain observed for KBAC. The aroma score did not show significant variation between generations from that of PB1121 except in backcross generations (BC1F1 and BC2F1). The linkage drag was delimited to the region of <0.5 mb upstream and <1.9 mb downstream the BB resistance gene Xa38 (Fig. 2). Finally, ten best BC2F5 families were isolated and evaluated for agronomic and cooking quality characters. The number of plants generated during each of the backcross generations, the selection differential obtained due to phenotypic selection and the background recovery realized are presented in Table 1.
Evaluation of NILs for Agronomic and Cooking Quality Performance
All the NILs tested were found to be at par with the recurrent parent PB1121 with some exceptions such as NIL Pusa1927-17-16 which was found to mature 2 days later with slightly taller stature but possessed higher spikelet fertility than that of PB1121 (Table 2). Interestingly, six NILs, namely Pusa1927-19-21, Pusa1927-14-77, Pusa1927-8-86, Pusa1927-13-55, Pusa1927-45-72 and Pusa1927-62-2, were found to possess significantly higher spikelet fertility as compared to PB1121. The NIL Pusa1927-8-86 was found to be significantly superior in yield as compared to PB1121 owing to its better spikelet fertility. With respect to grain and cooking quality, all NILs were at par with the recurrent parent PB1121 (Table 3). However, three NILs Pusa1927-19-21, Pusa1927-14-77 and Pusa1927-17-16 were found to be significantly inferior, while NIL Pusa1927-75-56 was significantly superior as compared to the recurrent parent PB 1121 for the trait kernel length after cooking (Table 3 and Fig. 3).
Comparative Analysis of Resistance Spectrum of BB Resistance Genes, xa13 + Xa21 and Xa38
The resistance imparted by the BB resistance genes, xa13 + Xa21 and Xa38 was compared by subjecting the NILs Pusa1927 (PB1121 + Xa38) and Pusa1718 (PB1121 + xa13 + Xa21) against six different Xoo isolates. The disease reaction observed in the recurrent parent PB1121, Pusa1927 (PB1121 + Xa38) and Pusa1718 (PB1121 + xa13 + Xa21) is presented in Table 4 and Fig. 4.
The recurrent parent PB1121 was found to be highly susceptible to all the six Xoo races, while Pusa1927 (PB1121 + Xa38) and Pusa 1718 (PB1121 + xa13 + Xa21) were found to be highly resistant against Xoo races 1, 2, 3 and 6. However, for race 5, Pusa1718 (PB1121 + xa13 + Xa21) was found to be moderately resistant to moderately susceptible with the lesion length ranging from 9.37 ± 0.26 cm to 10.30 ± 0.55 cm, while NIL Pusa1927 (PB1121 + Xa38) was highly resistant with the lesion length ranging from 0.77 ± 0.15 cm to 3.83 ± 0.64 cm. However, both Pusa1718 and Pusa1927 were found to be susceptible against race 4 of Xoo.
Discussion
Basmati is the specialty rice of India, acclaimed worldwide for its unprecedented cooking quality characters and appealing aroma. Being susceptible to most of the diseases and pests including BB disease, the most popular Basmati cultivar in India, PB1121, which enjoys an export contribution of about 60–70%, often entreats agro-management practices that include use of chemical pesticides. Clean production of Basmati rice under eco-friendly environments is a prime target for producing export standard rice. Arming the cultivars with in-built resistance is therefore inevitable in contemporary Basmati breeding. Several genes conforming resistance to diseases such as BB and blast have already been deployed in Basmati cultivars3,22,28.
In the current study, PB1121 was incorporated with the novel BB resistance gene Xa38 using the donor parent PR114-Xa38, an introgression line developed by crossing PR114 an elite rice cultivar of Punjab, India with a wild rice (O. nivara) acc. IRGC81825 25. PR114 is a semi-dwarf, stiff strawed variety narrow, dark green erect leaves and possess extra-long, clear translucent non-aromatic grains29. However, the grain and cooking quality of PR114 and PR114-Xa38 is quite inferior as compared to the recurrent parent PB1121. Using poor grain quality donor parent in Basmati rice breeding impairs the grain and cooking quality of derived line, which imposes a challenge of recovering the quality of recurrent parent. Therefore, maximization of RPG and RPP recovery in the derived NILs is important28.
The recovery of recurrent parent genome to the extent of 96.9% was achieved with two backcrosses through integration of foreground, phenotypic and background selection. Background selection using the SSR markers usually target the non-coding and heterochromatic regions and therefore may not quantify the recovery of functional part of the genome. However, the phenotypic selection, which indirectly targets functionally expressed region of the genome, was augmented for hastening the process of reconstruction of recurrent parent phenotype. It was interesting to note that the selection differential for the trait KLAC was progressive along the generations and the recombinants at par or superior to recurrent parent PB1121, could be obtained by BC2F4 generation. For the trait KLBC, the selection differential was positive till BC2F3 generation only; whereas, in case of aroma, the trait fixation was achieved in BC2F1 generation. The varying selection differential for different traits can be attributed to the number of genes governing them. The trait KLAC and KLBC are under polygenic control governed by additive, dominance and epistatic gene action. Hence, progressive response to selection in later generation was obtained. Similar phenotypic gain was obtained for other agronomic traits also (data not presented). Aroma is a key Basmati trait, which is mainly governed by two recessive genes badh1and badh2 30,31. Therefore, fixation for this trait was possible in the early backcross generations itself. The NILs carrying Xa38 gene, were agronomically similar to their recurrent parent PB1121 in most of the cases (Tables 2 and 3). Phenotypic selection prior to background selection has led to isolating the novel recombinant Pusa1927-75-56, which was similar to the recurrent parent but possessed significantly superior KLAC. The success of integrating phenotypic selection along with background selection in the MABB program for isolating superior recombinants and substantially reducing the number of plants for background selection has been demonstrated earlier by Ellur et al.3. In cases where limited backcrosses are given, NILs derived from a common backcross program differ with respect to their agronomic performance, grain and cooking quality traits because of the presence of minor donor segments on different chromosomes, which offer great opportunity to select NILs excelling the recurrent parent3,32. It is pertinent to mention that stringent selection for grain quality has resulted in near complete recovery of PB1121 grain quality traits in most of the NILs developed.
The efficacy of phenotypic selection in hastening the recovery of recurrent parent phenotype has been earlier demonstrated while developing Improved Pusa Basmati 1 carrying BB resistance genes xa13 + Xa2121,22 and Pusa 1609 carrying blast resistance genes32,33. In the current study, the recombinant selection on carrier chromosome resulted in delimiting the linkage drag to <0.5 mb upstream and <1.9 mb downstream of the Xa38 gene, which otherwise could have been a limiting step because of the donor parent PR114-Xa38, being a non –Basmati variety.
On artificial screening, when all the derived NILs of PB1121 were exposed to two virulent BB isolates along with their parents, the donor PR114-Xa38 and the PB1121 NILs produced incompatibility reaction, as against PB1121 that was highly susceptible. The BB isolates, IARI-Kaul and IARI-Ludhiana were sourced from Haryana and Punjab respectively, two major Basmati rice growing states of India. Both isolates were the most predominant pathogenic variants of BB in respective states3. The resistance imparted by Xa38 gene implied that the gene may be effective against pathogenic races available in the major Basmati growing regions of India. Effectiveness of Xa38 against all common pathogenic variants of Xoo in Punjab has been earlier reported by Vikal et al.34.
In a recent study, Mondal et al.14 grouped the Xoo isolates of India into six different races based on their reaction against 12 monogenic-NILs carrying different BB resistance genes. However, these monogenic lines did not include Xa38 gene. The combination of BB resistance genes, xa13 + Xa21 has been considered as one of the best combinations that can combat most of the virulent races of the Xoo. To establish the spectrum of BB resistance of Xa38 gene and to assess its efficacy in relation to xa13 + Xa21 combination, a set of Xa38 introgressed NILs of PB1121 were inoculated with the six Xoo races along with the recurrent parent PB1121. The BB resistance gene Xa38, as well as xa13 + Xa21, were effective against the Xoo races 1, 2, 3 and 6. Races 1 and 2 are predominantly distributed in the eastern India; race 3 is prevalent in Haryana; while race 6 is predominant in entire northwestern region. Race 5 reported to be sporadically distributed across India could not infect the NILs with Xa38 while it was virulent against the PB1121 NILs carrying xa13 + Xa21. Therefore, deployment of Xa38 into the genetic background of elite rice varieties is essential to effectively combat the Xoo races 1, 2, 3, 5 and 6. On the other hand, Xoo race 4, reported to be present in eastern and north-western states of India, was the most virulent race that could knock down xa13 + Xa21 combination as well as Xa38. Thus, further efforts are required to identify novel BB resistance genes or gene combinations that can provide resistance against the race 4 of Xoo.
To conclude, the study reports successful introgression of a highly effective resistance gene Xa38 into the genetic background of an elite Basmati rice variety PB1121 using a modified-MABB. The improved lines are now under evaluation in the National Basmati Trials for subsequent release. The study also highlights the importance of comparing the spectrum of resistance genes either singly or in combination against a wide range of pathogenic races, so that effective deployment of resistance genes can be planned to avoid future outbreak of disease.
Additional Information
How to cite this article: Ellur, R. K. et al. Marker-aided Incorporation of Xa38, a Novel Bacterial Blight Resistance Gene, in PB1121 and Comparison of its Resistance Spectrum with xa13 + Xa21. Sci. Rep. 6, 29188; doi: 10.1038/srep29188 (2016).
References
Singh, V. P., Singh, A. K., Atwal, S. S., Joseph, M. & Mohapatra, T. Pusa 1121: a rice line with exceptionally high cooked kernel elongation and basmati quality. International Rice Research Notes, 27(1), 25–26 (2002).
APEDA, India export of agro food products: Product group report/country wise—Basmati rice. Accessed online from: http://agriexchange.apeda.gov.inon 25 March 2016.
Ellur, R. K. et al. Improvement of Basmati rice varieties for resistance to blast and bacterial blight diseases using marker assisted backcross breeding. Plant Sci. 242, 330–341 (2016).
Jones, J. D. & Dangl, J. L. The plant immune system. Nature 444, 323–329 (2006).
Nurnberger, T., Brunner, F., Kemmerling, B. & Piater, L. Innate immunity in plants and animals: striking similarities and obvious differences. Immunol. Rev. 198, 249–66 (2004).
Alfano, J. R. & Collmer, A. Type III secretion system effector proteins: double agents in bacterial disease and plant defense. Annu. Rev. Phytopathol. 42, 385–414 (2004).
White, F. F. & Yang, B. Host and pathogen factors controlling the rice Xanthomonas oryzae interaction. Plant Physiol. 150, 1677–86 (2009).
Mudgett, M. B. New insights to the function of phytopathogenic bacterial TypeIII effectors in plants. Annu. Rev. Plant Biol. 56, 509–31 (2005).
Park, C. & Ronald, P. C. Cleavage and nuclear localization of the rice XA21 immune receptor. Nat. Commun. 3, 920, doi: 10.1038/ncomms1932 (2012).
Song, C. & Yang, B. Mutagenesis of 18 Type III effectors reveals virulence function of XopZpxo99 in Xanthomonas oryzae pv. oryzae . Mol. Plant Microbe Interact. 23, 893–902 (2010).
Yang, B. & White, F. F. Diverse members of the AvrBs3/PthA family of type III effectors are major virulence determinants in bacterial blight disease of rice. Mol. Plant Microbe Interact. 17, 1192–1200 (2004).
Yang, B., Sugio, A. & White, F. F. Avoidance of host recognition by alterations in the repetitive and C-terminal regions of AvrXa7, a type III effector of Xanthomonas oryzae pv. oryzae . Mol. Plant Microbe Interact. 18, 142–149 (2005).
Yang, B., Sugio, A. & White, F. F. (2006) Os8N3 is a host disease-susceptibility gene for bacterial blight of rice. PNAS. 103(27), 10503–8 (2006).
White, F. F., Potnis, N., Jones, J. B. & Koebnik, R. The type III effectors of Xanthomonas . Mol. Plant Pathol. 10(6), 749–66 (2009).
Mondal, K. K. et al. Pathotyping and genetic screening of type III effectors in Indian strains of Xanthomonas oryzae pv. oryzae causing bacterial leaf blight of rice. Physiol. and Mol. Plant Pathol. 86, 98–106 (2014).
Adhikari, T. B., Basnyat, R. C. & Mew, T. W. Virulence of Xanthomonas oryzae pv. oryzae on rice lines containing single resistance genes and gene combinations. Plant Dis. 83, 46–50 (1999).
Shanti, M. L. et al. Understanding the Bacterial blight pathogen- combining pathotyping and molecular marker studies. Int. J. Plant Pathol. 1 (2), 58–68 (2010).
Mishra, D. et al. Pathotype and genetic diversity amongst Indian isolates of Xanthomonas oryzae pv. oryzae . PLOS One. 8(11), e81996. doi: 10.1371/journal.pone.0081996 (2013).
Kim, S. M. et al. Identification and fine–mapping of a new resistance gene, Xa40, conferring resistance to bacterial blight races in rice (Oryza sativa L.). Theor. Appl. Genet. 128, 1933–1943 (2015).
Hutin, M., Sabot, F., Ghesquière, A., Koebnik, R. & Szurek, B. A knowledge-based molecular screen uncovers a broad-spectrum OsSWEET14 resistance allele to bacterial blight from wild rice. The Plant J. 84 (4), 694–703 (2015).
Joseph, M. et al. Combining bacterial blight resistance and Basmati quality characteristics by phenotypic and molecular marker-assisted selection in rice. Mol. Breed. 13, 377–387 (2004).
Gopalakrishnan, S. et al. Integrating marker assisted background analysis with foreground selection for identification of superior bacterial blight resistant recombinants in Basmati rice. Plant Breed. 127, 131–139 (2008).
Basavaraj, S. H. et al. Marker-assisted improvement of bacterial blight resistance in parental lines of PusaRH10, a superfine grain aromatic rice hybrid. Mol. Breed. 26, 293–305 (2010).
Cheema, K. K. et al. A novel bacterial blight resistant gene from Oryza nivara mapped to 38 kb region on chromosome 4L and transferred to Oryza sativa L. Genet. Res. Cambridge. 90, 397–307 (2008).
Bhasin, H. et al. New PCR-based sequence-tagged site marker for bacterial blight resistance gene Xa38 of rice. Mol. Breed. 30, 607–611 (2012).
Murray, M. G. & Thompson, W. F. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 8 (19), 4321–4325 (1980).
Kauffman, H. E., Reddy, A., Hsieh, S. P. Y. & Merca, S. D. An improved technique for evaluating resistance of varieties to Xanthomonas oryzae pv. oryzae . Plant Dis. Rep. 57, 537–541 (1973).
Khanna, A. et al. Development and evaluation of near–isogenic lines for major blast resistance gene(s) in Basmati rice. Theor. Appl. Genet. 128 (7), 1243–1259 (2015).
Dhaliwal, H. S. & Kular, J. S. Package of practices for the crops of Punjab (ed. Kular, J. S. ) 2–3 (Punjab Agricultural University, Ludhiana, 2014). Available at http://www.kvkfaridkot.com/pp_kharif_2014.pdf (Accessed: 10th November, 2015).
Singh, A. et al. SNP haplotypes of the BADH1 gene and their association with aroma in rice (Oryza sativa L.) Mol. Breed. 26, 325–338 (2010).
Bradbury, L. M. T., Fitzgerald, T. L., Henry, R. J., Jin, Q. S. & Waters, D. L. E. The gene for fragrance in rice. Plant Biotechnol. J. 3, 363–370 (2005).
Singh, V. K. et al. Marker-assisted simultaneous but stepwise backcross breeding for pyramiding blast resistance genes Pi2 and Pi54 into an elite Basmati rice restorer line PRR78. Plant Breed. 132 (5), 486–495 (2013).
Singh, V. K. et al. Incorporation of blast resistance into “PRR78”, an elite Basmati rice restorer line, through marker assisted backcross breeding. Field Crops Res. 128, 8–16 (2012).
Vikal, Y. et al. Identification of new sources of bacterial blight (Xanthomonas oryzae pv. oryzae) resistance in wild Oryza species and O. glaberrima . Plant Genet. Resour. C. 5, 108–112 (2007).
Acknowledgements
The research work was funded by Accelerated Crop Improvement Programme, Department of Biotechnology, Government of India. The study is part of the Ph.D. research of the first author.
Author information
Authors and Affiliations
Contributions
A.K.S. conceptualized the idea; K.S. provided the introgression line and generated crosses; R.K.E. and A.K. performed the experiments; R.K.E. and K.K.V. performed the statistical analysis; K.K.M. provided Xoo inoculum; K.K.M. and R.K.E. conducted blight inoculation experiment; R.K.E., A.K.S., G.K.S. and P.K.B. managed field trials; R.K.E., M.N., A.K.S. and G.K.S. managed the offseason field experiments; R.K.E. and A.K. formulated the manuscript; A.K.S., N.K.S. and K.V.P. supervised the experiments and revised the manuscript; All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ellur, R., Khanna, A., S, G. et al. Marker-aided Incorporation of Xa38, a Novel Bacterial Blight Resistance Gene, in PB1121 and Comparison of its Resistance Spectrum with xa13 + Xa21. Sci Rep 6, 29188 (2016). https://doi.org/10.1038/srep29188
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep29188
This article is cited by
-
Pyramiding resistance genes for bacterial leaf blight (Xanthomonas oryzae pv. Oryzae) into the popular rice variety, Pratikshya through marker assisted backcrossing
Molecular Biology Reports (2023)
-
Molecular profiling of BADH2 locus reveals distinct functional allelic polymorphism associated with fragrance variation in Indian aromatic rice germplasm
Physiology and Molecular Biology of Plants (2022)
-
Rice transcriptome upon infection with Xanthomonas oryzae pv. oryzae relative to its avirulent T3SS-defective strain exposed modulation of many stress responsive genes
3 Biotech (2022)
-
Marker Assisted Development and Characterization of Herbicide Tolerant Near Isogenic Lines of a Mega Basmati Rice Variety, “Pusa Basmati 1121”
Rice (2020)
-
Blast resistance gene Pi54 over-expressed in rice to understand its cellular and sub-cellular localization and response to different pathogens
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.