Abstract
Quorum sensing activation by signal pheromone (CSP) in Streptococcus mutans depends on the membrane-associated receptor ComD, which senses the signal and triggers the signaling cascade for bacteriocin production and other cell density-dependent activities. However, the mechanism of the signal recognition via the ComD receptor in this species is nearly unexplored. Here, we show that the membrane domain of the ComD protein forms six transmembrane segments with three extracellular loops, loopA, loopB and loopC. By structural and functional analyses of these extracellular loops, we demonstrate that both loopC and loopB are required for CSP recognition, while loopA plays little role in CSP detection. A deletion or substitution mutation of four residues NVIP in loopC abolishes CSP recognition for quorum sensing activities. We conclude that both loopC and loopB are required for forming the receptor and residues NVIP of loopC are essential for CSP recognition and quorum sensing activation in S. mutans.
Similar content being viewed by others
Introduction
Two-component signal transduction systems (TCSTSs) are the most prevalent form of signal transduction mechanisms in bacteria1. A typical TCSTS consists of a membrane-associated, histidine protein kinase (HPK), which senses a specific stimulus, and a cytoplasmic response regulator (RR), which enables the cell to respond to the stimulus, via regulation of gene expression2. The completed genomes show that each bacterial genome contains a dozen or several dozen of TCSTSs, which play important roles in signal transduction for various cellular processes, stress adaptation and virulence3,4. Many TCSTSs have been found to function as global regulators by initiating signaling cascades, in which large sets of genes can be switched on or off 5. These systems provide the major means by which bacteria communicate with each other and the outside world6. Although many TCSTSs are identified to regulate diverse physiological activities and virulence in bacteria7, relatively little is known of how a given TCSTS recognizes and transduces an extracellular signal to initiate the signaling cascade for gene expression. In many cases, signals sensed by TCSTSs are chemically undefined, so that signal recognition mechanisms of many TCSTSs remain poorly understood. Among many TCSTS systems, quorum-sensing signal pheromones are the best-studied signal molecules that can be specifically sensed by their cognate TCSTSs8,9,10,11,12. These TCSTS signal transduction systems may provide an excellent opportunity to study signal molecule-HPK receptor interactions in prokaryotic organisms.
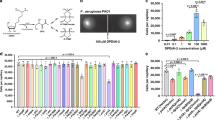
Streptococcus mutans, a leading cariogenic pathogen that can cause dental caries worldwide, has evolved a well conserved, signaling peptide-mediated quorum-sensing system, the ComCDE13,14. Quorum sensing activation through this signaling system depends on several gene products (Fig. 1). The comC encodes a signal peptide precursor, which is cleaved and exported to release a 21-residue peptide through a peptide-specific ABC transporter encoded by cslAB15,16. The 21-residue peptide is further modified by an extracellular protease SepM to remove the C-terminal 3 residues and generate an 18-residue functional peptide or competence-stimulating peptide (CSP)17. The comDE encode a two-component transduction system that specifically senses and responds to CSP. When it reaches a critical concentration, CSP interacts with the ComD histidine kinase receptor of the neighboring cells and activates its cognate response regulator, ComE, via autophospharylation. The phospharylated ComE in turn activates numerous downstream genes, triggering the signaling cascade to regulate bacteriocin production18, genetic competence14, biofilm formation9 and stress response19,20, which are all considered as the key virulence factors in the S. mutans pathogenesis. The quorum sensing circuit in S. mutans is the system in which the signal molecule is well studied in chemical details16,17,21. However, relatively little is known of the membrane-spanning receptor protein ComD and its interaction with the signal molecule.
The comC encodes a signal peptide precursor, which is cleaved and exported to release a 21-residue peptide through a peptide-specific ABC transporter encoded by cslAB. The 21-aa peptide is further modified by an extracellular protease SepM to remove the C-terminal 3 residues and generate an 18-residue functional peptide or competence-stimulating peptide (CSP). The comDE encode a two-component transduction system that specifically senses and responds to CSP. When it reaches a critical concentration, CSP interacts with the ComD receptor protein of the neighboring cells and activates its cognate response regulator, ComE, through autophospharylation. The phospharylated ComE in turn activates downstream genes, triggering the signaling cascade for bacteriocin production and other cell density-dependent activities.
Most known peptide pheromone receptors, with the exceptions of SpaK, ComP and NisK, fall into the HPK10 subfamily, which includes AgrC from Staphylococcus22,23, ComD from Streptococcus10 and PlnB from Lactobaccillus24. All members of the HPK10 subfamily belong to the orthodox histidine kinases that consist of a N-terminal transmembrane region representing the sensor domain and C-terminal transmitter domain containing the conserved histidine residues in the cytoplasm3,10,25. It has been predicted that membrane-associated regions in the HPK10 subfamily consist usually of 5–8 transmembrane segments (TMSs), which are contrast to the majority of prokaryotic HPKs with a N-terminal membrane domain consisting of two transmembrane segments flanking an extracellular loop3,10. By in silico analysis of ComD proteins from S. mutans strains, we predicted that the membrane-associated region of the ComD protein in this species likely forms six TMSs and three extracellular loops. We hypothesized that the extracellular loops of the ComD protein might act as the CSP receptor essential of signal recognition and quorum sensing activation. To test this hypothesis, we began to investigate the membrane topology of the ComD histidine kinase receptor protein. We then examined the effects of deletion or point mutations of the extracellular loops on signal recognition and quorum sensing activation in S. mutans.
Results
The ComD membrane topology
The ComD histidine protein kinase (HPK) of S. mutans is a membrane-associated protein consisting of 441 amino acid residues with a predicted molecular mass of 50.5 kDa and a pI value of 10.213. The sequence alignments indicate that ComD proteins from the fifteen genome-sequence completed S. mutans strains are highly conserved with 96.8–100% of identity3,26,27. However, ComD protein of S. mutans only shares 22% identity and 44% similarity with those of S. pneumoniae strains10. As the first step, we obtained a hypothetical topology model of ComD protein from S. mutans UA159 by combining several topology prediction methods, including SOSUI (http://bp.nuap.nagoya-u.ac.jp/sosui/), SMART (http://smart.embl-heidelberg.de/smart/), TMHMM (http://www.cbs.dtu.dk/services/TMHMM-2.0/) and PSIPRED (http://bioinf.cs.ucl.ac.uk/psipred/). Based the data from these methods, a hypothetical model of the ComD topology from S. mutans UA159 is presented in Fig. 2. As predicted by the topology model, the ComD protein consists of two hydropathically distinct regions, the N-terminal membrane-spanning region (1–220 residues) and the C-terminal hydrophilic region (221–441 residues) inside the cytoplasm. The membrane-spanning region of the ComD protein is predicted to form six transmembrane segments (TMSs) with three extracellular loops, designated as loopA, loopB and loopC, and two intracellular loops. Based on known peptide pheromone receptors in several species of Streptococcus10, we hypothesize that these extracellular loops likely form the receptor and contribute to CSP recognition, while the C-terminal region inside the cytoplasm is likely responsible for the signal transduction. To validate this hypothetical model, we constructed six comD-phoA-lacZ dual fusion reporters, which represented six in-frame insertion sites (L38, A70, T110, S150, P187, A224) of the membrane-spanning region of ComD protein (Fig. 2). The resulting fusion plasmids were transformed into an E. coli DH5α host, generating six comD-phoA-lacZ fusion reporter strains (Fig. 3A). These fusion strains along with two control strains were used for experimental determination of the ComD membrane topology.
The membrane-spanning domain of ComD protein from S. mutans UA159 is predicted to form six transmembrane segments (TMSs) with three extracellular loops, loopA, loopB and loopC, and two intracellular loops. An arrow indicates a potential cleavable side in loopA. Small open circles indicate insertion locations by a dual phoA-lacZ fusion reporter in frame after selected codons corresponding to the amino acid residues L38, A70, T110, S150, P187 and A224. Open rectangles indicate the amino acid residues of loopA, loopB and loopC involved in the construction of in-frame deletion or substitution mutants. The conserved histidine residue (H252) in the C-terminal domain of the ComD protein inside the cytoplasm is also indicated.
(A) A schematic diagram indicates the reporter fusion points at L38 (pKTop-ComD1-L38), A70 (pKTop-ComD1–A70), T110 (pKTop-ComD1–T110), S150 (pKTop-ComD1–S150), P187 (pKTop-comD1–P187) and A224 (pKTop-ComD1–A225) of the ComD membrane-spanning region. Blue boxes indicate extracellular locations, while pink boxes indicate cytosolic locations as predicted by the hypothetical model. (B) The strains expressing the ComDL38-Pho/Lac (GF-L38), ComDT110-Pho/Lac (GF-T110) and ComDP187-Pho/Lac (GF-P187) exhibit the higher levels of phosphatase activity (blue color), indicating the extracellular location of the reporter fusion points. The strains expressing the ComDA70-Pho/Lac (GF-A70), ComDS150-Pho/Lac (GF-S150) and ComDA225-Pho/Lac (GF-A224) exhibit the higher levels of β-galactosidase activity (pink color), indicating the cytosolic location of the reporter fusion points. E. coli DH5α without pKTop (negative control) shows no color, while E. coli DH5α with pKTop or GF-pKTop (positive control) also shows pink color.
We then examined these fusion strains for the reporter activities by growing them on LB agar plates containing dual indicators of a blue chromogenic substrate (X-Phos) for phosphatase activity (PhoA) and a red chromogenic substrate (Salman-Gal) for β-galactosidase activity (LacZ) using the methods as described previously28,29. The results showed that the strains expressing the ComDL38-Pho/Lac (L38), the ComDT110-Pho/Lac (T110) and the ComDP187-Pho/Lac (P187) exhibited the higher levels of phosphatase activities (blue color), indicating the periplasmic or extracellular location of these fusion points (Fig. 3B). In contrast, the strains expressing the ComDA70-Pho/Lac (A70), the ComDS150-Pho/Lac (S150) and the ComDA225-Pho/Lac (A224) exhibited the higher levels of β-galactosidase activity (pink color), indicating the cytosolic location of these fusion points. In control groups, E. coli DH5α without pKTop (plasmid negative control) showed no color, while E. coli DH5α with pKTop (plasmid positive control) showed pink (β-galactosidase activity). The results from the dual phoA-lacZ reporter assays clearly confirm the predicted membrane topology of the ComD protein, suggesting that three extracellular loops of the N-terminal membrane domain of ComD protein likely form the receptor for CSP recognition in S. mutans.
Confirmation of bacterial strains for investigating the extracellular loops in vivo
Having established the membrane topology of the ComD protein, we were particularly interested in the hypothesis that the ComD extracellular loops, loopA, loopB and loopC, might act as the receptor for CSP recognition and quorum-sensing activation. To test this hypothesis, we constructed three sets of S. mutans strains that allowed us to investigate the effects of individual extracellular loops on CSP recognition and quorum sensing activation. The first set of the strains included six in-frame deletion mutants and two substitution mutation mutants. The precise amino acid residues involved in the construction of the loopA, loopB and loopC mutants are highlighted in Fig. 4. All the constructs were generated from a plasmid template by a three-step genetic approach and confirmed by sequencing. After transformed into S. mutans UA159, the location and orientation of these constructs in the genome were further confirmed by a PCR strategy30. These mutants as well as control strains XT-C0 (ComD+) and ΔcomD (ComD−) were then transformed with two luxAB reporter plasmids, pGF-PcipB and pGF-PnlmAB respectively, generating a set of lux reporter strains for luciferase reporter activity assays (Table S1). To detect the mutant loopA, loopB and loopC proteins in the membrane fractions by Western blotting, we also constructed a set of S. mutans strains that had a chromosomally deleted comD (ΔcomD) but carried a shuttle vector pGF-Pldh-D-H that constitutively expressed a His-tagged loopA, loopB or loopC mutant protein. This shuttle vector was used to construct the strains, because it allowed not only rapid construction of the strains simply by replacing the wild copy comD between the Pldh promoter and 6His, but also constitutive expression of the mutant loopA, loopB and loopC proteins31. All the strains were then grown in THYE medium with addition of CSP to prepare the membrane proteins for Western blot analysis. The results showed that a strong reactive band (≈51-kDa) was detected in the membrane fractions of all the strains (Fig. 5), except the ComD− mutant. The positive reactive bands in the membrane fractions of these strains were consistent with the size (50.5 kDa) of the wild type ComD protein13,14. The results clearly demonstrated that a short deletion or substitution mutation of these extracellular loops did not affect the translocation or insertion of these mutant proteins into the cytoplasmic membrane. Thus, all the mutants as well as control strains should be valid for studying the effects of a deletion or mutation of these extracellular loops on CSP perception and quorum sensing activation.
All the strains were grown in THYE medium with addition of CSP to prepare the membrane proteins, which were then resolved on 10% SDS-PAGE gels and transferred onto PVDF membranes for Western blotting using the anti-His antibody. (A) Lanes 1–4, XT-D-H (ComD+), XT-A1H (loopA1−), XT-A2H (loopA2−) and XT-Pldh-H (ComD−); (B) Lanes 1–4, XT-D-H (ComD+), XT-Pldh-H (ComD−), XT-B1H (loopB1−) and XT-B2H (loopB2−); and (C) Lanes 1–4, XT-C1H (loopC1−), XT-C2H (loopC2−), XT-C3H (loopC3−) and XT-C4H (loopC4−). Arrow indicates 51-kDa proteins detected by Western blotting.
Effects of a deletion or mutation of the extracellular loops on CSP perception
Next, we determined whether a deletion or mutation of loopA, loopB or loopC affected CSP recognition and quorum sensing activation by examining specific luciferase reporter activities of the reporter strains in response to CSP. All the reporter strains were constructed in the mutant backgrounds and control strains (ComD+ and ComD−) by transforming a luxAB reporter plasmid, pGF-PcipB or pGF-PnlmAB into these strains. Therefore, each reporter strain carried a reporter plasmid containing a promoterless luxAB fused to the CSP-inducible promoter of two bacteriocin-encoding genes, cipB and nlmAB18. These two genes were chosen for constructing the reporter strains, because their promoters (PcipB and PnlmAB) contain the consensus ComE binding site and are directly controlled by the ComCDE quorum sensing system18,32. We then examined specific luciferase reporter activities of these strains in response to CSP. The results revealed that a deletion of loopA, either four residues SNVT (loopA1−) or eight residues SNVTLSKK (loopA2−), showed little effect on the responses of these two promoters to CSP, since the luciferase reporter activities in both loopA1− and loopA2− strains were similar to those in the ComD+ control strains (Fig. 6A,C). The results suggest that extracellular loopA appears to be not directly involved in CSP perception for quorum sensing activation. However, a deletion of four residues LDGT of loopB (loopB1−) resulted in a reduction in the luciferase reporter activities compared to the reporter activities in the ComD+ control strains. Interestingly, this was not the case when other four residues QGIV of loopB (loopB2−) were deleted, suggesting that residues LDGT but not QGIV of loopB largely participate in CSP recognition. Even more dramatically, a deletion of four residues NVIP of loopC (loopC1−) completely abolished the response of these two promoters to CSP, while a deletion of four residues TLKF of loopC (loopC2−) resulted in 50% of reduction in the lux reporter activities in response to CSP. This suggests that residues NVIP of loopC (loopC1−) may be essential for CSP recognition. The results strongly suggest that both extracellular loopC and loopB are required for CSP recognition and quorum sensing activation, while loopA appears to play little role in CSP detection.
Two CSP-inducible promoters, PcipB and PnlmAB, were monitored for luciferase reporter activities. (A) Luciferase reporter activities (RLU/OD590) of PcipB::luxAB reporter strains, XT-Lx20 (ComD+), XT-Lx21 (loopA1−), XT-Lx22 (loopA2−), XT-Lx23 (loopB1−), XT-Lx24 (loopB2−), and XT-Lx29 (ComD−), were assayed with addition of CSP (CSP+) and without CSP (CSP−). (B) Luciferase reporter activities (RLU/OD590) of PcipB::luxAB reporter strains, XT-Lx20 (ComD+), XT-Lx25 (loopC1−), XT-Lx26 (loopC2−), XT-Lx27 (loopC3−), XT-Lx28 (loopC4−) and XT-Lx29 (ComD−), were assayed under the same conditions. (C) Luciferase reporter activities (RLU/OD590) of PnlmAB::luxAB reporter strains, XT-Lx30 (ComD+), XT-Lx31 (loopA1−), XT-Lx32 (loopA2−), XT-Lx33 (loopB1−), XT-Lx34 (loopB2−) and XT-Lx39 (ComD−), were assayed under the same conditions. (D) Luciferase reporter activities (RLU/OD590) of PnlmAB::luxAB reporter strains, XT-Lx30 (ComD+), XT-Lx35 (loopC1−), XT-Lx36 (loopC2−), XT-Lx37 (loopC3−), XT-Lx38 (loopC4−), and XT-Lx39 (ComD−), assayed under the same conditions.
To further confirm the results that the residues NVIP of loopC (loopC1−) may be essential for CSP perception, we constructed two more loopC mutants that had an alanine substitution mutation in loopC, designated as loopC3− (AA/NV) and loopC4− (AA/IP). These mutants were then transformed with the lux reporter plasmids, generating four more reporter strains. By assaying the specific reporter activities, we found that both substitution mutants failed to respond to CSP for induction of the luciferase reporter activities (Fig. 6B,D), confirming that the four residues NVIP of loopC are truly essential for CSP recognition. Importantly, these substitution mutations did not affect detection of the mutant proteins by Western blotting (Fig. 5C, lanes 3–4), suggesting no effect on the translocation or insertion of these mutant proteins into the cytoplasmic membrane. Thus the results confirm that at least four residues, NVIP, of loopC are essential for CSP recognition and quorum sensing activation in S. mutans.
Natural point mutations in loopB or in both loopB and loopC on CSP perception
By sequence alignments of ComD proteins, we also identified several S. mutans strains that had one or two residue substitution mutations either within loopB, such as in strain GS-5, or within both loopB and loopC, such as strains KK23 and R221 (Fig. 7A). Compared with strain UA159, these strains have two amino acid residue substitutions of T110 (Threonine110) by N110 (Asparagine110) and G116 (Glycine116) by D116 (Aspartic acid116) within loopB. In addition to these two substitutions within loopB, strains KK23 and R221 also have one residue substitution of T178 (Threonine178) by V178 (Valine178) within loopC. We were curious to know whether these strains might be defective in perception and response to CSP for quorum sensing activation. Since both strains and their ComD deletion mutants were available in our lab, we directly used these strains to probe this question. We first transformed the lux reporter fusion plasmids, pGF-PcipB and pGF-PnlmAB, into these strains to generate two sets of new lux reporter strains (Table S1). We then assayed the luciferase reporter activities of these strains in response to CSP. Surprisingly, we found that all the strains showed the wild type levels of the luciferase reporter activities in response to CSP (Fig. 7B). There was no significant difference (P > 0.05) in the luciferase report activities between GS-5 or R221 and UA159 derived strains. In contrast, XT-D0GS-5 (ΔComDGS-5) and XT-D0R211 (ΔComDR211) derived reporter strains showed little induction in the luciferase report activities, suggesting that these mutants were unable to sense and respond to CSP, a pattern very similar to XT-D0UA159 (ComD−) derived reporter strains. The results suggest that the single residue mutations in loopB or in both loopB and loopC do not appear to affect CSP perception and quorum sensing activation in the S. mutans strains tested.
(A) A sequence alignment of loopA, loopB and loopC among four S. mutans strains showing point mutations in loopB (T110 >N110 and G116 >D116) in strain GS-5 or in both loopB (T110 >N110 and G116 >D116) and loopC (T178 >V178) in strains KK23 and R221. (B) Luciferase report activities (RLU/OD590) of S. mutans GS-5-derived strains XT-Lx40 (white, PcipB) and XT-Lx41 (black, PnlmAB), XT-D0GS5 (ΔComD)-derived strains XT-Lx42 and XT-Lx43, R221-derived strains XT-Lx44 and XT-Lx45, and XT-D0R211 (ΔComD)-derived strains XT-Lx46 and XT-Lx47. The luciferase report activities (RLU/OD590) of PcipB::luxAB and PnlmAB::luxAB reporter strains were assayed in THYE medium with addition of CSP.
Effects of a deletion or mutation of loopA, loopB or loopC on bacteriocin production
It is well known that CSP-mediated quorum sensing primarily regulates production of several bacteriocin-encoding genes, such as cipB (SMU.1904) and nlmAB (SMU.151/152) in S. mutans18. To determine the effects of a deletion or mutation of the extracellular loops on bacteriocin production, we examined CSP-inducible bacteriocin production of all the strains using a deferred antagonism assay. The results showed that except the loopA mutants that produced similar levels of bacteriocin to that by the ComD positive strain (ComD+), all the mutants produced reduced levels of bacteriocins compared to the ComD+ strain (Fig. 8). In particular, the loopC1, loopC3 and loopC4 mutants were significantly defective in producing bacterocins, which were consistent with the findings from the luciferase reporter activities. A minor inhibitory ring observed in the ComD deficient strains, including ΔcomD mutant, might result from a bacteriocin that was not controlled by the ComCDE quorum sensing system33. The results confirm that a deletion or mutation of loopB and loopC resulted in moderate or severe deficiency in CSP-mediated bacterocin production. We also examined bacteriocin production of S. mutans strains GS-5 and R211 in response to CSP, since both strains have a residue substitution mutation either in loopB or in both loopB and loopC based on the genome sequences (Fig. 7A). The results showed that both strains produced nearly equal levels of bacteriocins to that by UA159. In contrast, both GS-5- and R211-derived ComD deletion mutants (ΔComDGS-5 and ΔComDR211) were defective in the production of bacteriocins (Fig. 8C). The results suggest that the single substitution mutations either in loopB or loopC appear to have little effect on CSP-dependent production of bacteriocins. The results are highly consistent with the previous reports showing that both strains GS-5 and R211 are potent bacteriocin producers26,27.
A deferred antagonism assay was used to assess production of CSP-induced bacteriocins by S. mutans strains. The cell suspensions of each strain were stabled onto THYE agar plates and overlaid with an indicator strain S. sanguinin SK108 by mixing the cells (107 CFU/ml) in low-melting agarose. The overlaid plates were incubated anaerobically at 37 °C for 20 hours before examining bacteriocin production. (A) ComD+ (XT-C0), loopA1−, loopB1−, loopC1−, ComD− (ΔComD); (B) loopC1−, loopC2−, loopC3−, loopC4−; (C) UA159 (wt), R211 (wt), GS5 (wt), ΔComDR211 and ΔComDGS-5.
Discussion
In Gram-positive bacteria, signal peptide pheromone-activated histidine protein kinases from the HPK10 subfamily control several important physiological processes, including competence development, bacteriocin production and virulence expression3,10,11. These signaling regulatory systems, including the agr, com and pln regulons, have been extensively studied with the respect to the events following phosphorylation of their cognate response regulators10,11,23,24. However, relatively little is known of interactions between signal pheromones and their cognate receptor proteins. No report has directly described ComD receptor kinase proteins, which are widely distributed among the members of the Genus Streptococcus3,8,10. In this study, we began to investigate the membrane topology of the S. mutans ComD protein, with our focus on the structural analysis of the extracellular loops of the ComD that acts as the CSP receptor. We demonstrate by the dual phoA-lacZ reporter system that the membrane-spanning domain of the ComD protein forms six transmembrane segments (TMSs) with three extracellular loops, loopA, loopB and loopC. The most important conclusion from the topology studies is that we confirm the extracellular locations of loopA, loopB and loopC. The results show very good agreement with the in silico predicted topology model of the ComD (Fig. 2), thereby, validating the membrane topology of the ComD receptor.
Upon the establishment of the membrane topology of ComD protein, we further explored the contribution of the extracellular loops, loopA, loopB and loopC, to CSP recognition and quorum-sensing activation, since the data obtained may have important implications for the ligand-receptor interaction and design of quorum sensing inhibitors. One of the major considerations in mutational analysis of the ComD protein was whether a partial deletion or mutation of these extracellular loops affects translocation or insertion of the mutant proteins into the membrane. It was important to track the mutant proteins in the membrane fractions, otherwise, it would be difficult to interpret the results regarding the effects of a deletion or mutation of the extracellular loops on CSP recognition and quorum sensing activation. To detect the mutant ComD proteins in the membranes by Western blotting, we constructed a new set of the S. mutans strains that constitutively expressed a His-tagged mutant loopA, loopB or loopC in trans. This ensured constitutive expression of the loopA, loopB and loopC mutant proteins in the chromosomally deleted comD mutant. Western blot analysis showed that all the mutant proteins (≈51 kDa) were detectable in the membrane fractions of these strains, suggesting that the mutant proteins could adequately translocate and insert into the membrane. Thus, the results from the luciferase reporter assays should be valid to evaluate the effects of a deletion or mutation of these loops on CSP recognition. Our work demonstrate that the extracellular loopC and loopB most likely form the receptor for CSP recognition, since a deletion or mutation of either loop resulted in partial or complete deficiencies in CSP-dependent quorum sensing activation. In contrast, loopA appears to play little role in signal detection, because a deletion of up to eight residues of this loop caused little effect on quorum sensing activation. In addition, we have confirmed that at least four residues, NVIP, of loopC are essential for CSP recognition, since a deletion or mutation of these residues abolishes CSP recognition and quorum sensing activation.
Another interesting finding from this study is that by sequence alignments of the S. mutans ComD proteins we have identified several S. mutans strains that have one or two point substitution mutations either within loopB, such as in strain GS-5, or within both loopB and loopC, such as in strain R221. However, our experiments confirm that both of these strains can detect and respond to CSP as effectively as S. mutans UA159. The results reveal that these single substitution mutations, even at such important locations of the receptor, do not appear to affect CSP recognition and quorum sensing activation. Neither do these mutations significantly affect CSP-induced bacteriocin production, suggesting that the ComD receptor in S. mutans displays relatively low specificity to sense the signal for quorum sensing activation within the species. Such less constraint specificity in CSP-ComD interaction in S. mutans strains clearly differs from those in S. aureus and S. pneumoniae. It has been well recognized that quorum-sensing signaling molecules produced by many bacteria often induce species-specific or even strain-specific activities at nano-molar concentrations. This feature has been used to explore structure-activity relationships between a signal pheromone and its cognate receptor12,16,21,34. For example, the AIP-AgrC quorum sensing system in S. aureus is one of the best-studied model systems that show highly strain-specific activities to induce quorum-sensing response11,34. The sequence variations of the AIPs from different S. aureus strains have led to at least four specificity groups in S. aureus23,35. All strains within one group produce the same AIP, which only activates quorum sensing and the virulence within its own specificity group but not in other groups. In S. pneumoniae, extensive screening of pneumococcal isolates reveals two major CSP variants that are highly specific to interact with their respective receptors, ComD1 and ComD236,37. These studies have led to the proposal that S. pneumoniae strains can be divided into different pherotypes based on their quorum sensing pheromone specificity10,36. Similar studies have been carried out to identify quorum-sensing signaling peptide variants from S. mutans strains and clinical isolates. Seven comC alleles encoding three distinct mature CSPs are identified among 36 geographically diverse S. mutans strains15. In contrast to S. pneumoniae, however, all three CSP variants function equally well to induce quorum sensing, bacteriocin production and genetic competence15,17,21. There is no evidence showing quorum sensing signal pheromone pherotype in S. mutans. In fact, structural and functional divergences in ComCDE quorum sensing systems between S. mutans and S. pneumoniae have been recognized for some years38. Phylogenetic analysis of various streptococcal genomes reveals that ComCDE system in S. mutans is more closely related to BlpCRH quorum sensing system that directly controls bacteriocin production in S. pneumoniae38,39. Our sequence alignments also show that ComD proteins of S. mutans only shares 22% of identity and 44% of similarity with those of S. pneumoniae. This suggests that ComD proteins of S. mutans may not necessarily share the same membrane topology and signal recognition domains with ComD proteins in S. pneumoniae. Despite no experiments performed to test this difference, in silico analysis of ComD proteins of S. pneumoniae strains appears to support this suggestion10. The ComCDE system in S. mutans is suggested to combine the action of two orthologous systems, the ComCDE and BlpCRH in S. pneumoniae, which are well known to be involved in competence development and bacteriocin production, respectively38. What is the ecological implication of such less constraint specificity of the ComD-CSP interaction in S. mutans is unclear. However, we speculate that the less constraint specificity may provide S. mutans with more flexibility or even an advantage to sense the signal molecule for intra-species communication in densely packaged dental biofilms, where S. mutans needs to compete with closely related colonizers, such as various species of streptococci, by population-wide production of bacteriocins. This speculation regarding the ComCDE controlled activities of S. mutans in dental biofilms is currently under our investigation.
In summary, this study demonstrates that the membrane domain of the S. mutans ComD protein forms six transmembrane segments and three extracellular loops, loopA, loopB and loopC. Structural analysis of these extracellular loops reveals that both loopC and loopB are required for CSP recognition and quorum sensing activation, while loopA plays little role in CSP detection. However, single sequence variations in loopB and loopC exist, such as in S. mutans strains GS-5 and R211, but these strains do not show any detectable defects in CSP recognition. Thus, unlike other pheromone receptors in the HPK10 subfamily, the ComD receptor in S. mutans shows more flexibility to sense and transduce the signal for quorum sensing activation, which may provide S. mutans with an advantage for intra-species communication in its natural ecosystem, dental biofilms.
Methods
Bacterial strains, media and growth conditions
Bacterial strains and plasmids used in this study are listed in Tables S1 and S2, respectively. S. mutans wild-type strains UA159, GS-5 and R211 were grown on Todd-Hewitt medium supplemented with 0.3% yeast extract (THYE), whereas all the mutants, the lux reporter strains and other strains derived from S. mutans UA159, GS-5 and R211 were maintained on THYE medium supplemented with an appropriate antibiotic(s). Escherichia coli hosts and their derivatives generated by molecular cloning were grown in Luria-Bertani (LB) medium supplemented with an appropriate antibiotic(s).
In silico prediction of ComD topology
We combined several topology prediction methods, including SOSUI (http://bp.nuap.nagoya-u.ac.jp/sosui/), SMART (http://smart.embl-heidelberg.de/smart/), TMHMM (http://www.cbs.dtu.dk/services/TMHMM-2.0/) and PSIPRED (http://bioinf.cs.ucl.ac.uk/psipred/), to obtain a hypothetical topology model of ComD protein from S. mutans UA159. All the methods were used in single protein mode and user-adjustable parameters were left at their default values. Such in silico analyses of the ComD protein had led us to generate a hypothetical topology model (Fig. 2), which facilitated the design and genetic construction of dual reporter fusion strains and mutants. In addition, the sequence alignments of ComD proteins from many strains of S. mutans and S. pneumoniae were performed with MacVector 9.0 ClusterW. All the protein sequences of ComD proteins from these strains were obtained from the NCBI protein database (http://www.ncbi.nlm.nih.gov/protein/).
Construction of comD-phoA-lacZ fusion reporters
To study the membrane topology of ComD protein, we used a dual phoA-lacZ reporter system, pKTop, which consists of an E. coli alkaline phosphatase fragment PhoA22-472 fused in frame after the α-fragment of β-galactosidase LacZ4-6040, to construct a series of comD-phoA-lacZ fusion reporters in frame after selected comD codons, including L38 (pKTop-ComD1-L38), A70 (pKTop-ComD1–A70), T110 (pKTop-ComD1–T110), S150 (pKTop-ComD1–S150), P187 (pKTop-comD1–P187) and A224 (pKTop-ComD1–A225). The corresponding comD fragments were amplified from the S. mutans UA159 genomic DNA by PCR using appropriate primer pairs (Table S3) and subcloned into the BamHI and KpnI restriction sites of pKTop28. Positive transformants grown on LB plates supplemented with kanamycin (50 μg/ml) were selected for genetic confirmation by PCR and restriction analysis. For the membrane protein topology assays, E. coli DH5α were transformed with the resulted fusion plasmids to generate six pKTop derivatives expressing tripartite ComD-PhoA/LacZ. The phoA and lacZ genes missing their leader sequence were used as negative control for the report activity assays. The resulting reporter strains were then grown on LB plates containing dual indicators of a blue chromogenic substrate, 5-bromo-4-chloro-3-indoxyl phosphate disodium salt or X-Pho (Sigma) at a concentration of 80 μg/ml, and a red chromogenic substrate, 6-chloro-3-indoxyl-β-D-galactopyranoside or Salmon-Gal (Gold Biotech) at a concentration 120 μg/ml and IPTG (1 mM) with kanamycin (50 μg/ml). The periplasmic or cytosolic location was determined based on the locations of these fusion reporters. A periplasmic or extracellular location of a reporter fusion point should lead to the higher alkaline phosphatase activity (blue color), whereas a cytosolic location of a reporter fusion point should result in the higher β-galactosidase activity (pink color)28,40.
Construction of extracellular loopA, loopB and loopC mutants
To evaluate the contribution of the ComD extracellular loops to CSP recognition, we constructed eight comD mutants by a three-step genetic approach (Fig. S1). Each mutant constructed had an in-frame deletion or substitution mutation in loopA, loopB or loopC. In the first step, a 2064-bp DNA fragment containing the comD-coding sequence and its flanking regions was amplified by PCR against the genomic DNA of S. mutans UA159, and subcloned into pBlueScript SKII (Stratagene). The resulting plasmid was then used as a template to generate an amplicon (the entire plasmid) with two restriction sites of AscI and FseI by circular PCR from the location downstream of comD but within comC. The amplicon was digested and ligated to the same restriction sites of an erythromycin resistance cassette. The ligation product was cloned into an E. coli host, generating a new plasmid, pGF-C0 that contained a wild copy of comD (ComD+) but with comC (ComC−) inactivated by the insertion of the erm cassette. In the second step, this new plasmid was used as a template to generate six deletion constructs of loopA, loopB and loopC by an inverse PCR strategy and two alanine substitution mutant constructs corresponding to the target codons of loopC by a QuickChange II site-directed mutagenesis kit (Agilent Tech. Inc.). The resulting products were digested with BamH1, self-ligated and cloned into E. coli, generating new plasmids that harbored a mutant construct of loopA, loopB and loopC, respectively. All plasmids with a mutant construct were genetically confirmed by sequencing. In the last step, these plasmids were linearized and transformed into S. mutant UA159. Following double-crossover recombination, each of these constructs was integrated into the S. mutans genome, generating a number of ComD mutants (Table S1). Positive transformants were selected from THYE plates plus erythromycin (10 μg/ml) and confirmed by a PCR strategy using a combination of four primers specific to comD and the erm cassette30. A S. mutant strain with a full-length ComD but with comC inactivated by an insertion of the erm cassette (ComD+, ComC−, Ermr) was included as a positive control, while comD deletion mutant (ΔcomD) was used as a negative control. Since the comC was inactivated by the insertion of the erm cassette, all the strains constructed were unable to produce endogenous CSP.
Construction of S. mutans strains expressing a His-tagged, ComD mutant protein
To determine whether a deletion or point mutation of loopA, loopB or loopC affected translocation or insertion of the mutant proteins into the cytoplasmic membrane, we also constructed a set of S. mutans strains that carried a shuttle vector expressing a His-tagged ComD mutant protein under control of a constitutively expressed promoter of ldh, the gene encoding lactate dehydrogenase in S. mutans13,31. This allowed subsequent detection of His-tagged mutant proteins by Western blotting. To achieve this goal, we simply amplified each of the ComD mutant constructs (except stop codon) from constructed plasmids by primers ComD-BamHI-F and ComD-NotI-B. The PCR products were digested, purified and cloned into a vector pGF-Pldh-D-H by replacing the comD (Fig. S2). Each plasmid was constructed in such a way that the coding sequence was fused to the ldh promoter (Pldh) with its start codon and to 6His-tag with its end, generating a number of plasmids (Table S2). A plasmid pGF-Pldh-H (without comD) was also generated as a negative control by removing the comD from pGF-Pldh-D-H. The newly constructed plasmids were then confirmed by PCR and sequencing. The confirmed plasmids were transformed into the ΔcomD mutant, generating a new set of S. mutans strains that constitutively expressed a His-tagged mutant ComD protein that could be detected in the membrane fractions by Western blotting.
Cell fractionation and protein detection by Western blotting
The S. mutans strains that expressed a His-tagged mutant comD were grown in THYE medium. When reaching to the early-log phase (OD600 ≈ 0.3), all the cultures were added with 1 μM of CSP and incubated until the mid-log phase (OD600 ≈ 0.6). Aliquots of samples were taken to prepare the crude membrane proteins using a modified method as described previously41,42. Briefly, the cell pellets were re-suspended in 2 ml of TE buffer (50 mM Tris-HCl, 50 mM EDTA, pH 7.6, 200 μg/ml of lysozyme and 30 μg/ml of mutalysin) and incubated at 37 °C for 2 hrs. The cell lysates were prepared by sonication on ice at 30 s pulses for 3 times after added with an aliquot of a protease inhibitor cocktail (Sigma-Aldrich). The cell lysates were centrifuged at 30, 000× g at 4 °C for 1 h. The membrane fractions were resuspented in 100 μl of 50 mM ammonium bicarbonate (pH 11) and the proteins were extracted with 100 μl of trifluroethanol/chloroform (2:1 v/v)41,42. The samples were resuspended in 60 μl of solubilization buffer (7M urea, 2M thiourea, 2% Triton X-100, 0.5% ASB-14, 50 mM dithiothreitol, 0.2% Bio-Lytes). The proteins were resolved on SDS-PAGE gels and transferred to polyvinylidene difluoride (PVDF) membranes for Western blot analysis using an anti-His-tag antibody (1:3,000 dilution) by the method as described previously43. The membranes were washed and detected with an AP detection reagent (Novagen). The proteins on the membrane were analyzed using FluorChem SP image system (Alpha Innotech, Calif.).
Construction of luxAB transcriptional reporter strains and luciferase reporter assay
To determine the effects of a deletion or mutation of the extracellular loops on CSP perception and quorum sensing activation, we construct a number of transcriptional luxAB reporter strains in the loopA, loopB and loopC mutant backgrounds (Fig. S3). Each strain harbored a shuttle vector that carried a promoterless luxAB fused to the promoter of two CSP-inducible, bacteriocin-encoding gene, cipB and nlmAB13,18. The DNA fragments containing the promoter regions of these genes were generated by PCR, purified and cloned into a shuttle vector pGF-kan43, generating two luxAB reporter fusion plasmids (Table S2). The plasmids were genetically confirmed by PCR and sequencing. The confirmed plasmids, designated pGF-PcipB and pGF-PnlmAB, were transformed into the S. mutans mutants and control strains (ComD+ and ComD−), generating two groups of luxAB reporter strains that were then used to assay specific luciferase reporter activities of the promoters, PcipB and PnlmAB, in response to CSP.
To determine the effects of a deletion or mutation of loopA, loopB or loopC on CSP perception and quorum sensing activation, luciferase reporter activity assays were carried out to examine CSP-dependent quorum sensing activation. All the mutants that carried either pGF-PcipB or pGF-PnlmAB were grown to assay their specific luciferase report activities in response to CSP (SGSLSTFFRLFNRSFTQA), which was synthesized with 90% purity commercially (BioBasic Inc., Ontario). CSP was freshly dissolved in sterile distilled water at 1.0 mM (stock), which was further diluted to make working concentrations as required. S. mutans wild type background strains carrying the reporter plasmid were used as positive controls (ComD+), whereas the ΔcomD background strains carrying the plasmids were used as negative controls (ComD−). All the strains were grown in THYE broth and the cultures were added with an aliquot of CSP (1 μM) at O.D590 ≈ 0.3. Bacterial growth and luciferase reporter activities (O.D590) of the strains were assayed with a 96-well microplate reader (Synergy HT, Biotek, USA). One percent nonanal (Sigma-Aldrich) was added into the cultures, since LuxAB-catalyzed luciferase activity requires nonanal as a substrate31. Briefly, aliquots (200 μl) of cell suspension were transferred to wells of a pre-warmed (37 °C) microtiter plate. A 50 μl of the solution containing 1% nonanal in mineral oil in a volatile form were placed in the spaces of wells and the plate was covered with a lip. The microtiter plates were incubated and read in the prewarmed (37 °C) microplate reader at O.D590 at a time intervals of 15 min for 4 hours. The results were expressed in relative luminescent units (RLU) divided by cell density of the cultures. The data plotted represented the maximal RLU readings of each strain.
Deferred antagonism assay
To determine the effects of a deletion or mutation of loopA, loopB or loopC on CSP-induced bacteriocin production, all the S. mutans strains were grown for overnight. Aliquots of the cell suspension were stabbed onto THYE agar, incubated anaerobically at 37 °C for six hours and added on the surfaces of each inoculum with 5 μl of CSP (1 μM). The agar plates were then overlaid with a freshly grown indicator strain S. sanguinis SK108 by mixing the cell suspension (at 107 CFU/ml) with low-melting agarose (Bishop Canada Inc.). The overlaid plates were incubated anaerobically at 37 °C for additional 20 hours before inspection of bacteriocin production. S. mutans strains showing inhibitory zones around the stabs indicated positive production of bacteriocins21.
Additional Information
How to cite this article: Dong, G. et al. Membrane Topology and Structural Insights into the Peptide Pheromone Receptor ComD, A Quorum-Sensing Histidine Protein Kinase of Streptococcus mutans. Sci. Rep. 6, 26502; doi: 10.1038/srep26502 (2016).
References
Stock, A. M., Robinson, V. L. & Goudreau, P. N. Two-component signal transduction. Annu. Rev. Biochem. 69, 183–215 (2000).
Hoch, J. A. & Varughese, K. I. Keep signals straight in phosphorelay signal transduction. J. Bacteriol. 183, 4941–4949 (2001).
Grebe, T. W. & Stock, J. B. The histidine protein kinase superfamily. p139-227. In Poole, R. K. (ed) Advances in microbial physiology. (Academic Press, UK, 1999).
Jung, K., Fried, L., Behr, S. & Heermann, R. Histidine kinases and response regulators in networks. Curr. Opin. Microbiol. 15, 118–124 (2012).
Mascher, T., Helmann, J. D. & Unden, G. Stimulus perception in bacterial signal transduction histidine kinase. Microbiol. Mol. Biol. Rev. 70, 910–938 (2006).
West, A. H. & Stock, A. M. Histidine kinases and response regulator proteins in two-component signaling systems. Trends Biochem. Sci. 26, 369–76 (2001).
Stephenson, K. & Hoch, J. A. Two-component and phosphorelay signal-transduction systems as therapeutic targets. Curr. Opin. Pharmacol. 2, 507–512 (2002).
Kleerebezem, M., Quadri, L. E., Kuipers, O. P. & de Vos, W. M. Quorum sensing by peptide pheromones and two-component signal-transduction systems in Gram-positive bacteria. Mol Microbiol. 24, 895–904 (1997).
Li, Y. H. et al. A quorum sensing signaling system essential for genetic competence in Streptococcus mutans is involved in biofilm formation. J. Bacteriol. 184, 2699–2708 (2002).
Håvarstein, L. S. Intercellular communication in Gram-positive bacteria depends on peptide pheromones and their histidine kinase receptors. p341-363. In Inouye, M. & Dutta, R. (ed), Histidine kinases in signal transduction. (Academic Press, UK, 2003).
Novick, R. P. Autoinduction and signal transduction in the regulation of staphylococcal virulence. Mol, Microbiol. 48, 1429–1449 (2003).
Lyon, G. J. & Novick, R. P. Peptide signaling in Staphylococcus aureus and other Gram-positive bacteria. Peptides 25, 1389–403 (2004).
Ajdic, D. et al. Genome sequence of Streptococcus mutans UA159, a cariogenic dental pathogen. Proc. Natl. Acad. Sci. USA 99, 14434–14439 (2002).
Li, Y. H. et al. Natural genetic transformation of Streptococcus mutans growing in biofilms. J. Bacteriol. 183, 897–908 (2001).
Allan, E. et al. Genetic variation in comC, the gene encoding competence-stimulating peptide (CSP) in Streptococcus mutans . FEMS Microbiol. Lett. 268, 47–51 (2007).
Syvitski, R. T. et al. Structure-activity analysis of quorum sensing signaling peptides from Streptococcus mutans . J. Bacteriol. 189, 1441–1450 (2007).
Hossain, M. S. & Biswas, I. An extracellular protease, SepM, generates functional competence-stimulating peptide in Streptococcus mutans UA159. J. Bacteriol. 194, 5886–5896 (2012).
van der Ploeg, J. R. Regulation of bacteriocin production in Streptococcus mutans by the quorum-sensing system required for development of genetic competence. J. Bacteriol. 187, 3980–3989 (2005).
Lemme, A. et al. Subpopulation-specific transcriptome analysis of competence-stimulating-peptide-induced Streptococcus mutans . J. Bacteriol. 193, 1863–1877 (2011).
Leung, V. & Lévesque, C. M. A stress-inducible quorum sensing peptide mediates the formation of persister cells with non-inherited multidrug tolerance. J. Bacteriol. 194, 2265–2274 (2012).
Tian, X. L. et al. A method for structure-activity analysis of quorum-sensing signaling peptides from naturally transformable streptococci. Biol. Procedure Online 11, 207–226 (2009).
Novick, R. P. & Geisinger, E. Quorum sensing in Staphylococci. Ann. Rev. Genet. 42, 541–564 (2008).
Lina, G. et al. Transmembrane topology and histidine protein kinase activity of AgrC, the agr signal receptor in Staphylococcus aureus . Mol. Microbiol. 28, 655–662 (1998).
Johnsborg, O., Diep, D. B. & Nes, I. F. Structural analysis of the peptide pheromone receptor PlnB, a histidine protein kinase from Lactobacillus plantarum . J. Bacteriol. 185, 6913–6920 (2003).
Lévesque, C. M. et al. Systematic inactivation and phenotypic characterization of two-component systems in expression of Streptococcus mutans virulence properties. Lett. Appl. Microbiol. 45, 398–404 (2007).
Song, L. et al. A genome-wide study of two-component signal transduction system in eight newly sequenced mutants streptococci strains. BMC Genomics 13, 128 (2012).
Cornejo, O. E. et al. Evolutionary and population genomics of the cavity causing bacteria Streptococcus mutans . Mol. Biol. Evol. 30, 881–893 (2013).
Karimova, G., Robichon, C. & Ladant, D. Characterization of YmgF, a 72-residue inner membrane protein that associates with the Escherichia coli cell devision machinery. J. Bacteriol. 191, 333–346 (2009).
Falord, M., Karimova, G., Hiron, A. & Msadek, T. GraASR proteins interact with the VraFG ABC transporter to form a five-component system required for cationic antimicrobial peptide sensing and resistance in Staphylococcus aureus . Antimicrob. Agents Chemother. 56, 1047–1058 (2012).
Lau, P. C. Y. et al. PCR ligation mutagenesis in transformable streptococci: application and efficiency. J. Microbiol. Methods 49, 193–205 (2002).
Tian, X. T. et al. MecA protein acts as a negative regulator of genetic competence in Streptococcus mutans . J. Bacteriol. 195, 5196–5206 (2013).
Hung, D. C. et al. Characterization of DNA binding sites of the ComE response regulator from Streptococcus mutans . J. Bacteriol. 193, 3642–3652 (2011).
Yonezawa, H. & Kuramitsu, H. K. Genetic analysis of a unique bacteriocin, Smb, produced by Streptococcus mutans GS5. Antimicrob. Agents Chemother. 49, 541–548 (2005).
Jensen, R. O. et al. Differential recognition of Staphylococcus aureus quorum-sensing signals depends on both extracellular loops 1 and 2 of the transmembrane sensor AgrC. J. Mol. Biol. 381, 300–309 (2008).
Geisinger, E., Muir, T. W. & Novick, R. P. Agr receptor mutants reveal distinct modes of inhibition by staphylococcal autoinducing peptides. Proc. Natl. Acad. Sci. USA 106, 1216–1221 (2009).
Innelli, F., Oggioni, M. R. & Pozzi, G. Sensor domain of histidine kinase ComD confers competence pherotype specificity in Streptococcus pneumoniae . FEMS Microbiol. Lett. 252, 321–326 (2005).
Johnsborg, O., Kristiansen, P. E., Blomqvist, T. & Håvarstein, L. S. A hydrophobic patch in the competence-stimulating peptide, a pneumococcal competence pheromone, is essential for specificity and biological activity. J. Bacteriol. 188, 1744–1749 (2006).
Martin, B., Quentin, Y., Fichant, G. & Claverys, J.-P. Independent evolution of competence regulatory cascades in streptococci? Trend Microbiol. 14, 339–345 (2006).
Dawid, S., Roche, A. M. & Weiser, J. N. The blp bacteriocins of Streptococcus pneumoniae mediate intraspecies competition both in vitro and in vivo . Infect. Immun. 75, 443–451 (2007).
Alexeyev, M. F. & Winkler, H. H. Membrane topology of the Rickettsia prowazekii ATP/ADP translocase revealed by novel dual pho-lac reporters. J. Mol. Biol. 285, 1503–1513 (1998).
Zuobi-Hasona, K. & Brandy, L. J. Isolation and solubilization of cellular membrane proteins from bacteria. Methods Mol. Biol. 425, 287–293 (2008).
Cole, J. N., Djordjevic, S. P. & Walker, M. J. Isolation and solubilization of Gram-positive bacterial cell wall-associated proteins. Methods Mol. Biol. 425, 295–311 (2008).
Dong, G. F., Tian, X. L., Gomes, Z. A. & Li, Y. H. Regulated proteolysis of the alternative sigma factor SigX in Streptococcus mutans: implication in the escape from competence. BMC Microbiol. 14, 183–198 (2014).
Acknowledgements
We are grateful to Gouzel Karimova, Tarek Msadek and Daniel Ladant at Institute Pasteur, France, for the reporter plasmid pKTop. This work was supported in part by CIHR operating grant MOP-115007 and by NSERC discovery grant RGPIN311682. W.L. was the recipient of a NSERC Summer Studentship Award in 2012. K. C. was the recipient of a Faculty Summer Research Project Award in 2014.
Author information
Authors and Affiliations
Contributions
Y.-H.L., G.D. and X.-L.T. designed the experiments. G.D., X.-L.T., K.C., G.T. and T.L. performed the experiments. G.D., T.L., W.L. and Y.-H.L. did topology prediction, sequence alignments, primer design and data analysis. Y.-H.L. conceived the study and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Dong, G., Tian, XL., Cyr, K. et al. Membrane Topology and Structural Insights into the Peptide Pheromone Receptor ComD, A Quorum-Sensing Histidine Protein Kinase of Streptococcus mutans. Sci Rep 6, 26502 (2016). https://doi.org/10.1038/srep26502
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep26502
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.