Abstract
Metabolic markers in biofluids represent an attractive tool for guiding clinical management. The aim of this study was to identify metabolic mechanisms during the progress of pleural infection in children with Streptococcus pneumoniae pneumonia. Forty children diagnosed with pneumococcal pneumonia were enrolled and analysis of pleural fluid metabolites categorized by complicated parapneumonic effusions (CPE) and non-CPE was assessed by using 1H-NMR spectroscopy. Multivariate statistical analysis including principal components analysis (PCA) and partial least-squares discriminant analysis (PLS-DA) were performed. Metabolites identified were studied in relation to subsequent intervention procedures by receiver operating characteristic (ROC) curve analysis. Ten metabolites significantly different between CPE and non-CPE were identified. A significantly lower level of glucose for glycolysis was found in CPE compared to non-CPE. Six metabolites involving bacterial biosynthesis and three metabolites involving bacterial fermentation were significantly higher in CPE compared to non-CPE. Glucose and 3-hydroxybutyric acid were the metabolites found to be useful in discriminating from receiving intervention procedures. Metabolic profiling of pleural fluid using 1H-NMR spectroscopy provides direct observation of bacterial metabolism in the progress of pneumococcal pneumonia. An increase in the metabolism of butyric acid fermentation of glucose could potentially lead to the need of aggressive pleural drainage.
Similar content being viewed by others
Introduction
Pneumococcal pneumonia is a lung infection caused by Streptococcus pneumoniae and is the most common cause of community-acquired pneumonia (CAP)1. More than 40% of patients with bacterial pneumonia and 60% of patients with pneumococcal pneumonia develop parapneumonic effusions2. Infectious parapneumonic effusion accumulation is posited to be a continuing process of pleural inflammation resulting from the inflammatory response caused by pneumonia. Failing adequate therapy to control the pleural inflammation may progress from simple exudative effusion to empyema formation, requiring aggressive interventions3,4. An improved understanding the inflammatory process in the accumulated pleural fluid may provide a potential strategy for clinical management.
Biochemical analysis of infectious pleural fluid plays an important role in the management of parapneumonic effusions. Complicated parapneumonic effusions (CPE) are indicated by acidosis (pH ≤ 7.2) associated with raised lactate dehydrogenase (LDH ≥ 1000 U/L) and low glucose levels (glucose ≤ 40 mg/dL) and are more likely to require intervention procedures5,6. Despite this, proteins in pleural fluids originating from circulation or releasing locally from inflammatory or epithelial cells could also potentially be useful as markers of CPE for guiding clinical management7,8. A recent study in our laboratory has shown that a diagnostic model construction comprising three down-regulated biomarkers provides an alternative approach for discriminating CPE and subsequent surgical interventions9. The identification of a wide range of pleural fluid proteins related to inflammatory progress as biomarkers however is still challenging.
Inflammation in pleural fluids is characterized by a wide variety of cellular and molecular mediators resulting in a broad spectrum of possible metabolic products. Metabolomics using nuclear magnetic resonance (NMR) spectroscopy provides the advantage of studying a wide range of metabolites simultaneously and enables the discovery of small molecule metabolites by revealing any specific biomarker in human disease10,11. Metabolic markers in biofluids reflect alterations of metabolic fluxes of various organs and cells. Several studies have been reported to reveal metabolites in pleural fluids as biomarkers for exudative pleural effusions including lung cancer and pulmonary tuberculosis11,12. To date, there are, however, no studies addressing the metabolic changes in the host-microbe interaction of S. pneumoniae using infectious parapneumonic effusions.
The major aim of this study was to identify the metabolic profiles of infectious parapneumonic effusions by using 1H-NMR spectroscopy in children with pneumococcal pneumonia. The changes in pleural metabolites varies with inflammatory progress were assessed, and their relationships with subsequent intervention procedures were also examined.
Results
Population Characteristics
Forty children with pneumococcal pneumonia followed by parapneumonic effusions were consecutively enrolled into our study over a 4-year period. The mean age was 4.3 ± 2.7 years. The pleural fluid met the criteria of CPE (a pH ≤ 7.2, a LDH ≥ 1000 U/L and a glucose ≤40 mg/dL) were seen in 18 (45%) patients; whereas the fluid met the criteria of non-CPE were seen in 22 (55%) patients. The comparisons of pleural variables and characteristics of 40 children with pneumococcal pneumonia are shown in Table 1. A statistically significant higher rate of the identification of S. pneumoniase in pleural fluid (83% vs. 41%, P = 0.006) and the need of intervention procedures (100% vs. 32%, P < 0.001) were found in children with CPE compared to children with non-CPE.
Identification of Metabolite Sets between CPE and Non-CPE
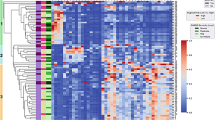
1H-NMR data of pleural fluid samples obtained were collected and analyzed. One thousand buckets varied across NMR spectra, 184 buckets of which corresponded to 25 of known metabolites (Supplementary Table S1). Unsupervised principal components analysis followed by examination of the first three principal components failed to reveal any clear separation between CPE and non-CPE. Metabolites contributing to the discrimination between CPE and non-CPE were identified by using supervised PLS-DA (Q2 = 0.15; R2 = 0.32). Metabolites selected by using the cutoff of PLS-DA VIP score >1.0 with a P value < 0.05 in the fold change of expression level between CPE and non-CPE are shown in Table 2. A more than 1.4-fold change in these metabolites was observed and volcano plot analysis was used to characterize the differential expressing metabolites between CPE and non-CPE (Supplementary Fig. S1A). These metabolites were further identified by comparison with reference spectra from the Human Metabolome Database (HMDB) and represented in a 2D 1H-13C NMR spectrum (Supplementary Fig. S1B). The two-dimensional graphs of PLS-DA score plots between CPE and non-CPE and a representative 600 MHz 1H-NMR spectra of selected metabolite signals are shown in Fig. 1. Figure 2 shows a heat map of these metabolites clustered using Hierarchical Clustering in the MetaboAnalyst 3.0. Among them, glucose was found to be significantly lower in CPE compared to non-CPE. In contrast, nine metabolites including tryptophan, thymine, phenylalanine, leucine/isoleucine and threonine in cluster 1, and lactic acid, 3-hydroxybutyric acid and succinic acid in cluster 2 were significantly higher in CPE compared to non-CPE.
(A) Two-dimensional scatter plot displays the model’s degree of separation between complicated parapneumonic effusions (CPE) and non-CPE. x axis, component 1 (% of total variance); y axis, component 2 (% of total variance) (B) Representative 600 MHz 1H-NMR spectra of pleural fluid showing the 10 metabolites with VIP scores greater than 1.0 with a P value < 0.05 by Mann-Whitney test (δ 0–8). x axis, parts per million (ppm); y axis, intensity (a.u.).
Metabolic Pathway and Function Analysis
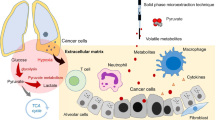
Metabolic pathway analysis with MetPA by filtering the dataset using a FDR-adjusted P value < 0.05 revealed that significantly different metabolites between CPE and non-CPE were important for amino acid and carbohydrate metabolisms (Table 3). Metabolites in cluster 1 were significantly associated with the biosynthesis of aminoacyl-tRNA and amino acids. Metabolites in cluster 2 were significantly related to carbohydrate metabolism responding for propanoate and butanoate metabolisms. Glucose in cluster 3 was predominantly associated with glycolysis or gluconeogenesis pathway (raw P < 0.05). Glucose dissimilation appeared to proceed via glycolysis and led to butyric acid and propionic acid fermentations, and biosynthesis of pyrimidines and amino acids. Figure 3 shows the metabolic pathways of metabolites significantly differentially expressed between CPE and non-CPE.
Metabolites for Discriminating CPE from Non-CPE and Subsequent Intervention Procedures
Table 4 shows the analysis of metabolites for discriminating CPE from non-CPE using receiver operating characteristic (ROC) curve analysis. Seven metabolites including glucose, lactic acid, succinic acid, 3-hydroxybutyric acid, thymine, threonine and tryptophan had the highest AUC significantly different from 0.5 (P < 0.05). After echo-guided thoracentesis, pleural metabolites of patients whose fever subsided (n = 15) and patients who received intervention procedures (n = 18) within 48 hours were compared and analyzed. Glucose (AUC = 0.781; 95% CI: 0.576–0.930; P = 0.008) and 3-hydroxybutyric acid (AUC = 0.734; 95% CI: 0.537–0.893; P = 0.032) were the metabolites found to be useful in discriminating from subsequent intervention procedures (Fig. 4).
Glucose (A); 3-Hydroxybutyric acid (B). AUC indicates the area under the curve and the dot refers to the cutoff value maximising sensitivity and specificity for the given samples. Box plots showing median and interquartile ranges of log transformed NMR intensity of metabolites over citric acid by subject groups.
Discussion
Infectious parapneumonic effusion is excess fluid that accumulates in the pleural cavity as a result of infections when lung tissues are damaged. With the progress of infection and inflammation in pleural fluids, metabolic changes in glucose associated with acidosis occur and require drainage for resolution. This study provides an overview of metabolic changes occurring in the infectious parapneumonic effusions and reveals potential metabolites for guiding management of infectious parapneumonic effusions in children with Streptococcus pneumonia pneumonia.
Complicated parapneumonic effusions (CPE) and empyema are well recognized complications in children with pneumococcal pneumonia13. Biochemical analysis of infectious pleural fluid, including pH, glucose, and LDH concentrations can significantly discriminate complicated from non-complicated parapneumonic effusions as in this study. In addition, the detection rate of S. pneumonia and number of neutrophils were found to be significantly higher in CPE compared to non-CPE. These findings indicate that CPE occur as a result of bacterial invasion into the pleural space that leads to an increased number of neutrophils and a series of metabolic processes.
Streptococcus pneumoniae is a Gram-positive diplococcus with a well-formed capsule and is recognized as a major cause of community-acquired pneumonia in children. The growth of pneumococci in the host depends on the catabolism of utilized carbon sources. The Embden-Meyerhof-Parnas (EMP) pathway for glucose dissimilation leading to pyruvate is the pathway at the center of metabolism in S. pneumonia14,15. The 13C isotopologue patterns in amino acids from labelled glucose have revealed the pathways to the de novo synthesized amino acids in S. pneumonia16. In this study, amino acids significantly increased during the progressive pleural inflammation were involved in the pathways to amino acid biosynthesis in S. pneumoniae, indicating that an increased pneumococcal invasion in CPE may lead to the aggressive metabolic processes of glucose consumption and subsequent biosynthesis of amino acids.
Streptococcus pneumoniae is a facultative anaerobe and fermentative microorganism. A facultative anaerobe is an organism that makes ATP by aerobic respiration if oxygen is present, but is capable of switching to fermentation or anaerobic respiration if oxygen is absent17. Under hypoxic conditions, pyruvate is mainly converted by LDH to lactic acid. Several studies have indicated that bacterial and cellular metabolism in pleural fluids cause the consumption of glucose and the excretion of lactate as in this study4,18,19. Increased propanoate and butanoate metabolisms along with significantly higher levels of succinic acid and 3-hydroxybutyric acid were also seen in CPE in this study. The anaerobic metabolic pathways for lactic acid, propionic and butyric acid fermentation of glucose leading to the excretion of protons and carbon dioxide (CO2) accumulation may result in a low pleural fluid glucose and pH.
Aggressive pleural drainage should be performed when the progression of CPE is not controlled and pleural fluid pH has been reported to be the best biochemical aid in predicting of CPE20. In this study, glucose and 3-hydroxybutyric acid were the metabolites useful for discriminating from CPE and subsequent intervention procedures, indicating that an increase in the metabolism of butyric acid fermentation of glucose with pleural fluid acidosis plays an important role in guiding management of infectious parapneumonic effusions caused by S. pneumonia. However, butyric acid is produced as end-product of a fermentation process solely performed by obligate anaerobic bacteria such as Clostridium spp21. Clinically, anaerobic bacteria contribute significantly to pleural infection, being identified as the sole or co-pathogen in 25–76% of pediatric cases22. In children with pneumococcal pneumonia, antibiotic choices should be informed by the results of blood and pleural fluid cultures and sensitivities; an anaerobic antibiotic choice may be considered for children with uncontrolled CPE and empyema.
A limitation of this study is the small sample size and a relatively low sensitivity of NMR-based metabolomic analysis than mass spectrometry-based methods. However, a combination of NMR and PLS-DA provides the advantage of studying a wide range of metabolites associated with bacterial and cellular pathways with accuracy. The overall comparability of pleural samples from non-CPE to CPE may result in the low R2 and Q2 scores in this study, further studies with a larger sample size with subgroup analysis by reducing within group variability may be needed to investigate more comprehensively. Despite this, a significant strength of the present study lies in its short time interval during which there were no major changes in surgical strategy for CPE. An age-matched comparison of children with CPE and non-CPE also makes the results demonstrated here are valid and potentially important.
In conclusion, metabolic profiling of pleural fluid using 1H-NMR spectroscopy provides new insight into the role of bacterial metabolism and physiology in the progress of pneumococcal pneumonia, and reveals putative biomarkers for requiring aggressive pleural drainage. An increased pneumococcal invasion in CPE may increase metabolic process of glucose consumption along with amino acid biosynthesis. A significantly increase in butyric acid fermentation of glucose in children with CPE receiving intervention procedures suggests that a mixed infection of anaerobic bacteria may contribute to the progress of pleural infection and antibiotics with anaerobic coverage should be considered to avoid further morbidity or mortality.
Methods
Study Population
The study population consisted of children who had pneumococcal pneumonia complicated by parapneumonic effusions requiring hospitalization. Streptococcus pneumoniae infection was defined by a positive result in blood or pleural fluid culture or the detection of antigens in the pleural fluid by latex agglutination testing23. Acute pneumococcal infection was also included for patients who had necrotic lung parenchyma with a positive urine test for S. pneumoniae (Binax, Portland, ME, USA). In general practice, antibiotic selection covered the likeliest organisms and adjusted after culture results were obtained. Intervention procedures for intercostal drainage including tube thoracostomy or video-assisted thoracoscopic surgery (VATS) were considered if patients had persistent fever >39 °C, dyspnea, and sepsis despite appropriate antibiotic therapy24. This study was approved by the Ethic Committee of Chang Gung Memory Hospital (No. 93-6299). All experiments in this study were performed in accordance with the relevant guidelines and regulations and written informed consent was obtained from the parents or guardians of all study subjects.
Pleural Fluid Collection
All subjects underwent a standard thoracocentesis procedure and pleural effusion samples were collected before intervention procedures. Pleural fluid was immediately analyzed for pH, total cell counts and differential cell count, and for glucose and LDH concentrations. Complicated parapneumonic effusions (CPE) were diagnosed if the pleural fluid met at least two of the following criteria: a pleural fluid pH of ≤7.2, a LDH level of ≥1000 U/L, and a glucose level of ≤40 mg/dL4,9. Otherwise, non-complicated parapneumonic effusions (non-CPE) were defined as patients whose pleural fluid met only one or none of the above criteria. Another 4 milliliters of specimen, if available, were mixed in Greiner Bio-One VACUETTE Coagulation Tubes with 3.2% sodium citrate anticoagulant solution (1 mL) in a ratio of 9:1 pleural fluid to citrate, which were immediately immersed in ice separately and centrifuged at 1,500 g for 10 min. The cell-free supernatant from each sample was stored at −80 °C until further use.
Sample Preparation
Pleural fluid samples collected from children with pneumococcal pneumonia were selected and examined. After thawing, to stabilize the pH value across samples prior to spectrum acquisition, 500 μL of pleural effusion was mixed with 250 μL of phosphate buffer (0.075 M Na2HPO4, pH 7.4) in deuterium water which containing 0.08% 3-(trimethylsilyl)-propionic-2,2,3,3-d4 acid sodium salt (TSP) as an internal chemical shift reference standard, 2 mM NaN3 as an inhibitor of bacterial contamination. Each sample was vortexed for 20 s and subsequently centrifuged at 12000 g at 4 °C for 30 min. After centrifugation, a 600 μL aliquot of the supernatant was transferred to a standard 5 mm NMR tube for analysis.
1H–Nuclear Magnetic Resonance (NMR) Spectroscopy
1H-NMR spectra were acquired on a Bruker Avance 600 MHz spectrometer (Bruker-Biospin GmbH, Karlsruhe, Germany) equipped with a 5 mm CPTCI 1H cryoprobe at Chang Gung Healthy Aging Research Center, Taiwan. Temperature was controlled at 300 K throughout the experiments. Low-power water pre-saturation pulse sequence was used for water signal suppression during the relaxation time of 4 s. For each spectrum, 64 scans were collected into 64K computer data points using a spectral width of 10,000 Hz (10 ppm). All 1D spectra were applied for analysis before Fourier transformation with zero-filled to exponential line-broadenings of 0.3 Hz. The acquired 1H-NMR spectra were manually phased, baseline-corrected, and referenced to the chemical shift of TSP (δ 0.0 ppm) using TopSpin 3.2 software (Bruker BioSpin, Rheinstetten, Germany).
NMR Data Processing and Analysis
The raw 1H-NMR spectra were imported into AMIX version 3.9.12 (Bruker BioSpin, Rheinstetten, Germany) for spectral bucket, spectral region exclusion and spectral normalization. 1H-NMR spectra were aligned on the TSP peak and normalized on the spectral area for calculating the concentration of each metabolite in the spectral peaks of each metabolite. The 1H-NMR spectra were subdivided into integrated regions of 0.01 ppm corresponding to the region of δ 0–10 ppm. Regions containing residual water (δ 4.825–4.725 ppm) were excluded from the data set to avoid spectral interference of residual water. Additional optimization of 1H-NMR spectra normalization was checked by using biochemical glucose concentration. The area of individual resonances of glucose metabolite was not significantly correlated with biochemical glucose concentration but the ratio of glucose/citric acid was (Supplementary Fig. S2). The spectra were therefore normalized to the integral of citric acid peak at δ 2.575–2.525 ppm to overcome the variation of pleural fluid amount when collecting samples. All metabolites in pleural fluid were identified using Chenomx NMR Suite 8.1 professional software (Chenomx Inc., Edmonton AB, Canada) with full resolution NMR data. A standard two-dimensional (2D) NMR experiment was conducted on a pooled pleural sample and metabolites were further assigned by comparison with reference spectra from the Human Metabolome Database (HMDB). For identifying metabolites contributing to the discrimination between groups, the normalized 1H-NMR bucket data were uploaded to MetaboAnalyst 3.0 (http://www.metaboanalyst.ca) for partial least squares-discriminant analysis (PLS-DA). All the NMR spectra were generalized log transformed (glog), method offered in MetaboAnalyst, to stabilize the variance across the spectral bins and to increase the weightings of the less intense peaks25. The spectral variables were mean-centered and scaled to unit variance, and 10-fold internal cross-validation was performed to evaluate the quality of the resulting statistical models by considering the diagnostic measures R2 and Q2 26. Potential metabolites were selected based on the Variable Importance in Projection (VIP) score greater than 1.0. The functional pathway analysis of potential biomarkers was based on the database source of the Kyoto Encyclopedia of Genes and Genomes (http://www.genome.jp/kegg/). Receiver Operating Characteristic (ROC) analysis of entire dataset was employed to measure the strength of metabolites for identifying potential biomarkers. The ROC curve analysis was performed using the online ROCCET (ROC curve explorer and tester, http://www.roccet.ca)27.
Statistical Analysis
Distribution of baseline characteristics and pleural variables of children with pneumococcal pneumonia categorized by CPE and non-CPE was done with univariable parametric and non-parametric tests such as Student’s t-test, Mann-Whitney test, χ2, and Fisher’s exact test. Hierarchical clustering was performed and heat maps were created based on the Pearson correlation and the Ward clustering algorithm. The fold changes, volcano plots, and statistical significance in metabolites between CPE and non-CPE were performed by the non-parametric Mann-Whitney test using MetaboAnalyst web server. Statistical analysis was performed by using the Statistical Program for Social Sciences (IBM SPSS Statistics for Windows, Version 20.0; IBM, Armonk, NY, USA). All statistical hypothesis tests were two tailed and a P value of less than 0.05 was considered to be significant.
Additional Information
How to cite this article: Chiu, C.-Y. et al. Metabolomic Profiling of Infectious Parapneumonic Effusions Reveals Biomarkers for Guiding Management of Children with Streptococcus pneumoniae Pneumonia. Sci. Rep. 6, 24930; doi: 10.1038/srep24930 (2016).
References
Tan, T. Q. Pediatric invasive pneumococcal disease in the United States in the era of pneumococcal conjugate vaccines. Clin Microbiol Rev 25, 409–419 (2012).
Lin, T. Y. et al. Etiology of empyema thoracis and parapneumonic pleural effusion in Taiwanese children and adolescents younger than 18 years of age. Pediatr Infect Dis J 32, 419–421 (2013).
Hamm, H. & Light, R. W. Parapneumonic effusion and empyema. Eur Respir J 10, 1150–1156 (1997).
Chiu, C. Y. et al. Proinflammatory cytokines, fibrinolytic system enzymes, and biochemical indices in children with infectious para-pneumonic effusions. Pediatr Infect Dis J 27, 699–703 (2008).
Heffner, J. E., Brown, L. K., Barbieri, C. & DeLeo, J. M. Pleural fluid chemical analysis in parapneumonic effusions. A meta-analysis. Am J Respir Crit Care Med 151, 1700–1708 (1995).
Light, R. W. A new classification of parapneumonic effusions and empyema. Chest 108, 299–301 (1995).
Nilsson, C. L., Puchades, M., Westman, A., Blennow, K. & Davidsson, P. Identification of proteins in a human pleural exudate using two-dimensional preparative liquid-phase electrophoresis and matrix-assisted laser desorption/ionization mass spectrometry. Electrophoresis 20, 860–865 (1999).
Tyan, Y. C., Wu, H. Y., Su, W. C., Chen, P. W. & Liao, P. C. Proteomic analysis of human pleural effusion. Proteomics 5, 1062–1074 (2005).
Chiu, C. Y. et al. The value of total protein in guiding management of infectious parapneumonic effusion by using matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. J Microbiol Immunol Infect 48, 483–489 (2015).
Muller, C. et al. Nuclear magnetic resonance spectroscopy with single spin sensitivity. Nat Commun 5, 4703 (2014).
Lam, C. W. & Law, C. Y. Untargeted mass spectrometry-based metabolomic profiling of pleural effusions: fatty acids as novel cancer biomarkers for malignant pleural effusions. J Proteome Res 13, 4040–4046 (2014).
Lam, C. W. & Law, C. Y. Pleural effusion lipoproteins measured by NMR spectroscopy for diagnosis of exudative pleural effusions: a novel tool for pore-size estimation. J Proteome Res 13, 4104–4112 (2014).
Antachopoulos, C. et al. Parapneumonic pleural effusions caused by Streptococcus pneumoniae serotype 3 in children immunized with 13-valent conjugated pneumococcal vaccine. Pediatr Infect Dis J 33, 81–83 (2014).
Hoskins, J. et al. Genome of the bacterium Streptococcus pneumoniae strain R6. J Bacteriol 183, 5709–5717 (2001).
Tettelin, H. et al. Complete genome sequence of a virulent isolate of Streptococcus pneumoniae. Science 293, 498–506 (2001).
Hartel, T. et al. Characterization of central carbon metabolism of Streptococcus pneumoniae by isotopologue profiling. J Biol Chem 287, 4260–4274 (2012).
Werno, A. M. & Murdoch, D. R. Medical microbiology: laboratory diagnosis of invasive pneumococcal disease. Clin Infect Dis 46, 926–932 (2008).
Cheng, D. S. et al. Comparison of pleural fluid pH values obtained using blood gas machine, pH meter, and pH indicator strip. Chest 114, 1368–1372 (1998).
Utine, G. E. et al. Childhood parapneumonic effusions: biochemical and inflammatory markers. Chest 128, 1436–1441 (2005).
Chiu, C. Y., Wong, K. S., Huang, Y. C., Lai, S. H. & Lin, T. Y. Echo-guided management of complicated parapneumonic effusion in children. Pediatr Pulmonol 41, 1226–1232 (2006).
Van Immerseel, F. et al. Butyric acid-producing anaerobic bacteria as a novel probiotic treatment approach for inflammatory bowel disease. J Med Microbiol 59, 141–143 (2010).
Rosenstengel, A. Pleural infection-current diagnosis and management. J Thorac Dis 4, 186–193 (2012).
Song, J. Y., Eun, B. W. & Nahm, M. H. Diagnosis of pneumococcal pneumonia: current pitfalls and the way forward. Infect Chemother 45, 351–366 (2013).
Bradley, J. S. et al. The management of community-acquired pneumonia in infants and children older than 3 months of age: clinical practice guidelines by the Pediatric Infectious Diseases Society and the Infectious Diseases Society of America. Clin Infect Dis 53, e25–76 (2011).
Durbin, B. P., Hardin, J. S., Hawkins, D. M. & Rocke, D. M. A variance-stabilizing transformation for gene-expression microarray data. Bioinformatics 18 Suppl 1, S105–110 (2002).
Westerhuis, J. A. et al. Assessment of PLSDA cross validation. Metabolomics 4, 81–89 (2008).
Xia, J., Broadhurst, D. I., Wilson, M. & Wishart, D. S. Translational biomarker discovery in clinical metabolomics: an introductory tutorial. Metabolomics 9, 280–299 (2013).
Acknowledgements
This work was supported by CMRPG2C0321-2 from the Chang Gung Memorial Hospital, Taiwan. We are extremely grateful for all support of the metabolomics analysis using 600 MHz NMR spectroscopy as tools carried out at the Metabolomics Core Laboratory, Healthy Aging Research Center (HARC), Chang Gung University and Clinical Phenome Center, Chang Gung Memorial Hospital.
Author information
Authors and Affiliations
Contributions
C.-Y.C., K.-S.W. and S.-Y.H. designed and supervised the study. G.L., M.-L.C. and M.-H.C. performed experiments and analyzed the data. M.-H.T. and S.-H.L. developed and provided reagents. All authors discussed the results and reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Chiu, CY., Lin, G., Cheng, ML. et al. Metabolomic Profiling of Infectious Parapneumonic Effusions Reveals Biomarkers for Guiding Management of Children with Streptococcus pneumoniae Pneumonia. Sci Rep 6, 24930 (2016). https://doi.org/10.1038/srep24930
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep24930
This article is cited by
-
Metabolomics in pediatric lower respiratory tract infections and sepsis: a literature review
Pediatric Research (2023)
-
Common pathogenic bacteria-induced reprogramming of the host proteinogenic amino acids metabolism
Amino Acids (2023)
-
Metabolomics profile in acute respiratory distress syndrome by nuclear magnetic resonance spectroscopy in patients with community-acquired pneumonia
Respiratory Research (2022)
-
Metabolic preference assay for rapid diagnosis of bloodstream infections
Nature Communications (2022)
-
Integrated metabolic and microbial analysis reveals host–microbial interactions in IgE-mediated childhood asthma
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.