Abstract
The H-type of atypical bovine spongiform encephalopathy (H-BSE) was serially passaged in bovinized transgenic (TgBoPrP) mice. At the fourth passage, most challenged mice showed a typical H-BSE phenotype with incubation periods of 223 ± 7.8 days. However, a different phenotype of BSE prion with shorter incubation periods of 109 ± 4 days emerged in a minor subset of the inoculated mice. The latter showed distinct clinical signs, brain pathology and abnormal prion protein profiles as compared to H-BSE and other known BSE strains in mice. This novel prion was transmitted intracerebrally to cattle, with incubation periods of 14.8 ± 1.5 months, with phenotypes that differed from those of other bovine prion strains. These data suggest that intraspecies transmission of H-BSE in cattle allows the emergence of a novel BSE strain. Therefore, the continuation of feed ban programs may be necessary to exclude the recycling of H-BSE prions, which appear to arise spontaneously, in livestock. Such measures should help to reduce the risks from both novel and known strains of BSE.
Similar content being viewed by others
Introduction
Prions cause transmissible spongiform encephalopathies (TSEs), which are characterized by a spongiform change in the central nervous system and the accumulation of an abnormal prion protein (PrPSc). PrPSc is a disease-associated isoform of the host-encoded prion protein1 and the conversion of the normal isoform (PrPC) to PrPSc is thought represent a central event in prion pathogenesis. PrPSc has been recognized as the major component of prions2 and variations in PrPSc are associated with different prion strains that, in turn, cause distinct disease phenotypes3.
Bovine spongiform encephalopathy (BSE) is a TSE of cattle. Most BSE cases show a unique phenotype that is thought to be caused by a single prion strain4. However, other atypical neuropathological and molecular phenotypes of BSEs (atypical BSEs) have been identified in aged animals5,6. Atypical BSEs have been classified into two groups, L-BSE and H-BSE, based on their biological and biochemical features5,6, which differ from those of classical BSE (C-BSE).
The worldwide occurrence of BSE is declining, primarily because of effective feed ban programs6. The emergence of atypical BSE cases has raised questions as to whether additional and/or modified control measures might be needed. Thus, characterization of atypical BSE prions is necessary for risk evaluation.
H-BSE was first reported in France7 and has been subsequently detected in several European countries and North America5,6. In one of two H-BSE cases detected in the U.S., a prion protein (PrP) amino-acid substitution that is associated with familial Creutzfeldt–Jakob disease in humans has been reported. This suggested the possibility that H-BSE might occur spontaneously or sporadically8. Regardless of its origin, previous studies have shown that H-BSE is transmissible to cattle9,10,11,12, as well as to wild type13,14, bovinized and ovinized PrP transgenic mice that express murine, bovine, or ovine PrPC, respectively15,16,17. Those studies also revealed different characteristics of H-BSE relative to C-BSE and L-BSE.
Serial passages of H-BSE in wild type18,19 and bovinized PrP transgenic mice20 can lead to the emergence of a C-BSE-like phenotype. This suggests that H-BSE-affected cattle harbor heterogeneous prions and that structural variants of H-BSE might generate a C-BSE-like phenotype. This, in turn, raises the possibility that C-BSE originated from H-BSE. To attempt to clarify the origin of C-BSE, we serially passaged H-BSE in bovinized PrP transgenic (TgBoPrP) mice. In contrast to previous studies, we did not detect C-BSE-like prions during serial passages. However, a novel BSE prion—different from C-BSE, L-BSE and H-BSE prions—was detected in a subset of inoculated mice. When this novel prion was inoculated intracerebrally into cattle, a novel BSE phenotype was confirmed. This study suggests that a novel BSE emerges during intraspecies transmission of H-BSE in cattle.
Results
Serial transmission of H-BSE in TgBoPrP mice
The results of serial transmission of the H-BSE isolate in TgBoPrP mice are shown in Table 1. All the H-BSE challenged mice developed progressive neurological disease with the incubation periods of 320.1 ± 12.2 days at primary passage. H-BSE-affected animals showed a distinctive clinical sign, namely, constant chewing of the bedding, as reported previously16. The incubation periods of the second and third passages were 226.9 ± 4.2 and 215.6 ± 5.0 days, respectively (Table 1)16. No clear differences were observed in their clinical signs, the banding pattern of PrPSc and histopathological features from the primary to third passage mice. At the fourth passage, mice from a single experimental group (#3), out of eight experiments, showed shorter incubation periods (108.8 ± 4.0 days) than the other groups. This group was challenged with brain homogenates of a mouse with 221-day incubation period. Group #3 animals showed weight loss, but no constant chewing of the bedding. This short incubation-type of BSE was designed BSE-SW (short incubation with weight loss) strain. BSE-SW was transmitted to TgBoPrP mice with 97.3 ± 3.7 day incubation periods and their clinical signs were identical to those of mice in the experimental group #3. The other mice in the fourth passage groups showed the symptomatic chewing of the bedding and their incubation periods were 223.3 ± 7.8 days (Table 1).
Molecular features of PrPcore of BSE-SW
Western blot analysis with monoclonal antibody (mAb) 6H4 revealed that the molecular mass of proteinase K (PK) digested PrPSc (PrPcore) of BSE-SW was lower than that of H-BSE, but similar to C-BSE (Fig. 1b). MAb P4, which recognizes the N-terminal region of PrPcore, reacted with H-BSE but not with BSE-SW or C-BSE (Fig. 1a). This revealed a partial similarity between PrPSc of BSE-SW and C-BSE. In contrast, truncated 12-kDa fragments (consistent with PrP 157/163–23114) were observed for H-BSE and BSE-SW (Fig. 1c). Deglycosylation analysis confirmed these results. Unglycosylated PrPcore with a molecular weight of approximately 17 kDa (PrPcore #114) was detected in C-BSE and BSE-SW, whereas a slightly higher molecular weight (approximately 19 kDa) of PrPcore #1 was detected in H-BSE. Unglycosylated PrPcore with a molecular weight of 12 kDa (PrPcore #214) was observed in H-BSE and BSE-SW (Fig. 1d). On the other hand, no difference was observed in the glycoprofiles of BSE-SW, H-BSE and C-BSE, as detected by mAb 6H4 (Fig. 1e). These biochemical properties of BSE-SW were maintained in the subsequent passages (data not shown).
Western blot analysis of PrPcore.
PrPcore from BSE prion-affected TgBoPrP mice was detected using mAbs P4 (a), 6H4 (b) and SAF84 (c). Lane 1, H-BSE; Lane 2, BSE-SW; Lane 3, C-BSE; Lane 4, L-BSE. All the samples were digested with 50 μg/ml PK at 37 °C for 1 h. Digested aliquots were treated with PNGaseF and probed with mAb SAF84 (d). Equivalents of 0.125 mg brain tissue were loaded. PrPcore of BSE-SW showed a different reaction with mAbs compared to H-BSE, as well as a distinct band pattern and molecular weight of PrPcore fragments. Molecular markers are shown on the left (kDa). Glycoform profile of PrPcore is given (e). Relative amounts of diglycosylated (solid black bar), monoglycosylated (gray bar) and unglycosylated (clear bar) forms of PrPcore from BSE-affected TgBoPrP mice. The signals were detected using mAb 6H4 and the results are presented as mean ± standard deviation of nine experiments. No significant differences were observed between H-BSE and BSE-SW in the glycoform ratio of PrPcore.
Neuropathological examination
The degree of brain vacuolation and neuroanatomical distribution patterns of PrPSc in TgBoPrP mice with BSE-SW were different from H-BSE and C-BSE (Fig. 2). The lesion scores of BSE-SW were lower than H-BSE scores in most brain areas, with the exception of dorsal medulla, cerebellar cortex and thalamus (Fig. 2a). Regarding the PrPSc deposition and distribution patterns in BSE-SW, punctuate and granular PrPSc deposition was observed in the septal nucleus, corpus callosum, habenular nucleus, hypothalamus, superior colliculus, mesencephalic tegmentum, vestibular nucleus, cochlear nucleus and reticular formation of the brainstem. Relatively heavy deposition of PrPSc, consisting mainly of intraglial and intraneuronal staining, was conspicuous in the white matter of the cerebellum of BSE-SW in comparison to H-BSE (Fig. 2e,f,k,l). The PrPSc staining was minimal in the cerebral and cerebellar cortex (Fig. 2c). PrP-plaques in the subcallosal region were less frequent in BSE-SW than in H-BSE (Fig. 2h,i). In contrast, the most conspicuous feature of PrPSc staining type in TgBoPrP mice infected with H-BSE comprised heavily deposited plaques and/or large aggregates, which mainly located in the subcallosal and adjacent periventricular area of the brain (Fig. 2h). Such plaque deposits were occasionally detected in the cerebral and cerebellar cortex, but were not present in the thalamus or the brainstem. The plaques displayed birefringence with Congo red under polarized light (data not shown). The minimal PrPSc staining in the cerebral and cerebellar cortex was similar to BSE-SW (Fig. 2b). Furthermore, the PrPSc staining pattern in C-BSE was completely different from that in BSE-SW (Fig. 2d,j,g,m). These results indicate that neuropathological properties of BSE-SW in TgBoPrP mice were apparently distinct from those of H-BSE and C-BSE.
Neuropathological analysis of TgBoPrP mice.
Lesion profiles (a), neuroanatomical distribution of PrPSc detected by PET-blot (b–g) and PrPSc staining types assessed by IHC (h–m) in the brains of BSE prion-affected TgBoPrP mice. Vacuolation in the following brain regions was scored, on a scale of 0–4 (mean values): 1, dorsal medulla; 2, cerebellar cortex; 3, superior cortex; 4, hypothalamus; 5, thalamus; 6, hippocampus; 7, septal nuclei; 8, cerebral cortex at the level of the hypothalamus and thalamus; and 9, cerebral cortex at the level of the caudate nuclei. The data are presented as mean ± standard deviation (n = 7). Black circles, H-BSE; Diamonds, BSE-SW; Black triangles, C-BSE. The degree of vacuolation in the brain of BSE-SW was different from H-BSE and C-BSE (a). PET blots with mAb SAF84 corresponding to the brain areas at the level of thalamus (b–d) and the level of medulla and cerebellum (e–g) are shown. IHC was performed with mAb F99/97.6.1. CC, corpus callosum; HB, habenular nucleus; PC, parietal cortex; TC, temporal cortex; H, hippocampus; T, thalamus; HT, hypothalamus; AM, amygdala; DMNV, dorsal motor nucleus of vagus nerve; NC, deep nuclei of the cerebellum. PrPSc deposits and distribution patterns in the BSE-SW brain were distinct from H-BSE and C-BSE.
Conformational stability studies
MAbs 6H4 and SAF84 were used to conduct conformation stability studies. Western blotting with mAb 6H4 showed that the [GdnHCl]1/2 values (see Methods Section) for PrPSc denaturation in the H-BSE and BSE-SW brains were 3.8 ± 0.4 M and 3.0 ± 0.2 M, respectively (Fig. 3e). This analysis detected PrPcore #1 signal of 17–27 kDa from BSE-SW and 19–29 kDa from H-BSE (Fig. 3a,c). PrPcore #1 from BSE-SW was more sensitive to GdnHCl treatments than that of H-BSE. A different result was observed with mAb SAF84, which detected both PrPcore #1 and PrPcore #2 (Fig. 3b,d). [GdnHCl]1/2 values of H-BSE and BSE-SW were 3.5 ± 0.3 M and 3.4 ± 0.3 M, respectively (Fig. 3f). These values were not significantly different. Signal intensity of the diglycosylated PrPcore #1 of BSE-SW was weaker than that of H-BSE during 3.0–3.5 M treatment (Fig. 3d). This result was consistent with the mAb 6H4 experiment (Fig. 3c). However, the lower three bands of PrPcore #2 showed similar signal intensities during 3.5–4.0 M treatment (Fig. 3b,d). This revealed that the conformational stability of PrPcore #1, but not PrPcore #2, is different in PrPSc from H-BSE and BSE-SW.
Conformational stability assay.
The conformation stability of PrPSc from BSE-SW and H-BSE. GdnHCl was added at the indicated concentrations and the samples subjected to PK digestion (20 μg/ml). PrPSc was detected with mAbs 6H4 and SAF84. Molecular weights are shown on the left (kDa). [GdnHCl]1/2 concentration (M) was calculated based on denaturation curves obtained from densitometric analysis of western blot data. The results are mean ± standard deviation from five experiments. The black and white bars indicate H-BSE and BSE-SW, respectively.
Transmission of BSE-SW to cattle
To examine whether the BSE-SW prion could become a threat as a novel prion disease in cattle, three Holstein calves were challenged intracerebrally. All the inoculated animals developed progressive neurological disease. The animals exhibited initial clinical signs of the disease between 11.5 and 12.5 months post-inoculation (mpi), which included disturbance, mild fear or anxiety, mild gait changes, and, occasionally, low head carriage. After one to two months of the initial clinical signs, the animals were leaning towards the floor and rested their heads against the wall, which was followed by ataxia of the hind limbs that progressed to difficulty in getting up without assistance at the clinically terminal stage of the disease. The bodily condition gradually worsened because of a loss of weight during the two to three month clinical duration. None of the animals exhibited anorexia, nervousness, or aggression and responded to visual, acoustic and tactile stimuli throughout the course of the disease. The cattle were eventually culled at 13.3 mpi, 15 mpi and 16.2 mpi before astasia, in accordance with the welfare guidelines for animal experiments. The incubation periods of cattle infected with BSE-SW (14.8 ± 1.5 mpi) were shorter than those for H-BSE, C-BSE and L-BSE (Table 2). Obex samples from these cattle were subjected to routine BSE confirmatory tests.
Histopathology and PrPSc immunohistochemistry of the obex of BSE-SW-affected cattle
Mild vacuolation of the extracellular neuropil was observed in the dorsal motor nucleus of the vagus nerve (DMNV), the solitary nucleus, the nucleus of trigeminal nerve spinal tract and the olivary nucleus in all animals. No intraneuronal vacuolation was seen. Spongy changes were not prominent in the gray matter of medulla oblongata at the obex (Fig. 4a). Immunolabeling of PrPSc with mAb F99/97.6.1 resulted in intraneuronal and intraglial patterns throughout the obex (Fig. 4b). Intraneuronal labeling was less common in DMNV and the hypoglossal nucleus compared to other nuclei. Fine and coarse granular PrPSc was sparsely distributed throughout the neuropil of reticular formation. No other extracellular types of PrPSc, such as plaque-like and stellate deposits, were identified in the obex region.
Molecular features of PrPcore of BSE-SW-affected cattle
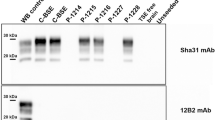
Western blot analysis detected PrPSc from the obex tissue of the challenged cattle. The molecular features of PrPcore of BSE-SW-affected cattle were distinctly different from C-BSE, L-BSE and H-BSE (Fig. 5). The molecular mass of PrPcore #1 of BSE-SW, as determined by mAb 6H4, was lower than H-BSE and similar to C-BSE (Fig. 5b). MAb P4 did not detect PrPcore of BSE-SW (Fig. 5a). PrPcore #2 was also observed in BSE-SW (Fig. 5c,d). These results revealed that the biochemical properties of BSE-SW have indeed been transmitted to cattle.
Western blot analysis of PrPcore from BSE-SW-affected cattle.
PrPcore from the affected cattle were detected using mAbs: P4 (a), 6H4 (b) and SAF84 (c). Lane 1, H-BSE; Lane 2, BSE-SW; Lane 3, C-BSE; Lane 4, L-BSE. All the samples were digested with 40 μg/ml PK at 37 °C for 1 h. Digested aliquots were treated with PNGaseF and were probed using mAb SAF84 (d). Equivalents of 0.5 mg brain tissues were loaded. Molecular markers are shown on the left (kDa).
Discussion
Our previous reports have revealed the usefulness of TgBoPrP mice for characterizing BSE prions16,21,22. In this study, a novel BSE, BSE-SW, was detected using this mouse model. The incubation periods of H-BSE, L-BSE and C-BSE prions in the TgBoPrP mice were approximately 215 days, 150 days and 190 days, respectively16,21,22. The BSE-SW prion showed the shortest incubation period (approximately 90 days) among the known BSE prions. We have previously performed several transmission experiments of sheep scrapie to TgBoPrP mice, but their incubation periods were over 170 days23 and we have not observed any prions with ~90-day incubation periods in these mice. In addition, the biochemical and biological properties of PrPSc from BSE-SW were clearly different from C-BSE, L-BSE, H-BSE and sheep scrapie (data not shown). PrPSc of BSE-SW has some similarity to H-BSE on the account of the presence of truncated 12-kDa fragments (PrPcore #2). Fig. 6 shows the putative PK digestion site of PrPSc from BSE-SW, as assessed by immunoreactivity with mAbs P4, 6H4 and SAF84. These results argue against the possibility that the BSE-SW prion resulted from a contamination of other laboratory prion strains.
Schematic representation of PrPcore of BSE prions.
Putative PK cleavage sites and PrPcore are shown, as estimated by immunoreactivity with mAbs P4, 6H4 and SAF84. PrPcore #1 is an unglycosylated PrPcore with a molecular weight of 17–19 kDa. PrPcore #2 is an unglycosylated PrPcore with a molecular weight of 12 kDa.
It is known that sheep scrapie comprises different prion strains24,25 and some affected sheep harbor these mixed scrapie prion strains3. Numerous scrapie strains had emerged in the course of several passage histories26. We have also previously isolated distinct scrapie strains from the brain of a scrapie-affected sheep after primary passage in wild type mice27. The different scrapie prion strains appeared after primary passage in sheep scrapie cases, which was considered to be due to prion strain selection. For H-BSE prions, French and Polish cases were reported, which transformed into a C-BSE-like phenotype during mouse passages18,19,20, revealing their potential heterogeneity. However, the BSE-SW prions described herein appear to have emerged by a different manner than our previous scrapie case27. The average incubation period of H-BSE in TgBoPrP mice was 320 days at first passage and then shortened to 227 days and 216 days at secondary and third passages, respectively. Based on this observation, H-BSE prions were adapted to TgBoPrP mice at second passage. By the third passage, no multiple prion strains in the H-BSE material were evident. BSE-SW prion with a shorter incubation period emerged after H-BSE has adapted to TgBoPrP mice. The results of this study suggest that BSE-SW emerged through a conformational rearrangement. Alternatively, the BSE-SW prion was only present in a very small titer in H-BSE and required several passages in sensitive animals to emerge after selection from a mixture of preexisting prion strains. The Canadian H-BSE sample that was used in this study was also used to inoculate wild type mice, but we did not observe the emergence of a C-BSE-like phenotype in those circumstances (unpublished data). The underlying mechanisms of the emergence of new prion strains are important to elucidate prion heterogeneity. This novel BSE strain has never been observed in field BSE cases, but our experiments reveal the potential risk associated with H-BSE.
It has been suggested that different conformations of PrPSc are involved in the prion strain diversity28 and that rearrangement of PrPSc from a uniform conformation causes the emergence of new host-adapted PrPSc 3. PrPSc of BSE-SW exhibited different conformational stability from H-BSE. It is also known that strain “mutation or transformation” may occur upon intraspecies transmission, where the PrP amino acid sequences of the host and the donor are identical3. These finding are consistent with the conclusion that PrPSc of BSE-SW has a different conformation than H-BSE.
Furthermore, our new strain was successfully transmitted to cattle. Standard diagnostic testing for BSE confirmed the presence of spongiform changes associated with PrPSc accumulation in the obex and the challenged cattle fulfilled BSE criteria (Figs 4 and 5). The disease phenotype and features of PrPSc, different from the known types of BSE, indicated that this prion could cause a novel type of atypical BSE. The shorter incubation periods in cattle were consistent with the relative incubation periods in TgBoPrP mice and indicate high virulence of this novel prion. Further analysis of diseased cattle is necessary to clarify its characteristics. Such studies could help to elucidate the mechanisms of conformational change in PrPSc, which lead to the propagation of new prion strains.
The ban on meat-and-bone meal in livestock feed has contributed to the decline in C-BSE occurrences6. Recently, easing of the BSE-related regulations and control measures has been discussed. The origin of atypical BSE remains unknown, but it has been proposed to be spontaneous or sporadic29. H-BSE has been reported to transform into a C-BSE-like phenotype during animal passages18,19,20. Furthermore, we have shown here that the sequential transmission of H-BSE in TgBoPrP mice, i.e., to a homologous bovine PrP context, generated a novel type of BSE. Considering these observations, a continuous feed ban program may be necessary even after C-BSE is eradicated. Prohibiting the recycling of spontaneously occurring H-BSE prions in cattle should help to prevent both re-emerging and emerging types of BSEs.
Methods
Ethics statement
Procedures involving animals have been approved by the Animal Care and Use Committee at the National Institute of Animal Health (approval ID: 11-008, 13-005). Animal experiments were performed in accordance with the Guidelines for Animal Transmissible Spongiform Encephalopathy Experiments of the Ministry of Agriculture, Forestry and Fisheries of Japan.
Transgenic mice
TgBoPrP mice overexpressing the bovine PrP gene (encoding BoPrP) in a mouse PrP deficient background were used. These mice harbored the cattle PrP gene containing six copies of the octarepeat sequence (EMBL, X55882) and produced approximately eight times more BoPrP per gram of protein than found in the cattle brain30.
BSE material
Brain samples of Canadian H-BSE cattle, courteously provided by Dr. S. Czub, were used in this study31. Mouse-passaged L-BSE and C-BSE prions were also used. These prions were routinely maintained by serial passaging into TgBoPrP mice, as described previously21,22. C-BSE and L-BSE prions were adapted to TgBoPrP mice by serial passaging and their incubation periods were approximately 190 days and 150 days, respectively. Brain samples from C-BSE, L-BSE and H-BSE-affected cattle were also used10,32,33.
Mouse transmission study
Brain tissues from BSE-affected animals were homogenized in nine volumes of phosphate buffered saline (PBS) using a multi-bead shocker (Yasui Kikai) and centrifuged at 1,000 × g for 5 min at room temperature (RT). Three-week-old female TgBoPrP mice were inoculated intracerebrally with 20 μl supernatant. Following inoculation, clinical status of the mice was monitored daily to assess the onset of neurological signs. The brains of diseased mice were removed and stored at -80 °C for biochemical analysis or fixed for histopathology. For the fourth passage, eight diseased mice brains from the third passage were used to independently challenge to TgBoPrP mice.
Cattle transmission study
Three female 3-4-month-old Holstein calves were challenged intracerebrally with 1 ml of 10% brain homogenate of BSE-SW-affected TgBoPrP mice, as described previously10. Animals were euthanized before ataxia deterioration. The brains of diseased cattle were removed and stored at −80 °C for western blotting analysis or fixed for pathological examination (i.e., BSE confirmatory tests).
Extraction of PrPSc from BSE-affected TgBoPrP mice and cattle
Brain tissues were homogenized in PBS (20% homogenate, w/v) using a multi-bead shocker. The brain homogenate (125 μl) was mixed with an equal volume of buffer containing 4% (w/v) Zwittergent 3–14 (Calbiochem), 1% (w/v) Sarkosyl, 100 mM NaCl and 50 mM Tris-HCl (pH 7.6) and incubated with 6.25 μl of 40 mg/ml collagenase solution. The samples (50 μg/ml for mice brain tissues and 40 μg/ml for cattle brain tissues) were then subjected to PK (Roche Diagnostic) digestion at 37 °C for 1 h to detect PK-resistant PrPSc fragments (PrPcore). PK digestion was terminated with 2 mM 4-(2-aminoethyl)benzenesulfonyl fluoride hydrochloride (Pefabloc; Roche Diagnostic). The samples were mixed with equal volumes of 2-butanol:methanol mixture (5:1) and centrifuged at 20,000 × g for 10 min. The pellets were resuspended in gel-loading buffer containing 2% (w/v) SDS and were then boiled for 10 min before western blotting. After PK treatment, some samples were deglycosylated with N-glycosidase F (PNGaseF; New England Biolabs), following the manufacturer’s instruction.
Antibodies
The following monoclonal antibodies (mAbs) against PrP were used in this study: P4 (R-Biopharm), 6H4 (Prionics), SAF84 (SPI-bio) and F99/97.6.1 (VMRD). MAb P4 recognizes amino acid residues 101–107 of bovine PrP sequence34. MAbs 6H4 and SAF84 recognize amino acid residues 156–16335 and 175–18036, respectively, of the bovine PrP sequence. F99/97.6.1 recognizes C-terminus of PrP, amino acid residues 229–23210.
Western blot analysis
Samples were separated by SDS-PAGE and blotted electrically onto a PVDF membrane (Millipore). The blotted membrane was incubated with mAbs P4, 6H4 and SAF84 at RT for 1 h. MAb binding was detected by horseradish peroxidase-conjugated anti-mouse IgG. Signals were developed with a chemiluminescent substrate (SuperSignal; Pierce Biotechnology).
Neuropathology, immunohistochemistry and PET-blot analysis
Histopathological analysis of TgBoPrP mice and cattle was performed according to a previously described method10,16,21. Briefly, the brains were fixed in 10% buffered formalin solution (pH 7.4) containing 10% methanol. The formalin-fixed brains were immersed in 98% formic acid and embedded in paraffin wax. Sections (4 μm thick) were cut and stained with hematoxylin and eosin (HE). The lesion profile was determined by scoring the vacuolar changes in nine standard grey matter areas, as previous described37. For PrPSc immunohistochemistry (IHC), sections were incubated with mAbs SAF84 or F99/97.61, followed by incubation with anti-mouse universal immunoperoxidase polymer (Histofine Simple Stain MAX-PO (M), Nichirei) as the secondary antibody and visualized using 3,3′-diaminobenzidine tetrachloride as the chromogen. Finally, the sections were counterstained with hematoxylin. Paraffin embedded tissue (PET) blot was performed as described previously21. Dewaxed membranes were treated with 80 μg/ml of PK for 30 min at 37 °C and then denatured using 3 M guanidine thiocyanate for 10 min at RT. The blotted membranes were incubated with mAb SAF84 for 90 min at RT. Signals were detected using Histofine MAX-AP (M) Kit (Nichirei) with 5-bromo 4-chloro-3-indolyl phosphate and nitro blue tetrazolium (BCIP/NBT; Roche Diagnostics) as substrates.
Conformational stability assay
Conformation stability assay was performed according to a previously described method with minor modification38. Briefly, 50 μl of 10% brain homogenate were added to an equal volume of guanidine hydrochloride (GdnHCl), concentration range 0-8 M. Mixed samples were incubated at 37 °C for 1 h. The samples were diluted by the addition of 850 μl Tris buffer containing 10 mM Tris-HCl (pH 8.0), 150 mM NaCl, 0.5% Triton-X and 0.5% deoxycholate. Following this, 50 μL GdnHCl were added to each sample to obtain 0.4 M final concentration. Next, the samples were digested with 20 μg/ml PK at 37 °C for 1 h. PrPSc concentration and western blot analysis were carried out as described above. Conformational stability was examined using mAbs 6H4 and SAF84. Denaturation curves were obtained by densitometric analysis using Fluorochem software (Alpha Innotech Co.). GdnHCl concentration at half maximal denaturation ([GdnHCl]1/2) was used as a measure of the relative conformational stability of PrPSc. [GdnHCl]1/2 was calculated based on the denaturation curves.
Additional Information
How to cite this article: Masujin, K. et al. Emergence of a novel bovine spongiform encephalopathy (BSE) prion from an atypical H-type BSE. Sci. Rep. 6, 22753; doi: 10.1038/srep22753 (2016).
References
Prusiner, S. B. Molecular biology of prion diseases. Science 252, 1515–1522 (1991).
Bolton, D. C., McKinley, M. P. & Prusiner, S. B. Identification of a protein that purifies with the scrapie prion. Science 218, 1309–1311 (1982).
Collinge, J. & Clarke, A. R. A general model of prion strains and their pathogenicity. Science 318, 930–936 (2007).
Bruce, M. et al. Transmission of bovine spongiform encephalopathy and scrapie to mice: strain variation and the species barrier. Philos Trans R Soc Lond B Biol Sci 343, 405–411 (1994).
Jacobs, J. G. et al. Molecular discrimination of atypical bovine spongiform encephalopathy strains from a geographical region spanning a wide area in Europe. J Clin Microbiol 45, 1821–1829 (2007).
Ducrot, C., Arnold, M., de Koeijer, A., Heim, D. & Calavas, D. Review on the epidemiology and dynamics of BSE epidemics. Vet Res 39, 15 (2008).
Biacabe, A. G., Laplanche, J. L., Ryder, S. & Baron, T. Distinct molecular phenotypes in bovine prion diseases. EMBO Rep 5, 110–114 (2004).
Nicholson, E. M., Brunelle, B. W., Richt, J. A., Kehrli, M. E. J. & Greenlee, J. J. Identification of a heritable polymorphism in bovine PRNP associated with genetic transmissible spongiform encephalopathy: evidence of heritable BSE. PLoS ONE 3, e2912 (2008).
Balkema-Buschmann, A. et al. Experimental challenge of cattle with German atypical bovine spongiform encephalopathy (BSE) isolates. J Toxicol Environ Health A 74, 103–109 (2011).
Okada, H. et al. Experimental H-type bovine spongiform encephalopathy characterized by plaques and glial- and stellate-type prion protein deposits. Vet Res 42, 79–88 (2011).
Priemer, G., Balkema-Buschmann, A., Hills, B. & Groschup, M. H. Biochemical characteristics and PrP distribution pattern in the brains of cattle experimentally challenged with H-type and L-type atypical BSE. PLoS One 8, e67599 (2013).
Konold, T. et al. Experimental H-type and L-type bovine spongiform encephalopathy in cattle: observation of two clinical syndromes and diagnostic challenges. BMC Vet Res 8, 22 (2012).
Baron, T. G., Biacabe, A. G., Bencsik, A. & Langeveld, J. P. Transmission of new bovine prion to mice. Emerg Infect Dis 12, 1125–1128 (2006).
Biacabe, A. G., Jacobs, J. G., Bencsik, A., Langeveld, J. P. & Baron, T. G. H-type bovine spongiform encephalopathy: complex molecular features and similarities with human prion diseases. Prion 1, 61–68 (2007).
Buschmann, A. et al. Atypical BSE in Germany–proof of transmissibility and biochemical characterization. Vet Microbiol 117, 103–116 (2006).
Okada, H. et al. Experimental Transmission of H-type Bovine Spongiform Encephalopathy to Bovinized Transgenic Mice. Vet Pathol 48, 942–947 (2010).
Beringue, V. et al. Isolation from cattle of a prion strain distinct from that causing bovine spongiform encephalopathy. PLoS Pathog 2, e112 (2006).
Baron, T. et al. Emergence of classical BSE strain properties during serial passages of H-BSE in wild-type mice. PLoS One 6, e15839 (2011).
Bencsik, A., Leboidre, M., Debeer, S., Aufauvre, C. & Baron, T. Unique properties of the classical bovine spongiform encephalopathy strain and its emergence from H-type bovine spongiform encephalopathy substantiated by VM transmission studies. J Neuropathol Exp Neurol 72, 211–218 (2013).
Torres, J. M. et al. Classical bovine spongiform encephalopathy by transmission of H-type prion in homologous prion protein context. Emerg Infect Dis 17, 1636–1644 (2011).
Masujin, K. et al. Biological and biochemical characterization of L-type-like bovine spongiform encephalopathy (BSE) detected in Japanese black beef cattle. Prion 2, 123–128 (2008).
Masujin, K., Miwa, R., Okada, H., Mohri, S. & Yokoyama, T. Comparative analysis of Japanese and foreign L-type BSE prions. Prion 6, 89–93 (2012).
Yokoyama, T. et al. Intraspecies prion transmission results in selection of sheep scrapie strains. PLoS One 5, e15450 (2010).
Kimberlin, R. H., Walker, C. A. & Fraser, H. The genomic identity of different strains of mouse scrapie is expressed in hamsters and preserved on reisolation in mice. J Gen Virol 70, 2017–2025 (1989).
Bruce, M. E. Scrapie strain variation and mutation. Br Med Bull 49, 822–838 (1993).
Kimberlin, R. H. & Walker, C. A. Evidence that the transmission of one source of scrapie agent to hamsters involves separation of agent strains from a mixture. J Gen Virol 39, 487–496 (1978).
Masujin, K. et al. Isolation of two distinct prion strains from a scrapie-affected sheep. Arch Virol 154, 1926–1932 (2009).
Telling, G. C. et al. Evidence for the conformation of the pathologic isoform of the prion protein enciphering and propagating prion diversity. Science 274, 2079–2082 (1996).
Seuberlich, T., Heim, D. & Zurbriggen, A. Atypical transmissible spongiform encephalopathies in ruminants: a challenge for disease surveillance and control. J Vet Diagn Invest 22, 823–842 (2010).
Scott, M. R. et al. Identification of a prion protein epitope modulating transmission of bovine spongiform encephalopathy prions to transgenic mice. Proc Natl Acad Sci USA 94, 14279–14284 (1997).
Dudas, S. et al. Molecular, biochemical and genetic characteristics of BSE in Canada. PLoS One 5, e10638 (2010).
Fukuda, S. et al. Intraspecies transmission of L-type-like Bovine Spongiform Encephalopathy detected in Japan. Microbiol Immunol 53, 704–707 (2009).
Fukuda, S. et al. Neuroanatomical distribution of disease-associated prion protein in experimental bovine spongiform encephalopathy in cattle after intracerebral inoculation. Jpn J Infect Dis 65, 37–44 (2012).
Harmeyer, S., Pfaff, E. & Groschup, M. H. Synthetic peptide vaccines yield monoclonal antibodies to cellular and pathological prion proteins of ruminants. J Gen Virol 79, 937–945 (1998).
Korth, C. et al. Prion (PrPSc)-specific epitope defined by a monoclonal antibody. Nature 390, 74–77 (1997).
Feraudet, C. et al. Screening of 145 anti-PrP monoclonal antibodies for their capacity to inhibit PrPSc replication in infected cells. J Biol Chem 280, 11247–11258 (2005).
Fraser, H. & Dickinson, A. G. The sequential development of the brain lesion of scrapie in three strains of mice. J Comp Pathol 78, 301–311 (1968).
Legname, G. et al. Strain-specified characteristics of mouse synthetic prions. Proc Natl Acad Sci USA 102, 2168–2173 (2005).
Acknowledgements
We thank Dr. Stefanie Czub for providing the Canadian H-BSE sample. We thank Dr. Byron Caughey for critical review of this manuscript and Drs. Morikazu Shinagawa and Shirou Mohri for their encouragement. We are also grateful to Naoko Tabeta, Ritsuko Miwa, Naomi Furuya and Noriko Shimozaki for their technical help and Junko Yamada, Tomoaki Yamamura and the animal laboratory staff at the National Institute of Animal Health for maintaining the mouse colony and the cattle. This study was supported by a research project for improving food safety and animal health (Ministry of Agriculture, Forestry and Fisheries of Japan), a research project for atypical BSE diagnosis (Ministry of Health, Labour and Welfare of Japan) and partially supported by basic research fund of NARO.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: K. Masujin, H.O. and T.Y. Performed the experiments: K. Masujin, H.O., K. Miyazawa, Y. Matsuura, M.I., Y.I. and Y. Murayama. Analyzed the data: K. Masujin, H.O., K. Miyazawa. Contributed reagents/materials/analysis tools: K. Masujin, H.O. Wrote the paper: K. Masujin, T.Y.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Masujin, K., Okada, H., Miyazawa, K. et al. Emergence of a novel bovine spongiform encephalopathy (BSE) prion from an atypical H-type BSE. Sci Rep 6, 22753 (2016). https://doi.org/10.1038/srep22753
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep22753
This article is cited by
-
Prion protein PrP nucleic acid binding and mobilization implicates retroelements as the replicative component of transmissible spongiform encephalopathy
Archives of Virology (2020)
-
Interspecies transmission to bovinized transgenic mice uncovers new features of a CH1641-like scrapie isolate
Veterinary Research (2018)
-
Divergent prion strain evolution driven by PrPC expression level in transgenic mice
Nature Communications (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.