Abstract
We recently discovered a novel population of stem cells from the injured murine skeletal muscle. These injury induced muscle-derived stem cell-like cells (iMuSCs) are partially reprogrammed from differentiated myogenic cells and display a pluripotent-like state. The iMuSCs exhibit stem cell properties including the ability to differentiate into multiple lineages, such as neurogenic and myogenic differentiations; they also display a superior migration capacity that demonstrating a strong ability of muscle engraftment in vivo. IMuSCs express several pluripotent and myogenic stem cell markers; have the capability to form embryoid bodies and teratomas and can differentiate into all three germ layers. Moreover, blastocyst microinjection showed that the iMuSCs contributed to chimeric embryos but could not complete germline transmission. Our results indicate that the iMuSCs are in a partially reprogrammed state of pluripotency, which are generated by the microenvironment of injured skeletal muscle.
Similar content being viewed by others
Introduction
Tissue repair after injury is a complex biological process, which involves the activation of tissue-resident precursors and stem cells and a variety of infiltrating cells responding to local and systemic signals. Mammalian skeletal muscle regeneration relies on the activation and proliferation of the resident muscle precursor cells1,2 including satellite cells and muscle stem cells (MuSCs), which are populations of mononucleated cells located between the basal lamina and sarcolemma of muscle fibers. MuSCs are a functionally heterogeneous population of cells and have variable proliferation rates, marker expression profiles, self-renewal capacities, clonogenicity and differentiation capacities2,3. We have previously discovered that among MuSCs, a small population of iMuSCs exist and can be isolated from injured murine skeletal muscles using a Cre-LoxP system established in our laboratory4. We have shown that iMuSCs not only express CD34, Sca1 (Stem cell antigen-1) and Pax7 (Paired box protein 7), but also presented strong myogenic differentiation and muscle regeneration abilities in vivo5. In addition, we demonstrated that iMuSCs demonstrate stem cell behaviors and are capable of differentiating into non-myogenic lineages, such as CD31+ endothelial-like cells in the healed skeletal muscle4. Here, we further investigate the unique nature of the iMuSCs, focusing on their morphology, marker expression profile, pluripotency, migratory abilities and differentiation potential.
Results
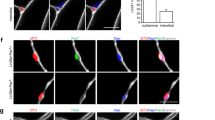
By applying our established cell isolation method (Fig. 1a) iMuSCs were successfully isolated from the injured murine tibialis anterior (TA) muscle. Three days after the cell isolation, proliferating iMuSCs (~0.1% of the entire muscle cell population) appeared in the culture dishes; however, no cells were present in the cultures that established from the control, uninjured muscles (Fig. 1b). Microscopic evaluation revealed that representative iMuSCs were 5–7 μm in diameter, contained relatively large nuclei and a narrow rim of cytoplasm. Their nuclei were Hoechst 33342 positive and incorporated BrdU (Bromodeoxyuridine) with Msx1 (Msh homeobox 1) expression (Supplementary Fig. S1a). The freshly isolated, or early population of iMuSCs contained a high percentage of Msx1 and Cxcr4 (C-X-C chemokine receptor type 4) positive cells, among which were a few cells expressing Pax7 and Sca1 (Fig. 1c). Gene expression analysis of the entire biopsied injured TA muscles showed that Msx1, Oct4 (also called Pou5f1), Sox2 (SRY-box 2) and Nanog expression were up-regulated compared to the control uninjured old TA muscle (Fig. 1d and Supplementary Fig. S1b). Freshly isolated iMuSCs expressed myogenic stem cell related markers, i.e. Sca1, Pax7 and CD34 and the core pluripotency marker genes, i.e. Oct4, Sox2 and Nanog (Fig. 1e and Supplementary Fig. S1c). Cultured iMuSCs were expanded in vitro in muscle growth medium with an average cell population doubling time of 13 hours. Cytogenetic analysis revealed that iMuSCs had a normal female karyotype; however, chromosomal aberrations appeared during long-term culture (passage 33), resulting in trisomy for chromosome 5 (Supplementary Fig. S1d). We also discovered that iMuSCs had remarkable migration properties. Data from a time-lapse motility assay confirmed that iMuSCs migrated longer distances with higher velocities compared to the control mouse myoblast cell line, C2C12 and MuSCs isolated from control uninjured muscle (Fig. 1f). Moreover, iMuSCs expressed high levels of β-Catenin and several Cadherins at the mRNA level (Fig. 1g).
IMuSCs display stemness and exhibit improved migration ability.
(a) Schematic of iMuSCs isolation method from injured murine TA muscles. (b) Bright field images of uninjured and injured cultures. 3 days after the cell isolation no cells appeared in the control uninjured cultures, but iMuSCs were present in the injured cultures. 7 days after cell isolation, the proliferation of iMuSCs was apparent. Scale bar = 10 μm. (c) Msx1 (green), Pax7 (red), Cxcr4 (green) and Sca1 (red) expression of iMuSCs. Nuclei were stained with DAPI (blue). Scale bar = 100 μm. (d) qPCR analysis of whole biopsied TA muscles and (e), fresh isolated iMuSCs. (f) Single cell migration pathways of iMuSCs and the control C2C12 and MuSCs. The migration paths of 20 individual cells from different experimental groups captured in a time-lapse motility assay. Data was pooled from 3 independent experiments. Graphs show the calculated accumulated distance and velocity of the cells. Data are represented as the mean ±SEM of 60 individual cells from 3 biological replicates. **P < 0.01. (g) qPCR analysis of β-Catenin, E-Cadherin, M-Cadherin, N-Cadherin expression of iMuSCs. Data are represented as the mean ±SEM of 5 biological replicates.
In vitro multipotent differentiation assays showed that iMuSCs were able to fuse with MyHC+ (Myosin heavy chain) myotubes in muscle differentiation medium with a similar fusion index as control MuSCs and C2C12 myoblasts (Fig. 2a). The iMuSCs were also capable of differentiating into osteogenic lineages (Supplementary Fig. S2) within osteogenic medium with BMP2. The iMuSCs could also be easily and effectively induced into a neurogenic lineage via neurosphere formation once cultured in neural stem cell medium (see Method) for one week (Fig. 2b), whereas the control primary myoblasts and MuSCs showed no sign of forming these structures. The iMuSCs-induced neurospheres exhibited a neural phenotype and expressed Nestin, CNPase and Nefm (Neurofilament) (Fig. 2b). After three weeks, the neurospheres when re-plated in a laminin/polyornithine coated monolayer culture in neural differentiation medium, could differentiate into the three major neural lineages (neurons, astrocytes and oligodendrocytes) and they expressed Mtap2, β-Tubulin III, Nefm, Nestin and Olig1/2 (Oligodendrocyte transcription factor 1/2) (Fig. 2b,c).
Multiple differentiation and muscle engraftment of iMuSCs.
(a) Induced myotube formation of iMuSCs. Myotubes expressed MyHC (red). The fusion index was similar to the control C2C12 and MuSCs. (b) Representative bright-field picture shows iMuSCs-formed neurospheres floating in suspension. Immunofluorescence staining of cryosectioned neurospheres shows Nestin (green), CNPase (red) and Nefm (red) positive cells. Plated 21 day differentiated neurospheres in ND medium show neural phenotype; β-Tubulin III (red) and Nefm (green). Nuclei were stained with DAPI. Scale bar = 10 and 100 μm. (c) Gene expression kinetics of Mtap2 and β-Tubulin III and Olig1 and Olig2 in the neural differentiating iMuSCs analyzed by qPCR. Data were compared to undifferentiated iMuSCs and are presented as the mean ±SEM of 5 biological replicates. (d) Engraftment of iMuSCs after intramuscular cell implantation. IF staining shows Utrophin+ (green) and Dystrophin+ (red) muscle engraftment of control MuSCs and iMuSCs in mdx/scid mice 2 weeks after cell injection. Scale bar = 100 μm. Quantification of Dystrophin+ myofibers. **P < 0.01.
To further investigate the origin of the iMuSCs, we performed in vivo intramuscular transplantation studies. Equal numbers of iMuSCs and control MuSCs were injected into the TA muscles of six 6–8 week-old male mdx/SCID mice (Jackson Lab, USA). Two and three weeks after cell implantation, we detected Utrophin and Dystrophin (Fig. 2d) expression in the host TA muscles and observed that the iMuSCs formed larger and more robust Dystrophin+ muscle grafts compared to the control MuSCs (Fig. 2d).
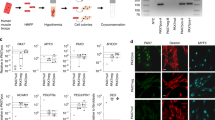
We also performed quantitative real-time polymerase chain reaction (qPCR) and immunohistochemistry analysis to elucidate the gene and protein expression profile of the iMuSCs and compared these to embryonic stem cells (ESCs) and myogenic stem cells (C2C12 and MuSCs). The iMuSCs expressed Oct4, Ssea1 (Stage-specific embryonic antigen 1), Sox2, Cxcr4, Msx1, Pax7 and Sca1 (Fig. 3a and Supplementary Fig. S3a,b), similar to the ESCs, but at a lower expression level. QPCR analysis revealed that the iMuSCs expressed the majority of pluripotency marker genes, with the exception of Esg1 and Dax1 (Fig. 3b); however, unlike the ESCs, the iMuSCs expressed myogenic marker genes and interestingly some of the primordial germ-cell-related markers, e.g. Blimp1 and Fragilis and did not express other lineage related genes, such as CD45 or CD90 (Fig. 3c). Moreover, the iMuSCs were positive for alkaline phosphatase (Fig. 3a). These results indicate that the iMuSCs are similar to, but not identical to the ESCs, since they maintain their myogenic memory (e.g., high expression of myogenic genes when compared to the ESCs and are easily induced to differentiate into a myogenic lineage in vitro and in vivo).
Skeletal muscle injury induced iMuSCs fulfil several in vitro criteria of pluripotency.
(a) Representative immunofluorescent images of iMuSCs expressing Ssea1 (red), Oct4 (green), Sox2 (red), Cxcr4 (green), Pax7 (red), Msx1 (green), Sca1 (red) and alkaline phosphatase. Nuclei were stained with DAPI (blue). Scale bars = 10 and 200 μm. (b) qPCR analysis of iMuSCs for pluripotent marker genes and (c) myogenic marker genes. Data are represented as the mean ±SEM of 5 biological replicates. (d) Bright-field image of formed embryoid bodies (EBs) in suspension and immunofluorescence staining of cryosectioned EBs. EBs contained cells positive for Sox2 (red), Oct4 (green) and Nanog (red). Nuclei were stained with DAPI (blue). Scale bar = 100 μm. (e) qPCR data showed upregulation of Oct4, Nanog, Sox2 and no change of Nestin expression compared to control undifferentiated iMuSCs. (f) Immunofluorescence images of differentiated EBs expressing αSMA (red), β-Tubulin III (green), Brachyury (red) and β-Tubulin III β (red) with MyHC (green). Nuclei were stained with DAPI (blue). Scale bar = 100 μm. (g) qPCR data of 7 days differentiated EBs show Nestin, Mtap2, Desmin and Myogenin upregulation and Nanog downregulation compared to control undifferentiated iMuSCs. All qPCR data was analyzed by the delta-delta Ct method and are represented as the mean ±SEM of 5 biological replicates.
To clarify the pluripotent potential of the iMuSCs, we performed differentiation assays6,7 in vitro that showed that the iMuSCs were able to form embryoid bodies (EBs) in a petri dish (Fig. 3d,e). After seven days in suspension culture, EBs were expanded and initiated spontaneous differentiation into a variety of ectodermal and mesodermal germ layer derivatives and after an additional two weeks in culture, attached EBs formed contracting multinucleated myotubes encompassed with neural-like structures (Fig. 3f,g). We further examined the pluripotency of the iMuSCs by teratoma formation in vivo. When grafted into SCID-beige mice (Jackson Lab, USA) for seven weeks, the iMuSCs formed teratomas (90%, n = 7) containing representative tissues of the three germ layers (Fig. 4a). Histological examination revealed that the iMuSCs differentiated into neural, muscle and adipose tissues and epithelium. To verify that the teratomas were formed directly from the implanted cells, the iMuSCs were pre-labelled with β-gal before injection, we detected all three germ layer derivatives in the teratomas contained the β-gal+ cells when stained with LacZ (Fig. 4b).
Skeletal muscle injury induced iMuSCs fulfil several in vivo criteria of pluripotency.
(a) Teratoma formation assay of iMuSCs. Hematoxylin and eosin staining showed differentiated structures of all three germ layers: skeletal muscle (mesoderm, arrows), cartilage (mesoderm, asterisks), blood vessel (mesoderm, asterisk), fat (mesoderm, asterisk), gut epithelium (endoderm, asterisk), neural rosettes (ectoderm, arrow) and neuroepithelium (ectoderm, asterisks) within the same sample. (b) LacZ staining indicated LacZ pre-labelled iMuSC-differentiated structures of adipose, muscle and neural rosettes contain β-gal+ signals (dark blue dots). Scale bar = 100 μm. (c) Contribution of iMuSCs to mouse embryonic development. Embryos at E14 were analyzed by GFP and LacZ staining, (d) and by qPCR analysis of GFP and LacZ marker gene expression in normally and abnormally developed embryos. (e) The E14 embryos were sectioned and stained with anti-GFP antibody (dark purple). Cells were counterstained with eosin (pink): skin and under-skin, heart, lung, cartilage, gastrointestinal tract and neural tube. Scale bar = 200 μm. (f) qPCR analysis of GFP and LacZ marker gene expression in the born P21 white pups. Data were pulled together from 6 pups.
To evaluate whether the iMuSCs could give rise to chimeric mice, a blastocyst injection assay was performed (Fig. 4c). We transferred undifferentiated β-gal+ and GFP-pre-labelled iMuSCs as single cells into BALB/c (Jackson Lab, USA) blastocysts by microinjection following standard procedures8. We obtained eight embryos at E14, six of which developed properly and demonstrated the contribution of GFP+ iMuSCs to the embryo. A high-to-moderate contribution of β-gal and GFP-expressing cells could be seen in these E14 chimeric embryos (Fig. 4c,d and Supplementary Fig. S4a). Histological analysis confirmed that the iMuSCs contributed to all three germ layers (Fig. 4e and Supplementary Fig. S4b). Offspring derived from the iMuSCs-injected blastocysts were born and developed normally. After repeating this experiment 3 times, we obtained 23 pups, all born with a white coat (Supplementary Table S1). Although their hair did not display iMuSCs germline transmission, immunostaining and qPCR analysis revealed the presence of LacZ+ and GFP+ iMuSCs in several tissues of the pups, such as skin, muscle, heart, lung, kidneys, spleen and brain (Fig. 4f and Supplementary Fig. S4c).
Discussion
The existence of pluripotent-like cells in adult tissues has been a matter of debate for years, since inconsistent results have been reported by various groups9,10,11,12,13,14,15; however, no study thus far has proven that such pluripotent stem cells can arise from differentiated somatic tissues. In this study we reveal that cellular reprogramming can be initiated by the strong stimuli that occurs when skeletal muscle is injured; thus, we were able to isolate reprogrammed iMuSCs from the injured skeletal muscle.
Collectively, our findings demonstrate that iMuSCs represent a unique, very sensitive population of cells that possess characteristics (morphology, size and a gene expression profiles) that differs from all cell types studied so far. IMuSCs not only display several characteristics typical of ESCs (e.g. a large nucleus surrounded by a narrow rim of cytoplasm, high nuclear/cytoplasmic ratio, open chromatin, unstructured nucleoplasm and diploid number of chromosomes) (Table 1), but also express several pluripotency marker genes while maintaining a high expression level of myogenic genes. Moreover, the most remarkable discovery of this study was that iMuSCs fulfilled several in vitro and in vivo criteria for pluripotency; however, we could not obtain iMuSCs with germline transmission after blastocyst microinjection. This may be due to the fact that iMuSCs have a lower gene expression profile of the pluripotency markers (e.g., Oct4, Nanog, and Sox2) and lack Esg1 and Dax1 expression when compared to ESCs. It is also plausible that the relatively high expression of Blimp1, Fragilis and myogenic marker genes by the iMuSCs may contribute to this observation. These results indicate that iMuSCs do not regress completely to pluripotency and possibly hold an epigenetic memory of their myogenic tissue origin. Further manipulation of the iMuSCs, such as the inhibition of DNA methylases or Nanog overexpression, could potentially push the iMuSCs to achieve full pluripotency.
Thus, the key conclusion from our study is that changes in microenvironmental factors, such as skeletal muscle with injuries, can partially reprogram terminally differentiated myogenic cells into a pluripotent-like state.
Methods
Animal studies
The animal experiments and all experimental protocols were approved by the Center for Laboratory Animal Medicine and Care at The University of Texas Health Science Center at Houston (protocol No: AWC-13-134). All methods were carried out in accordance with the approved relevant guidelines and regulations. Mouse strains utilized in this study: C57BL/6J, BALB/c, SCID-beige, and mdx/SCID were purchased from the Jackson Laboratory, USA.
Cell isolation and maintenance
Mouse iMuSCs were isolated from the injured TA muscles of C57BL/6J (3–8-week-old female; Jackson Lab, USA) mice four days after laceration injury, while control MuSCs were isolated from uninjured TA muscles. The iMuSCs were separately cultured in ESGRO Complete PLUS Clonal Grade Medium (Millipore, USA) on 12-well tissue culture plates (Corning, USA) for 3 weeks. The medium was then replaced with normal muscle growth medium [Dulbecco’s Modified Eagle’s Medium (DMEM) supplemented with 20% Fetal Bovine Serum (FBS), 10% Horse Serum (HS), 1% chicken embryo extract (CEE; Accurate Chemical Co., UK) and 1% Penicillin-Streptomycin antibiotics; unless otherwise mentioned, all from Gibco, USA] and iMuSCs were further cultured and expanded on collagen type IV-coated flasks at 5% CO2 at 37 °C. C2C12 primary mouse myoblasts (purchased from ATTC, USA) and MuSCs were used as controls and were cultured on collagen type IV-coated flasks in growth medium in 5% CO2 at 37 °C. Characterization of iMuSCs was performed by applying standard in vitro and in vivo assays.
Generation and analysis of chimeric mice
Undifferentiated single cells of β-gal- and GFP-pre-labelled iMuSCs were transferred into blastocysts of BALB/c mice (Jackson Lab, USA) by microinjection following standard procedures8. Pregnant mice were sacrificed and E14 embryos were collected and dehydrated in a serial dilution of 30, 20 and 10% sucrose and then fixed in 4% paraformaldehyde (Sigma, USA) and embedded in paraffin. Sections were stained with anti-GFP antibody, counterstained with eosin and visualized by fluorescent microscopy (Nikon).
Teratoma formation assay
IMuSCs were suspended at 1 − 2 × 106 cells/ml in Phosphate Buffered Saline (PBS). SCID-beige mice (Jackson Lab, USA) were anesthetized with diethyl ether and injected with 500 μl of the cell suspension subcutaneously into the dorsal flank. Seven weeks after the cell injection, tumours were surgically dissected from the mice. Samples were weighed, fixed in 4% formaldehyde and embedded in paraffin. Sections were stained with hematoxylin and eosin.
In vitro differentiation
In vitro embryoid body (EB) formation of iMuSCs was induced by applying the routinely used ‘hanging-drop’ technique6,16. Single cell suspensions of iMuSCs (3.75 × 104 cells/ml) were placed as 20 μl micro-drops of ESGRO Complete PLUS Clonal Grade Medium (Millipore, USA) on the lids of 100 mm dishes. Bottom plates of the dishes were filled with sterile PBS to avoid desiccation of samples. Lids were then placed on the bottom plates for 4 days to achieve hanging drop cultures. During this time, single iMuSCs formed EBs, which were harvested and further cultured in suspension for 3 additional days. For spontaneous in vitro differentiation, EBs were cultured on collagen type IV-coated 24-well plates in EB Medium [DMEM supplemented with 20% FBS, 2 mM Glutamax, 1% non-essential amino acids and 1% antibiotics; all from Gibco (USA)] for 2 weeks with medium changed every other day. Additionally, multiple differentiations of iMuSCs were assessed by myogenic and osteogenic differentiation assays following standard protocols5,17,18.
Singe cell migration
Target cell populations and control cells (MuSCs and C2C12 cells) were plated separately on collagen type IV-coated 6-well plates, 10,000 cells/well, in growth medium, 48 hours prior to the time-lapse experiment. Time lapse images were acquired with an Andor IXon3 885 EMCCD camera (Andor, USA) on an Olympus IX-81 (Olympus, USA) microscope fitted with a microscope enclosure (Precision Plastics, USA) and images of single cell migration were taken for six hours at three minute intervals; three different fields/well were chosen for the recording. Proper environmental conditions were maintained in a micro incubator at 37 °C with 5% CO2. A series of images were analyzed using NIH ImageJ analysis software to track the centroid positions (x, y) of cell nuclei (which were assumed to be the representations of cell-bodies). Migration paths were plotted and analyzed by the Chemotaxis and Migration Tool v2.0 from Ibidi. Net translocation distance was measured as the distance between the starting point and the end point of cells after six hours. Migration speed was calculated as total length of the migration path during the six-hour period.
Immunohistochemistry
Samples were fixed with 4% paraformaldehyde (Sigma, USA) for 20 minutes at room temperature. After permeabilization with 0.2% Triton X-100 (Sigma, USA) for 30 minutes, nonspecific binding of antibodies was blocked for one hour with 10% BSA and 5% HS (Sigma, USA) in PBS, at room temperature. The primary antibodies (Supplementary Table 2) were applied overnight at 4 °C. After the overnight incubation, the cells were incubated for one hour at room temperature with the appropriate fluorescence-conjugated secondary antibodies (Extended Data Table 2). The nuclei were revealed using 4′, 6′-diamidino-2-phenylindole (DAPI) staining (Sigma, USA) and fluorescent microscopy (Nikon) was used to visualize the results. Quantitative image analysis was performed using the NIH ImageJ Software.
Karyotype analysis
Karyotypes were determined using conventional G-Banding analysis at the Texas Children’s Hospital Cytogenetic Core Lab, Houston, TX, USA.
Population doubling assay and histochemical staining for alkaline phosphatase
To estimate the growth of cultures, a standard protocol was followed for population doubling assays. Briefly, 5000 iMuSCs were plated on collagen type IV-coated 6-well plates in growth medium for four days and every day the well content was collected and counted. The approximate population doubling time (PDT) was determined as:  , where, t = time period (24 h), N0 = initial cell number, Nt = number of cells at the time period.
, where, t = time period (24 h), N0 = initial cell number, Nt = number of cells at the time period.
Histological staining for alkaline phosphatase activity was carried out using a commercially available kit (Sigma-Aldrich, USA) following the manufacturer’s instructions.
Quantitative real-time PCR
Total RNA was isolated using the RNeasy Plus Mini Kit (Quiagen, USA) and cDNA was synthesized from 1 μg of RNA via a High Capacity RNA-to-cDNA Kit (Life Technologies, USA) following the manufacturer’s instructions. Gene expression was analyzed by quantitative real-time PCR (qPCR) using MyiQ real time PCR System (Bio-Rad, USA). The applied primers (Supplementary Table 3) were designed by Oligo software (Oligo Perfect Designer, Invitrogen, USA). Reactions were measured in duplicates using custom 2× Syber Green Master Mix based on the hot-start Jumpstart Taq DNA Polymerase enzyme (Sigma, USA). The amplification was done for 40 cycles (95 °C 20 sec, 60 °C 20 sec, 72 °C 40 sec). To verify the PCR products, melting curves and negative controls were carried out in each reaction. Relative quantification of mRNA was determined by the ΔΔCt method (2−ΔΔCt formula)19 by using the expression profile of the corresponding control samples as reference.
Data processing and statistical analysis
Prism 6.0 (GraphPad Software, USA) was used for data plotting, non-linear regression and statistical analysis. Data are given as mean ± S.E.M. Comparisons between two groups were performed by using Student’s t-test assuming two-tailed distribution and unequal variances. For multiple comparisons, ANOVA or Kruskal-Wallis test was applied. Statistical significance was considered at p < 0.05.
Additional Information
How to cite this article: Vojnits, K. et al. Characterization of an Injury Induced Population of Muscle-Derived Stem Cell-Like Cells. Sci. Rep. 5, 17355; doi: 10.1038/srep17355 (2015).
References
Ceafalan, L. C., Popescu, B. O. & Hinescu, M. E. Cellular players in skeletal muscle regeneration. Biomed Res Int 2014, 957014 (2014).
Relaix, F. & Zammit, P. S. Satellite cells are essential for skeletal muscle regeneration: the cell on the edge returns centre stage. Development 139, 2845–2856 (2012).
Brack, A. S. & Rando, T. A. Tissue-specific stem cells: lessons from the skeletal muscle satellite cell. Cell Stem Cell 10, 504–514 (2012).
Mu, X., Peng, H., Pan, H., Huard, J. & Li, Y. Study of muscle cell dedifferentiation after skeletal muscle injury of mice with a Cre-Lox system. PLoS One 6, e16699 (2011).
Mu, X. et al. Slow-adhering stem cells derived from injured skeletal muscle have improved regenerative capacity. Am J Pathol 179, 931–941 (2011).
Potter, S. W. & Morris, J. E. Development of mouse embryos in hanging drop culture. Anat Rec 211, 48–56 (1985).
Keller, G. M. In vitro differentiation of embryonic stem cells. Curr Opin Cell Biol 7, 862–869 (1995).
Nagy, A. Manipulating the mouse embryo: a laboratory manual. 3rd edn, (Cold Spring Harbor Laboratory Press, 2003).
Jiang, Y. et al. Pluripotency of mesenchymal stem cells derived from adult marrow. Nature 418, 41–49 (2002).
D’Ippolito, G. et al. Marrow-isolated adult multilineage inducible (MIAMI) cells, a unique population of postnatal young and old human cells with extensive expansion and differentiation potential. J Cell Sci 117, 2971–2981 (2004).
Kucia, M. et al. A population of very small embryonic-like (VSEL) CXCR4(+)SSEA-1(+)Oct-4+ stem cells identified in adult bone marrow. Leukemia 20, 857–869 (2006).
Kuroda, Y. et al. Unique multipotent cells in adult human mesenchymal cell populations. Proc Natl Acad Sci USA 107, 8639–8643 (2010).
Obokata, H. et al. The potential of stem cells in adult tissues representative of the three germ layers. Tissue Eng Part A 17, 607–615 (2011).
Lengner, C. J., Welstead, G. G. & Jaenisch, R. The pluripotency regulator Oct4: a role in somatic stem cells? Cell Cycle 7, 725–728 (2008).
Berg, J. S. & Goodell, M. A. An argument against a role for Oct4 in somatic stem cells. Cell Stem Cell 1, 359–360 (2007).
Murry, C. E. & Keller, G. Differentiation of embryonic stem cells to clinically relevant populations: lessons from embryonic development. Cell 132, 661–680 (2008).
Zuk, P. A. et al. Multilineage cells from human adipose tissue: implications for cell-based therapies. Tissue Eng 7, 211–228 (2001).
Zheng, B., Cao, B., Li, G. & Huard, J. Mouse adipose-derived stem cells undergo multilineage differentiation in vitro but primarily osteogenic and chondrogenic differentiation in vivo. Tissue Eng 12, 1891–1901 (2006).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408 (2001).
Martin, G. R. Isolation of a pluripotent cell line from early mouse embryos cultured in medium conditioned by teratocarcinoma stem cells. Proc Natl Acad Sci USA 78, 7634–7638 (1981).
Evans, M. Discovering pluripotency: 30 years of mouse embryonic stem cells. Nat Rev Mol Cell Biol 12, 680–686 (2011).
Evans, M. J. & Kaufman, M. H. Establishment in culture of pluripotential cells from mouse embryos. Nature 292, 154–156 (1981).
Pillarisetti, A. et al. Mechanical phenotyping of mouse embryonic stem cells: increase in stiffness with differentiation. Cell Reprogram 13, 371–380 (2011).
Sainz, J., Garcia-Alcalde, F., Blanco, A. & Concha, A. Genome-wide gene expression analysis in mouse embryonic stem cells. Int J Dev Biol 55, 995–1006 (2011).
Pereira Fde, A., Tavares, R. L., Camargos, A. F. & da Silva Filho, A. L. Telomerase activity alterations in sequential passages of mouse embryonic stem cells. Cell Biol Int 36, 755–757 (2012).
Niwa, H. Molecular mechanism to maintain stem cell renewal of ES cells. Cell Struct Funct 26, 137–148 (2001).
Bernstein, B. E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, 315–326 (2006).
Shim, J. H. et al. Proteomic analysis of time-dependent difference of protein expression profile changes during neuronal differentiation of mouse embryonic stem cells. Mol Cells 29, 239–244 (2010).
Chung, S. et al. Genetic engineering of mouse embryonic stem cells by Nurr1 enhances differentiation and maturation into dopaminergic neurons. Eur J Neurosci 16, 1829–1838 (2002).
Okabe, S., Forsberg-Nilsson, K., Spiro, A. C., Segal, M. & McKay, R. D. Development of neuronal precursor cells and functional postmitotic neurons from embryonic stem cells in vitro. Mech Dev 59, 89–102 (1996).
Kim, P. T. & Ong, C. J. Differentiation of definitive endoderm from mouse embryonic stem cells. Results Probl Cell Differ 55, 303–319 (2012).
Kim, P. T. et al. Differentiation of mouse embryonic stem cells into endoderm without embryoid body formation. PLoS One 5, e14146 (2010).
Wang, P. et al. A molecular signature for purified definitive endoderm guides differentiation and isolation of endoderm from mouse and human embryonic stem cells. Stem Cells Dev 21, 2273–2287 (2012).
Sachinidis, A. et al. Cardiac specific differentiation of mouse embryonic stem cells. Cardiovasc Res 58, 278–291 (2003).
Chen, M., Lin, Y. Q., Xie, S. L., Wu, H. F. & Wang, J. F. Enrichment of cardiac differentiation of mouse embryonic stem cells by optimizing the hanging drop method. Biotechnol Lett 33, 853–858 (2011).
Bradley, A., Evans, M., Kaufman, M. H. & Robertson, E. Formation of germ-line chimaeras from embryo-derived teratocarcinoma cell lines. Nature 309, 255–256 (1984).
Acknowledgements
We thank Dr. E. Zsigmond and A. Domozhirov for their help in the blastocyst microinjection assay and for the discussion of the results. We also thank to Dr. Zhengmei and Dr. A. L. Hazen for their continuous help and advices for the immunohistochemistry and FACS analysis and to Justin Pham for his help in karyotype analysis. We also thank Mr. James H. Cummins for his editorial assistance.
Author information
Authors and Affiliations
Contributions
K.V. performed experiments and wrote the manuscript. H.P. performed or assisted with some of the experiments, including the iMuSCs’ isolation. X.M. involved in the project discussion and assisted some of the experiments. Y.L. designed the project and wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Vojnits, K., Pan, H., Mu, X. et al. Characterization of an Injury Induced Population of Muscle-Derived Stem Cell-Like Cells. Sci Rep 5, 17355 (2015). https://doi.org/10.1038/srep17355
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep17355
This article is cited by
-
Kinins: Locally formed peptides during inflammation with potential use in tissue regeneration
Inflammation Research (2023)
-
DOCK6 promotes chemo- and radioresistance of gastric cancer by modulating WNT/β-catenin signaling and cancer stem cell traits
Oncogene (2020)
-
Emerin anchors Msx1 and its protein partners at the nuclear periphery to inhibit myogenesis
Cell & Bioscience (2019)
-
Functional Neuronal Differentiation of Injury-Induced Muscle-Derived Stem Cell-Like Cells with Therapeutic Implications
Scientific Reports (2017)
-
Novel Regenerative Therapies Based on Regionally Induced Multipotent Stem Cells in Post-Stroke Brains: Their Origin, Characterization, and Perspective
Translational Stroke Research (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.