Abstract
mRNA poly(A) tails are important for mRNA stability and translation and enzymes that regulate the poly(A) tail length significantly impact protein profiles. There are eleven putative deadenylases in mammals and it is thought that each targets specific transcripts, although this has not been clearly demonstrated. Nocturnin (NOC) is a unique deadenylase with robustly rhythmic expression and loss of Noc in mice (Noc KO) results in resistance to diet-induced obesity. In an attempt to identify target transcripts of NOC, we performed “poly(A)denylome” analysis, a method that measures poly(A) tail length of transcripts in a global manner and identified 213 transcripts that have extended poly(A) tails in Noc KO liver. These transcripts share unexpected characteristics: they are short in length, have long half-lives, are actively translated and gene ontology analyses revealed that they are enriched in functions in ribosome and oxidative phosphorylation pathways. However, most of these transcripts do not exhibit rhythmicity in poly(A) tail length or steady-state mRNA level, despite Noc’s robust rhythmicity. Therefore, even though the poly(A) tail length dynamics seen between genotypes may not result from direct NOC deadenylase activity, these data suggest that NOC exerts strong effects on physiology through direct and indirect control of target mRNAs.
Similar content being viewed by others
Introduction
Poly(A) tails are hallmarks of most eukaryotic mRNAs, serving to protect the mRNA from degradation and promote circularization of the RNA to allow efficient translation initiation1. Changes in poly(A) length occur throughout the life of the mRNA and long poly(A) tails of ~150–250 nt are initially acquired during the 3′ end processing of the nascent transcripts in the nucleus. After transcripts are translocated into the cytoplasm, they subsequently become shortened by deadenylases, a group of ribonucleases specific for homopolymeric tracts of adenosines1. Deadenylation is a rate-limiting step for mRNA degradation that occurs in both mRNA- and context- specific manners and can result in RNA de-stabilization or translational silencing2.
There are 11 putative deadenylases currently identified in mammals based on their sequence homology and most of them have been shown to play important roles in regulating diverse processes, ranging from general viability of organisms to bone formation, cell growth and metabolism2,3,4,5,6,7,8. All known deadenylases are magnesium-dependent and belong to one of two superfamilies: The DEDD (Asp-Glu-Asp-Asp) superfamily contains POP2 (or CAF1), CAF1Z, PARN and PAN2 families, whereas the second superfamily includes members related to a class of Exonucleases, Endonucleases and Phosphatases, known as the EEP (or CCR4) superfamily that includes CCR4, NOCTURNIN, ANGEL, 2′PDE2,9. It still remains unclear, however, why there are so many deadenylases, whether they are functionally redundant or have distinct roles and whether these deadenylases act on specific target mRNAs. As the deadenylation process generally occurs in a biphasic manner in mammals, in which long poly(A) tails are first gradually shortened by the PAN2-PAN3 complex to ~100 nt, followed by the CCR4-CAF1-NOT complex to 8–12 nt10, it is possible that multiple deadenylases act on the same mRNA with discrete but overlapping functions. However, given the difference in temporal and spatial expression patterns of the deadenylases5,8,10,11 and the different phenotypes caused by disrupting specific deadenylases3,4,5,6,7, it is more likely that each deadenylase targets a specific set of transcripts, although this has not been clearly demonstrated.
Among these deadenylases, Noc (Gene name; Ccrn4l) is unique, in that it exhibits a robustly rhythmic expression pattern driven by the biological clock, peaking during the night with particularly high amplitude rhythms in liver12,13. Noc is also unique in that it is an immediate early gene (IEG), showing acute responses to several stimuli including serum shock, phorbol ester, lipopolysaccharide (LPS) and rosiglitazone, a peroxisome proliferator-activated receptor γ (PPARγ) agonist14,15,16. NOC has a conserved catalytic domain in the C-terminus, with sequence similarities to other CCR4 family members, but it has a significantly divergent N-terminus and lacks the leucine-rich repeat region required for yeast Ccr4p and mammalian CCR4a and CCR4b to interact with Caf1 and other proteins in the major CCR4–NOT complex10,11,17. These differences in expression pattern and structure suggest that NOC has a function distinct from other members of this protein family.
Mice lacking Noc (Noc KO) are resistant to diet-induced obesity and hepatic steatosis, yet do not have reduced food intake, increased activity, or measureable changes in whole body energy expenditure when fed a High-Fat Diet (HFD)3. This phenotype is due, at least in part, to abnormal dietary lipid trafficking in intestinal enterocytes, preventing efficient energy intake from diet18. Noc is also one of the differentiation switches of mesenchymal stromal cells (MSCs), supporting a shift towards adipogenesis rather than osteogenesis15,19. However it is unknown whether the deadenylase function of NOC contributes to these phenotypes and if so, which specific transcripts NOC targets for its deadenylase activity.
Because Noc exhibits a unique expression pattern that is different from other deadenylases8,16, we hypothesized that NOC would exert its enzymatic activity on a specific set of transcripts and shorten their poly(A) tail length and a lack of such regulation would ultimately lead to the phenotypes observed in Noc KO mice. In order to attempt to identify these transcripts, we used our recently developed technique, a genome-wide screen which we call “Poly(A)denylome” analysis to assess the poly(A) tail length of mRNAs from WT and Noc KO livers in an unbiased manner. Using this analysis, we identified 213 transcripts that have longer poly(A) tail lengths in Noc KO mouse liver. Despite Noc’s robust rhythmicity, however, the great majority of these transcripts do not exhibit rhythmicity in poly(A) tail length or in their steady-state mRNA level in mouse liver. Instead, these transcripts share surprising yet interesting characteristics: they are generally short in length and have relatively long half-lives. Gene ontology analysis of these transcripts, as well as other mRNAs that have altered abundance in the Noc KO (both up- and down-regulated), suggests that NOC-regulated transcripts are enriched in those involved in ribosome functions and in oxidative phosphorylation. Most of the transcripts that have altered tail length in the Noc KO livers do not fit our expectations for direct targets of NOC and this suggests that NOC acts, at least in part, through a novel activity or via indirect control of target mRNA metabolism that ultimately contribute to the phenotypes observed in Noc KO mice.
Results and Discussion
Identification of transcripts that have different poly(A) tail lengths in Noc KO liver
In order to identify mRNAs that have altered poly(A) tail lengths in the Noc KO mice, we performed “poly(A)denylome” analysis8, a method that can measure poly(A) tail length of transcripts in an unbiased and global manner. Mouse livers were collected at Zeitgeber Time 4 (ZT 4; where ZT 0 is defined as time (hours) of lights on and ZT 12 is defined as time of lights off) and ZT 16 (Fig. 1a), time points when the NOC expression is at its nadir and peak, respectively (Supplemental Fig. S2)12,13. RNAs were then extracted and fractionated by oligo(dT) chromatography into populations that contained either short (<75 nt) or long (60–250 nt) poly(A) tails by varying the salt concentrations in the elution step8,20 (Fig. 1b). Non-fractionated mRNAs (the total poly(A)+ pools) were also obtained as a reference. These RNA samples were subjected to microarray analysis using the Affymetrix Gene ST 1.0 chips that do not require oligo(dT)-based sample preparation to avoid bias from the different poly(A) lengths in the different fractionated samples. Since the correlation of gene expression between each RNA pool was quite strong (Supplemental Fig. 1a–d)8, we used the long/short ratio from the microarray analyses as an indicator of poly(A) tail length. Indeed, this normalization largely eliminated the correlation of the expression levels in each fraction (Supplemental Fig. 1d) and we have previously shown that this ratio provides reliable quantitative information on the relative poly(A) tail length8.
Poly(A)denylome analysis to identify target mRNAs of NOC.
(a) Shown is the tissue sampling scheme in which mouse livers were harvested at ZT 4 and ZT 16 under 12:12 light:dark (12L:12D) conditions from both WT and Noc KO mice (n = 3 per time point). (b) The bulk poly(A) tail length (nt) of each fraction was tested by 3′-end labeling assay. Representative gel images are shown. Oligo(dT) chromatography was used to separate total RNA into fractions with either short or long poly(A) tails by varying salt concentrations in the elution8. (c,d) The correlation of the long/short ratio between genotypes (c) and time points (d). (e) Validation of transcripts that have differences in their poly(A) tail length between WT and Noc KO. Shown are long-short ratios (e) and direct measurements of tail-length by LM-PAT or PAL-PAT assay (f). Representative gel images are shown on top and poly(A) tail length distribution calculated by Image J is shown on the bottom. TVN is an oligonucleotide that predominantly recognizes A12, serving as an internal control. The horizontal lines across distribution plot depict the expected location of poly(A)12. Each lane on gel consists of pooled samples (n = 3 for each time point). All the graphs represent mean ± SEM, *p < 0.05 (Student’s t-test).
As was previously reported, there are dynamic circadian changes in poly(A) tail length in WT mouse liver8. Since NOC expression is higher during the night (i.e. ZT 16) compared to the daytime (i.e. ZT 4), we expected that the effect of NOC would be stronger and the number of transcripts with longer poly(A) tail in Noc KO would be higher at ZT 16. However, the correlation of the long/short ratio between WT and Noc KO was weaker at ZT 4 than at ZT 16, indicating there is more variability between two genotypes at ZT 4 (Fig. 1c). We also observed that the long/short ratio was more variable between the two time points in Noc KO than in WT (Fig. 1d), despite the known circadian rhythms in tail length in the WT8.
In order to identify potential NOC target transcripts, we compared the long/short ratio between WT and Noc KO samples and identified transcripts that had significantly different ratios (i.e. tail lengths) in the absence of NOC. Using stringent criteria in which we only examined transcripts that had raw expression values of greater than 200 and a fold-change ratio of greater than 1.5, we identified 309 transcripts that exhibited a statistically significant difference (p < 0.05) at ZT 4 and 10 transcripts at ZT 16 (Supplementary Table 1). All 10 transcripts that were found to differ at ZT 16 had longer tails (higher long/short ratios) in the Noc KO samples, while at ZT 4, 203 transcripts had longer tails and 106 transcripts had shorter tails in the Noc KO liver. Although it remains unclear why these 106 transcripts had shorter poly(A) tails in Noc KO liver, it may be that other deadenylases (all expressed in liver8) over-compensate for the lack of Noc. It is likely that these differences underestimate the number of transcripts that have altered poly(A) tail length in the KO due to the stringent criteria we used to call a difference in the long/short ratio. In addition, our method does not capture mRNAs with extremely short tails, or lacking tails altogether, which may represent a significant population of mRNAs20,21,22,23,24.
In order to independently validate the microarray results, the poly(A) tail lengths of randomly chosen transcripts were directly measured using the ligation-mediated poly(A) tail length (LM-PAT) assay25 as well as Poly(A) Length (PAL) assay26 (Fig. 1e,f). These assays revealed that the poly(A) tail length of the mRNAs we tested indeed exhibited changes between WT and Noc KO, as predicted from microarray analyses (Fig. 1e). Due to the heterogeneous nature of poly(A) tail lengths, signals are often detected as smears, therefore, the distribution of sizes detected by densitometry analysis was used to compare the poly(A) tail length between genotypes (Fig. 1f). In addition, these PAT assays also displayed multiple differently-sized PCR products for some mRNAs (such as Bloc1s1 or Hspb1 in Fig. 1f), either possibly arising from alternative polyadenylation which occurs in more than 70% of all genes27 or representing two populations of transcripts: newly synthesized RNAs with long poly(A) tails and old RNAs that have shortened poly(A) tails.
Changes in poly(A) tail length do not correlate with changes in mRNA abundance
Because the poly(A) tail length is an important factor in determining the translation initiation and mRNA stability, it has long been thought that depletion of deadenylases would lead to the stabilization of mRNAs, hence increased mRNA abundance1. Recent studies, however, have challenged this idea and shown that the loss of a deadenylase can have only small effects on changes in mRNA abundance and changes in mRNA stability have an inverse correlation with mRNA abundance28,29. In addition, there is accumulating evidence that mRNA decay and transcription are coupled to buffer mRNA abundances and to maintain RNA homeostasis and increased mRNA stability is not predictive of increased abundance30,31,32. Because “mRNA abundance” (i.e. the steady-state mRNA levels) represents the sum of RNA degradation and de novo RNA synthesis, longer half-lives may not always result in higher mRNA abundance. We therefore tested whether this is also true in Noc KO mouse livers by analyzing the changes in mRNA abundance, obtained from the poly(A)+ (non-fractionated) pool and comparing this with changes in poly(A) tail length between WT and Noc KO. Interestingly, we did not observe any correlation at either time point (Fig. 2a), supporting the idea that these NOC-dependent changes in poly(A) tail length do not necessarily correlate with the changes in mRNA abundance.
Changes in the poly(A) tail length do not correlate with changes in mRNA abundance.
(a) Correlation of changes in the long/short ratio vs mRNA abundance between WT and Noc KO at each time point. The degree of correlation is shown in the upper right hand corner of each graph. (b) Changes in mRNA abundance for transcripts that had longer (left) or shorter (right) poly(A) tail length in Noc KO. (c) Changes in mRNA abundance of the same four candidate transcripts whose poly(A) tail length difference between WT and Noc KO were validated in Fig. 1. mRNA abundance was analyzed by microarray (upper) and qPCR normalized by the expression of Rplp0 (36B4) (lower). N = 3 for each sample. All the graphs represent mean ± SEM, *p < 0.05, **p < 0.01, ***p < 0.005 (Student’s t-test). (d) mRNA stability in WT and Noc KO MEFs. MEFs were treated with 5 ug/ml Actinomycin D for 0, 0.5, 3, 6, 9 hrs and cells were harvested at each time point. The expression of each transcript was measured by qPCR. The expression level of time 0 was set to 1 and all the other data were normalized accordingly. Graphs represent mean ± SEM (n = 3–4).
Since we expected NOC targets to have increased tail lengths in the KO, we also examined the relationship between mRNA abundance and poly(A) tail length specifically among the 213 transcripts (from both time points) that had longer poly(A) tail length in Noc KO liver. We found that 29 of these transcripts (13.6%) had higher mRNA abundance, but all other transcripts (86.4%) had no changes in mRNA abundance in the Noc KO samples. Among the 106 transcripts that had shorter poly(A) tails in Noc KO, 1 transcript (0.9%) had lower mRNA abundance and another transcript (0.9%) had higher mRNA abundance in Noc KO, while all other transcripts (98.2%) were present at equal levels in both genotypes (Fig. 2B). The changes of mRNA abundance from the microarray data was further confirmed by qPCR for several transcripts that had altered poly(A) tail length (Fig. 2c). In addition to confirming the general lack of correlation between mRNA abundance and poly(A) tail length, in some cases even small differences that had been observed in abundance in the microarrays (i.e. Bloc1s1 and Cd52), did not reach the level of significance when re-examined by quantitative RT-PCR (Fig. 2c). These results suggest that tail shortening by NOC does not usually result in changes in mRNA abundance, but may have some other function.
We also measured the mRNA stability of several transcripts in mouse embryonic fibroblasts (MEFs) derived from both WT and Noc KO mice. We observed small changes in half-life in some of the mRNAs that we tested, but for most of the mRNAs we examined, there was no difference in half-life in the Noc KO MEFs (Fig. 2d).
Characteristics of transcripts that have longer poly(A) tails in the Noc KO
The 213 transcripts whose poly(A) tail length (i.e. long/short ratio) was longer in Noc KO were of particular interest, because these transcripts fit the pattern expected for bona fide target transcripts of NOC. Therefore, we further hypothesized that NOC target transcripts would share common characteristics that would distinguish themselves as targets. In general, the fate of an mRNA is largely determined by the composition and timing of the interaction between trans-acting factors (i.e. miRNAs and RNA-binding proteins) and cis-elements commonly found in 5′ or 3′ untranslated regions (UTRs) of mRNAs. Therefore, we first attempted to identify cis-elements commonly found in either UTR of the 213 mRNAs that had longer tails in the KOs. However, there was no enrichment of any specific RNA sequence found by HOMER RNA Motif Analysis (http://homer.salk.edu/homer/motif/rnaMotifs.html) (data not shown), although this does not rule out the possibility that NOC recognizes a cis motif defined by RNA secondary structure, rather than RNA sequence.
Because we have previously shown that many mRNAs have poly(A) tail lengths that change over the daily cycle and because Noc is expressed with robust rhythms in mouse liver, we hypothesized that NOC might contribute to this circadian control of poly(A) tail length and therefore NOC targets would be enriched for transcripts with rhythmic poly(A) tails. However, this does not appear to be the case, as only 3 (1.4%) of the 213 putative NOC targets (Lims2 (peak ZT 16.4), Alkbh7 (ZT 8.6), Hspb1 (ZT 16.2) were found to have rhythmic poly(A) tails in our previous analysis8, a significant underrepresentation compared to all mRNAs, of which 2.3% had rhythmic poly(A) tail lengths (Fig. 3a). None of the transcripts with shorter poly(A) tail length in Noc KO had rhythmic poly(A) tails (Fig. 3a).
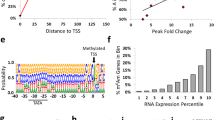
mRNAs with longer tails in the Noc KO are short in length, have longer mRNA half-lives and are actively translated.
(a) Poly(A) rhythmicity (%)8 (ALL; n = 10217, Long(A); n = 148, Short(A); n = 95), (b,c) Percentage of transcripts that were shown to be rhythmic from circadian transcriptome analyses in mouse liver. (b) mature RNA expression34 (ALL; n = 20069, Long(A); n = 213, Short(A); n = 106) and c) exon expression33 (ALL; n = 28661, Long(A); n = 213, Short(A); n = 106). (d–g) The length of 5′UTR (d) (ALL; n = 33560, Long(A); n = 114, Short(A); n = 77), CDS (e) (ALL; n = 29620, Long(A); n = 119, Short(A); n = 78), 3′UTR (f) (ALL; n = 33562, Long(A); n = 116, Short(A); n = 80) and gene (g) (ALL; n = 31118, Long(A); n = 127, Short(A); n = 78). (h-i) mRNA half-lives (hr) in mESCs35 (h) (ALL; n = 15717, Long(A); n = 93, Short(A); n = 63) and NIH3T3 cells32 (i) (ALL; n = 4356, Long(A); n = 33, Short(A); n = 16). (j) Ribosome footprint density as indicated by the numbers of transcripts bound to polysome fractions in mESCs36 (ALL; n = 20055, Long(A); n = 118, Short(A); n = 74). All the graphs represent mean ± SEM, *p < 0.05, **p < 0.01, ***p < 0.005 (Student’s t-test). All analysis was done on 213 transcripts pooled from both the ZT 4 and ZT 16 time points.
In addition, we had originally expected that NOC’s rhythmic expression pattern would contribute to rhythmic mRNA profiles by destabilizing target transcripts during the night. However, this was not the case, because only 333 (Alkbh7 (peak ZT 21.8), Safb (ZT 21.7), Hist1h1c (ZT 22.1)) or 634 (Saa1 (peak ZT1.4), Hist1h1c (ZT 0.8), Arid3a (ZT 22.4), Cd52 (ZT 6.4), Dct (ZT 3.2), Hspb1 (ZT 20.8)) of the 213 transcripts were determined to be rhythmic in abundance, based on the two recent datasets of extensively analyzed circadian mRNA profiles at the steady-state level in mouse liver33,34, a much smaller percent than the percent of total rhythmic mRNAs (Fig. 3b,c).
A recent study also made an interesting observation that the intrinsic poly(A) tail length does not correlate with open reading frame (ORF), untranslated region (UTR) and mRNA length in mouse liver26, although in yeast there is a correlation between poly(A) tail length and UTR, mRNA and ORF length21. In order to gain insight into whether this group of mRNAs with extended tails in the Noc KO show any such correlation in gene/transcript length, we analyzed the length of 5′- or 3′-UTRs, ORF and the entire gene. Interestingly, all these parameters were significantly shorter in the transcripts that had longer poly(A) tails in Noc KO, as compared to all the transcripts included in the dataset (ALL) (Fig. 3d–g).
It has long been thought that one of the major roles of poly(A) tails is to determine mRNA stability1. Therefore, despite the lack of correlation with overall abundance, we wondered whether the mRNAs with longer tails in the Noc KO were generally long- or short-lived mRNAs. To this end, we utilized genome-wide datasets of mRNA half-life measurements from mouse embryonic stem cells (mESCs) and NIH3T3 cells32,35 and examined reported half-lives of this group of mRNAs. Our in silico analyses using these datasets revealed that the transcripts with longer tails in the Noc KO have an average half-live that was significantly longer than the average half-live of both the entire group of all mRNAs and those with shorter poly(A) tails in Noc KO. This was true for both the mESCs and NIH3T3 cells (Fig. 3h,i).
Another important function of poly(A) tail length is to regulate translation initiation, as mRNAs form a “closed-loop” circular structure in order to initiate translation and the presence of longer poly(A) tails facilitates the formation of this structure1, although this relationship has recently been challenged in some situations26. Thus, in order to clarify whether NOC may function in translation, we next examined a ribosomal profiling dataset from mES cells36, an indicator of translation status. In this dataset, the Noc target transcripts are associated with larger numbers of ribosomes, compared to all transcripts and those with shorter tails in the KO (Fig. 3j), indicating that Noc target transcripts may belong to a more actively translated group of mRNAs at least in mES cells.
Since poly(A) tail length impacts translation initiation and NOC targets are generally translationally active (Fig. 3j), we hypothesize that the NOC-dependent changes may be reflected in translational competence and therefore intrinsic protein levels. Recent proteomic analyses from mouse liver over the circadian cycle have provided datasets on rhythmic protein levels37,38, but the proteins encoded by most of the 213 candidate NOC targets were not represented in these datasets, likely due to the sensitivity limits of the mass spectroscopy.
Together, these results suggest that NOC acts either directly or indirectly on specific sets of target mRNAs to shorten their poly(A) tails but this tail shortening does not generally affect the RNA stability or overall abundance of these mRNAs and NOC does not seem to significantly regulate rhythmicity of poly(A) tail length or mRNA abundance. Consistent with this, these target mRNAs are generally quite stable, suggesting that tail-shortening by NOC may not be important for regulating mRNA half-life. Nevertheless, it still remains a possibility that NOC regulates mRNA stability, regardless of the changes in poly(A) tail length, until direct NOC-targets can be identified. It should be noted that we have previously reported that NOC stabilizes the iNOS mRNA although it is still unclear whether this is a direct effect14.
Nocturnin’s role in ribosome and mitochondrial oxidative phosphorylation
In order to determine whether the transcripts that have longer poly(A) tails in Noc KO liver belong to particular biological pathways and functions, we performed gene ontology analyses using DAVID39,40. Pathways significantly enriched in transcripts with longer poly(A) tail length in Noc KO were: Ribosomes, Cytosol, Oxidative Phosphophorylation and Laminin and these keywords were not enriched in transcripts with shorter poly(A) tail (Supplemental Table S3). Similar results were also obtained when we used JEPETTO41, a different algorithm that enabled visualization of the network analysis data (Fig. 4a,b,d,e).
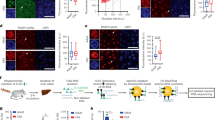
Gene ontology analyses extracted “ribosome” and “oxidative phosphorylation” as biological pathways affected by a loss of NOC.
(a,d) Transcripts that have longer poly(A) tail (i.e. higher long/short ratio) involved in ribosome cluster (DAVID: p < 0.001, JEPETTO: q < 0.001) (a) and oxidative phosphorylation cluster (DAVID: p < 0.05, JEPETTO: q = 0.234) (d). Transcripts that had altered mRNA abundance involved in ribosome cluster (DAVID: p < 0.01, JEPETTO: p = 0.084) (b) and oxidative phosphorylation cluster (DAVID: p < 0.01, JEPETTO: p = 0.106) (e). Data visualization was performed with JEPETTO41. Blue nodes represent genes overlapped between target dataset and each process, while green nodes indicate genes specific to each process and grey nodes depict target dataset specific genes. Venn diagrams on the right (c,f) denote the numbers of transcripts extracted by DAVID39 that are known to function in ribosomes (c) or oxidative phosphorylation pathways (f). Orange circle represents the number of transcript that had higher long/short ratio in Noc KO, while blue circle represents the number of transcript that had altered mRNA abundance in Noc KO liver. (g) Hepatic ATP levels (WT: n = 14, Noc KO: n = 10). (h) Serum beta-hydroxybutyrate levels (WT: n = 9, Noc KO: n = 9). (i) Relative copy number of mitochondrial DNA measured by the ratio of mtDNA (mtCoI) and nuclear DNA (18srRNA) in liver (WT: n = 18, Noc KO: n = 18). (j) The mRNA expression of mitochondrial-encoded RNAs in liver measured by qPCR (WT: n = 18, Noc KO: n = 18). Expression was normalized by nuclear-encoded Rplp0 expression and the relative expression level of WT was set as 1. All the graphs represent mean ± SEM, *p < 0.05, ***p < 0.005 (Student’s t-test).
We also performed gene ontology analyses using transcripts that had altered mRNA abundance in Noc KO, independent of whether or not their poly(A) tail length changed and also found the pathways “Mitochondrion” and “Ribosome” to be significantly enriched and this enrichment was particularly strong in a set of genes that had increased mRNA abundance in Noc KO (Supplemental Table 4). Interestingly, this enrichment of transcripts in ribosomal and mitochondrial functions in both datasets is not due to the same sets of transcripts present in both poly(A) and mRNA abundance datasets. For example, among 31 transcripts (out of a total of 213 transcripts) that have differences in their poly(A) tail length (Supplementary Table 1) and 53 transcripts (out of a total of 196 transcripts) that have altered mRNA abundance (Supplementary Table 2) between the two genotypes that are involved in regulating mitochondrial functions, only 9 transcripts (Atp5l, Bloc1s1, Mrps24, Mrps28, Ndufa2, Romo1, Tmem256, Timm13, Uqcc2, Uqcr) were included in both datasets (Fig. 4f). Similarly, among 41 (poly(A) tail length) and 40 (mRNA abundance) transcripts that display functions in ribosome, only 6 transcripts (Mrps24, Mrps28, Rpl35, Rps29, Snrpd2, Timm13) were present in both datasets (Fig. 4c). This suggests that NOC plays a prominent function (direct or indirect) in regulation of these biological processes/pathways, through both poly(A) tail length shortening and via changes in mRNA abundance. Because of NOC’s known role in metabolism, we were particularly interested in a potential role for NOC in mitochondrial function. Therefore, we measured hepatic ATP levels as well as serum beta-hydroxybutyrate levels and found that both are decreased in Noc KO (Fig. 4g,h), despite that the total number of mitochondria is unchanged (Fig. 4i). Since the function of oxidative phosphorylation is to produce ATP and beta-hydroxybutyrate predominantly in liver from acetyl-CoA by oxidizing fatty acids, these data suggest that the mitochondrial functions are altered in the Noc KO. To support this idea, the levels of Nd4/4L, Nd5, Nd6, CoxI and CoxII, all of which are mitochondrial RNAs that encode proteins that are involved in Complex I and Complex IV of oxidative phosphorylation chains, are decreased in Noc KO (Fig. 4j), further suggesting a role for NOC in regulating the oxidative phosphorylation pathway.
In conclusion, our Poly(A)denylome analysis identified transcripts that have altered poly(A) tail length in Noc KO. A significant portion of these transcripts are related to mitochondrial function as revealed by gene ontology analyses (Supplemental Table 3-4) and may contribute to the metabolic phenotypes observed in the Noc KO mice3,42. However, transcripts that have longer poly(A)tails in Noc KO liver are relatively stable and the majority of these transcripts do not exhibit rhythmicity in poly(A) tail length or in their steady-state mRNA level in mouse liver, even though the expression of Noc is robustly rhythmic. Therefore, although the poly(A) tail lengths are altered, these may not be direct target transcripts of NOC deadenylase activity. Further supporting this idea is the finding that the number of transcripts that have longer poly(A) tails is higher at ZT 4 when the expression of Noc hits nadir, as compared to ZT 16 that the level of Noc reaches its peak. In fact, we have not been able to detect any direct physical interaction between NOC and possible target transcripts, even though we have attempted several different techniques with various materials (i.e. cell lines vs tissues, endogenous vs overexpressed NOC). It may be that because NOC does not have an RNA binding domain outside of its catalytic pocket, it only associates with its targets transiently, making it difficult to trap the targets. Together these findings suggest that NOC may be regulating tail length through an indirect mechanism, or through some novel activity. One possibility is that NOC competes against some other deadenylase activity in a rhythmic manner. We have previously shown that the deadenylase activity of NOC is moderate, compared to that of PARN16, therefore, slow or incomplete deadenylation by NOC may block complete deadenylation by another more active deadenylase. It will be of interest to explore whether NOC has a molecular function other than a deadenylase.
Materials and Methods
Poly(A)denylome analysis
RNA fractionation and the 3′-end labeling assay were performed as described previously8. Briefly, mouse liver total RNAs were isolated and resuspended in PolyATract GTC extraction buffer (Promega). RNAs were then mixed with Biotinylated Oligo(dT) Probe (Promega) in dilution buffer (6xSSC, 10 mM Tris-HCl (pH 7.6), 1mM EDTA, 0.25% SDS, 1% β-mercaptoethanol) and incubated for 10 min at 70 °C. After centrifugation at 12,000 g for 10 min at room temperature, supernatant was mixed with Streptavidin MagneSphere Paramagnetic Particles (Promega) and incubated for 15 min at room temperature while nutating. After three washes with 0.5x SSC at room temperature, short poly(A) RNAs were eluted by 0.075x SSC and subsequently long poly(A) RNAs were eluted by DEPC-treated water. Alternatively, total Poly(A)+ RNAs were eluted by DEPC-treated water immediately after the washing step. For bulk poly(A) tail length analysis, RNAs were 3′-end labeled with [32P]-pCp with T4 RNA ligase (EPICENTRE) overnight at 4 °C, followed by RNaseA/T1 (Fermentas) digestion to remove the bodies of the mRNAs. The non-digested poly(A) tails were then resolved by 7.5% denaturing PAGE and detected on X-ray film. Fractionated or non-fractionated RNAs were further purified using RNeasy MinElute Cleanup Kit (QIAGEN) for microarray that was performed at Molecular Biology Core Facilities at Dana-Farber Cancer Institute. For each time point at ZT 4 and ZT 16, three biological replicas from WT and Noc KO mouse liver were each hybridized to individual microarray chips (Affymetrix Mouse Gene ST 1.0), according to the manufacturer’s instruction. Raw data were analyzed and normalized by the quantile method through the dChip software. Probe sets were annotated using the Affymetrix annotation file as of February 2009.
Poly(A) tail length measurement
LM-PAT assays were performed as described previously8,25 with a slight modification. In brief, 50 ng of poly(A)+ enriched RNAs extracted using PolyATract System 1000 (Promega), were first incubated with 5′-phosphorylated oligo(dT)15 in the presence of T4 DNA ligase for 30 min at 42 °C to anneal with poly(A) tails of RNAs, followed by an excess amount of anchor primer with oligo(dT)12 (5′-GCGAGCTCCGCGGCCGCGTTTTTTTTTTTT-3′) to anneal at the end of poly(A) tails. This was further incubated for 2 h at 12 °C to complete ligation between oligo(dT)s. These oligo(dT)-annealed RNAs were then subjected to reverse transcription reaction using SuperScript III (Life Technolgoies) for cDNA synthesis.
PAL-PAT assays were performed based on the PAL-seq library construction method previously reported26. In brief, 1 ug of poly(A)+ enriched RNAs extracted using PolyATract System 1000 (Promega), were first incubated with an adapter (5′-Phos.AGA TCG GAA GAG CGT CGT GTA GGG AAA GAG TGT AGA CAC ATA C-3′) and a splint (5′-TTC CGA TCT TTT TTT TT-3′) in the presence of RNA ligase 2 (NEB) at 18 °C for overnight to attach the splint adapter at the 3′-end of poly(A)+ RNAs. These adapter-ligated RNAs were then partially digested by RNaseT1 (Life Technologies) for 30 min at room temperature, followed by reverse transcription reaction using SuperScript III (Life Technolgoies) with solexa_rt_primer (5′-AAT GAT ACG GCG ACC ACC GAG ATC TAC ACT CTT TCC CTA CAC G -3′) after inactivating RNaseT1 by Precipitation/inactivation buffer (Life Technologies).
Aliquots of this cDNA were used as templates for PCR reactions with message-specific primers of our mRNAs of interest. Then, PCR products were digested by a restriction enzyme to confirm the specificity. Resulting DNA fragments were visualized by Gel-Doc system (Bio-Rad) and poly(A) tail length distribution was visualized by Image J. Primer sequences and restriction enzymes used in both PAT assays can be found in Table S5.
Animals, cells, qRT-PCR and DNA extraction
Male mice of WT and Noc KO3 were maintained on a 12:12 LD cycle and fed ad libitum. All the procedures were performed in accordance with the Guideline of Institutional Animal Care and Use Committee (IACUC) of UT Southwestern Medical Center and were approved by the IACUC of UT Southwestern Medical Center. Mouse liver samples for Poly(A)denylome, qPCR and Western blot analyses were taken from three biological replicas of WT and Noc KO at ZT 4 and ZT 16. Immediately after mice were sacrificed, livers were macrodissected and then snap-frozen in liquid nitrogen. Frozen livers were stored at −80C until RNA, DNA, proteins were extracted within one year.
Frozen mouse livers were mechanically homogenized (Kinematica) to extract RNA and protein. Total RNAs were extracted using TRIZOL reagent (Life Technologies) and then poly(A) enriched RNAs were extracted by Poly(A)Tract System (Promega). After the RNA quantification by NanoDrop (Thermo Scientific), 50 ng of poly(A) enriched RNAs were subjected to cDNA synthesis (total volume 20 ul) with Oligo(dT)12-18 primer (Life Technologies) using 1 ul/sample of SuperScript II (Life Technologies) according to manufacturer’s instructions. DNase I treatment was not performed at any point. Quantitative PCR (qPCR) was performed using ABI7900 (Applied Biosystems) with SYBR Power Green (Applied Biosystems) with a total volume of 10 ul/well in 384 well plate including 1ul of cDNA (diluted in 1/50 after cDNA synthesis) using relative standard curve method. To quantify each gene expression, a standard curve was generated for each primer set with serial 5-fold dilutions (linear dynamic range 1/10-1/6250) using the WT mouse liver cDNAs and the limit of detection was empirically determined. Cycling parameters were as follows: holding at 50 °C for 2 min and 95 °C for 10 min, then 40 cycles of 95 °C for 15 sec and 60 °C for 1 min. Subsequently, melting curves were drawn to check the target specificity of each reaction. Primers were designed to flank intron sequences and include all the isoforms whenever possible and their specificity was tested by BLAST. Primer concentrations were determined based on the test run to yield the most reproducible and reliable results at the lowest concentration and there was no obvious PCR inhibition observed with any primer pair. No primer pairs yielded any signals for Non-Template (DNA) control under our conditions. Primer information used in qPCR analyses can be found in Table S5 and the parameters for standard curves for each gene can be found in Table S6. Data were analyzed by SDS v2.3 (Applied Biosystems) and each Ct was determined automatically. All the data were normalized to the expression level of Rplp0 (36B4), as Rplp0 has been shown to have no fluctuation in the gene expression around the circadian clock in mouse liver43 and we did not omit any data as outliers. cDNA synthesis and qPCR was performed once using three biological replicas, all of which were run as duplicates in qPCR.
Nuclear and mitochondrial DNAs were extracted from frozen liver samples using DNeasy Blood and Tissue Kit (QIAGEN) according to the manufacturer’s instructions. Isolated DNAs were subjected to qPCR analysis to quantify the relative mt copy number by measuring the relative amount of nuclear-DNA (18srRNA) and mitochondria-DNA (mtCoxI).
Primary mouse embryonic fibroblasts (MEFs) from both WT and Noc KO14 were immortalized by introducing aP53 expressing retrovirus44. These immortalized MEFs were treated with 5 ul/mg actinomycin D (Sigma) for 0, 0.5, 3, 6 and 9 hrs before RNA was extracted.
Hepatic ATP and serum beta-hydroxybutyrate measurements
Hepatic ATP concentration was measured as previously described45. Briefly, snap frozen aliquots of liver were homogenized in ice-cold DMEM supplemented with 5% perchloric acid. After centrifuging at 16,000 g for 10 min at 4°C, the homogenates were neutralized by 10N NaOH and 1 M Tris-HCl (pH 7.4), ATP concentration was measured using CellTiter-Glo Luminescent Cell Viability Assay (Promega) according to the manufacturer’s instructions. The results were normalized by measuring protein concentration of each sample. Serum beta-hydroxybutyrate concentration was measured by the Mouse Metabolic Phenotyping Centers (National Mouse Metabolic Phenotyping Centers).
Additional Information
How to cite this article: Kojima, S. et al. Changes in poly(A) tail length dynamics from the loss of the circadian deadenylase Nocturnin. Sci. Rep. 5, 17059; doi: 10.1038/srep17059 (2015).
References
Kuhn, U. & Wahle, E. Structure and function of poly(A) binding proteins. Biochim Biophys Acta 1678, 67–84, 10.1016/j.bbaexp.2004.03.008 (2004).
Goldstrohm, A. C. & Wickens, M. Multifunctional deadenylase complexes diversify mRNA control. Nat Rev Mol Cell Biol 9, 337–344, 10.1038/nrm2370 (2008).
Green, C. B. et al. Loss of Nocturnin, a circadian deadenylase, confers resistance to hepatic steatosis and diet-induced obesity. Proc Natl Acad Sci USA 104, 9888–9893, 10.1073/pnas.0702448104 (2007).
Molin, L. & Puisieux, A. C. elegans homologue of the Caf1 gene, which encodes a subunit of the CCR4-NOT complex, is essential for embryonic and larval development and for meiotic progression. Gene 358, 73–81, 10.1016/j.gene.2005.05.023 (2005).
Morita, M. et al. Depletion of mammalian CCR4b deadenylase triggers elevation of the p27Kip1 mRNA level and impairs cell growth. Mol Cell Biol 27, 4980–4990, 10.1128/MCB.02304-06 (2007).
Morris, J. Z., Hong, A., Lilly, M. A. & Lehmann, R. twin, a CCR4 homolog, regulates cyclin poly(A) tail length to permit Drosophila oogenesis. Development 132, 1165–1174, 10.1242/dev.01672 (2005).
Washio-Oikawa, K. et al. Cnot7-null mice exhibit high bone mass phenotype and modulation of BMP actions. Journal of bone and mineral research : the official journal of the American Society for Bone and Mineral Research 22, 1217–1223, 10.1359/jbmr.070411 (2007).
Kojima, S., Sher-Chen, E. L. & Green, C. B. Circadian control of mRNA polyadenylation dynamics regulates rhythmic protein expression. Genes Dev 26, 2724–2736, 10.1101/gad.208306.112 (2012).
Godwin, A. R., Kojima, S., Green, C. B. & Wilusz, J. Kiss your tail goodbye: The role of PARN, Nocturnin and Angel deadenylases in mRNA biology. Biochim Biophys Acta 1829, 571–579, 10.1016/j.bbagrm.2012.12.004 (2013).
Yamashita, A. et al. Concerted action of poly(A) nucleases and decapping enzyme in mammalian mRNA turnover. Nat Struct Mol Biol 12, 1054–1063, 10.1038/nsmb1016 (2005).
Wagner, E., Clement, S. L. & Lykke-Andersen, J. An unconventional human Ccr4-Caf1 deadenylase complex in nuclear cajal bodies. Mol Cell Biol 27, 1686–1695, 10.1128/MCB.01483-06 (2007).
Kojima, S., Gatfield, D., Esau, C. C. & Green, C. B. MicroRNA-122 modulates the rhythmic expression profile of the circadian deadenylase Nocturnin in mouse liver. PLoS One 5, e11264, 10.1371/journal.pone.0011264 (2010).
Wang, Y. et al. Rhythmic expression of Nocturnin mRNA in multiple tissues of the mouse. BMC Dev Biol 1, 9 (2001).
Niu, S. et al. The circadian deadenylase Nocturnin is necessary for stabilization of the iNOS mRNA in mice. PLoS One 6, e26954, 10.1371/journal.pone.0026954 (2011).
Kawai, M. et al. A circadian-regulated gene, Nocturnin, promotes adipogenesis by stimulating PPAR-gamma nuclear translocation. Proc Natl Acad Sci USA 107, 10508–10513, 10.1073/pnas.1000788107 (2010).
Garbarino-Pico, E. et al. Immediate early response of the circadian polyA ribonuclease nocturnin to two extracellular stimuli. RNA 13, 745–755, 10.1261/rna.286507 (2007).
Draper, M. P., Liu, H. Y., Nelsbach, A. H., Mosley, S. P. & Denis, C. L. CCR4 is a glucose-regulated transcription factor whose leucine-rich repeat binds several proteins important for placing CCR4 in its proper promoter context. Mol Cell Biol 14, 4522–4531 (1994).
Douris, N. et al. Nocturnin regulates circadian trafficking of dietary lipid in intestinal enterocytes. Curr Biol 21, 1347–1355, 10.1016/j.cub.2011.07.018 (2011).
Guntur, A. R. et al. An essential role for the circadian-regulated gene nocturnin in osteogenesis: the importance of local timekeeping in skeletal homeostasis. Ann N Y Acad Sci 1237, 58–63, 10.1111/j.1749-6632.2011.06213.x (2011).
Meijer, H. A. et al. A novel method for poly(A) fractionation reveals a large population of mRNAs with a short poly(A) tail in mammalian cells. Nucleic Acids Res 35, e132, 10.1093/nar/gkm830 (2007).
Beilharz, T. H. & Preiss, T. Widespread use of poly(A) tail length control to accentuate expression of the yeast transcriptome. RNA 13, 982–997, 10.1261/rna.569407 (2007).
Choi, Y. H. & Hagedorn, C. H. Purifying mRNAs with a high-affinity eIF4E mutant identifies the short 3' poly(A) end phenotype. Proc Natl Acad Sci USA 100, 7033–7038, 10.1073/pnas.1232347100 (2003).
Peng, J. & Schoenberg, D. R. mRNA with a <20-nt poly(A) tail imparted by the poly(A)-limiting element is translated as efficiently in vivo as long poly(A) mRNA. RNA 11, 1131–1140, 10.1261/rna.2470905 (2005).
Wu, Q. et al. Poly A- transcripts expressed in HeLa cells. PLoS One 3, e2803, 10.1371/journal.pone.0002803 (2008).
Salles, F. J. & Strickland, S. Analysis of poly(A) tail lengths by PCR: the PAT assay. Methods Mol Biol 118, 441–448, 10.1385/1-59259-676-2:441 (1999).
Subtelny, A. O., Eichhorn, S. W., Chen, G. R., Sive, H. & Bartel, D. P. Poly(A)-tail profiling reveals an embryonic switch in translational control. Nature 508, 66–71, 10.1038/nature13007 (2014).
Tian, B. & Manley, J. L. Alternative cleavage and polyadenylation: the long and short of it. Trends Biochem Sci 38, 312–320, 10.1016/j.tibs.2013.03.005 (2013).
Lee, J. E. et al. The PARN deadenylase targets a discrete set of mRNAs for decay and regulates cell motility in mouse myoblasts. PLoS Genet 8, e1002901, 10.1371/journal.pgen.1002901 (2012).
Mittal, S., Aslam, A., Doidge, R., Medica, R. & Winkler, G. S. The Ccr4a (CNOT6) and Ccr4b (CNOT6L) deadenylase subunits of the human Ccr4-Not complex contribute to the prevention of cell death and senescence. Molecular biology of the cell 22, 748–758, 10.1091/mbc.E10-11-0898 (2011).
Haimovich, G. et al. Gene expression is circular: factors for mRNA degradation also foster mRNA synthesis. Cell 153, 1000–1011, 10.1016/j.cell.2013.05.012 (2013).
Bernstein, J. A., Khodursky, A. B., Lin, P. H., Lin-Chao, S. & Cohen, S. N. Global analysis of mRNA decay and abundance in Escherichia coli at single-gene resolution using two-color fluorescent DNA microarrays. Proc Natl Acad Sci USA 99, 9697–9702, 10.1073/pnas.112318199 (2002).
Schwanhausser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342, 10.1038/nature10098 (2011).
Koike, N. et al. Transcriptional Architecture and Chromatin Landscape of the Core Circadian Clock in Mammals. Science, 10.1126/science.1226339 (2012).
Menet, J. S., Rodriguez, J., Abruzzi, K. C. & Rosbash, M. Nascent-Seq reveals novel features of mouse circadian transcriptional regulation. eLife 1, e00011, 10.7554/eLife.00011 (2012).
Sharova, L. V. et al. Database for mRNA half-life of 19 977 genes obtained by DNA microarray analysis of pluripotent and differentiating mouse embryonic stem cells. DNA Res 16, 45–58, 10.1093/dnares/dsn030 (2009).
Ingolia, N. T., Lareau, L. F. & Weissman, J. S. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147, 789–802, 10.1016/j.cell.2011.10.002 (2011).
Mauvoisin, D. et al. Circadian clock-dependent and -independent rhythmic proteomes implement distinct diurnal functions in mouse liver. Proc Natl Acad Sci USA 111, 167–172, 10.1073/pnas.1314066111 (2014).
Robles, M. S., Cox, J. & Mann, M. In-vivo quantitative proteomics reveals a key contribution of post-transcriptional mechanisms to the circadian regulation of liver metabolism. PLoS Genet 10, e1004047, 10.1371/journal.pgen.1004047 (2014).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature protocols 4, 44–57, 10.1038/nprot.2008.211 (2009).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 37, 1–13, 10.1093/nar/gkn923 (2009).
Winterhalter, C., Widera, P. & Krasnogor, N. JEPETTO: a Cytoscape plugin for gene set enrichment and topological analysis based on interaction networks. Bioinformatics 30, 1029–1030, 10.1093/bioinformatics/btt732 (2014).
Stubblefield, J. J., Terrien, J. & Green, C. B. Nocturnin: at the crossroads of clocks and metabolism. Trends in endocrinology and metabolism: TEM 23, 326–333, 10.1016/j.tem.2012.03.007 (2012).
Yang, X. et al. Nuclear receptor expression links the circadian clock to metabolism. Cell 126, 801–810, 10.1016/j.cell.2006.06.050 (2006).
Ossovskaya, V. S. et al. Use of genetic suppressor elements to dissect distinct biological effects of separate p53 domains. Proc Natl Acad Sci USA 93, 10309–10314 (1996).
Chavin, K. D. et al. Obesity induces expression of uncoupling protein-2 in hepatocytes and promotes liver ATP depletion. J Biol Chem 274, 5692–5700 (1999).
Acknowledgements
We would like to thank Dr. Edward Fox at the Microarray Core Facility of Dana-Farber Cancer Institute for performing microarray analyses. Our mouse colony was managed by Hyun Jeung Sung and Laura Weseman and the authors thank all the Green lab members for helpful discussions and comments, especially Dr. Danielle Hyman for her technical assistance and Jeremy Stubblefield for harvesting tissue samples. This work was supported by National Institutes of Health Grants GM090247, GM076626, GM111387 to C.B.G. and Tomizawa Jun-ichi & Keiko Fund of Molecular Biology Society of Japan for Young Scientist and the Brain and Behavior Research Foundation (NARSAD) to S.K.
Author information
Authors and Affiliations
Contributions
S.K., K.L.G., E.L.S. and P.G. performed experiments and analyzed data. S.K. and C.B.G. wrote the main manuscript and S.K. prepared all the figures.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Kojima, S., Gendreau, K., Sher-Chen, E. et al. Changes in poly(A) tail length dynamics from the loss of the circadian deadenylase Nocturnin. Sci Rep 5, 17059 (2015). https://doi.org/10.1038/srep17059
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep17059
This article is cited by
-
Stage-specific requirement for METTL3-dependent m6A modification during dental pulp stem cell differentiation
Journal of Translational Medicine (2022)
-
ANGEL2 phosphatase activity is required for non-canonical mitochondrial RNA processing
Nature Communications (2022)
-
Circadian- and UPR-dependent control of CPEB4 mediates a translational response to counteract hepatic steatosis under ER stress
Nature Cell Biology (2017)
-
Molecular modulators of the circadian clock: lessons from flies and mice
Cellular and Molecular Life Sciences (2017)
-
Post-transcriptional control of the mammalian circadian clock: implications for health and disease
Pflügers Archiv - European Journal of Physiology (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.