Abstract
Middle East respiratory syndrome coronavirus (MERS-CoV) originated in bats and spread to humans via zoonotic transmission from camels. We analyzed the evolution of the spike (S) gene in betacoronaviruses (betaCoVs) isolated from different mammals, in bat coronavirus populations, as well as in MERS-CoV strains from the current outbreak. Results indicated several positively selected sites located in the region comprising the two heptad repeats (HR1 and HR2) and their linker. Two sites (R652 and V1060) were positively selected in the betaCoVs phylogeny and correspond to mutations associated with expanded host range in other coronaviruses. During the most recent evolution of MERS-CoV, adaptive mutations in the HR1 (Q/R/H1020) arose in camels or in a previous host and spread to humans. We determined that different residues at position 1020 establish distinct inter- and intra-helical interactions and affect the stability of the six-helix bundle formed by the HRs. A similar effect on stability was observed for a nearby mutation (T1015N) that increases MERS-CoV infection efficiency in vitro. Data herein indicate that the heptad repeat region was a major target of adaptive evolution in MERS-CoV-related viruses; these results are relevant for the design of fusion inhibitor peptides with antiviral function.
Similar content being viewed by others
Introduction
Middle East respiratory syndrome coronavirus (MERS-CoV), a newly emerged virus that can cause severe lower respiratory tract infection in humans, was first identified in Saudi Arabia in 20121. Since then, 945 laboratory-confirmed cases of MERS-CoV infection, leading to at least 348 related deaths, have been reported to the WHO (as of January 5th, 2015) (http://www.who.int/csr/don/05-january-2015-mers-jordan/en/).
MERS-CoV belongs to the clade c of betacoronaviruses (betaCoVs)2, which also includes two bat sister species, namely Ty-BatCoV HKU4 and Pi-BatCoV HKU5, isolated from the lesser bamboo bats (Tylonycteris pachypus) and Japanese pipistrelles (Pipistrellus abramus), respectively3. Additional viruses related to MERS-CoV have been described in bats (BtCoV/KW2E-F93, BtCoV/133) and hedgehogs (Erinaceus coronavirus, EriCoV)4,5. Recently, a virus belonging to the same species as MERS-CoV was isolated in Neoromicia bats (NeoCoV), supporting a bat-origin for MERS-CoV6. This hypothesis received further confirmation by the identification of dipeptidyl-peptidase 4 (DPP4, also known as CD26) as the cellular receptor used by both Ty-BatCoV HKU4 and MERS-CoV7,8,9.
Notably, high titers of MERS-CoV neutralizing antibodies have been detected in camels from various countries and MERS-CoV has been isolated from these animals in Saudi Arabia and Qatar10. Genetic diversity is slightly higher for camel-derived viruses compared to human MERS-CoV isolates, suggesting camel to human transmission rather than vice versa10. Therefore, the most likely scenario envisages that a bat-derived MERS-CoV spread to humans via the zoonotic transmission from dromedary camels.
Coronaviruses use their spike (S) protein to bind a host receptor and to promote membrane fusion. The spike protein assembles as a trimer on the viral surface and belongs to the class I fusion protein family11. Class I fusion proteins are found in many other virus genera including retroviruses, orthomyxoviruses, paramyxoviruses and filoviruses and share similar domain organization, as well as common functional properties12.
Host proteases cleave the CoV spike protein into two functionally distinct domains: the N-terminal region (usually referred to as the S1 subunit) contains the receptor binding domain (RBD), whereas the C-terminal portion (S2 subunit) includes the fusion peptide, two heptad repeats (HR1 and HR2) and the transmembrane (TM) domain12 (see Fig. 1A). Following receptor binding, membrane fusion is mediated by a major conformational rearrangement that exposes the fusion peptide and results in the formation of a six-helix bundle (6HB)13,14. The core of the 6HB is a triple-stranded coiled coil formed by the HR1s of the three spike subunits forming the trimer; the HR2 elements pack within the grooves of the coiled coil in an antiparallel direction13,14.
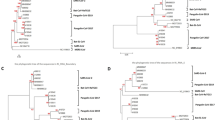
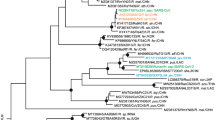
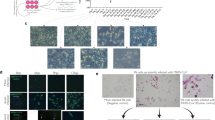
Positive selection at the spike gene of MERS-CoV-related coronaviruses.
(A) Cartoon representation of the MERS-CoV spike protein with distinct domains in different colors (SP, signal peptide; NTD, N-terminal domain; RBD, receptor binding domain; FP, fusion peptide, HR1 and HR2, heptad repeat 1 and 2; TM, transmembrane domain; CP, cytoplasmic domain). The location of positively selected sites detected in MERS-CoV related sequences is shown in red. A positively selected residue in MERS-CoV isolated from humans and camels is in orange. The positively selected site and recombination breakpoints in Pi-BatCoV HKU5 sequences are shown in blue and black, respectively. Furin cleavage sites are depicted as triangles56. Two alignment portions are shown; positions that alter virus host range or tropism in SARS-CoV, MHV and Beaudette strain IBV (IBV Beau CK) are highlighted in green25,26,27. A functional mutation which arose during tissue-culture adaptation of MERS-CoV (strain EMC2012) is highlighted in grey31. (B) Bayesian phylogenies for the S1 (left) and S2 (right) sequences6; branch length is proportional to dS. Branches in red were set as foreground lineages in independent branch-site tests. Thick branches yielded statistically significant evidence of positive selection. The bracket denotes a subset of sequences that were used for analysis of positive selection in the RBD. (C) OmegaMap results for Pi-BatCoV HKU5 spike genes. The hatched red line corresponds to a posterior probability of selection equal to 0.95.
Because of its central role in membrane fusion, a number of antiviral peptides that interfere with the 6HB formation have been developed as potential therapeutic compounds against HIV15, Ebola virus16, SARS-CoV17,18 and MERS-CoV14.
Although the RBD of spike proteins is generally considered the major determinant of host range, several reports have suggested that variation in the C-terminal portion of spike proteins, particularly in the HR1 and HR2, determine host range expansion12. Moreover, recent works indicated that MERS-CoV and Ty-BatCoV HKU4 bind DPP4 both of human and of bat origin7,8,9. In particular, although MERS-CoV binds human DPP4 with higher affinity than Ty-BatCoV HKU4, which shows a preference for the bat receptor, the RBDs of the two viruses engage human DPP4 via a similar binding mode9. These observations suggest that MERS-CoV and related viruses have the potential to shift host range with little adaptation of the RBD.
Motivated by the notion that evolutionary analyses can provide information on the molecular events that underlie host shifts and, more generally, host-pathogen interactions19, we investigated the evolutionary history of S proteins in MERS-CoV and related betaCoVs. Specifically, we aimed to determine whether natural selection drove the evolution of specific regions and sites that may contribute to variation in host range or replication efficiency. Thus, using different strategies, we analyzed MERS-CoV strains isolated from human and camels, as well as MERS-CoV-related viruses from other mammals. Data indicate the HR1 to HR2 region as a major target of adaptive evolution in these viruses.
Results
Positive selection shaped the evolution of clade c betaCoV spike protein
We first investigated whether positive selection drove the evolution of MERS-CoV-related coronavirus spike proteins. Previous phylogenetic analyses of S genes of viruses isolated from humans/camels (MERS-CoV), bats and hedgehogs (Supplementary Table. S1) indicated that the S1 and S2 regions display different tree topologies (Fig. 1B), possibly as a result of recombination6. Because recombination can inflate estimates of positive selection20, we separately analyzed the two regions.
We pruned the S1 and S2 multiple sequence alignments (MSAs) from unreliably aligned codons and we screened them for evidence of additional recombination events using GARD (Genetic Algorithm Recombination Detection)21, which detected no breakpoint.
The saturation of substitution rates represents a major problem in the detection of positive selection among distantly related sequences. Computation of the nonsynonymous (dN) and synonymous (dS) substitution rates over whole phylogenies allows breaking of long branches, resulting in improved rate estimation. Thus, evidence of episodic positive selection was searched for by using the codeml branch-site test22, which is relatively insensitive to the saturation of substitution rates23. In the S1 and S2 regions, 3 and 4 branches yielded statistically significant evidence of positive selection under different codon frequency models (Fig. 1B, Table 1 and Supplementary Table S3). Positively selected sites along these branches were detected using the BEB (Bayes empirical Bayes) procedure and validated using the Mixed Effects Model of Evolution (MEME)24.
One positively selected site was found in the S1 region, 7 sites in the S2 subunit (Fig. 1A, Table 1). The R652 selected site (in S1) almost corresponds to two mutations that independently arose in the SARS-CoV spike gene as a result of in vitro adaptation of zoonotic strains to primate cells25 (Fig. 1A). In S2, most selected sites are located in the HR1, HR2 and in the intervening linker. Remarkably, position 1060 is the almost exact counterpart of aminoacid changes that expand the host range or cell-type tropism of infectious bronchiolitis virus (IBV, L857F) and murine hepatitis virus (MHV, E1035D) (Fig. 1A)26,27.
No positively selected site was found to be located in the RBD (Fig. 1A). Nevertheless, the pruning of unreliably aligned codons operated on the MSA left a minority of RBD sites available for analysis. We thus repeated the branch-site test on a subset of more closely related sequences (Fig. 1B). This procedure decreased divergence and pruning in the RBD and allowed analysis of most codons; even with this procedure, no positively selected site was detected (not shown).
Minor effect of positive selection for the S genes of Ty-BatCoV HKU4 and Pi-BatCoV HKU5
Previous analysis of Ty-BatCoV HKU4 and Pi-BatCoV HKU5 viruses isolated in Hong Kong indicated that positive selection targeted the spike protein and particularly its S1 region28. Nevertheless, recombination was not accounted for in that analysis. We thus analyzed the sequence alignments of the Hong Kong isolates for the presence of recombination breakpoints using GARD. The algorithm detected 9 recombination breakpoints for the Pi-BatCoV HKU5 alignment (Fig. 1A) and none for Ty-BatCoV HKU4. The Ty-BatCoV HKU4 spike gene was therefore analyzed using the codeml site models, which test the hypothesis that a subset of codons evolve with dN/dS > 1. No evidence of positive selection was found, even using the relatively non-conservative model M7/model M8 comparison (Supplementary Table S4). As for the Pi-BatCoV HKU5 spike gene, rampant recombination prevents application of a similar approach. We thus resorted to the simultaneous estimation of selection and recombination using omegaMap29. This analysis confirmed high recombination along the whole gene (Supplementary Fig. S1) and detected a single positively selected site in the S1 region (Fig. 1C). The same site (codon 198, position relative to the MERS-CoV spike sequence) was also detected by MEME, which was run by incorporating the alternative phylogenies detected by GARD. Most of the previously reported selected sites28 were not detected using other methods that account for recombination (Supplementary Table S5). We thus consider that robust inference of positive selection in Pi-BatCoV HKU5 spike genes can only be made for position 198, which lies outside the RBD (Fig. 1A).
Positive selection in the MERS-CoV heptad repeat 1
We next wished to determine whether positive selection occurred at the spike gene of MERS-CoV viruses circulating in the recent outbreak. A previous study suggested that positive selection drove the evolution of two codons in the MERS-CoV spike gene (positions 509 and 1020)30. Nevertheless, in that study only one method was used to infer selection and sequences isolated from camels were not included.
We thus retrieved 54 fully or almost fully spike sequences of MERS-CoV isolated from camels or humans (Supplementary Table S1). Alignments for the S1 and S2 regions were separately analyzed and screened for the presence of recombination. No breakpoint was detected and the codeml site models were applied. For the S2 region, two models of gene evolution that allow a class of codons to evolve with dN/dS > 1 (NSsite models M2a and M8) showed better fit to the data than the null models (NSsite models M1a and M7), strongly supporting the action of positive selection (Table 2, Supplementary Table S6). No evidence of selection was detected for the S1 portion (Table 2, Supplementary Table S6). In S2, both BEB and MEME detected one selected site: position 1020 in the HR1 (Fig. 1A). Interestingly, three different residues are observed at this site in both camel- and human-derived viruses (Fig. 1A). MERS-CoV is thought to have spread from camels to humans; the presence of the three alternative residues in viruses isolated from camels suggests that adaptive evolution at this site occurred prior to the infection of humans.
Interestingly, the 1020 variant is in proximity to a mutation (T1015N) (Figs 1A and 2A) that arose during tissue-culture adaptation of MERS-CoV (strain EMC2012) and increases replication efficiency31.
Analysis of variation in the MERS-CoV HR1.
(A) Ribbon representation of MERS-CoV HR region. Positively selected sites in betaCoVs are shown in red; Q1020 (orange) and M1266 (yellow) are shown as sticks. T1015 is in blue. (B) Detail of inter- and intra-helical interactions for residue 1020. Interactions are shown for Q1020 (left), R1020 (middle) and H1020 (right). Hydrogen bonds are shown as hatched yellow lines; color codes: carbon, white; oxygen, red; nitrogen, blue; sulphur, yellow. Hydrogens have been omitted for clarity. (C) Stability analysis for HR1 positions 1020 (left) and 1015 (right). ΔΔG in kcal/mol for mutations of Q1020 and T1015 to all other 19 aminoacids are reported. Aminoacid residues observed in MERS-CoV sequences are circled. Results are shown for FoldX, PopMuSiC and I-Mutant.
Analysis of MERS-CoV HR1 variation
The structure of the 6HB of MERS-CoV has been solved13,14; through its side chain, the Q1020 residue forms hydrogen bonds with D1024 and interacts with M1266 in HR2 (Fig. 2A,B). Replacement of the glutamine residue with histidine or arginine (observed in the camel- and human-derived viruses) results in loss of side chain interactions with M1266 and variably affects hydrogen bonds with D1024 (Fig. 2B).
To gain further insight into the effect of adaptive evolution at position 1020, we performed a stability computational analysis after in silico mutagenesis. Q1020 was replaced with all other possible aminoacids: even if changes of different magnitude in ΔG were obtained using three different methods32,33,34, trends were very consistent (Fig. 2C). In particular, replacement with histidine or arginine residues resulted in mild destabilization (Fig. 2C). As a comparison, the same analysis was performed for position 1015. Replacement of the threonine residue with asparagine, which was previously associated with increased replication efficiency in vitro31, resulted in a similar level of destabilization as observed for Q1020H/R (Fig. 2C).
Discussion
We analyzed the evolution of the S protein in betaCoVs, in bat Ty-BatCoV HKU4 and Pi-BatCoV HKU5 viral populations, as well as in MERS-CoV isolates from the current outbreak. Different strategies were applied, as appropriate depending on divergence and recombination.
Results indicated that several adaptive changes are located in the S2 region, with fewer in the S1 domain and none of these within the RBD. It should be noted, though, that in all analyses we applied quite conservative approaches and we intersected two different methods to declare a site as positively selected. Whereas this approach was meant to limit the false positive rate it may have yielded some false negative results. In particular, the branch-site test we used to analyze the S1 and S2 regions of betaCoVs is robust to saturation issues and has a minimal false positive rate, but lacks power23. Moreover, due to the high divergence and the consequent need of alignment pruning, analysis of the RBD was performed on a shallower phylogeny compared to the other regions. This procedure is expected to reduce power, but is nonetheless necessary. In fact, alignment errors, together with unrecognized recombination, inflate estimates of positive selection and represent major sources of false positive results in evolutionary analyses20,35. Consistently, when we accounted for recombination in Pi-BatCoV HKU5 sequences most previously described selection signals disappeared, including those in the RBD28. Thus, whereas we cannot exclude that adaptive variants in the RBD of betaCoVs were missed by our approach, we conclude that the more recent evolution of MERS-CoV, Ty-BatCoV HKU4 and Pi-BatCoV HKU5 was not driven by positive selection in this domain. Conversely, as previously shown for SARS-CoV25, our data support a role for the S1 region separating the RBD and fusion peptide as a determinant of betaCoV host range expansion.
Analysis of both betaCoVs and MERS-CoV strains revealed evidence of positive selection in the S2 region. Most positively selected sites were found to be located either in the heptad repeats or in the intervening linker. Among these sites, position 1060 is particularly interesting, as it almost corresponds to substitutions that modify the host range and/or cell tropism in MHV and IBV26,27. Both viruses belong to the coronavirus genus. The Beaudette strain of IBV has been adapted to embryonated chicken eggs; following passages in culture, the strain was further adapted to infect Vero cells (from African Green monkey) and primary chicken kidney cells26. The L857F mutation was shown to represent a major determinant of the fusogenic activity in these cell types26. As far as MHV is concerned, the E1035D mutation was recovered after passages in mouse liver and was shown to contribute significantly to the hepatotropism and hepatic virulence of a previously attenuated strain (MHV-A59)27. Additional variants in the HR1 and fusion peptide of MHV strains cooperate with changes in the S1 region, resulting in a broadening of receptor usage (to heparan sulfate) and, consequently, an extension of the host range36. Finally, in the MHV-A51 strain the ability to bind human CAECAM receptors is strongly influenced by mutant residues located in the fusion peptide, HR1 and HR1/HR2 linker37. Similar observations have been reported for viruses that do not belong to the coronavirus genus, but that use a class I fusion protein. For instance, one single mutation in the HR1 of a simian-human immunodeficiency virus (SHIV) strain (KB9) increases by two- to three-fold infection efficiency in cells expressing the marmoset cellular receptors38, whereas a nearby HR1 change in SIV contributes to macrophage tropism39. Overall, these findings pinpoint the relevance of changes in the HR1 and HR2 as modifiers of host range and cell-type tropism.
The molecular mechanisms underlying the altered phenotype of HR1 and HR2 mutants remain to be determined in all these instances, although changes in conformational structure have been suggested as a possible explanation. Unfortunately, no coronavirus S protein fusion intermediate or pre-fusion state has been solved to date, hampering investigation of molecular interactions. We therefore analyzed the effect of variation at the 1020 position of MERS-CoV on the stability of the 6HB in the post-fusion conformation14. The presence of a Q1020 had previously been suggested to confer higher stability to the MERS-CoV 6HB compared to SARS-CoV14. Indeed, replacement of this residue results in loss of intra- and inter-helical interactions. In line with these observations, three different methods used for stability analysis were concordant in showing that the alternative arginine and histidine residues at position 1020 result in a moderate and similar level of destabilization. Although the observed ΔΔG is relatively small, it was calculated on the single monomer and is expected to be multiplied in the trimer. The observation whereby mildly destabilizing variants are favored by selection may seem counterintuitive. Nonetheless, we show that a similar level of destabilization is observed for a mutation in MERS-Cov HR1 (T1015N) that increases infection efficiency, at least in vitro31. Mutagenesis of HR1 in retroviral type I fusion proteins has indicated that, whereas strong destabilization of the 6HB (as measured by circular dichroism) almost inevitably results in reduced infectivity, a minor stability decrease is not necessarily associated with defects in cell fusion and infection efficiency40,41,42. For instance, different aminoacid replacements at the same HR1 position in HIV-1 gp41 result in decreased stability, but unaffected or even increased infectivity40. In the case of EIAV (equine infectious anemia virus), destabilized HR1 mutants were found to display different infection phenotypes depending on temperature42. This observation may be extremely interesting in the context of MERS-CoV, as both bats and dromedary camels display adaptive heterothermy (i.e. sensible daily or season variation in body temperature).
Coronavirus spike proteins are highly exposed on the virus surface and represent major targets for antibody response43, raising the possibility that adaptive evolution of the S protein is driven by the host immune system. This hypothesis is difficult to address due to the paucity of information concerning the specific MERS-CoV epitopes recognized by human antibodies. Recently, different studies identified human antibodies againts MERS-CoV from non-immune human antibody libraries: all of them were directed against the RBD, suggesting that this region represents a major target for the host immune system43. Nevertheless, data on the humoral immune response to MERS-CoV in infected subjects are presently lacking, whereas such information is richer for SARS-CoV. Indeed, analysis of antibodies derived from a patient who recovered from SARS indicated that some of them recognize epitopes in the HR2 region44. Their neutralizing effect was ascribed to interference of the interaction between HR2 and HR1. Whether antibodies against the S2 region also arise in human subjects (or other mammalian hosts) infected with MERS-CoV and related betaCoVs remains to be determined; if this were the case, some of the selected sites we identified may be under selective pressure to evade recognition.
Finally, it is worth noting that HRs have been studied in different viruses because synthetic peptides interfering with 6HB formation are promising antiviral molecules15,16,17,18. This is also the case for MERS-CoV and HR2-like peptides were recently shown to be effective in vitro14. These peptides were tested against a MERS-CoV strain carrying Q1020 and all include the interacting M1266 residue14. These antivirals may display decreased activity depending on the MERS-CoV strain and its aminoacid status at the selected 1020 position.
Materials and Methods
Sequences and alignments
Virus sequences were retrieved from the NCBI database and a list of accession numbers is provided as Supplementary Table S1. Sequences of Ty-BatCoV HKU4 and Pi-BatCoV HKU5 isolated in Hong Kong were derived from a previous work28.
Errors in the inferred multiple sequence alignment (MSA), which may be common when highly divergent sequences are analyzed, can inflate estimates of positive selection. We therefore used PRANK45 for building the MSA and GUIDANCE46 for filtering unreliably aligned codons (i.e. we masked codons with a score <0.90), as suggested35.
Detection of recombination and positive selection
To detect positive selection at the S gene of clade c betaCoVs we applied the branch-site test from the PAML suite22. The test compares a model (MA) that allows positive selection on one or more lineages (foreground lineages) with a model (MA1) that does not allow such positive selection. Twice the difference of likelihood for the two models (ΔlnL) is then compared to a χ2 distribution with one degree of freedom22. Specifically, the internal branches of previously reconstructed6 Bayesian phylogenies of the S1 and S2 regions were set as the foreground lineages in independent tests. A false discovery rate (FDR) correction was applied to account for multiple hypothesis testing (i.e. we corrected for the number of tested branches), as suggested47.
Positively selected sites were identified through the BEB analysis (with a p value cutoff of 0.90), which calculates the posterior probability that each site belongs to the site class of positive selection on the foreground branch(es). Sites were validated using MEME (with the default cutoff of 0.1), which allows the distribution of dN/dS (also referred to as ω) to vary from site to site and from branch to branch at a site, therefore allowing the detection of episodic positive selection24.
The site models implemented in PAML were applied -independently- for the analysis of HKU4 and MERS-CoV sequences, which display very limited divergence and do not suffer from saturation problems. To detect selection, site models that allow (M2a, M8) or disallow (M1a, M7) a class of sites to evolve with ω > 1 were fitted to the data48. Trees were generated by maximum-likelihood using the PhyML program49 with a GTR model of nucleotide substitution and γ distributed rates. Positively selected sites were identified using the BEB analysis (from model M8)50. Again, sites were validated using MEME.
To assure consistency, all models were run using the F3 × 4 and the F61 codon frequency models.
MSAs were screened for the presence of recombination using GARD. Recombination breakpoints were considered significant if the HK (Kishino-Hasegawa) p value was <0.01.
Simultaneous inference of selection and recombination for analysis of positive selection was performed using omegaMap29, a program for detecting natural selection and recombination based on a model of population genetics and molecular evolution. The model uses a population genetic approximation to the coalescent with recombination. This latter is estimated from patterns of linkage disequilibrium assuming that recombination events occur only between codons and not within them. OmegaMap applies reversible-jump Markov Chain Monte Carlo (MCMC) to perform Bayesian inferences of both ω and the recombination parameter ρ, allowing both parameters to vary along the sequence. An average block length of 10 and 30 codons was used to estimate ω and ρ, respectively. To determine the influence of the choice of the priors on the posteriors, analyses were repeated with alternative sets of priors (Supplementary Table S2). Three independent omegaMap runs, each with 500,000 iterations and a 50,000 burn-in iteration, were compared to assess convergence and merged to obtain the posterior probability estimate.
The REL (random effects likelihood) analysis models variation in both dN and dS across sites according to a predefined distribution with different rate classes; positively selected sites are identified through an empirical Bayes method51. The default criterion of a Bayes Factor >50 was used to identify positively selected sites.
For GARD, MEME and REL the nucleotide substitution models were chosen using a Genetic Algorithm implemented in the dataMonkey suite52. All analyses were performed through the DataMonkey server53 (http://www.datamonkey.org).
In silico analysis of HR1 variants
The crystal structure of MERS-CoV HR1 and HR2 region was obtained from PDB (PDB ID: 4MOD).
Histidine or arginine residues were introduced at positions 1020 and suitable rotamers were sampled through the rapid torsion scan utility in Maestro (Maestro. 9.1; Schrodinger). Intraprotein interactions were calculated with PIC (protein interaction calculator)54.
Because programs that calculate stability changes achieve only moderate accuracy55, we used three different methods to assure reliability. These approaches are based on different principles. Specifically, PoPMuSiC uses statistical potentials and takes into account amino acid volume variation upon mutation33; FoldX uses an empirical force field and evaluates the energetic effect of point mutations and the interactions contributing to the stability of proteins32. Finally, I-Mutant 2.0 is based on a neural network approach to evaluate the free energy change after a single point mutation with incorporation of information on the three-dimensional structure of the protein34.
In FoldX and I-Mutant the ΔΔG values are calculated as follows: ΔΔG = ΔGmutant − ΔGwild-type. In FoldX and I-Mutant ΔΔG values > 0 kcal/mol indicate mutations that decrease protein stability, whereas in PoPMuSiC ΔΔG values > 0 kcal/mol are mark of mutations increasing protein stability. Therefore, PoPMuSiC ΔΔG values were multiplied by −1 to obtain homogeneous results.
In the analysis carried out with FoldX 3D, the three-dimensional structure of the protein was repaired using the <RepairPDB> command. Mutations were introduced using the <BuildModel> command with <numberOfRuns> set to 5 and <VdWdesign> set to 0. Temperature (298K), ionic strength (0.05 M) and pH (7) were set to default values and the force-field was used to predict the water molecules on the protein surface.
Additional Information
How to cite this article: Forni, D. et al. The heptad repeat region is a major selection target in MERS-CoV and related coronaviruses. Sci. Rep. 5, 14480; doi: 10.1038/srep14480 (2015).
References
Zaki, A. M., van Boheemen, S., Bestebroer, T. M., Osterhaus, A. D. & Fouchier, R. A. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N. Engl. J. Med. 367, 1814–1820, 10.1056/NEJMoa1211721 (2012).
de Groot, R. J. et al. Middle East respiratory syndrome coronavirus (MERS-CoV): announcement of the Coronavirus Study Group. J. Virol. 87, 7790–7792, 10.1128/JVI.01244-13 (2013).
van Boheemen, S. et al. Genomic characterization of a newly discovered coronavirus associated with acute respiratory distress syndrome in humans. MBio 3, 10.1128/mBio.00473-12 (2012).
Corman, V. M. et al. Characterization of a novel betacoronavirus related to middle East respiratory syndrome coronavirus in European hedgehogs. J. Virol. 88, 717–724, 10.1128/JVI.01600-13 (2014).
Tang, X. C. et al. Prevalence and genetic diversity of coronaviruses in bats from China. J. Virol. 80, 7481–7490, doi: 10.1128/JVI.00697-06 (2006).
Corman, V. M. et al. Rooting the phylogenetic tree of middle East respiratory syndrome coronavirus by characterization of a conspecific virus from an African bat. J. Virol. 88, 11297–11303, 10.1128/JVI.01498-14 (2014).
Lu, G. et al. Molecular basis of binding between novel human coronavirus MERS-CoV and its receptor CD26. Nature 500, 227–231, 10.1038/nature12328 (2013).
Yang, Y. et al. Receptor usage and cell entry of bat coronavirus HKU4 provide insight into bat-to-human transmission of MERS coronavirus. Proc. Natl. Acad. Sci. USA. 111, 12516–12521, 10.1073/pnas.1405889111 (2014).
Wang, Q. et al. Bat origins of MERS-CoV supported by bat coronavirus HKU4 usage of human receptor CD26. Cell. Host Microbe 16, 328–337, 10.1016/j.chom.2014.08.009 (2014).
Al-Tawfiq, J. A. & Memish, Z. A. Middle East respiratory syndrome coronavirus: transmission and phylogenetic evolution. Trends Microbiol. 22, 573–579, 10.1016/j.tim.2014.08.001 (2014).
Jiang, S., Lu, L., Du, L. & Debnath, A. K. A predicted receptor-binding and critical neutralizing domain in S protein of the novel human coronavirus HCoV-EMC. J. Infect. 66, 464–466, 10.1016/j.jinf.2012.12.003 (2013).
Graham, R. L. & Baric, R. S. Recombination, reservoirs and the modular spike: mechanisms of coronavirus cross-species transmission. J. Virol. 84, 3134–3146, 10.1128/JVI.01394-09 (2010).
Gao, J. et al. Structure of the fusion core and inhibition of fusion by a heptad repeat peptide derived from the S protein of Middle East respiratory syndrome coronavirus. J. Virol. 87, 13134–13140, 10.1128/JVI.02433-13 (2013).
Lu, L. et al. Structure-based discovery of Middle East respiratory syndrome coronavirus fusion inhibitor. Nat. Commun. 5, 3067, 10.1038/ncomms4067 (2014).
Jiang, S., Lin, K., Strick, N. & Neurath, A. R. HIV-1 inhibition by a peptide. Nature 365, 113, 10.1038/365113a0 (1993).
Watanabe, S. et al. Functional importance of the coiled-coil of the Ebola virus glycoprotein. J. Virol. 74, 10194–10201 (2000).
Liu, S. et al. Interaction between heptad repeat 1 and 2 regions in spike protein of SARS-associated coronavirus: implications for virus fusogenic mechanism and identification of fusion inhibitors. Lancet 363, 938–947, 10.1016/S0140-6736(04)15788-7 (2004).
Bosch, B. J. et al. Severe acute respiratory syndrome coronavirus (SARS-CoV) infection inhibition using spike protein heptad repeat-derived peptides. Proc. Natl. Acad. Sci. USA. 101, 8455–8460, 10.1073/pnas.0400576101 (2004).
Longdon, B., Brockhurst, M. A., Russell, C. A., Welch, J. J. & Jiggins, F. M. The evolution and genetics of virus host shifts. PLoS Pathog. 10, e1004395, 10.1371/journal.ppat.1004395 (2014).
Anisimova, M., Nielsen, R. & Yang, Z. Effect of recombination on the accuracy of the likelihood method for detecting positive selection at amino acid sites. Genetics 164, 1229–1236 (2003).
Kosakovsky Pond, S. L., Posada, D., Gravenor, M. B., Woelk, C. H. & Frost, S. D. Automated phylogenetic detection of recombination using a genetic algorithm. Mol. Biol. Evol. 23, 1891–1901, 10.1093/molbev/msl051 (2006).
Zhang, J., Nielsen, R. & Yang, Z. Evaluation of an improved branch-site likelihood method for detecting positive selection at the molecular level. Mol. Biol. Evol. 22, 2472–2479, 10.1093/molbev/msi237 (2005).
Gharib, W. H. & Robinson-Rechavi, M. The branch-site test of positive selection is surprisingly robust but lacks power under synonymous substitution saturation and variation in GC. Mol. Biol. Evol. 30, 1675–1686, 10.1093/molbev/mst062 (2013).
Murrell, B. et al. Detecting individual sites subject to episodic diversifying selection. PLoS Genet. 8, e1002764, 10.1371/journal.pgen.1002764 (2012).
Rockx, B. et al. Synthetic reconstruction of zoonotic and early human severe acute respiratory syndrome coronavirus isolates that produce fatal disease in aged mice. J. Virol. 81, 7410–7423, 10.1128/JVI.00505-07 (2007).
Yamada, Y., Liu, X. B., Fang, S. G., Tay, F. P. & Liu, D. X. Acquisition of cell-cell fusion activity by amino acid substitutions in spike protein determines the infectivity of a coronavirus in cultured cells. PLoS One 4, e6130, 10.1371/journal.pone.0006130 (2009).
Navas-Martin, S., Hingley, S. T. & Weiss, S. R. Murine coronavirus evolution in vivo: functional compensation of a detrimental amino acid substitution in the receptor binding domain of the spike glycoprotein. J. Virol. 79, 7629–7640, 10.1128/JVI.79.12.7629-7640.2005 (2005).
Lau, S. K. et al. Genetic characterization of Betacoronavirus lineage C viruses in bats reveals marked sequence divergence in the spike protein of pipistrellus bat coronavirus HKU5 in Japanese pipistrelle: implications for the origin of the novel Middle East respiratory syndrome coronavirus. J. Virol. 87, 8638–8650, 10.1128/JVI.01055-13 (2013).
Wilson, D. J. & McVean, G. Estimating diversifying selection and functional constraint in the presence of recombination. Genetics 172, 1411–1425, 10.1534/genetics.105.044917 (2006).
Cotten, M. et al. Spread, circulation and evolution of the Middle East respiratory syndrome coronavirus. MBio 5, 10.1128/mBio.01062-13 (2014).
Scobey, T. et al. Reverse genetics with a full-length infectious cDNA of the Middle East respiratory syndrome coronavirus. Proc. Natl. Acad. Sci. USA. 110, 16157–16162, 10.1073/pnas.1311542110 (2013).
Schymkowitz, J. et al. The FoldX web server: an online force field. Nucleic Acids Res. 33, W382–8, 10.1093/nar/gki387 (2005).
Dehouck, Y., Kwasigroch, J. M., Gilis, D. & Rooman, M. PoPMuSiC 2.1: a web server for the estimation of protein stability changes upon mutation and sequence optimality. BMC Bioinformatics 12, 151-2105-12-151, 10.1186/1471-2105-12-151 (2011).
Capriotti, E., Fariselli, P. & Casadio, R. I-Mutant2.0: predicting stability changes upon mutation from the protein sequence or structure. Nucleic Acids Res. 33, W306–10, 10.1093/nar/gki375 (2005).
Privman, E., Penn, O. & Pupko, T. Improving the performance of positive selection inference by filtering unreliable alignment regions. Mol. Biol. Evol. 29, 1–5, 10.1093/molbev/msr177 (2012).
de Haan, C. A. et al. Cooperative involvement of the S1 and S2 subunits of the murine coronavirus spike protein in receptor binding and extended host range. J. Virol. 80, 10909–10918, 10.1128/JVI.00950-06 (2006).
McRoy, W. C. & Baric, R. S. Amino acid substitutions in the S2 subunit of mouse hepatitis virus variant V51 encode determinants of host range expansion. J. Virol. 82, 1414–1424, 10.1128/JVI.01674-07 (2008).
Pacheco, B., Basmaciogullari, S., Labonte, J. A., Xiang, S. H. & Sodroski, J. Adaptation of the human immunodeficiency virus type 1 envelope glycoproteins to new world monkey receptors. J. Virol. 82, 346–357, 10.1128/JVI.01299-07 (2008).
Mori, K., Rosenzweig, M. & Desrosiers, R. C. Mechanisms for adaptation of simian immunodeficiency virus to replication in alveolar macrophages. J. Virol. 74, 10852–10859 (2000).
Eggink, D. et al. Detailed mechanistic insights into HIV-1 sensitivity to three generations of fusion inhibitors. J. Biol. Chem. 284, 26941–26950, 10.1074/jbc.M109.004416 (2009).
Suntoke, T. R. & Chan, D. C. The fusion activity of HIV-1 gp41 depends on interhelical interactions. J. Biol. Chem. 280, 19852–19857, 10.1074/jbc.M502196200 (2005).
Du, J. et al. Structural and biochemical insights into the V/I505T mutation found in the EIAV gp45 vaccine strain. Retrovirology 11, 26-4690-11-26, 10.1186/1742-4690-11-26 (2014).
Ying, T., Li, H., Lu, L., Dimitrov, D. S. & Jiang, S. Development of human neutralizing monoclonal antibodies for prevention and therapy of MERS-CoV infections. Microbes Infect. 17, 142–148, 10.1016/j.micinf.2014.11.008 (2015).
Rockx, B. et al. Structural basis for potent cross-neutralizing human monoclonal antibody protection against lethal human and zoonotic severe acute respiratory syndrome coronavirus challenge. J. Virol. 82, 3220–3235, 10.1128/JVI.02377-07 (2008).
Loytynoja, A. & Goldman, N. An algorithm for progressive multiple alignment of sequences with insertions. Proc. Natl. Acad. Sci. USA. 102, 10557–10562, 10.1073/pnas.0409137102 (2005).
Penn, O. et al. GUIDANCE: a web server for assessing alignment confidence scores. Nucleic Acids Res. 38, W23–8, 10.1093/nar/gkq443 (2010).
Anisimova, M. & Yang, Z. Multiple hypothesis testing to detect lineages under positive selection that affects only a few sites. Mol. Biol. Evol. 24, 1219–1228, 10.1093/molbev/msm042 (2007).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591, 10.1093/molbev/msm088 (2007).
Guindon, S., Delsuc, F., Dufayard, J. F. & Gascuel, O. Estimating maximum likelihood phylogenies with PhyML. Methods Mol. Biol. 537, 113–137, 10.1007/978-1-59745-251-9_6 (2009).
Anisimova, M., Bielawski, J. P. & Yang, Z. Accuracy and power of bayes prediction of amino acid sites under positive selection. Mol. Biol. Evol. 19, 950–958 (2002).
Kosakovsky Pond, S. L. & Frost, S. D. Not so different after all: a comparison of methods for detecting amino acid sites under selection. Mol. Biol. Evol. 22, 1208–1222, 10.1093/molbev/msi105 (2005).
Delport, W. et al. CodonTest: modeling amino acid substitution preferences in coding sequences. PLoS Comput. Biol. 6, 10.1371/journal.pcbi.1000885 (2010).
Delport, W., Poon, A. F., Frost, S. D. & Kosakovsky Pond, S. L. Datamonkey 2010: a suite of phylogenetic analysis tools for evolutionary biology. Bioinformatics 26, 2455–2457, 10.1093/bioinformatics/btq429 (2010).
Tina, K. G., Bhadra, R. & Srinivasan, N. PIC: Protein Interactions Calculator. Nucleic Acids Res. 35, W473–6, 10.1093/nar/gkm423 (2007).
Khan, S. & Vihinen, M. Performance of protein stability predictors. Hum. Mutat. 31, 675–684, 10.1002/humu.21242 (2010).
Millet, J. K. & Whittaker, G. R. Host cell entry of Middle East respiratory syndrome coronavirus after two-step, furin-mediated activation of the spike protein. Proc. Natl. Acad. Sci. USA. 111, 15214–15219, 10.1073/pnas.1407087111 (2010).
Acknowledgements
NAD is supported by the Deanship of Scientific Research, Prolific Research Group Program (PRG-1436-15), Vice Rectorate for Graduate Studies and Scientific Research in King Saud University (KSU), Riyadh, Saudi Arabia.
Author information
Authors and Affiliations
Contributions
M.S. and D.F. conceived the study; M.S. and M.C. supervised the project; D.F., R.C., U.P. and N.A.D. performed the evolutionary analysis; L.D.G. and G.F. performed the in silico studies; D.F. and G.F. produced the figures, with input from all authors; U.P. provided support during the bioinformatic analyses; M.S. wrote the manuscript, with critical input from M.C. and from the remaining authors.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Forni, D., Filippi, G., Cagliani, R. et al. The heptad repeat region is a major selection target in MERS-CoV and related coronaviruses. Sci Rep 5, 14480 (2015). https://doi.org/10.1038/srep14480
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep14480
This article is cited by
-
MERS-CoV: epidemiology, molecular dynamics, therapeutics, and future challenges
Annals of Clinical Microbiology and Antimicrobials (2021)
-
Genomic epidemiology of SARS-CoV-2 in Esteio, Rio Grande do Sul, Brazil
BMC Genomics (2021)
-
Molecular aspects of MERS-CoV
Frontiers of Medicine (2017)
-
Evolutionary Dynamics of MERS-CoV: Potential Recombination, Positive Selection and Transmission
Scientific Reports (2016)
-
Polyphyletic origin of MERS coronaviruses and isolation of a novel clade A strain from dromedary camels in the United Arab Emirates
Emerging Microbes & Infections (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.