Abstract
Bacillus thuringiensis has been globally used as a microbial pesticide for over 70 years. However, information regarding its various adaptions and virulence factors and their roles in the entomopathogenic process remains limited. In this work, we present the complete genomes of two industrially patented Bacillus thuringiensis strains (HD-1 and YBT-1520). A comparative genomic analysis showed a larger and more complicated genome constitution that included novel insecticidal toxicity-related genes (ITRGs). All of the putative ITRGs were summarized according to the steps of infection. A comparative genomic analysis showed that highly toxic strains contained significantly more ITRGs, thereby providing additional strategies for infection, immune evasion and cadaver utilization. Furthermore, a comparative transcriptomic analysis suggested that a high expression of these ITRGs was a key factor in efficient entomopathogenicity. We identified an active extra urease synthesis system in the highly toxic strains that may aid B. thuringiensis survival in insects (similar to previous results with well-known pathogens). Taken together, these results explain the efficient entomopathogenicity of B. thuringiensis. It provides novel insights into the strategies used by B. thuringiensis to resist and overcome host immune defenses and helps identify novel toxicity factors.
Similar content being viewed by others
Introduction
Bacillus thuringiensis, a member of the Bacillus cereus sensu lato group, is a natural insecticidal bacterium. The bacterium produces parasporal crystals when it sporulates in limited nutrient conditions. These crystals consist of crystal proteins (Cry proteins, also named delta-endotoxins) and exhibit insecticidal activity against a range of invertebrates, including members of the orders Lepidoptera, Diptera, Coleoptera, some nematodes, mites and protozoa1,2. For over 70 years, B. thuringiensis products, which contain a mixture of spores and insecticidal crystals, have been the most important biopesticides in fields such as agriculture and health. Compared with chemical pesticides, they have the advantages of specificity and high efficiency and are environmentally safe3.
However, the thousands of B. thuringiensis isolated strains show considerable variety in their insecticidal toxicity and spectra. Only highly toxic strains are used for biopesticide production. B. thuringiensis serovar kurstaki HD-1 is the original strain used in the microbial insecticide Dipel. Since its isolation in 1970, the products from strain HD-1 and similar strains (Btk) have been used as effective biopesticides. Strain HD-1 has also been designated as the primary U.S. reference standard strain for toxicity evaluation in all commercial B. thuringiensis formulations. This strain has enjoyed the greatest commercial success for microbial control worldwide4.
To understand the mechanism of B. thuringiensis insecticidal toxicity, early research primarily focused on the pore-forming mechanism of the parasporal crystal proteins (Cry and Cyt proteins) in insect gut epithelial cells5. However, controversy remained regarding whether B. thuringiensis was a toxin-producing soil bacterium (such as Streptomyces species) or an insect pathogen6,7. Subsequently, studies demonstrated that in addition to its role as an efficient insecticidal microorganism, B. thuringiensis could also produce virulent, pathogenic and adaption factors during the infection process8. These results suggested that B. thuringiensis possessed a complicated pathogenic mechanism. Due to the rapid development of high-throughput sequencing technology, many B. thuringiensis genomes have been completed in recent years9,10,11,12,13,14,15,16. However, with the exception of two draft genomes of Btk17,18, no complete genomes of the biopesticide production strains (especially Btk HD-1 and similar strains) are currently available. Furthermore, there is a lack of systematic explanations on the effective insecticidal toxicity phenotype of these B. thuringiensis strains at the genomic, transcriptomic, or omic levels.
In this study, we completed the genome sequence of B. thuringiensis insecticide standard strain HD-1 and the related strain YBT-1520, which is also widely used for Bt biopesticide production (product trademark “Mianfeng”) in China in an attempt to obtain a comprehensive understanding of B. thuringiensis pathogenesis. Additionally, we performed a comparative study with other B. cereus group genomes. A comparative genomic analysis revealed that the Btk strains possess a larger number and more types of insecticidal toxicity-related genes (ITRGs) compared with other B. cereus group bacteria with characteristic chromosomal features. Interestingly, the comparative transcriptomic analysis (a microarray of strain YBT-1520) indicated that the ITRGs in Btk are not only all actively transcribed but are also transcribed at a substantially higher level than general B. thuringiensis. This work explains the efficient entomopathogenicity of B. thuringiensis, especially regarding the increased transcription of ITRGs in highly toxic strains.
Results
General genome feature: larger genome size and more plasmids
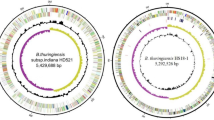
The chromosome of strain HD-1 and strain YBT-1520 are circular molecules that are 5,631,672 bp and 5,602,565 bp in length, respectively, with the same average G + C content (35.3%) (Table 1). There were no significant differences between the chromosomes of strain HD-1 and strain YBT-1520; therefore, the genes discussed below should have common names. Strain HD-1 contained 13 plasmids with a combined size of 1,135 kb, whereas strain YBT-1520 contained 11 plasmids with a combined size of approximately 978 kb (Supplementary Table S1). Most of the plasmids had significantly lower G + C contents than the chromosome, with the exception of pBMB46 and pBMBLin15 (35.4% and 40.1%) in strain HD-1, which are a putative circular prophage and a linear prophage, respectively. The chromosome of strain HD-1 contained 5,864 genes, 41 rRNA genes and 94 tRNA, whereas strain YBT-1520 contained 5,830 genes, 39 rRNA genes and 98 tRNA (Table 1). The plasmids of strain HD-1 and strain YBT-1520 contained 1,064 and 890 genes, respectively.
Because the genome data available for the B. cereus sensu lato group in NCBI include 50 B. thuringiensis, 161 B. cereus and 93 B. anthracis strains annotated according to previously reported insecticidal activities and serotypes, we divided the 50 B. thuringiensis strains into three groups: highly, generally and weakly toxic strains (12, 29 and 9 strains, respectively, Table 1 and Supplementary Table S2). The comparison showed that the B. thuringiensis strains with high insecticidal toxicity10,12,19 had significantly larger genome sizes and were richer in plasmid content than the weakly toxic B. thuringiensis strains and typical B. cereus, B. anthracis strains which are not toxic to insects (P < 0.05, T-test, Fig. 1A,B).
Comparison of the whole genome size and plasmid content of the B. cereus group strains.
The 304 B. cereus group strains were downloaded from the NCBI genome database. The B. thuringiensis strains listed in Table 1 and Supplementary Table S2 were sorted on the basis of their insecticidal toxicity level from high to low.
A chromosomal inversion with novel insecticidal toxicity-related genes (ITRGs)
The genomes of the B. cereus group strains possessed high sequence homology, protein identity and conserved synteny, with the exception of a laboratory B. anthracis mutant strain CDC684 that had a 3.3 Mb chromosomal inversion20. Compared with the other B. cereus group genomes, B. thuringiensis serovar kurstaki strains (including HD-1, YBT-1520, HD739 and T03a00121) all had a massive and rare inversion with a 740 kb fragment on the chromosome relative to the replication origin.
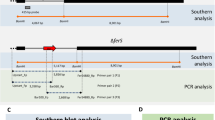
The inversion in the HD-1 chromosome is located between the base-pair coordinates 2,644 kb and 3,380 kb; in the YBT-1520 chromosome, the inversion is located between 2,613 kb and 3,351 kb. In contrast to B. anthracis strain CDC684, the genome rearrangement events did not occur between homologous lambda-like prophage regions20 but instead occurred between two reversed IS232A elements (Fig. 2A,B). Interestingly, we found that most of the genes that flank the two IS232A regions are unique genes in the Btk strains, such as a novel HBL gene operon (hblIII2) and a putative pectate lyase family gene (pel2). Notably, Btk strain HD73, which is listed in the “general” toxicity group, has the same chromosomal inversion and these novel virulence related genes. It is a special case in Btk because that ,as previous work reported9, the “general” toxic strain HD73 only contains one parasporal crystal protein gene cry1Ac and protein Cry1Ac is the primary toxin for the insecticidal phenotype. Additionally, a comparative analysis of the completed B. cereus group genomes showed a large number of functional genes in the chromosomal inversion region in the Btk strains, including amino acid transporters, various resistant proteins and regulators, which were not highly conserved in the other strains (Fig. 2A).
Chromosome alignments of B. cereus group members.
(A) The diagram represents the BLASTn results of a circular chromosome comparison using the BRIG program59. Each genome is color-coded as indicated by the legend. Relative shading density (from darker to lighter) within each circle represents relative levels of nucleotide homology. White regions indicate regions with no identity to the reference. (B) Chromosome inversion region comparison. Genes are represented by colored arrows. Well-conserved segments of the chromosomes are paired using shaded regions; dark gray indicates 62 to 100% nucleotide identity. The non-shaded regions lack homology between the B. cereus group strains. The outer scale is given in kilobases.
Highly toxic B. thuringiensis strains encode more ITRGs
B. thuringiensis encodes a large number of pathogenic factors in its genome and is an insecticidal bacterium that has been used as a biopesticide for many years. Using the reported virulence-associated genes8,22,23 and our unpublished data, a whole-genome InterproScan analysis of strain HD-1 and strain YBT-1520 were used to identify and divide the insecticidal toxicity-related genes and gene clusters (ITRGs) into three groups: insecticidal and other virulence synergic genes, pathogenic and virulence assistant genes and genes for saprophytic colonization; these genes including 33 families and approximately 100 ITRGs (Supplementary Table S3). Significantly, the results indicated that Btk HD-1 and YBT-1520 have 20.0%–87.5% more genes and a 22.2% higher variety of ITRGs than generally, weakly toxic B. thuringiensis and other B. cereus group bacteria (an operon or a gene cluster counted as one gene, Fig. 3, Supplementary Table S3).
The parasporal crystal protein is the primary insecticidal toxin of B. thuringiensis8. Strain HD-1 encodes six crystal protein genes, including cry1Aa, cry1Ab, cry1Ac, cry1Ia, cry2Aa, and cry2Ab. Four of these proteins (cry1Aa, cry1Ia, cry2Aa, and cry2Ab) are located on the large plasmid pBMB299 with the vegetative insecticidal protein gene vip3Aa where they form a pathogenicity island (PAI); two additional proteins are located on plasmids pBMB95 (cry1Ac) and pBMB65 (cry1Ab). Strain YBT-1520 has nearly identical cry genes with strain HD-1 with the exception of cry1Ab.
Zwittermycin A (ZmA) is a very important virulence synergic factor that has a broad spectrum of antimicrobial activity and the ability to enhance the insecticidal activity of the Cry protein24,25. Interestingly, we found a highly conserved bacteriocin PAI in both large plasmids (pBMB431/pBMB422) of strain HD-1 and strain YBT-1520. This PAI includes a ZmA biosynthetic gene cluster26,27 and two bacteriocin biosynthetic gene clusters (Thurin and Thuricin). And the PAI is flanked by insertion sequences (IS elements). Thurin is a putative novel bacteriocin of unknown function; the gene cluster includes a regulator gene (thrR), a precursor gene (thrA), a modifier gene (thrM) and a transporter gene (thrT). The Thuricin gene cluster encodes ten biosynthetic-related genes (Supplementary Table S3) and shows 99% identity with Thuricin H and 1728,29. These two bacteriocins were divided into saprophytic and colonization groups based on their potential antibacterial activity.
The genes encoding hemolysin and non-hemolysin enterotoxin represent the major important factor involved in the pathogenic and virulence assistant genes group. Although they are conserved among the B. cereus group bacteria, B. thuringiensis was proven to be safe for use in biological pest control30. A comparative genomic analysis revealed that strain HD-1 and strain YBT-1520 possessed a novel copy of the hemolysin BL (HBL) gene operon, which is located in the insecticidal PAI described above (hblIII1, Supplementary Table S3). Notably, these two Btk also encode five novel proteins belonging to the hemolysin XhlA family, which is a key factor for the full virulence of the insect pathogen Xenorhabdus nematophila31 (Supplementary Table S3). Additionally, two novel pore-forming toxin genes are located in the large plasmids (pft and nep1, Supplementary Table S3). The Pft protein showed 33% identity with 213 amino acids (aa) of the parasporin-2 from B. thuringiensis32 and 38% identity with 222 aa of the hydralysin from Hydra vulgaris33. This finding suggested that these two genes may play a very important role in pathogenicity. B. thuringiensis also encodes a large number of pathogenic assistant factors in the genome, such as phospholipase C (plc), sphingomyelinase C (spl) and immune inhibitor A (inhA)23,34 (Supplementary Table S3).
The comparative genomic analysis indicated that in addition to the conserved factors among the B. cereus group, highly toxic B. thuringiensis strains encode more genes related to degradation, adhesion and recognition. These genes could aid in the effective utilization of insect tissues during the saprophytic colonization stage, especially the enhancins (bel), collagenase (colA), pectate lyase (pel), cell wall hydrolases (cwl), extracellular metalloprotease (mpr) and cell enveloped S-layer anchor proteases (P < 0.05, T-test, Fig. 3, Supplementary Table S3). These enzymes could potentially introduce competitive advantages to the highly toxicity strains, thereby enabling them to avoid the host immune system and degrade and utilize insect tissues.
The extra nitrogen metabolism system could make highly toxic B. thuringiensis strains better adapted for survival in the host
Most of the B. cereus group bacteria only possess the nitrate-nitrite reduction gene cluster for nitrogen source utilization. Previous works have demonstrated that some B. cereus group strains lack the nitrate-nitrite reduction gene cluster but use a nine-gene urease synthesis cluster instead in a manner similar to that of B. cereus ATCC1098735. This study utilized the NCBI database and determined that, in addition to the conserved genes involved in the nitrate-nitrite reduction, all of the Btk genomes encoded an extra system for nitrogen source utilization; this system was composed of a nine-gene urease synthesis cluster (ureA, ureB, ureC, ureE, ureF, ureG, ureD, ureI, and ureT, BTK_19245-BTK_19285/YBT1520_19325-YBT1520_19365, Fig. 4). The amino acids had up to 94% identity compared with a specific B. cereus strain ATCC10987 isolated from cheese spoilage that only contains the urease synthesis gene cluster but not the nitrate-nitrite reduction system36. The urease synthesis cluster in the Btk genomes also had homology with some pathogens, including Helicobacter pylori (Fig. 4), that use urease for acid acclimation during gastric colonization and acid exposure37. There were no insert sequences or transposons flanking the gene cluster, indicating that this extra nitrogen source metabolism system is not a result of horizontal gene transfer but is a specific feature of Btk.
Comparative transcriptomic analysis
To further understand the B. thuringiensis-insect interaction, a microarray strategy was used to identify the transcription modulation of the ITRGs during the different growth phases of Btk YBT-1520 under culture. Of the 6,668 genes in strain YBT-1520 examined by microarray analysis, 4,998, 4,471, 4,650 and 3,484 genes (75.0%, 67.1%, 69.7% and 52.2%) were transcribed in the four growth phases, respectively. It showed that the transcriptional activity decreased with time, but still a half of the genes in YBT-1520 transcribed in the late stationary growth phase. Additionally, B. thuringiensis serovar chinensis strain CT-43 was used for comparative transcriptomic analysis in this work. This strain has a similar Cry protein composition to that of Btk (especially the same insecticidal PAI). It is also a similar strain to the serovar thuringiensis, which was used in early products with less potency than Btk. Using the gatB (YBT1520_22885) and rpoA (YBT1520_00670) genes as reference genes38,39, the transcriptomic data from the strain YBT-1520 microarray and RNA sequencing from strain CT-4340 were normalized for comparison.
The transcription of ITRGs in B. thuringiensis YBT-1520: genes in the same family are transcribed during different phases
A total of 37.4% of the ITRGs reached the highest transcriptional level in the middle logarithmic growth phase (ML); these ITRGs included cry1Ia, 12 of 21 hemolysin family genes, the phospholipase family genes plc and spl, alveolysin (alo), bel1, four of seven neutral proteinase genes, a subtilase gene, three metallo-protease genes, four of six collagenase genes and 12 of 20 S-layer anchor protease genes (Supplementary Fig. S1A, S1B and S1C). The hemolysin family genes were nearly always transcribed at a high level during the log growth phase34; interestingly, the novel hblIII1 operon in the insecticidal PAI had a higher transcription level than the other hemolysin family genes (P < 0.05, T-test, Supplementary Fig. S1B). Additionally, the immune inhibitor A gene inhA1 was transcribed at a very high level that was close to the transcribed level of the cry genes during the forespore formatting phase (P < 0.05, T-test, Supplementary Fig. S1B). Bacterial growth changed from logarithmic growth to stationary growth (early stationary growth phase, ES); a total of 31.3% of the ITRGs reached their highest transcriptional levels during this phase. Notably, the bacteriocin gene clusters (genes encoding Vip3Aa, camelysin, four neutral proteinases, chitinase, four different S-layer anchor proteins and the pathogenic-related QS regulators PlcR and NprR) showed extra transcriptional activity during this phase (Supplementary Fig. S1A, S1B and S1C). This indicated that during the logarithmic growth phase, B. thuringiensis could express various factors (especially the hemolysin BL compomers) to attack the host while also expressing many cofactors (primarily immune inhibitor A and collagenases) to escape host immunity and degrade tissue barriers, as well as antimicrobial peptides to inhibit other competitive bacteria and fungi.
During the middle and late stationary growth phases (MS and LS), the main transcriptional signals of the ITRGs were the Cry protein genes (cry1Aa, cry1Ac, cry2Aa, and cry2Ab, P < 0.05, T-test). Interestingly, some pathogenic factors were also expressed, such as hemolysin III1, XhlA3, XhlA4, four subtilases, pectase, metalloproteases and some S-layer anchor proteases (Supplementary Fig. S1B and S1C). The high levels of expression of the larger number of ITRGs in Btk suggested that the bacteria have additional strategies for infection, immune evasion and cadaver utilization.
We also found that ITRGs in the same protein families were not transcribed during the same phases. The transcription of three immune inhibitor A genes (inhA) serves as an example: inhA1 was up-regulated during the ML phase (5.2-fold compared with the LS phase), but inhA2 and inhA3 were up-regulated during the ES phase (2.8-fold compared with the MS and 1.9-fold compared with the LS phases, respectively, Supplementary Fig. S1B and Table S4). Similarly, two enhancin genes (bel) were also up-regulated during different phases: bel1 was up-regulated during the ML phase, whereas bel2 was up-regulated during the ES phase (6.3- and 3.2-fold, respectively, compared with the LS phase, Supplementary Fig. S1C and Table S4). Most of the saprophytic colonization-related genes were also up-regulated more than three- during the log growth phase (ML and ES), especially the genes from the degradation enzyme families (Supplementary Fig. S1C). The 20 cell envelope S-layer anchor genes could be divided into four families. The three internalin protein genes were regulated at different phases: ilsa1 was increased in the ML phase (6.1-fold compared with the LS phase), ilsa2 was increased in the MS phases (52.0-fold compared with the ES phase) and ilsa3 was increased in the ES phase (1.8-fold compared with the ML phase, Supplementary Fig. S1C and Table S4). The transcriptional pattern of the eight peptidoglycan hydrolase-related genes also differed. Six of these genes were increased in the ML phase (12.5-, 11.3-, 12.0-, 3.4-, 297.4- and 6.1-fold compared with the ES phase), whereas the remaining two were up-regulated in the LS phase (3.4- and 1.9-fold compared with the ML phase). The other S-layer anchor genes were primarily up-regulated during the ML and ES phases. The different transcriptional phases of the genes in the same protein family suggested that they play a similar role during different infection steps.
In addition, the microarray data analysis showed that all 26 novel unique genes around the 740 kb chromosome inversion ends were actively transcribed, including hblIII2, pel2, xhlA3, a regulator gene pagR, a reverse transcriptase and a hypothetical protein gene (Supplementary Table S4). And remarkably, the main urease synthesis-related genes were not silenced as previously reported in B. cereus35 but were actively transcribed during the ES phase even though the transporter genes were up-regulated during the LS phases (Fig. 5, Supplementary Table S4).
The ITRGs are transcribed more actively and for longer periods of time in the highly toxic strains
A comparative transcriptomic analysis showed that 8.2% of ITRGs were not transcribed during any phases; 47.6% were not transcribed during the spore formation phase (LS) in strain YBT-1520. In strain CT-43, these values were 19.0% and 79.8%, respectively (Supplementary Fig. S2). The primary transcribed genes in strain CT-43 included cry1B, cry1Aa and cry2Aa; strain YBT-1520 also expressed cry1Ia, cry2Ab, and vip3Aa in addition to the highly transcribed cry1Aa, cry1Ac, and cry2Aa genes, which may provide diversity and a wider spectrum of insect toxicity (Supplementary Table S4). The cry and vip3Aa genes in strain YBT-1520 showed significantly more activity than the genes in strain CT-43 during the growth phases; the relative expressions of the former were more than tenfold those of the latter (P < 0.05, T-test, Fig. 6). Specifically, the ratio of the cry2Ab gene in strain YBT-1520 was increased by more than six-fold at all times (LS, P < 0.05, T-test, Fig. 6). Interestingly, one prevalent explanation for the high toxicity and broad host range spectrum phenotype of B. thuringiensis is that there are striking differences in the expression of the conserved cry genes between various strains.
The relative gene expression ratio of conserved ITRGs between B. thuringiensis strain YBT-1520 and strain CT-43.
The heat map represents the Log2 (fold change) of the expression level of the three groups of conserved ITRGs and the global virulence regulator genes between strain YBT-1520 and strain CT-43. ML, ES, MS and LS are defined in Fig. 5.
Many ITRGs in the pathogenic- and virulence-assisted gene group were conserved in the B. cereus group and were highly transcribed in strain YBT-1520 but sparsely transcribed in strain CT-43. These ITRGs included most of the hemolysin family genes, inhA and virulence factor alveolysin (Fig. 6, Supplementary Table S4). By contrast, nearly all of the protease genes in the saprophytic colonization group in strain YBT-1520 (especially the S-layer anchor-related genes) showed a more than twofold increase in transcription during the three phases compared with strain CT-43 (P < 0.05, T-test, Fig. 6). Specifically, the important pathogenic protease gene camelysin, which can significantly enhance the hemolytic activity of Cyt proteins41, was primarily transcribed during the ES phase (Supplementary Table S4) in strain YBT-1520 and the corresponding relative expression (YBT-1520 vs CT-43) in the ES phase was increased two-fold (P < 0.05, T-test, Fig. 6).
Moreover, the ITRGs in the saprophytic colonization group were transcribed for a substantially longer period than those in strain CT-43 during the growth phases. A comparative transcriptomic analysis showed that nearly all of the conserved saprophytic colonization-related protease genes (46 genes in total) in strain YBT-1520 were transcribed during all four growth phases (Supplementary Fig. S1C and S2). However, according to the CT-43 RNA-seq data, 25 of these were only transcribed in one or two growth phases, including the 12 genes that did not exhibit ratios in any of the four growth phases. These genes included a variety of protease families, such as chitinase, neutral protease, subtilisin, collagenase and several S-layer anchor proteases. This result indicated that highly toxic B. thuringiensis strains could express more proteases for a longer period compared with general B. thuringiensis, thereby enabling more efficient larval body utilization during the infection process of high toxicity strains.
Comparative transcriptomic analysis of global virulence regulator genes
To explain the substantially higher transcriptional activity and longer continuous transcription of ITRGs, we compared the transcription of the conserved global virulence regulator genes (primarily the quorum-sensing system (QS) regulators, which regulate several microbial processes, i.e., sporulation, virulence, biofilm formation, conjugation and the production of extracellular enzymes34,42). The QS systems were substantially more active in strain YBT-1520 than strain CT-43. During the vegetative growth phase (ML to MS), the relative expression of the QS regulator genes plcR and nprR were more than sevenfold higher (P<0.05, T-test, Fig. 6). This result is consistent with the higher transcriptional activity detected for many ITRGs (such as the hemolysin genes, which were demonstrated to be controlled by these regulators34,42).
codY of strain YBT-1520, an important global transcriptional regulator that primarily represses the sporulation phase genes, maintained a steady transcription level during the log and stationary growth phases (15.9-, 12.1- and 20.5-fold compared with the LS phase, Supplementary Table S4) and suddenly decreased during the LS growth phase. However, codY had an increasing trend during the log and stationary growth phases in strain CT-43 (2.0-, 5.3- and 18.0-fold compared with the LS phase, P < 0.05, T-test). These results implied that Btk has a different regulatory mechanism for development from other B. thuringiensis strains.
A global virulence regulator mga-like gene (inpR, YBT1520_32436) was found upstream of the vip3Aa and cry genes in the insecticidal PAI. The protein InpR has two Mga helix-turn-helix DNA-binding domains. A BLASTP analysis showed that the 471 aa InpR protein had 25% identity with 213 aa of Mga, which is one of the key global regulators of virulence in Group A Streptococcus43. InpR also showed 20% identity with 469 aa of the anthrax toxin expression trans-acting positive regulator AtxA from B. anthracis plasmid pXO1 and 26% identity with 151 aa of the capsule synthesis trans-acting positive regulator AcpA from pXO2. A phylogenetic analysis indicated that InpR is closely related to the regulators AtxA and AcpA (Supplementary Fig. S3). Moreover, the analysis suggested that the genes in the insecticidal PAI may have their own transcriptional controls. Further analysis showed that the inpR in strain YBT-1520 was rarely transcribed during the growth phases; however, in CT-43, inpR was transcribed during all four phases. Furthermore, the transcriptional trend was on the rise prior to the release of the spores and parasporal crystals from the mother cell (15.6-, 28.5- and 55.2-fold compared with the LS phase, P < 0.05, T-test). Interestingly, as mentioned above, the relative expression of the important toxins downstream of the inpR gene in the PAI, such as cry1Ia and vip3Aa, were all up-regulated. These results provided an important clue for further study on the function of InpR.
Functional confirmation of microarray data with qRT-PCR
Quantitative real-time RT-PCR (qRT-PCR) was performed as a complementary approach to verify the results of the comparative transcriptomic analysis. The qRT-PCR assessments were conducted on RNA samples obtained from the four growth phases of strains YBT-1520, HD-1, CT-43 and YBT-020 under the same culture conditions using primers for plcR, nprR, codY, inpR, and inhA1. Strain HD-1 exhibited an up-regulated trend that was similar to that of strain YBT-1520. The relative expression of plcR from strain YBT-1520 to strain CT-43 was found to be 4.5-fold in the ML phase and the up-regulation was 370.7-fold compared with strain YBT-020 (P < 0.05, T-test, Fig. 7A,B). The relative expression of nprR was up-regulated 2.3- and 4.7-fold during the ES and MS phases, respectively (strain YBT-1520/CT-43, P < 0.05, T-test). The relative expression of codY was up-regulated 10.0- and 5.2-fold in the ES and MS phases, respectively (strain YBT-1520/CT-43); compared with strain YBT-020, codY was up-regulated 5.6- and 13.5-fold in the ML and ES phases, respectively (P < 0.05, T-test). The relative expression of inpR was down-regulated 0.1- and 0.4-fold in the ML and LS phases, respectively and remained at 1.8-fold during the MS phase (strain YBT-1520/CT-43, P < 0.05, T-test). The relative expression of the conserved inhA1 gene was increased 1.5-, 4.3- and 1.7-fold during the ML to MS phases (strain YBT-1520/CT-43); compared with YBT-020, inhA1 was up-regulated 10.5- and 4.0-fold during the ES and MS phases, respectively (P < 0.05, T-test). These qRT-PCR results were consistent with the comparative transcriptomic results described above. They also confirmed that the transcriptional levels of the global regulator and conserved pathogenic genes in B. thuringiensis strains were significantly different under culture conditions.
The qRT-PCR analysis of the virulence-related regulators and inhA gene in B. thuringiensis strains YBT-1520, CT-43 and YBT-020. ML, ES, MS and LS were defined in Fig. 5.
(A) The relative gene expression between strain YBT-1520 and strain CT-43. (B) The relative gene expression between strain YBT-1520 and strain YBT-020.
Discussion
B. thuringiensis usually shows substantial variations in insecticidal toxicity. Thus, more efficient insecticidal strains are used for biopesticide production and in studies of host-pathogen interactions. Remarkably, these strains always contain a large number of plasmids, which makes it difficult to complete genome sequencing using a single high-throughput sequencing strategy. In this study, a multiple sequencing strategy was chosen: after obtaining data via high-throughput technologies, Optical Mapping44 was used for chromosome assembly to correct and further assemble the scaffold. Because the various plasmids have no appropriate reference sequence, a genomic BAC library was constructed to determine the relationship between the scaffold and the short contigs. Additionally, a manual analysis of the read data also assisted in closing the gaps. The results confirmed that the “multiple” strategy was effective and practical for complete genome sequencing of complicated bacteria.
Interestingly, comparative genomic analyses indicated that highly insecticidal strains have larger genome sizes and more complicated genome constitutions. This finding is significantly different from that for other pathogens in which the pathogenic strains usually have a smaller genome size than the nonpathogenic strains45,46. Our results implied that the genome size of B. thuringiensis increases as genome complexity increases due to mechanisms involving plasmid content and chromosomal inversion. And we also found a lot of novel ITRGs and other unique genes in these plasmids and the inversion region. This provides the ability of B. thuringiensis to adapt to various insect and nematode hosts.
The infection process of B. thuringiensis generally involves Cry proteins. These proteins destroy the epithelial cells in the midgut and the spores germinate and proliferate using the degraded products from the insect cadaver7,8. Another example includes Btk spores (which are not toxic to nematodes alone), which can cause accelerated killing when fed to C. elegans with the pore-forming protein Cry5B47. These results suggested that B. thuringiensis is an insect pathogen but not a toxin-producing soil bacterium. In this study, the insecticidal toxicity-related genes (ITRGs) were summarized and categorized into three groups corresponding to the steps of infection and necrotrophism to investigate the pathogenic mechanisms during infection and the necrotrophic process. Based on the results of the comparative genomic analysis, a hypothesis was proposed to explain the efficient pathogenicity phenotype: 1) The large number of ITRGs could help B. thuringiensis effectively kill and utilize nutrient substances from the host; 2) A complicated genome that includes chromosomal inversion rearrangement and a larger number of mobile elements (plasmids, insertion sequences and introns) not only harbor novel ITRGs but also may provide the evolutionary potential to adapt to a broad spectrum of host environments (insect and nematode) in a manner similar to that of other pathogens; and 3) An active extra nitrogen utilization system may help the bacteria survive and adapt in the host.
Based on this hypothesis, another equally important issue was whether the ITRGs were transcribed or silenced. The microarray analysis showed that the ITRGs were all transcribed to various degrees during the four growth phases. A total of 47.6% and 25.2% of the ITRGs exhibited maximal transcriptional values during the ML and ES growth phases, respectively. In the middle and late stationary growth phases (MS and LS), the main transcriptional signals were the cry genes and several pathogenic factors. This finding illustrated the most effective period for the expression of different virulence and pathogenicity-related genes (especially protease genes in the same family), which could be used to improve the model of the B. thuringiensis infection process.
Furthermore, this work revealed that ITRGs and conserved global virulence regulator genes in the highly toxic strains were more transcriptionally active than those in the general strain, especially the hemolysins, immune inhibitor A, chitinase and QS regulators PlcR and NprR. Hence, a fourth explanation should be added to our hypothesis: high expressions of the ITRGs, including ITRGs unique to highly toxic strains and conserved ITRGs among the B. cereus group, accelerate the infection process and prevent possible host resistance. Moreover, because the global regulators do not control all of the ITRGs, the high expression of these ITRGs may be under the control of an unknown regulation mechanism. Thus, the different expression levels of ITRGs in these strains may represent an unknown strategy used by B. thuringiensis to adapt to various hosts.
Materials and Methods
Bacterial strains and culture conditions
B. thuringiensis serovar kurstaki strain NRRL HD-1 (serotypes H3a, 3b and 3c) was obtained from the U.S. Department of Agriculture’s Agricultural Research Service (ARS) in Peoria, IL. The bacterium was compared with strain 4D1 (BGSC code for HD-1) from the Bacillus Genetic Stock Center (BGSC) in Columbus, USA and the similar Btk strain YBT-1520 was isolated from soil by our laboratory in China. This strain has been used to produce the Mianfeng biopesticide in China48 because it showed higher toxicity to the larvae of lepidopteran pests (Helicoverpa armigera and Plutella xylostella) compared with the technical standardization strain HD-1 or other B. thuringiensis biopesticide strains. All of the Escherichia coli and B. thuringiensis strains were maintained on Luria-Bertani (LB) agar plates (1% tryptone, 0.5% yeast extract, 0.5% NaCl and 1.5% agar) supplemented with appropriate antibiotics at 37 and 28 °C, respectively.
Whole genome sequencing and assembly
Total genomic DNA of B. thuringiensis strains HD-1 and YBT-1520 was extracted from the cells according to the procedure of Andrup et al.49. Due to the complexity of the B. thuringiensis genome sequence (especially the abundant and varied plasmids contents), a multiple-step strategy was used for HD-1 whole genome sequencing: 1) After performing a series of strict filtering steps to remove artificial duplication, adapter contamination and low-quality reads, we obtained 65,899,485 Paired-End (PE) clean reads from different length libraries using the Ilumina Genome Analyzer System (GAIIx) at BGI-Shenzhen, China, representing 1300-fold coverage; 2) a total of 83,060 PE clean reads (14-fold coverage) were obtained using the Roche 454 Genome Sequencer FLX platform (454 GS-FLX) at the Roy J. Carver Biotechnology Center, University of Illinois at Urbana-Champaign, Urbana, IL, USA. A total of 46 scaffolds (>500 bp) of strain HD-1 were obtained using the assembly software SOAP50: the N50 length was 1,861,364 bp, the N90 length was 461,121 bp, the maximum length was 2,497,856 bp and the minimum length was 242,258 bp. The genome alignment generated using the software MAUVE51 showed that 15 of the reads overlapped the entire chromosome, whereas the rest may belong to natural plasmids or prophage fragments. Following the analysis of the 454 GS-FLX reads and PCR verifications, we obtained five large scaffolds that overlapped the chromosome of strain HD-1. All reads were mapped onto the contigs for scaffold building by utilizing the paired-end information. After Optical Mapping (OM)52 of the HD-1 chromosome was performed at BGI-Shenzhen, the number of scaffolds was reduced to three and some errors were redressed. The HD-1 chromosome was finished by closing the remaining gaps with PCR. At this point, 21 scaffolds belonged to strain HD-1 plasmids, including some shorter than 500 bp. Thus, we used a whole genome BAC library of strain HD-1 with an approximately 50-kb insert DNA length for complete plasmid assembly. The construction process of the BAC library followed our previous works on B. thuringiensis strains YBT-1765 and CT-4353,54.
For strain YBT-1520, we obtained 8,403,791 clean reads (200-fold coverage) via GAIIx and 86,428 clean reads (14-fold coverage) via 454 GS-FLX. Using strain HD-1 as a reference, the initial assembly produced 1 scaffold that overlapped the chromosome and 15 scaffolds that belonged to plasmids. Following PCR verification, the entire genome of strain YBT-1520 was completed. We obtained a draft genome of strain YBT-1520 using a Sanger Shotgun sequencing strategy in a previous study; this enabled us to complete most of the plasmids55,56,57. The complete genome of strain YBT-1520 in this work was essentially in agreement with the previous draft genome.
Genome annotation and comparative genomics analysis
The genome annotation was performed by the NCBI Prokaryotic Genomes Automatic Annotation Pipeline Group (http://www.ncbi.nlm.nih.gov/genomes/static/Pipeline.html). The comparative analysis of the chromosomes was performed using BRIG and EasyFig58,59. A maximum likelihood phylogenomic tree was created using the concatenated amino acid sequences with the program MEGA 5.160.
Transcriptomic analysis
Harvested bacterial cells were ground in liquid nitrogen with TRIzol and transferred to tubes (109 cells/ml TRIzol) for shaking. RNA isolation and purification were performed according to the TRIzol method61. For microarray slide preparation, 6,668 probes were designed according to the whole genomic strain YBT-1520 ORFs and 2,057 ORFs of the B. cereus group plasmids in the NCBI database; the microarray was synthesized using the Combimatrix CustomArray platform. cDNA synthesis, labeling and purification were performed by Advanced Throughput Inc, USA. Four time points within 30 hours of growth were chosen to focus on the differential expression of the pathogenic factors: the 6th, 9th, 18th and 27th hours represent the middle logarithmic growth phase (ML), early stationary growth phase (ES), middle stationary growth phase with forespore formatting (MS) and late stationary growth phase with some spores and parasporal crystals release (LS), respectively (Supplementary Fig. S4). The R platform, Cluster 3.0 and Tree View 1.1.6 were used for filtering, normalization and cluster analysis. P-values were computed using the false discovery rate correction of 0.05.
Quantitative real-time PCR
Prior to reverse transcription (RT), total RNA was subjected to PCR using primers specific for 16S rDNA and 23S rDNA to exclude the possibility of genomic DNA contamination. RNA (40 ng) was converted into cDNA using the QuantiTec reverse transcription kit (Qiagen) as recommended by the manufacturer. Quantitative real-time RT-PCRs (qRT-PCRs) were performed using the QuantiFast SYBR green PCR kit (Qiagen) according to the manufacturer’s protocol and were run in a LightCycler instrument (ABI). The entire experiment was repeated two times on RNA samples extracted from the four growth phases of the B. thuringiensis strains. Changes in mRNA levels were calculated using the 2−ΔΔCT method62.
Additional Information
Accession numbers: The genome sequences of strain HD-1 and strain YBT-1520 have been submitted to the GenBank of NCBI (http://www.ncbi.nlm.nih.gov/) under accession nos CP004870-CP004883 and CP004858-CP004869, respectively. Microarray data for strain YBT-1520 are available in the ArrayExpress database (www.ebi.ac.uk/arrayexpress) under accession number E-MTAB-2162.
How to cite this article: Zhu, L. et al. Genomic and transcriptomic insights into the efficient entomopathogenicity of Bacillus thuringiensis. Sci. Rep. 5, 14129; doi: 10.1038/srep14129 (2015).
References
van Frankenhuyzen, K. Insecticidal activity of Bacillus thuringiensis crystal proteins. J Invertebr Pathol 101, 1–16, 10.1016/j.jip.2009.02.009 (2009).
Schnepf, E. et al. Bacillus thuringiensis and its pesticidal crystal proteins. Microbiol Mol Biol Rev 62, 775–806 (1998).
Roh, J. Y., Choi, J. Y., Li, M. S., Jin, B. R. & Je, Y. H. Bacillus thuringiensis as a specific, safe and effective tool for insect pest control. J Microbiol Biotechnol 17, 547–559 (2007).
Dulmage, H. T. B. Thuringiensis U. S. Assay Standard: Report on the Adoption of a Primary U.S. Reference Standard for Assay of Formulations Containing the delta-Endotoxin of Bacillus thuringiensis. Bull. Entomol. Soc. Am. 19, 200–202 (1973).
Soberon, M., Gill, S. S. & Bravo, A. Signaling versus punching hole: How do Bacillus thuringiensis toxins kill insect midgut cells? Cell Mol Life Sci 66, 1337–1349, doi: 10.1007/s00018-008-8330-9 (2009).
Broderick, N. A. et al. Contributions of gut bacteria to Bacillus thuringiensis-induced mortality vary across a range of Lepidoptera. BMC Biol 7, 11, doi: 10.1186/1741-7007-7-11 (2009).
Raymond, B., Johnston, P. R., Nielsen-LeRoux, C., Lereclus, D. & Crickmore, N. Bacillus thuringiensis: an impotent pathogen? Trends Microbiol 18, 189–194, 10.1016/j.tim.2010.02.006 (2010).
Nielsen-Leroux, C., Gaudriault, S., Ramarao, N., Lereclus, D. & Givaudan, A. How the insect pathogen bacteria Bacillus thuringiensis and Xenorhabdus/Photorhabdus occupy their hosts. Curr Opin Microbiol 15, 220–231, 10.1016/j.mib.2012.04.006 (2012).
Liu, G. et al. Complete genome sequence of Bacillus thuringiensis subsp. kurstaki strain HD73. Genome announcements 1, e0008013, 10.1128/genomeA.00080-13 (2013).
Murawska, E., Fiedoruk, K., Bideshi, D. K. & Swiecicka, I. Complete genome sequence of Bacillus thuringiensis subsp. thuringiensis strain IS5056, an isolate highly toxic to Trichoplusia ni. Genome announcements 1, e0010813, 10.1128/genomeA.00108-13 (2013).
Sheppard, A. E., Poehlein, A., Rosenstiel, P., Liesegang, H. & Schulenburg, H. Complete Genome Sequence of Bacillus thuringiensis Strain 407 Cry. Genome announcements 1, e00158–00112, 10.1128/genomeA.00158-12 (2013).
Guan, P. et al. Complete genome sequence of Bacillus thuringiensis serovar Sichuansis strain MC28. J Bacteriol 194, 6975, 10.1128/JB.01861-12 (2012).
He, J. et al. Complete genome sequence of Bacillus thuringiensis mutant strain BMB171. J Bacteriol 192, 4074–4075, 10.1128/JB.00562-10 (2010).
He, J. et al. Complete genome sequence of Bacillus thuringiensis subsp. chinensis strain CT-43. J Bacteriol 193, 3407–3408, 10.1128/JB.05085-11 (2011).
Zhu, Y. et al. Complete genome sequence of Bacillus thuringiensis serovar finitimus strain YBT-020. J Bacteriol 193, 2379–2380, 10.1128/JB.00267-11 (2011).
Han, C. S. et al. Pathogenomic sequence analysis of Bacillus cereus and Bacillus thuringiensis isolates closely related to Bacillus anthracis. J Bacteriol 188, 3382–3390, 10.1128/JB.188.9.3382-3390.2006 (2006).
Day, M., Ibrahim, M., Dyer, D. & Bulla, L., Jr. Genome Sequence of Bacillus thuringiensis subsp. kurstaki Strain HD-1. Genome announcements 2, e00613–00614, 10.1128/genomeA.00613-14 (2014).
Zwick, M. E. et al. Genomic characterization of the Bacillus cereus sensu lato species: backdrop to the evolution of Bacillus anthracis. Genome Res 22, 1512–1524, 10.1101/gr.134437.111 (2012).
Wang, P. et al. Complete genome sequence of Bacillus thuringiensis YBT-1518, a typical strain with high toxicity to nematodes. J Biotechnol 171, 1–2, 10.1016/j.jbiotec.2013.11.023 (2014).
Okinaka, R. T. et al. An attenuated strain of Bacillus anthracis (CDC 684) has a large chromosomal inversion and altered growth kinetics. BMC Genomics 12, 477, doi: 10.1186/1471-2164-12-477 (2011).
Zwick, M. E. et al. Genomic characterization of the Bacillus cereus sensu lato species: backdrop to the evolution of Bacillus anthracis. Genome Res 22, 1512–1524, 10.1101/gr.134437.111 (2012).
Luo, X. et al. Bacillus thuringiensis metalloproteinase Bmp1 functions as a nematicidal virulence factor. Appl Environ Microbiol 79, 460–468, 10.1128/AEM.02551-12 (2013).
Fang, S. et al. Bacillus thuringiensis bel protein enhances the toxicity of Cry1Ac protein to Helicoverpa armigera larvae by degrading insect intestinal mucin. Appl Environ Microbiol 75, 5237–5243, 10.1128/AEM.00532-09 (2009).
Broderick, N. A., Goodman, R. M., Raffa, K. F. & Handelsman, J. Synergy Between Zwittermicin A and Bacillus thuringiensis subsp. kurstaki Against Gypsy Moth (Lepidoptera: Lymantriidae). Environmental Entomology 29, 101–107, doi: 10.1603/0046-225X-29.1.101 (2000).
Silo-Suh, L. A., Stabb, E. V., Raffel, S. J. & Handelsman, J. Target range of zwittermicin A, an aminopolyol antibiotic from Bacillus cereus. Current microbiology 37, 6–11 (1998).
Zhao, C., Song, C., Luo, Y., Yu, Z. & Sun, M. L-2,3-diaminopropionate: one of the building blocks for the biosynthesis of Zwittermicin A in Bacillus thuringiensis subsp. kurstaki strain YBT-1520. FEBS Lett 582, 3125–3131, 10.1016/j.febslet.2008.07.054 (2008).
Luo, Y. et al. Validation of the intact zwittermicin A biosynthetic gene cluster and discovery of a complementary resistance mechanism in Bacillus thuringiensis. Antimicrob Agents Chemother 55, 4161–4169, 10.1128/AAC.00111-11 (2011).
Lee, H., Churey, J. J. & Worobo, R. W. Biosynthesis and transcriptional analysis of thurincin H, a tandem repeated bacteriocin genetic locus, produced by Bacillus thuringiensis SF361. FEMS Microbiol Lett 299, 205–213, doi: 10.1111/j.1574-6968.2009.01749.x (2009).
Lee, K. D. et al. The class IId bacteriocin thuricin-17 increases plant growth. Planta 229, 747–755, doi: 10.1007/s00425-008-0870-6 (2009).
Raddadi, N. et al. Characterization of polyvalent and safe Bacillus thuringiensis strains with potential use for biocontrol. J Basic Microbiol 49, 293–303, 10.1002/jobm.200800182 (2009).
Cowles, K. N. & Goodrich-Blair, H. Expression and activity of a Xenorhabdus nematophila haemolysin required for full virulence towards Manduca sexta insects. Cell Microbiol 7, 209–219, 10.1111/j.1462-5822.2004.00448.x (2005).
Akiba, T. et al. Crystal structure of the parasporin-2 Bacillus thuringiensis toxin that recognizes cancer cells. J Mol Biol 386, 121–133, 10.1016/j.jmb.2008.12.002 (2009).
Zhang, M., Fishman, Y., Sher, D. & Zlotkin, E. Hydralysin, a novel animal group-selective paralytic and cytolytic protein from a noncnidocystic origin in hydra. Biochemistry 42, 8939–8944, 10.1021/bi0343929 (2003).
Gohar, M. et al. The PlcR virulence regulon of Bacillus cereus. PLoS One 3, e2793, 10.1371/journal.pone.0002793 (2008).
Mols, M. & Abee, T. Role of ureolytic activity in Bacillus cereus nitrogen metabolism and acid survival. Appl Environ Microbiol 74, 2370–2378, 10.1128/AEM.02737-07 (2008).
Rasko, D. A. et al. The genome sequence of Bacillus cereus ATCC 10987 reveals metabolic adaptations and a large plasmid related to Bacillus anthracis pXO1. Nucleic Acids Res 32, 977–988, 10.1093/nar/gkh258 (2004).
Mobley, H. L. The role of Helicobacter pylori urease in the pathogenesis of gastritis and peptic ulceration. Aliment Pharmacol Ther 10 Suppl 1, 57–64 (1996).
Salvetti, S., Faegri, K., Ghelardi, E., Kolsto, A. B. & Senesi, S. Global gene expression profile for swarming Bacillus cereus bacteria. Appl Environ Microbiol 77, 5149–5156, 10.1128/AEM.00245-11 (2011).
Reiter, L., Kolsto, A. B. & Piehler, A. P. Reference genes for quantitative, reverse-transcription PCR in Bacillus cereus group strains throughout the bacterial life cycle. J Microbiol Methods 86, 210–217, 10.1016/j.mimet.2011.05.006 (2011).
Wang, J. et al. The metabolic regulation of sporulation and parasporal crystal formation in Bacillus thuringiensis revealed by transcriptomics and proteomics. Molecular & cellular proteomics: MCP 12, 1263–1276, 10.1074/mcp.M112.023986 (2013).
Nisnevitch, M., Sigawi, S., Cahan, R. & Nitzan, Y. Isolation, characterization and biological role of camelysin from Bacillus thuringiensis subsp. israelensis. Curr Microbiol 61, 176–183, doi: 10.1007/s00284-010-9593-6 (2010).
Dubois, T. et al. Necrotrophism Is a Quorum-Sensing-Regulated Lifestyle in Bacillus thuringiensis. PLoS Pathog 8, e1002629, 10.1371/journal.ppat.1002629 (2012).
Hondorp, E. R. et al. Characterization of the Group A Streptococcus Mga virulence regulator reveals a role for the C-terminal region in oligomerization and transcriptional activation. Mol Microbiol 83, 953–967 (2012).
Riley, M. C., Lee, J. E., Lesho, E. & Kirkup, B. C., Jr. Optically mapping multiple bacterial genomes simultaneously in a single run. PLoS One 6, e27085, 10.1371/journal.pone.0027085 (2011).
Ochman, H. & Davalos, L. M. The nature and dynamics of bacterial genomes. Science 311, 1730–1733, 10.1126/science.1119966 (2006).
Rahman, S. A. et al. Comparative analyses of nonpathogenic, opportunistic and totally pathogenic mycobacteria reveal genomic and biochemical variabilities and highlight the survival attributes of Mycobacterium tuberculosis. MBio 5, e02020, 10.1128/mBio.02020-14 (2014).
Kho, M. F. et al. The pore-forming protein Cry5B elicits the pathogenicity of Bacillus sp. against Caenorhabditis elegans. PLoS One 6, e29122, 10.1371/journal.pone.0029122 (2011).
Sun, M., Liu, Z. & Yu, Z. [Characterization of the insecticidal crystal protein genes of Bacillus thuringiensis YBT-1520]. Wei Sheng Wu Xue Bao 40, 365–371 (2000).
Andrup, L., Damgaard, J. & Wassermann, K. Mobilization of small plasmids in Bacillus thuringiensis subsp. israelensis is accompanied by specific aggregation. J Bacteriol 175, 6530–6536 (1993).
Li, R., Li, Y., Kristiansen, K. & Wang, J. SOAP: short oligonucleotide alignment program. Bioinformatics 24, 713–714, 10.1093/bioinformatics/btn025 (2008).
Darling, A. E., Mau, B. & Perna, N. T. progressiveMauve: multiple genome alignment with gene gain, loss and rearrangement. PLoS One 5, e11147, 10.1371/journal.pone.0011147 (2010).
Neely, R. K., Deen, J. & Hofkens, J. Optical mapping of DNA: single-molecule-based methods for mapping genomes. Biopolymers 95, 298–311, 10.1002/bip.21579 (2011).
Liu, X. et al. Construction of an Escherichia coli to Bacillus thuringiensis shuttle vector for large DNA fragments. Appl Microbiol Biotechnol 82, 765–772, doi: 10.1007/s00253-008-1854-y(2009).
Wang, L., Guo, S., Huang, J., Yu, Z. & Sun, M. [Cloning and physical map construction of a large plasmid pBMB165 in Bacillus thuringiensis]. Wei Sheng Wu Xue Bao 48, 15–20 (2008).
Chao, L. et al. Complete nucleotide sequence of pBMB67, a 67-kb plasmid from Bacillus thuringiensis strain YBT-1520. Plasmid 57, 44–54, 10.1016/j.plasmid.2006.06.002 (2007).
Sun, M., Wei, F., Liu, Z. D. & Yu, Z. N. [Cloning of plasmid pBMB2062 in Bacillus thuringiensis strain YBT-1520 and construction of plasmid vector with genetic stability]. Yi Chuan Xue Bao 27, 932–938 (2000).
Zhong, C. et al. Determination of plasmid copy number reveals the total plasmid DNA amount is greater than the chromosomal DNA amount in Bacillus thuringiensis YBT-1520. PLoS One 6, e16025, 10.1371/journal.pone.0016025 (2011).
Sullivan, M. J., Petty, N. K. & Beatson, S. A. Easyfig: a genome comparison visualizer. Bioinformatics 27, 1009–1010, 10.1093/bioinformatics/btr039 (2011).
Alikhan, N. F., Petty, N. K., Ben Zakour, N. L. & Beatson, S. A. BLAST Ring Image Generator (BRIG): simple prokaryote genome comparisons. BMC Genomics 12, 402, doi: 10.1186/1471-2164-12-402 (2011).
Kumar, S., Nei, M., Dudley, J. & Tamura, K. MEGA: a biologist-centric software for evolutionary analysis of DNA and protein sequences. Briefings in bioinformatics 9, 299–306, 10.1093/bib/bbn017 (2008).
Rio, D. C., Ares, M., Jr., Hannon, G. J. & Nilsen, T. W. Purification of RNA using TRIzol (TRI reagent). Cold Spring Harbor protocols 2010, pdb prot5439, 10.1101/pdb.prot5439 (2010).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408, 10.1006/meth.2001.1262 (2001).
Zhu, L. et al. Complete genome sequence of Bacillus thuringiensis serovar galleriae strain HD-29, a typical strain of commercial biopesticide. J Biotechnol 195, 108–109, 10.1016/j.jbiotec.2014.12.021 (2015).
Doggett, N. A. et al. Complete Genome Sequence of Bacillus thuringiensis Serovar Israelensis Strain HD-789. Genome announcements 1, e01023-01013, 10.1128/genomeA.01023-13 (2013).
Challacombe, J. F. et al. The complete genome sequence of Bacillus thuringiensis Al Hakam. J Bacteriol 189, 3680–3681, 10.1128/JB.00241-07 (2007).
Ivanova, N. et al. Genome sequence of Bacillus cereus and comparative analysis with Bacillus anthracis. Nature 423, 87–91, 10.1038/nature01582 (2003).
Ravel, J. et al. The complete genome sequence of Bacillus anthracis Ames “Ancestor”. J Bacteriol 191, 445–446, 10.1128/JB.01347-08 (2009).
Acknowledgements
We thank the Institute of Plant Protection from the Chinese Academy of Agricultural Sciences and BGI Hangzhou for their previous work, as well as Dr. D. Zeigler in BGSC for providing B. thuringiensis 4D1 (BGSC code for HD-1) for comparison. This research was supported by the National Programs for High Technology Research and Development of China (Grant No. 2011AA10A203), China 948 Program of Ministry of Agriculture (2011-G25), the National Natural Science Foundation of China (Grants No. 31170047 and 31171901) and the National Basic Research Program (973 Program) of China (Grant No. 2009CB118902).
Author information
Authors and Affiliations
Contributions
J.H., Z.N.Y. and M.S. conceived and designed the research. L.Z., Y.Y.W., W.X.Y., C.M.Z. and D.M.H. performed the experiments. J.S.Z. and C.G. helped with the data analysis. L.Z. analyzed the data and wrote the manuscript. Y.Y.W., D.H.P. and L.F.R. helped with manuscript revision.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Zhu, L., Peng, D., Wang, Y. et al. Genomic and transcriptomic insights into the efficient entomopathogenicity of Bacillus thuringiensis. Sci Rep 5, 14129 (2015). https://doi.org/10.1038/srep14129
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep14129
This article is cited by
-
Whole genome sequencing of a novel Bacillus thuringiensis isolated from Assam soil
BMC Microbiology (2023)
-
Distribution and Genetic Diversity of Genes from Brazilian Bacillus thuringiensis Strains Toxic to Agricultural Insect Pests Revealed by Real-Time PCR
Microbial Ecology (2023)
-
Genomic–proteomic analysis of a novel Bacillus thuringiensis strain: toxicity against two lepidopteran pests, abundance of Cry1Ac5 toxin, and presence of InhA1 virulence factor
Archives of Microbiology (2023)
-
Thurincin H Is a Nonhemolytic Bacteriocin of Bacillus thuringiensis with Potential for Applied Use
Probiotics and Antimicrobial Proteins (2023)
-
The CRISPR-Cas systems were selectively inactivated during evolution of Bacillus cereus group for adaptation to diverse environments
The ISME Journal (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.