Abstract
Non - specific PCR amplification and DNA contamination usually accompany with PCR process, to overcome these problems, here we establish a sensor for thrombin by sequence - specific recognition of the PCR product with molecular beacon through triplex formation. Probe A and probe B were designed for the sensor, upon addition of thrombin, two probes hybridized to each other and the probe B was extended in the presence of Klenow Fragment polymerase and dNTPs. The PCR amplification occurred with further addition of Taq DNA Polymerase and two primers, the PCR product was recognized by molecular beacon through triplex formation. The fluorescence intensity increased with the logarithm of the concentration of thrombin over the range from 1.0 × 10−12 M to 1.0 × 10−7 M, with a detection limit of 261 fM. Moreover, the effect of DNA contamination and non - specific amplification could be ignored completely in the proposed strategy.
Similar content being viewed by others
Introduction
The polymerase chain reaction (PCR) technology is an essential tool for gene research, because the exponential amplification efficiency enables the detectionof small numbers of nucleic acid1. To extend the scope of PCR to high sensitive detection of protein, Sano et al. established the immuno - PCR (IPCR) method by combines antibody recognition and PCR amplification2,3,4,5,6, which could lead to about 100 - 10000 fold increase of sensitivity over the conventional ELISA for protein7,8,9. Owing to the exponential amplification and high sensitivity of PCR, it has been widely applied in detection of gene mutation10,11,12,13, pharmacogenetics14,15, gene expression analysis16,17,18, recognition of methylation specific loci19,20,21, protein22,23,24,25,26, detection of metal ion27 and constructing of DNA nano - structures28. However, non - specific amplification is inevitable during the process of PCR amplification. Therefore, the sensitivity and selectivity of PCR technique is controlled by detection method for PCR product. There are mainly five kinds of methods. Firstly, gel electrophoresis, it is generally cumbersome and time - consuming, which has been successfully used in the IPCR29,30,31. Secondly, Detection electrochemical signal of guanine nucleobase of PCR product after separation32. Thirdly, detection of PCR products with the TaqMan probes33. Fourthly, PNA or LNA modified molecular beacon through displacement hybridization34. Fifthly, intercalative molecules, such as Ethdium bromide and SYBR Green I etc., their fluorescence increase upon addition of double - stranded DNA, but they cannot distinguish PCRproduct from non - specific amplification product such as primer - dimers35. It is still a challenge to sequence - specifically recognize the duplex structure of PCR product.
Homopurine·Homopyrimidine duplex DNA can be sequence - specifically recognized by homopyrimidine Oligonucleotide (or homopurine Oligonuceotide) through triplex formation36,37. Study of triplex DNA mainly focus on following aspect: firstly, biological function of triplex DNA, such as their effects on the process of gene translation38, DNA transcription, replication and cleavage39,40; Secondly, effect of hole transport41,42,43,44; Thirdly, pH controlled switchable conformation change between duplex DNA and triplex DNA45,46,47,48; Fourthly, sequence - specific recognition of double - stranded DNA49,50,51,52,53,54; Fifthly, biosensing for other molecules55,56,57,58,59,60. However, there had little biosensor that based upon triplex formation and PCR amplification.
To overcome the drawback of PCR - based biosensor and explore new direction for the study of triplex DNA, here we developing a novel biosensor for protein by combining the amplification property of PCR and sequence - specific recognition ability of triplex formation for DNA duplex. As a proof of concept, here a sensitive fluorescent sensor for thrombin is being established with molecular beacon based upon triplex formation and PCR.
Results and Discussion
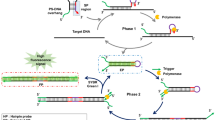
The scheme of the biosensor is shown in Fig. 1. Oligonucleotide 5′-AAT ACC CGA TTG CAG TAC GAC TCT CCA CAA GCC TTT TTT TTT TTT TTT TTT TTT TTT TTT TTT TTG GTT GGT GTG GTT GG -3′ (probe A) and Oligonucleotide5′-CGA GTC CGT GGT AGG GCA GGT TGG GGT GAC TAA AGA GAA GGA AGA GAA GAA GAA AGA AAA GAA AAA GTT TTT TTG GCT TGT G-3′ (probe B) are designed for the sensor, the underlined sequence are aptamers for thrombin, the T32 in probe A and T7 in probe B are designed as spacer. Oligonucleotide5′-AAT ACC CGA TTG CAG TAC GAC TC-3′ (P1) and 5′-GAG TCC GTG GTA GGG CAG GT-3′ (P2) are designed as primers. Upon addition of thrombin, the red and italic sequences of two probes hybridize to each other and the probe B is extended in the presence of Klenow Fragment polymerase and dNTPs. Then PCR amplification is carried out with further addition of dNTPs, Taq DNA Polymerase and two primers. Green sequence of Probe B is homopurine strand, it is designed to create homopurine•homopyrimidine double strand during the process of PCR amplification, which can be recognized by molecular beacon with the sequence of 5′-6-FAM-GTG GAG TTT CTC TTC CTT CTC TTC TTC TTT CTT TTC TTT TTC CTC CAC-BHQ-1-3′ (MB) through triplex formation49,50,51,52,53,54. As shown in Fig. 2A, the fluorescence intensity of PCR product and molecular beacon was only about 60 in the absence of thrombin, it increased to about 155 upon addition of 1.0 × 10−7 M thrombin. These results confirmed the probability of the proposed strategy.
(A) Fluorescence spectroscopy of molecular beacon after hybridization with different cycle of PCR product. The PCR amplification was carried out under the condition of 30 s denaturation at 94 °C, 30 s of annealing temperature at 60 °C and 10 s of extending temperature at 72 °C. MB: 200 nM, spermine: 0.4 mM, pH: 5.0. (B) Circular dichroism (CD) spectroscopy of Oligonucleotides under different conditions. a. MB: 1.0 × 10−5 M; b. R1·R2: 1.0 × 10−5 M; c. Mixture of 1.0 × 10−5 M MB and 1.0 × 10−5 M R1·R2. Spermine: 0.4 mM, pH: 5.0.
The numbers of PCR cycle affect the fluorescence intensity of sensor. As shown in Fig. 2A, the fluorescence intensity increased with the numbers of PCR cycle and reached a maximum at 45 cycles. Thus 45 cycles of PCR reaction cycle was used.
Circular dichroism (CD) spectroscopy was a powerful tool and was usually applied into the study of DNA structure61. It was reported that the negative peak of 210 nm was the marker for triplex DNA62, thereby CD spectroscopy was used to investigate the structure information of Oligonucleotide5′-CGA GTC CGT GGT AGG GCA GGT TGG GGT GAC TAA AGA GAA GGA AGA GAA GAA GAA AGA AAA GAA AAA GTT TTT TTG GCT TGT GGA GAG TCG TAC TGC AAT CGG GTA TT-3′ (R1) and Oligonucleotide5′-AAT ACC CGA TTG CAG TAC GAC TCT CCA CAA GCC AAA AAA ACT TTT TCT TTT CTT TCT TCT TCT CTT CCT TCT CTT TAG TCA CCC CAA CCT GCC CTA CCA CGG ACT CG-3′ (R2), which were two sequences of the PCR product. As shown in Fig. 2B, there were no (or little) negative peak at 210 nm for CD spectroscopy of R1·R2 duplex and MB respectively, while there occurred a strong negative peak at 210 nm for the mixture of R1·R2 and MB, which indicated that there was triplex formation between R1-R2 (PCR product) and MB. These results was in accordance with that of literature49,50,51,52,53,54.
The sensitivity of the proposed strategy was investigated under the conditions of 200 nM MB, 0.4 mM of spermine, pH 5.0 and 90 minutes of hybridization time for fluorescence measurement and 45 circles of PCR. As demonstrated in Fig. 3, the fluorescence intensity was proportional to the logarithm of the concentration of thrombin over the range from 1.0 × 10−12 M to 1.0 × 10−7 M. The linear regression equation was I = 15.12 log10C + 243.9 (C: M, r = 0.995, C represents the concentration of thrombin, I denotes the fluorescence intensity), with a detection limit of 261 fM, which was obtained from the equation of DL = 3б/slope. The comparison of the proposed method with others was listed in Table 1, it had merit of wide linear range.
Calibration curve for the biosensor.
45 cycles of PCR amplification was carried out by using 30 s of denaturation at 94 °C, 30 s of annealing temperature at 60 °C and 10 s of extending temperature at 72 °C. MB: 200 nM, spermine: 0.4 mM, pH:5.0. Each point was the mean of three measurements. The error bars are the standard deviation.
Non - specific PCR amplification usually accompanied with the PCR amplification process. DNA contamination might easily occurred during PCR amplification process as well. Both DNA contamination and non - specific amplification PCR product were double - stranded DNA, which was similar to that of PCR product, so it was difficult to distinguish the signal for non - specific amplification and DNA contamination from that for PCR product and then they usually affect the detecting result. To investigate the effect of non - specific PCR amplification and DNA contamination on the proposed strategy, the fluorescence intensity of the PCR product were measured with the proposed strategy and the common probe (SYRB - Green) for real - time PCR respectively. Here lambda - DNA was used to act as non - specific amplification product and DNA contamination due to its duplex structure. As shown in Fig. 4, the fluorescence intensity increased with concentration of lambda - DNA by using the probe of SYRB - Green, while the fluorescence intensity had little change when the concentration of lambda - DNA increased from 0 to 1.6 μg/mL by using the proposed strategy. These results indicated that the proposed strategy could overcome the effect of non - specific PCR amplification and DNA contamination completely and the effect of non - specific amplification could be ignored in the proposed method. These results mainly due to the stringent sequence - specific recognition between PCR product and MB through triplex formation49,50,51,52,53,54.
Effect of lambda - DNA on the fluorescence of the PCR product of 1.0 × 10−8 M thrombin.
(A) Detecting PCR product by using SYRB - Green; SYRB - Green: 1 × , spermine: 0.4 mM, pH:5.0. (B) Detecting PCR product with the proposed method (detecting PCR product through triplex formation); MB: 200 nM, spermine:0.4 mM, pH:5.0.
In order to test the selectivity of the proposed sensor, the effect of other possible interferences were investigated. 1.0 × 10−7 M of lysozyme, hemoglobin and apo - transferrin human were used to replace the 1.0 × 10−8 M of thrombin for PCR amplification respectively, As demonstrated in Fig. 5A, each of their fluorescence intensity was almost at the same level of the blank. Moreover, the existence of 10-folded lysozyme, hemoglobin and apo -transferrin human had little effect on the fluorescence intensity of thrombin. To further certificate the selectivity of the sensor, electrophoresis photograph of PCR product was investigated, as demonstrated in Fig. 5B, there were no bands with addition of lysozyme, hemoglobin and apo-transferrin human, while there was the correct size band in the thrombin, which indicated that these proteins had no effect on detection of thrombin. These results indicated that the proposed biosensor had good selectivity.
Selectivity of the sensor and effect of interference molecules.
(A) a: blank; b: 1.0 × 10−7 M of lysozyme; c: 1.0 × 10−7 M of hemoglobin; d: 1.0 × 10−7 M of apo-transferrin human; e: 1.0 × 10−8 M of thrombin; f: mixture of 1.0 × 10−8 M of thrombin and 1.0 × 10−7 M of lysozyme; g: mixture of 1.0 × 10−8 M of thrombin and 1.0 × 10−7 M of hemoglobin; h: mixture of 1.0 × 10−8 M of thrombin and 1.0 × 10−7 M of apo-transferrin human. Every point was the mean of three measurements, error bar was the standard deviation. (B) Electrophoresis photograph of PCR product. 1. 500 bp DNA ladder; 2. Blank; 3. 1.0 × 10−8 M of thrombin; 4. 1.0 × 10−7 M of lysozyme; 5. 1.0 × 10−7 M of hemoglobin; 6. 1.0 × 10−7 M of apo-transferrin human; 7. mixture of 1.0 × 10−8 M of thrombin and1.0 × 10−7 M of lysozyme, hemoglobin, apo-transferrin human; EB: 0.5 μg/mL; Agarose: 3%.
To assess the analytical application of the sensor, the method was used to detect thrombin in human sera. Since no thrombin was found from the human sera, addition and recovery experiment was performed to estimate the application of the assay in complex sample. As demonstrated in Table 2, 5.0 × 10−11 – 5.0 × 10−8 M of thrombin was added into each human sera, the recovery ranges from 87.6% to 112.6% and the relative standard deviation values were in the ranges of 4.3%–9.8%, which indicated that the method had good analytical application in human sera.
In summary, we have developed an ultrasensitive and selective sensor for thrombin by combining the PCR amplification and triplex formation. The fluorescence intensity was proportional to the logarithm of the concentration of thrombin over the range of 1.0 × 10−12 M–1.0 × 10−7 M, with a detection limit of 261 fM. The most important contribution is that the proposed strategy can overcome the effect of non - specific amplification and DNA contamination completely, which was realized by detecting PCR product with MB through triplex formation. Moreover, the proposed strategy may open a new avenue for the application of triplex DNA, it may be extend to immune - PCR and other aptasensor in the future.
Methods
Materials and Reagents
All Oligonucleotides were purchased from Sangon Bioengineering Technology and Services Co. Ltd. (Shanghai, China). Human thrombin, lysozyme (LYS), apo-transferrin human (ATH), hemoglobin (HB), spermine were purchased from Sigma-Aldrich (USA). Klenow Fragment (exo-) polymerase was purchased from Thermo scientific Inc (USA). Taq DNA Polymerase and dNTPs were obtained from Sangon Bioengineering Technology and Services Co. Ltd. (Shanghai, China). SYBR Green I (20×) was purchased from Bio Teke Corporation (Beijing, China). Nanopure water (18.1 MΩ) was obtained from a 350 Nanopure water system (Guangzhou Crystalline Resource Desalination of Sea Water and Treatment Co. Ltd.) and was used in all experiments. All chemicals were analytical grade unless stated.
Tris-HAc buffer (20 mM, pH 7.9, 50 mM KAc, 10 mM Mg (Ac)2 and 1.0 mM DTT) was used throughout the recognition of thrombin with aptamer. Phosphate buffered solution (PBS) (pH 5.0, 0.1 M Na2HPO4, 0.1 M NaH2PO4 and 0.1 M NaAc) was used for triplex formation between PCR product and MB.
Apparatus
The PCR amplification was performed on a ETC 811 PCR Instrument (Eastwin Life Sciences, Inc., China). The fluorescence signal was measured on a RF-5301PC spectrofluorimeter (Shimadzu, Japan). The circular dichroism (CD) spectroscopy was obtained from a J-810-150S spectropolarimeter (JASCO International Co. Ltd., Japan).
Recognition of thrombin and preparation of PCR template
Probe A and probe B were denatured at 90 °C for 5 min, then cooled to 0 °C rapidly. 2 μL of thrombin solution was mixed with 2 μL 4.0 × 10−10 M probe A and probe B in Tris-HAc buffer, kept at 37 °C for 30 min. After that, 0.5 U Klenow Fragment (exo-) polymerase and 0.9 μL 2.5 mM dNTPs were added and the total mixture volume reached 10 μL, kept the mixture at 37 °C for 30 min, along with heating to 95 °C to inactivate the Klenow Fragment (exo-) polymerase. Then the inactivated mixture was acted as PCR template for following PCR amplification.
PCR amplification
The PCR amplification was carried out in a 200 μL PCR tube that containing 5.0 μL PCR template, 2.0 μL 10 μM of two primers, 5 μL of 10 × PCR Buffer, 1 μL 5 U/μL of Taq DNA Polymerase, 3 μL 25 mM of MgCl2, 1 μL 10 mM of dNTPs and 31 μL of Nanopure H2O. The thermal program was comprised of an initial denaturation at 95 °C for 10 min, 45 cycles of PCR amplification was carried out by using 30 s of denaturation at 94 °C, 30 s of annealing temperature at 60 °C and 10 s of extending temperature at 72 °C. All the reactions were run in triplicates and the control experiment was carried out with same reaction mixture in the absence of thrombin.
Measurement of fluorescence spectrum
10 μL of 10 μΜ MB were mixed with 490 μL PBS buffer solution which contained different amounts of PCR product. After 1.5 hours of incubation(25 °C), the fluorescence signal was measured with spectrofluorimeter. Slit widths were both 5.0 nm and the excitation and emission wavelengths were set at 495 and 518 nm, respectively.
Measurement of CD spectroscopy
The circular dichroism (CD) spectroscopy was measured at room temperature and performed over the wavelength range from 200 to 300 nm in 0.1 cm path length cuvettes. The result was obtained by averaging 3 scans at the scanning rate of 100 nm per minute with a response time of 1.0 s and the bandwidth of 1.71 nm.
Additional Information
How to cite this article: Li, Y. et al. Triplex DNA: A new platform for polymerase chain reaction - based biosensor. Sci. Rep. 5, 13010; doi: 10.1038/srep13010 (2015).
References
Saiki, R. K. et al. Enzymatic amplification of beta-globin genomic sequences and restriction site analysis for diagnosis of sickle cell anemia. Science 230, 1350–1354 (1985).
Sano, T., Smith, C. L. & Cantor, C. R. Immuno-PCR: very sensitive antigen detection by means of specific antibody-DNA conjugates. Science 258, 120–122 (1992).
Malou, N. & Raoult, D. Immuno-PCR: a promising ultrasensitive diagnostic method to detect antigens and antibodies. Trends. Microbiol. 19, 295–302 (2011).
Zhang, H. et al. Ultrasensitive assays for proteins. Analyst 132, 724–737 (2007).
Adler, M., Wacker, R. & Niemeyer, C. M. Sensitivity by combination: immuno-PCR and related technologies. Analyst 133, 702–718 (2008).
Niemeyer, C. M., Adler, M. & Wacker, R. Immuno-PCR: high sensitivity detection of proteins by nucleic acid amplification. Trends. Biotechnol. 23, 208–216 (2005).
Ruzicka, V. et al. University expedition to study the natives of centralaustralia. Science 260, 260–261 (1993).
Fredriksson, S. et al. Protein detection using proximity-dependent DNA ligation assays. Nat. Biotechnol. 20, 473–477 (2002).
Niemeyer, C. M., Adler, M. & Wacker, R. Detecting antigens by quantitative immuno-PCR. Nat. Protoc. 2, 1918–1930 (2007).
Wang, J. et al. Quantifying EGFR alterations in the lung cancer genome with nanofluidic digital PCR arrays. Clin. Chem. 56, 623–632 (2010).
Dias-Santagata, D. et al. Rapid targeted mutational analysis of human tumours: a clinical platform to guide personalized cancer medicine. EMBO Mol. Med. 2, 146–158 (2010).
Totoki, Y. et al. High-resolution characterization of a hepatocellular carcinoma genome. Nat. Genet. 43, 464–469 (2011).
Day, E., Dear, P. H. & McCaughan, F. Digital PCR strategies in the development and analysis of molecular biomarkers for personalized medicine. Methods 59, 101–107 (2013).
Ouahchi, K., Lindeman, N. & Lee, C. Clinical guidelines for psychiatrists for the use of pharmacogenetic testing for CYP450 2D6 and CYP450 2C19. Pharmacogenomics. 7, 25–29 (2006).
Wang, L., McLeod, H. L. & Weinshilboum, R. M. Genomics and drug response. N. Engl. J. Med. 364, 1144–1153 (2011).
Reiter, M. et al. Quantification noise in single cell experiments. Nucleic Acids Res. 39, e124 (2011).
Rottiers, V. & Naar, A. M. MicroRNAs in metabolism and metabolic disorders. Nat. Rev. Mol. Cell Biol. 13, 239–250 (2012).
Petriv, O. I. et al. Comprehensive microRNA expression profiling of the hematopoietic hierarchy. Proc. Natl. Acad. Sci. USA. 107, 15443–15448 (2010).
Baylin, S. B. & Jones, P. A. A decade of exploring the cancer epigenome - biological and translational implications. Nat. Rev. Cancer 11, 726–734 (2011).
Weisenberger, D. J. et al. DNA methylation analysis by digital bisulfite genomic sequencing and digital MethyLight. Nucleic Acids Res. 36, 4689–4698 (2008).
Cipriany, B. R. et al. Single molecule epigenetic analysis in a nanofluidic channel. Anal. Chem. 82, 2480–2487 (2010).
Fredriksson, S. et al. Protein detection using proximity-dependent DNA ligation assays. Nat. Biotechnol. 20, 473–477 (2002).
Wang, X. L. et al. Ultrasensitive detection of protein using an aptamer-based exonuclease protection assay. Anal. Chem. 76, 5605–5610 (2004).
Schallmeiner, E. et al. Sensitive protein detection via triple-binder proximity ligation assays. Nat. Methods. 4, 135–137 (2007).
Xue, L. Y., Zhou, X. M. & Xing, D. Highly sensitive protein detection based on aptamer probe and isothermal nicking enzyme assisted fluorescence signal amplification. Chem. Commun. 46, 7373–7375 (2010).
Ma, C. P. et al. The real-time PCR for sensitive protein detection by target-induced intermolecular hybridization. Chem. Commun. 47, 11303–11305 (2011).
Lee, S. Y. et al. Rapid pathogen detection with bacterial-assembled magnetic mesoporous silica. Biosens. Bioelectron. 53, 123–128 (2014).
Kuang, H. et al. Nanoscale superstructures assembled by polymerase chain reaction (PCR): programmable construction, structural diversity and emerging applications. Acc. Chem. Res. 46, 2341–2354 (2013).
Furuya, D. et al. Immuno-PCR assay for homodimeric osteoprotegerin. Clin. Chem. 47, 1475–1477 (2001).
Komatsu, M. et al. Tumor necrosis factor-alpha in serum of patients with inflammatory bowel disease as measured by a highly sensitive immuno-PCR. Clin. Chem. 47, 1297–1301 (2001).
Chye, S. M. et al. Immuno-PCR for detection of antigen to Angiostrongylus cantonensis circulating fifth-stage worms. Clin. Chem. 50, 51–57 (2004).
Xiang, Y. et al. Ultrasensitive label-free aptamer-based electronic detection. Angew. Chem. Int. Ed. 46, 9054–9056 (2007).
Kutyavin, I. V. et al. 3′-minor groove binder-DNA probes increase sequence specificity at PCR extension temperatures. Nucleic Acids Res. 28, 655–661 (2000).
Costa, J. M. et al. Chimeric LNA/DNA probes as a detection system for real-time PCR. Clin. Biochem. 37, 930–932 (2004).
Zipper, H. et al. Investigations on DNA intercalation and surface binding by SYBR Green I, its structure determination and methodological implications. Nucleic Acids Res. 32, e103 (2004).
Doluca, O., Withers, J. M. & Filichev, V. V. Molecular engineering of guanine-rich sequences: Z-DNA, DNA triplexes and G-quadruplexes. Chem. Rev. 113, 3044–3083 (2013).
Ling, L. S., Butt, H. J. & Berger, R. Rupture force between the third strand and the double strand within a triplex DNA. J. Am. Chem. Soc. 126, 13992–13997 (2004).
Knudsen, H. & Nielsen, P. E. Antisense properties of duplex- and triplex-forming PNAs. Nucleic Acids Res. 24, 494–500 (1996).
Vasquez, K. M., Narayanan, L. & Glazer, P. M. Specific mutations induced by triplex-forming oligonucleotides in mice. Science 290, 530–533(2000).
Maine, I. P. & Kodadek, T. Efficient unwinding of triplex DNA by a DNA helicase. Biochem. Biophys. Res. Commun. 204, 1119–1124 (1994).
Wang, J., Sun, L. X. & Bu, Y. X. Negative dissociation energy phenomenon of metastable H-bonds as revealed in triplex DNA hole migration. J. Phys. Chem. B. 114, 1144–1147 (2010).
Tanabe, K. et al. Electrochemical evaluation of alternating duplex-triplex conversion effect on the anthraquinone-photoinjected hole transport through DNA duplex immobilized on a gold electrode. J. Am. Chem. Soc. 128, 692–693 (2006).
Dohno, C., Nakatani, K. & Saito, I. Guanine of the third strand of C.G*G triplex serves as an effective hole trap. J. Am. Chem. Soc. 124, 14580–14585 (2002).
Lewis, F. D. et al. DNA-mediated electron transfer across synthetic T:A.T triplex structures. Angew. Chem. Int. Ed. 41, 3485–3487 (2002).
Idili, A., Vallee-Belisle, A. & Ricci, F. Programmable pH-triggered DNA nanoswitches. J. Am. Chem. Soc. 136, 5836–5839 (2014).
Amodio, A. et al. Rational design of pH-controlled DNA strand displacement. J. Am. Chem. Soc. 136, 16469–16472 (2014).
Idili, A. et al. Thermodynamic basis for engineering high-affinity, high-specificity binding-induced DNA clamp nanoswitches. Acs Nano. 7, 10863–10869 (2013).
Li, T. et al. Input-dependent induction of oligonucleotide structural motifs for performing molecular logic. J. Am. Chem. Soc. 134, 3508–3516 (2012).
Patterson, A. et al. Using triplex-forming oligonucleotide probes for the reagentless, electrochemical detection of double-stranded DNA. Anal. Chem. 82, 9109–9115 (2010).
Feng, L. Y. et al. Graphene platform used for electrochemically discriminating DNA triplex. ACS Appl. Mater. Interfaces. 6, 3513–3519 (2014).
McKenzie, F., Faulds, K. & Graham, D. LNA functionalized gold nanoparticles as probes for double stranded DNA through triplex formation. Chem. Commun. 20, 2367–2369 (2008).
Miao, X. M. et al. Dynamic-light-scattering-based sequence-specific recognition of double-stranded DNA with oligonucleotide-functionalized gold nanoparticles. Chem. - Eur. J. 17, 11230–11236 (2011).
Xiao, Z. Y., Guo, X. T. & Ling, L. S. Sequence-specific recognition of double-stranded DNA with molecular beacon with the aid of Ag(+) under neutral pH environment. Chem. Commun. 49, 3573–3575 (2013).
Miao, X. M. et al. Electrochemical molecular beacon biosensor for sequence-specific recognition of double-stranded DNA. Biosens. Bioelectron. 59, 54–57 (2014).
Ihara, T. et al. Silver ion unusually stabilizes the structure of a parallel-motif DNA triplex. J. Am. Chem. Soc. 131, 3826–3827 (2009).
Han, M. S., Lytton-Jean, A. K. R. & Mirkin, C. A. A gold nanoparticle based approach for screening triplex DNA binders. J. Am. Chem. Soc. 128, 4954–4955 (2006).
Xi, D. M. et al. Detection of cancer cells using triplex DNA molecular beacons based on expression of enhanced green fluorescent protein (eGFP). Chem. Commun. 50, 9547–9549 (2014).
Zhu, D. S. et al. Sensitive detection of transcription factors using an Ag(+)-stabilized self-assembly triplex DNA molecular switch. Chem. Commun. 50, 14987–14990 (2014).
Du, Y. et al. An integrated sensing system for detection of DNA using new parallel-motif DNA triplex system and graphene-mesoporous silica-gold nanoparticle hybrids. Biomaterials 32, 8584–8592 (2011).
Bi, S. et al. Exponential amplification for chemiluminescence resonance energy transfer detection of microRNA in real samples based on a cross-catalyst strand-displacement network. Anal. Chem. 83, 3696–3702 (2011).
Baase, W. A. & Johnson, Jr, W. C. Circular dichroism and DNA secondary structure. Nucleic Acids Res. 6, 797–814 (1979).
Manzini, G. et al. Triple helix formation by oligopurine-oligopyrimidine DNA fragments. Electrophoretic and thermodynamic behavior. J. Mol. Biol. 213, 833–843 (1990).
Yuan, Y. L. et al. Hemin/G-quadruplex simultaneously acts as NADH oxidase and HRP-mimicking DNAzyme for simple, sensitive pseudobienzyme electrochemical detection of thrombin. Chem. Commun. 48, 4621–4623 (2012).
Lin, X. X. et al. A portable microchip for ultrasensitive and high-throughput assay of thrombin by rolling circle amplification and hemin/G-quadruplex system. Biosens. Bioelectron. 56, 71–76 (2014).
Hu, J. et al. Electrostatic Interaction Based Approach to Thrombin Detection by Surface-Enhanced Raman Spectroscopy. Anal. Chem. 81, 87–93 (2009).
Zhou, W. J. et al. Near Infrared Surface Plasmon Resonance Phase Imaging and Nanoparticle-Enhanced Surface Plasmon Resonance Phase Imaging for Ultrasensitive Protein and DNA Biosensing with Oligonucleotide and Aptamer Microarrays. Anal. Chem. 84, 440–445 (2012).
Acknowledgements
The work was supported by National Natural Science Foundation of China (No: 21375153, 21305053), the Fundamental Research Funds for the Central Universities (No: 13lgzd05) and open project of Beijing National Laboratory for Molecular Sciences.
Author information
Authors and Affiliations
Contributions
L.S.Ling and X.M.Miao designed the experiments. Y.B.Li performed experiments and data analysis. L.S.Ling and Y.B.Li wrote the main text. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Li, Y., Miao, X. & Ling, L. Triplex DNA: A new platform for polymerase chain reaction – based biosensor. Sci Rep 5, 13010 (2015). https://doi.org/10.1038/srep13010
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep13010
This article is cited by
-
Applications of triplex DNA nanostructures in sensor development
Analytical and Bioanalytical Chemistry (2022)
-
Switchable DNA tweezer and G-quadruplex nanostructures for ultrasensitive voltammetric determination of the K-ras gene fragment
Microchimica Acta (2019)
-
Aptamer-involved fluorescence amplification strategy facilitated by directional enzymatic hydrolysis for bioassays based on a metal-organic framework platform: Highly selective and sensitive determination of thrombin and oxytetracycline
Microchimica Acta (2017)
-
Homogenous assay for protein detection based on proximity DNA hybridization and isothermal circular strand displacement amplification reaction
Analytical and Bioanalytical Chemistry (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.