Abstract
The high mobility group box 1 (HMGB1), which is a highly conserved and evolutionarily non-histone nuclear protein, has been shown to associate with a variety of biological important processes, such as transcription, DNA repair, differentiation and extracellular signalling. High HMGB1 expression has been reported in many cancers, such as prostate, kidney, ovarian and gastric cancer. However, there have been few studies of the function of HMGB1 in the malignant biological behaviour of bladder urothelial carcinoma (BUC) and the potential mechanism of HMGB1 in the pathogenesis of BUC remains unclear. Thus, in this study, we constructed plasmid vectors that are capable of synthesizing specific shRNAs targeting HMGB1 and transfected them into BUC cells to persistently suppress the endogenous gene expression of HMGB1. The expression of HMGB1, the bioactivity of BUC cells, including proliferation, apoptosis, cell cycle distribution, migration and invasion and the effects of HMGB1 knockdown on downstream signalling pathways were investigated. Our data suggest that HMGB1 promotes the malignant biological behaviour of BUC and that this effect may be partially mediated by the NF-κB signalling pathway. HMGB1 may serve as a potential therapeutic target for BUC in the future.
Similar content being viewed by others
Introduction
Bladder carcinoma (BC) is the most common malignancy of the urinary tract and the seventh most common cancer in men and the 17th in women worldwide1. In the United States, it has been estimated that nearly 74,690 new cases of BC were diagnosed in 2014, with approximately 15,580 deaths2. Despite the development of surgical techniques and instruments, patients with non-muscle invasive BC still experience a high risk of recurrence and 1/3 of these patients will progress to muscle-invasive BC3,4. The clinical outcome of muscle-invasive BC is also unfavourable, with an overall survival rate of 48% to 67% within 5 years5. Bladder urothelial carcinomas (BUCs) represent nearly 90% of BCs that arise from an epithelial origin. Therefore, it is essential to study the mechanism of BUC development and progression and to identify more effective treatment strategies for this malignancy.
High mobility group box 1 (HMGB1), a nuclear non-histone protein, has been associated with a variety of biological important processes, such as transcription, DNA repair, V(D)J recombination, differentiation and extracellular signalling6,7,8. Acting as a chromatin-binding factor, HMGB1 exerts its key functions within the nucleus by binding the minor groove of DNA and facilitating the assembly of site-specific DNA-binding proteins, which regulate the transcription of a number of genes9,10,11,12. In addition to its nuclear role, HMGB1 can be actively secreted by inflammatory cells and passively released from necrotic cells into the local microenvironment, acting as an extracellular signalling molecule that binds individual surface receptors, including the receptor for advanced glycation end products (RAGE) and Toll-like receptors (TLRs) –2, –4 and –9 during inflammation, cell migration, cell differentiation and cancer metastasis13,14. The overexpression of HMGB1 has been confirmed in a variety of cancers, such as prostate cancer15, renal cell carcinoma16, bladder cancer17, hepatocellular carcinoma18, gastric cancer19, colorectal cancer20 and lung cancer21. Furthermore, overexpression of HMGB1 is associated with all the hallmarks of cancer, including limitless replicative potential, evasion of apoptosis, angiogenesis, inflammatory microenvironment and tissue invasion and metastasis, indicating that HMGB1 might be a new potential therapeutic target for the treatment of human malignancies22.
There have been few studies of the function of HMGB1 in the malignant biological behaviour of BUC. Thus, in this study, we constructed HMGB1 shRNA plasmid vectors, which can be processed to generate small interfering RNAs (siRNAs) and transfected them into BUC cells to persistently suppress the endogenous gene expression of HMGB1. The expression of HMGB1, the bioactivity of BUC cells, including proliferation, apoptosis, cell cycle distribution, migration and invasion and the effects of HMGB1 knockdown on downstream signalling pathways were investigated to obtain further insights into the role of HMGB1 in the pathogenesis of BUC and the probable mechanism of this role.
Results
HMGB1 expression is up-regulated in BUC tissues and cell lines
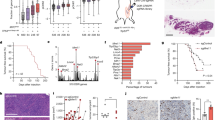
Fifteen BUC tissues and paired adjacent non-tumour tissues were used to detect the expression of HMGB1. Quantitative real-time PCR (qRT-PCR) showed that the HMGB1 mRNA level was significantly higher in BUC tissues compared with paired adjacent non-tumour tissues (5.390 ± 3.329 vs. 2.373 ± 1.485, P < 0.05, Fig. 1A1, A2). Similarly, western blotting (WB) analyses showed that the expression of HMGB1 protein was higher in BUC tissues compared with paired adjacent non-tumour tissues (0.262 ± 0.109 vs. 0.129 ± 0.111, P < 0.05, Fig. 1B1, B2). Next, the expression and subcellular localization of HMGB1 protein were detected by immunohistochemistry (IHC) in a series of 30 BUC tissues and paired adjacent non-tumour tissues. According to the criteria described below, high expression of HMGB1 was observed in 56.7% (17/30) of BUC tissues and only 10.0% (3/30) of adjacent non-tumour tissues (P < 0.05, Fig. 1C1, C2 and C3). Additionally, expression of HMGB1 in 5637, BIU-87, T24 and SV-HUC-1 cells was detected by qRT-PCR and WB. As shown in Fig. 1D, E1 and E2, HMGB1 mRNA and protein were highly expressed in BUC cell lines compared to the SV-HUC-1 normal urothelial cell line (all P < 0.001). Furthermore, among these BUC cells, the expression level of HMGB1 was highest in T24 cells and lowest in 5637 cells.
HMGB1 expression is up-regulated in BUC tissues and cell lines.
Figure 1A1 and A2: qRT-PCR was performed to evaluate the levels of HMGB1 mRNA in 15 paired fresh BUC and adjacent non-tumour tissues. *P < 0.05, compared with adjacent non-tumour tissue. Figure 1B1 and B2: The overexpression of HMGB1 protein was detected by WB in 15 BUC and paired adjacent non-tumour tissues (T: BUC tissue; N: adjacent non-tumour tissue). *P < 0.05, compared with adjacent non-tumour tissue. Figure 1C1, C2 and C3: The expression of HMGB1 in BUC and adjacent non-tumour tissues was examined by IHC (×200). HMGB1 staining was much weaker in the majority of adjacent non-tumour tissues (Fig. 1C1). Among BUC samples, low expression of HMGB1 was detected only in 13/30 (43.3%) of BUC tissues (Fig. 1C2), while high expression of HMGB1 was apparent in 17/30 (56.7%) of BUC tissues (Fig. 1C3). Figure 1D, E1 and E2: qRT-PCR and WB were performed to measure the levels of HMGB1 mRNA and protein in different BUC cell lines. Pairwise significant differences were detected among them, with the HMGB1 expression levels all significantly higher than in the SV-HUC-1 cell line (all P < 0.001). Furthermore, among these BUC cells, the expression level of HMGB1 was highest in T24 cells and the lowest in 5637 cells. The display of cropped gels is used to improve the clarity and conciseness of the presentation and all the cropped gels have been run under the same experimental conditions.
RNA interference (RNAi) effectively inhibits HMGB1 expression in BUC cells
For the following experiments, three BUC cell lines (T24, BIU-87 and 5637) were divided into three groups: the CON group (non-transfected cells), the NC group (shNC-transfected cells) and the shRNA group (HMGB1 shRNA-transfected cells). WB and qRT-PCR were performed to measure HMGB1 expression after RNAi (Fig. 2A1, A2 and A3). The results showed different expression levels of HMGB1 among the three groups of each cell line. HMGB1 shRNA plasmids significantly inhibited the expression of HMGB1 mRNA and protein compared with the CON and NC groups (all P < 0.001). There were no significant differences in the expression of HMGB1 mRNA and protein between the CON and NC groups (all P > 0.05). The results of qRT-PCR and WB demonstrated that the RNAi was effective.
Knockdown of HMGB1 expression inhibits the biological behaviour of BUC cell lines.
Figure 2A1, A2 and A3: HMGB1 expression of BUC cell lines was detected in groups after different transfections, which resulted in the differential expression of HMGB1 among the groups in each BUC cell line. The results indicated that the HMGB1 shRNA plasmids significantly inhibited the expression of HMGB1 compared with the CON and NC groups (all P < 0.001). No significant differences were found between CON and NC in three BUC cell lines (all P > 0.05). Figure 2B1, B2 and B3: Cell proliferation of BUC cell lines evaluated by MTT assay showed that knockdown of HMGB1 expression (shRNA group) resulted in lower cell proliferation than in the CON and NC groups (all P < 0.05). Figure 2C1 and C2: Cell apoptosis assays for BUC cell lines indicated that knockdown of HMGB1 expression induced cell apoptosis compared with the CON and NC groups (all P < 0.05). Figure 2D1, D2, D3 and D4: Cell cycle phase assays for BUC cell lines demonstrated that knockdown of HMGB1 expression led to an increase in the percentage of cells in G0/G1 phase, with a significantly lower proliferation index (PI) than the other groups (all P < 0.05). Figure 2E1, E2, F1 and F2: Cell migration and invasion assays confirmed that knockdown of HMGB1 expression reduced the migration and invasive ability of BUC cell lines in vitro compared with the CON and NC groups (all P < 0.001).
Knockdown of HMGB1 expression inhibits BUC cell proliferation
Following knockdown of HMGB1 gene expression with shRNA, the proliferation of transfected BUC cells was measured using the MTT assay at 24 h, 48 h, 72 h and 96 h post-transfection (Fig. 2B1, B2 and B3). The cell proliferation in the shRNA groups was significantly reduced 24 h after transfection compared with both the CON and NC groups (all P < 0.05, Supplementary Table S1). There were no significant differences in cell proliferation between the CON and NC groups in BUC cells (all P > 0.05, Supplementary Table S1).
Knockdown of HMGB1 expression enhances BUC cell apoptosis
As demonstrated in Fig. 2C1 and C2, early and late apoptosis were higher in the cells transfected with HMGB1 shRNA plasmids (shRNA group) than in the CON and NC groups (all P < 0.05). No significant differences in cell apoptosis between the CON and NC groups were observed in the BUC cells (all P > 0.05).
Knockdown of HMGB1 expression induces BUC cell cycle arrest
To investigate the mechanism by which the proliferation of BUC cells was suppressed by the down-regulation of HMGB1 expression, flow cytometry was used to detect the specific phase of the cell cycle. The data in Fig. 2D1, D2 D3 and D4 demonstrate that knocking down HMGB1 expression (shRNA group) induced BUC cell arrest in the G0/G1 phase, with a proliferation index (PI) that was significantly lower than the CON and NC groups (all P < 0.05). No significant differences were found between the CON and NC groups in the cell cycle assay (all P > 0.05).
Knockdown of HMGB1 expression inhibits BUC cell migration and invasion
As shown in Fig. 2E1, E2, F1 and F2, the number of migrating and invading BUC cells in the shRNA group were significantly lower compared with the CON and NC groups. In addition, the BUC cells in which HMGB1 had been knocked down had lower migration and invasive abilities (all P < 0.001), while no significant differences between cell migratory and invasive ability were found for the CON and NC groups (all P > 0.05).
Effects of HMGB1 down-regulation on NF-κB/p65, IκBα and VEGF-C in T24 cells
To further understand the probable mechanism of HMGB1 shRNA-mediated biological changes, the expression levels of NF-κB/p65, IκBα and VEGF-C were detected by WB and qRT-PCR. In addition, the nucleic localization and DNA-binding activity of NF-κB/p65 were determined by immunofluorescence and electrophoretic mobility shift assay (EMSA), respectively. As shown in Fig. 3A1, A2 and A3, the protein and mRNA levels of NF-κB/p65 and VEGF-C were significantly lower in cells transfected with HMGB1 shRNA plasmids than in those transfected with shNC plasmids or untransfected (all P < 0.05). On the contrary, IκBα expression showed the opposite tendency. There were no significant differences in NF-κB/p65, IκBα and VEGF-C expression between the CON and NC groups (all P > 0.05). At 72 h post-transfection, the NF-κB/p65 proteins were stained green and the nuclei of the T24 cells were stained blue. As shown in Fig. 3B, the knockdown of HMGB1 expression by shRNA-mediated RNAi inhibited the translocation of NF-κB/p65 from the cytoplasm to the nucleus. Furthermore, the EMSA assay suggested that the DNA-binding activity of NF-κB/p65 in T24 cells was decreased by HMGB1 knockdown (Fig. 3C).
Effects of HMGB1 down-regulation on NF-κB/p65, IκBα and VEGF-C in T24 cells.
Figure 3A1, A2 and A3: The expression of NF-κB/p65 and VEGF-C in the shRNA group was lower than in the other two groups transfected with shNC plasmids or the untransfected controls (all P < 0.05). On the contrary, the expression of IκBα showed an opposite tendency, while no significant differences in the expression of NF-κB/p65, IκBα and VEGF-C in T24 cells were found between the CON and NC groups (all P > 0.05). The display of cropped gels is used to improve the clarity and conciseness of the presentation and all the cropped gels have been run under the same experimental conditions. Figure 3B: The blue areas indicate nuclei stained using 4, 6-diamidino-2-phenylindole (DAPI) and the green areas indicate the nuclear translocation of NF-κB/p65 in T24 cells transfected with shNC plasmids or untransfected and cytoplasmic localization of NF-κB/p65 in cells transfected with shRNA plasmids. The results showed that knockdown of HMGB1 expression inhibited the translocation of NF-κB/p65 from the cytoplasm to the nucleus. Figure 3C: EMSA revealed that the DNA-binding activity of NF-κB/p65 in T24 cells was decreased by HMGB1 knockdown.
Discussion
In the nucleus, HMGB1 is a DNA-binding protein that serves as a structural component to facilitate the formation of nucleoprotein complexes. Outside the cell, HMGB1 can act as a cytokine, which can be released into the extracellular environment by tumour or inflammatory cells undergoing necrosis, or in response to many distinct triggers, including hypoxia, nutrient deprivation, shortage of essential growth factors, or application of radiation and chemotherapy23. Thus, it has been demonstrated that HMGB1 plays an important role in a variety of biological processes, including transcription, DNA repair, differentiation, the activation of endothelial cells and the initiation of inflammation24,25. Previous studies have confirmed that HMGB1 is overexpressed in a variety of cancers15,16,17,18,19,20,21. Furthermore, HMGB1 has been implicated in many hallmarks of cancer, including apoptosis, angiogenesis, invasion, metastasis and inflammatory microenvironment22. In this study, we confirmed that both mRNA and protein levels of HMGB1 were significantly higher in BUC tissues and cell lines than in non-tumour cells. To investigate whether HMGB1 could be used as a potential therapeutic target, we constructed specific HMGB1 shRNA plasmids and transfected them into BUC cells. The results showed that shRNA-mediated RNAi efficiently suppressed HMGB1 expression in BUC, which suggests that plasmid-based shRNA targeting HMGB1 expression in BUC cells in vitro can be used as a model system for further functional assays.
Using MTT and flow cytometry assays, we found that knockdown of HMGB1 suppressed the proliferation and induced the apoptosis of BUC cells. However, it also increased the number of cells in the G0/G1 phase and reduced the number of cells in the S phase. These results indicate that HMGB1 is important for tumour growth. Chen RC et al.26 have reported that anti-HMGB1 neutralizing antibody could reduce the cell viability of HCCLM3 hepatocellular cancer cells, while this effect could be reversed by rhHMGB1, which indicated that HMGB1 might play an important role in cell proliferation. Chen J et al.27 constructed a lentivirus vector with HMGB1 shRNA to specifically suppress the expression of HMGB1 in ovarian cancer cells. They found that the knockdown of HMGB1 expression could decrease the expression of PCNA and Cyclin D1, which controls cell cycle transit, induces cell cycle G0/G1 arrest and inhibits cell proliferation. These results indicated that HMGB1 might have an effect on cell proliferation by regulating the expression of some cell cycle proteins. Gnanasekar M et al.28 demonstrated that knockdown of HMGB1 by shRNA plasmids in LNCaP prostate cancer cells could induce apoptosis via a caspase-3 dependent pathway. Volp K et al.29 found that HMGB1 overexpression increased NF-κB activity and led to c-IAP2 up-regulation in colon carcinoma, which could inhibit apoptosis via suppressing caspase-3 and caspase-9 activity. Brezniceanu M et al.30 demonstrated that HMGB1 protected mammalian cells against different death stimuli, including caspase-8, CD 95, ultraviolet radiation, TRAIL and Bax-induced apoptosis.
Metastasis is considered one of the primary causes of death in patients with malignant tumours. Previous studies have demonstrated that HMGB1 may be involved in cancer progression, including angiogenesis, mobility, inflammatory microenvironment, invasion and metastasis31,32. In this study, we concluded that the down-regulation of HMGB1 could suppress the migration and invasion of BUC cells, indicating that HMGB1 is involved in the metastasis of BUC. Yan W et al.33 found that HMGB1 played a pivotal role in hepatocellular carcinoma invasion and metastasis by activating caspase-1, with the subsequent production of multiple mediators, including IL-1β and IL-18. Lin L et al.16 demonstrated that HMGB1 binding to RAGE initiated the signalling pathway and activated ERK1/2, which promoted gene expression, protein synthesis and the migration and invasion of renal cell carcinoma cells. In lung cancer, Wang C et al.34 confirmed that HMGB1 binding to RAGE and TLR-4 was critical for the up-regulation of matrix metalloproteinase (MMP) –2, 9 in 95D cells, which could enhance the invasive potential of 95D cells.
NF-κB complexes are composed of homo- or heterodimers formed from the multi-gene family of p65, p50, p52, c-Rel and Rel B35. In most cell types, inactive NF-κB is predominantly sequestered in the cytoplasm bound to its inhibitors (IκBs). Upon activation, IκBs are degraded, thus allowing NF-κB translocation to the nucleus, where it acts as a sequence-specific DNA-binding transcription factor involved in the tumourigenesis of many cancers35,36. Of the five subunits of NF-κB, p65 has the highest transcriptional activity and has been widely studied37. As one of the subunits in the VEGF family, VEGF-C has been demonstrated to promote not only angiogenesis but also lymphangiogenesis in cancer progression by binding to its receptors, which makes it more important in tumour progression38. Furthermore, the VEGF family is a downstream target gene of NF-κB activation39,40. In this study, we demonstrated that knockdown of HMGB1 expression significantly inhibited the expression levels of NF-κB/p65 and VEGF-C, up-regulated IκBα expression and suppressed the nuclear translocation and DNA-binding activity of NF-κB/p65, which indicated that HMGB1 might regulate VEGF-C expression in the development of BUC via the NF-κB signalling pathway.
In conclusion, our results reveal that HMGB1 is overexpressed in BUC. The down-regulation of HMGB1 by shRNA plasmids can specifically and effectively inhibit the proliferation, migration and invasion of cells and induce apoptosis and G0/G1 arrest in BUC cells. The action of HMGB1 in the progression of BUC may regulate VEGF-C via the NF-κB signalling pathway. Because of its high efficiency and specificity in knockdown gene expression, silencing HMGB1 gene expression by specific shRNA plasmids could be a potential therapeutic strategy against BUC based on the targeting of the extracellular HMGB1 protein41. However, in vivo experiments are needed to elucidate the effects and risks of using shRNA-mediated RNAi in the HMGB1 gene. Meanwhile, the precise mechanism by which HMGB1 is involved in the progression and development of BUC needs further study.
Methods
Ethics statement
Approval for this study was obtained from the Institutional Research Ethics Committee of the Second Xiangya Hospital, Central South University. Prior consent was obtained from all patients for the use of tissue samples. All samples were harvested and made anonymous according to ethical and legal standards.
Tissue samples
Samples of BUC tissues and matched adjacent non-tumour tissues were obtained from BUC patients at our department. None of the patients had received either radiotherapy or chemotherapy before surgery and all the diagnoses were confirmed by histological examination.
Cell lines and cell culture
The 5637, BIU-87, T24 and SV-HUC-1 cell lines were purchased from Xiangya Medical College (Changsha, China). The cells were maintained in DMEM (Invitrogen, Carlsbad, USA) supplemented with 10% foetal bovine serum (FBS) at 37 °C in a humidified atmosphere containing 5% CO2.
Construction of shRNA plasmids and cell transfection
To minimize the off-target effects of RNAi, we constructed three potential sequences targeting the human HMGB1 gene and selected the shRNA sequence (5′-CACCGCGAAGAAACTGGGAGAGATGTTCAAGAGACATCTCTCCCAGTTTCTTCGCTTTTT TG-3′) for the subsequent experiments (Supplementary Table S2 and Figure S1). The selected HMGB1 shRNA sequence and a scrambled sequence (shNC) with no homology to any known human genes were synthesized and ligated into pGPU6/GFP/Neo plasmid vectors. All constructs were identified by sequence analysis.
The cells were seeded at a density of 5 × 104/well in a 6-well plate 24 h before transfection to achieve more than 70% confluence. For transfection, 7.5 μl of each plasmids and 5 μl of Lipofectamine 2000 (Invitrogen, Carlsbad, USA) were added to 250 μl Opti-MEM I (Invitrogen, Carlsbad, USA) and then mixed gently. The cells were transfected with the different mixtures according to the manufacturer’s instructions. The plate was incubated at 37 °C for 48–72 h until the transfection efficiency was more than 80% and was then used in the experiments described below.
Western blotting
Total protein extracts were prepared with the Total Protein Extraction Kit (ProMab Biotechnologies, Inc., CA, USA) and nuclear protein was extracted using the NucBusterTM Protein Extraction Kit (Merck Biosciences, Germany) in accordance with the manufacturer’s protocols. The protein concentration was confirmed using a BCA Kit (Beyotime, China). Equal amounts of protein were separated by 12% SDS-PAGE and then electrotransferred to polyvinylidene fluoride membranes (Millipore, Bedford, USA). The membranes were blocked with 5% non-fat dried milk in PBS (37 °C overnight) and incubated for 2 h at 37 °C with specific primary antibodies. The specific antibodies were Mouse HMGB1 antibody (Santa Cruz Biotechnology, Santa Cruz, USA; 1:500 dilution), Rabbit NF-κB/p65 antibody (Boster Biological Technology, Wuhan, China; 1:100 dilution), Rabbit IκBα antibody (Abcam, Cambridge, USA; 1:500 dilution), Rabbit VEGF-C antibody (Boster Biological Technology, Wuhan, China; 1:100 dilution) and Mouse GAPDH antibody (Santa Cruz Biotechnology, Santa Cruz, USA; 1:800 dilution). After washing, the membranes were incubated with horseradish peroxidase-conjugated anti-mouse (1:2000 dilution) and anti-rabbit (1:5000 dilution) antibodies for 1 h at room temperature and the blots were detected using Enhanced Chemiluminescence. The film was scanned and the band intensity was analysed with Gel-Pro Analyzer Software 4.0 (Media Cybernetics, Bethesda, MD) to calculate the integral optical density (IOD). The relative levels of target protein were evaluated using the ratio IOD/IOD GAPDH.
Quantitative real-time PCR
Total RNA was isolated from tissue samples and BUC cell lines using TRizol reagent (Invitrogen, Carlsbad, USA). The quality of the extracted RNA was assessed by the OD260/OD280 ratio. RNA was reverse transcribed to cDNA using the RevertAidTM H Minus First Strand cDNA Synthesis Kit (Fermentas, Vilnius, Lithuania) according to the manufacturer’s protocols. The qRT-PCR assay was carried out using an ABI PRISM 7900HT Sequence Detection System (Applied Biosystems, Foster City, USA). Each well (25 μl reaction volume) contained 12.5 μl of SYBR Green PCR Master Mix (Applied Biosystems, Foster City, USA), 1 μl of template, 1 μl of each primer and 10.5 μl ddH2O. The PCR reaction conditions included an initial denaturation for 5 min at 95 °C followed by 40 cycles each of denaturation for 20 sec at 94 °C, annealing for 20 sec at 61 °C, extension for 20 sec at 72 °C and a final extension for 5 min at 72 °C. All experiments were repeated in triplicate. The specificity of amplified products was determined by melting curve analysis. The relative expression levels of mRNA were calculated with the 2–ΔΔCt method42 and expressed as the RQ value normalized to β-actin. The primers used in the study were as follows: HMGB1 forward (5′-CTGTCCATTGGTGATGTTGC-3′) and reverse (5′-CTTCCTCCTCCTCCTCATCC -3′); NF-κB/p65 forward (5′-GCGAGAGGAGCACAGATACC-3′) and reverse (5′- CTGATAGCCTGCTCCAGGTC-3′); IκBα forward (5′-ACCTGGTGTCACTCCTGTTGA-3′) and reverse (5′-CTGCTGCTGTATCCGGGTG-3′); VEGF-C forward (5′-GGAAAGAAGTTCCACCACCA-3′) and reverse (5′-TTTGTTAGCATGGACCCACA-3′); and β-actin forward (5′-CATTAAGGAGAAGCTGTGCT-3′) and reverse (5′-GTTGAAGGTAGTTTCGTGGA-3′).
Cell proliferation assay (MTT)
The cells were seeded in 96-well plates at a density of 1 × 104/well and then transfected with shRNA plasmids and incubated at 37 °C in a humidified atmosphere. At 0 h, 24 h, 48 h, 72 h and 96 h post-transfection, 50 μl of MTT solution (5 mg/ml) was added to each well and incubated for 4 h at 37 °C until the formazan crystals formed. Then, the medium was removed and 150 μl DMSO was added to dissolve the formazan crystals. The OD (570 nm) was measured using a microplate reader (Bio-Rad, Hercules, CA, USA). All experiments were performed in triplicate. The cell growth curves were calculated as the mean OD values of each group at 5 different time points.
Cell apoptosis assay
Annexin V-FITC/propidium iodide was used to evaluate cell apoptosis. At 72 h post-transfection, the cells were trypsinized, washed, collected and re-suspended in 200 μl binding buffer and 5 μl Annexin V-FITC for 15 min in the dark at a concentration of 1 × 106 cell/ml. Then, 10 μl propidium iodide and 300 μl binding buffer were added to each sample. The cells were evaluated by flow cytometry (BD Biosciences, USA) and the results were analysed as previously described43.
Cell cycle assay
Seventy-two hours after transfection, the cells were collected, washed and centrifuged at 1000 rpm and then fixed with 75% cold ethanol at 4 °C overnight. After staining in 10 μg/ml propidium iodide and 10 mg/ml RNase A at room temperature for 30 min in the dark, the percentage of cells in each cell cycle phase was determined with Cell Modifit (Verity Software House, Topsham, USA) by a flow cytometer.
Cell migration and invasion assays in vitro
Cell invasion assays were performed in a 6-well transwell chamber (Corning, Cambridge, USA; pore size 8 μm) precoated with Matrigel (BD Biotechnology, USA). After 72 h of transfection, the cells were trypsinized, collected, washed and suspended in a serum-free medium containing 5% BSA at a cell density of 5 × 104 /ml. Then, the lower chamber was filled with 10% FBS and a 2 ml cell suspension was added to the upper chamber. After incubation with 5% CO2 for 24 h at 37 °C, cells on the lower surface were fixed with 95% ethanol, stained with hematoxylin-eosin and counted. Five fields were randomly selected to calculate the invading cells (×400). Similarly, the migration assays were performed as described above but without Matrigel.
Immunofluorescence
The cells were fixed in 4% formaldehyde and incubated in 0.3% Triton X-100 (Sigma, USA) for 1 h at room temperature. After incubation with a specific primary antibody for NF-κB/p65 at 4 °C overnight, the cells were treated with anti-rabbit IgG (H + L) antibody labelled with Alexa 488 (Molecular Probes, USA; 1:200 dilution) for 1 h at room temperature in the dark. Then, the cells were washed three times in PBS after each treatment described above. A confocal microscope (Olympus, Japan) was used to observe the cells.
Electrophoretic mobility shift assay
The nuclear extracts were prepared as previously described. After determining the concentrations of protein, the biotin end-labelled probe (sense sequence: 5′-biotin- AGTTGAGGGGACTTTCCCAGGC-3′) corresponding to the consensus of NF-κB/p65 was developed. EMSA was performed with a non-radioactive EMSA Kit (Viagene, USA) according to the manufacturer’s protocol. The biotin-labelled duplex DNA (0.5 μl) was then incubated with 3.0 μl nuclear protein extracts in a binding buffer. The reaction mixtures were separated by electrophoresis on 6.5% polyacrylamide gel in 0.5 × TBE, transferred to a nylon membrane, UV cross-linked and probed with streptavidin-HRP conjugate.
Immunohistochemistry
The IHC assay was performed as previously described44. Formalin-fixed and paraffin-embedded tissue sections (3 μm) were dewaxed in xylene, rehydrated with ethanol in a descending concentration and immersed in 3% hydrogen peroxide for 10 min to block endogenous peroxidase activity. Antigen retrieval was carried out by submerging sections into 0.01 mol/L citrate buffer (PH 6.0) in a microwave oven for 20 min. After incubation with 10% normal goat serum at room temperature for 15 min, the sections were incubated with rabbit polyclonal HMGB1 antibody (Abcam, Cambridge, USA; 1:200 dilution) overnight at 4 °C. Then, the horseradish peroxidase-conjugated secondary antibody was used to incubate the sections at room temperature for 30 min. Finally, the sections were stained with 3, 3-diaminobenzidine (DAB), counterstained with hematoxylin, dehydrated and mounted. The negative controls included incubation in PBS without primary antibody.
Each section was evaluated and scored according to the staining intensity and extent as described previously45. We evaluated the intensity of staining and grouped them into the following four categories: 0 (no staining), 1 (weak staining), 2 (moderate staining) and 3 (intense staining). Meanwhile, the extent of staining was ranked into the following five categories according to the percentage of positive staining areas: 0 (0%), 1 (1–25%), 2 (26–50%), 3 (51–75%) and 4 (76–100%). The index was obtained by multiplying the intensity and extent of the scores (0–7). The sections with an index of ≥3 were defined as HMGB1 high expression and <3 were classified as low expression.
Statistical analysis
All statistical analyses were performed using SPSS 17.0 (SPSS Inc., Chicago, USA). The quantitative data are shown as the mean ± standard deviation (SD). Comparisons between two groups were performed using the student’s t-test for continuous variables and the Fisher’s exact test for categorical variables. One-way ANOVA was used to assess the significant differences among multiple groups under various treatments. For all tests, P values of <0.05 were considered statistically significant.
Additional Information
How to cite this article: Huang, Z. et al. Down-regulation of HMGB1 expression by shRNA constructs inhibits the bioactivity of urothelial carcinoma cell lines via the NF-κB pathway. Sci. Rep. 5, 12807; doi: 10.1038/srep12807 (2015).
References
Babjuk, M. et al. EAU guidelines on non-muscle-invasive urothelial carcinoma of the bladder: update 2013. Eur. Urol. 64, 639–653, 10.1016/j.eururo.2013.06.003 (2013).
Siegel, R., Ma, J., Zou, Z. & Jemal, A. Cancer statistics, 2014. CA. Cancer J. Clin. 64, 9–29, 10.3322/caac.21208 (2014).
Parekh, D. J., Bochner, B. H. & Dalbagni, G. Superficial and muscle-invasive bladder cancer: principles of management for outcomes assessments. J. Clin. Oncol. 24, 5519–5527, 10.1200/jco.2006.08.5431 (2006).
Kaufman, D. S., Shipley, W. U. & Feldman, A. S. Bladder cancer. Lancet 374, 239–249, 10.1016/s0140-6736(09)60491-8 (2009).
Malkowicz, S. B. et al. Muscle-invasive urothelial carcinoma of the bladder. Urology 69, 3–16, 10.1016/j.urology.2006.10.040 (2007).
Czura, C. J., Wang, H. & Tracey, K. J. Dual roles for HMGB1: DNA binding and cytokine. J. Endotoxin Res. 7, 315–321 (2001).
Bianchi, M. E. & Beltrame, M. Upwardly mobile proteins. Workshop: the role of HMG proteins in chromatin structure, gene expression and neoplasia. EMBO Rep. 1, 109–114, 10.1093/embo-reports/kvd030 (2000).
Muller, S. et al. New EMBO members’ review: the double life of HMGB1 chromatin protein: architectural factor and extracellular signal. EMBO J. 20, 4337–4340, 10.1093/emboj/20.16.4337 (2001).
Bianchi, M. E. & Agresti, A. HMG proteins: dynamic players in gene regulation and differentiation. Curr. Opin. Genet. Dev. 15, 496–506, 10.1016/j.gde.2005.08.007 (2005).
Brickman, J. M., Adam, M. & Ptashne, M. Interactions between an HMG-1 protein and members of the Rel family. Proc. Natl. Acad. Sci. USA 96, 10679–10683 (1999).
Imamura, T. et al. Interaction with p53 enhances binding of cisplatin-modified DNA by high mobility group 1 protein. J. Biol. Chem. 276, 7534–7540, 10.1074/jbc.M008143200 (2001).
Onate, S. A. et al. The DNA-bending protein HMG-1 enhances progesterone receptor binding to its target DNA sequences. Mol. Cell. Biol. 14, 3376–3391 (1994).
Ellerman, J. E. et al. Masquerader: high mobility group box-1 and cancer. Clin. Cancer Res. 13, 2836–2848, 10.1158/1078-0432.CCR-06-1953 (2007).
Sims, G. P., Rowe, D. C., Rietdijk, S. T., Herbst, R. & Coyle, A. J. HMGB1 and RAGE in inflammation and cancer. Annu. Rev. Immunol. 28, 367–388, 10.1146/annurev.immunol.021908.132603 (2010).
Ishiguro, H. et al. Receptor for advanced glycation end products (RAGE) and its ligand, amphoterin are overexpressed and associated with prostate cancer development. Prostate 64, 92–100, 10.1002/pros.20219 (2005).
Lin, L., Zhong, K., Sun, Z., Wu, G. & Ding, G. Receptor for advanced glycation end products (RAGE) partially mediates HMGB1-ERKs activation in clear cell renal cell carcinoma. J. Cancer Res. Clin. Oncol. 138, 11–22, 10.1007/s00432-011-1067-0 (2012).
Yang, G. L. et al. Increased expression of HMGB1 is associated with poor prognosis in human bladder cancer. J. Surg. Oncol. 106, 57–61, 10.1002/jso.23040 (2012).
Jiang, W., Wang, Z., Li, X., Fan, X. & Duan, Y. High-mobility group box 1 is associated with clinicopathologic features in patients with hepatocellular carcinoma. Pathol. Oncol. Res. 18, 293–298, 10.1007/s12253-011-9442-3 (2012).
Akaike, H. et al. Expression of high mobility group box chromosomal protein-1 (HMGB-1) in gastric cancer. Anticancer Res. 27, 449–457 (2007).
Yao, X. et al. Overexpression of high-mobility group box 1 correlates with tumor progression and poor prognosis in human colorectal carcinoma. J. Cancer Res. Clin. Oncol. 136, 677–684, 10.1007/s00432-009-0706-1 (2010).
Liu, P. L. et al. High-mobility group box 1-mediated matrix metalloproteinase-9 expression in non-small cell lung cancer contributes to tumor cell invasiveness. Am. J. Respir. Cell Mol. Biol. 43, 530–538, 10.1165/rcmb.2009-0269OC (2010).
Guo, Z. S., Liu, Z., Bartlett, D. L., Tang, D. & Lotze, M. T. Life after death: targeting high mobility group box 1 in emergent cancer therapies. Am J Cancer Res 3, 1–20 (2013).
Kepp, O. et al. Immunogenic cell death modalities and their impact on cancer treatment. Apoptosis 14, 364–375, 10.1007/s10495-008-0303-9 (2009).
Rovere-Querini, P. et al. HMGB1 is an endogenous immune adjuvant released by necrotic cells. EMBO Rep. 5, 825–830, 10.1038/sj.embor.7400205 (2004).
Scaffidi, P., Misteli, T. & Bianchi, M. E. Release of chromatin protein HMGB1 by necrotic cells triggers inflammation. Nature 418, 191–195, 10.1038/nature00858 (2002).
Chen, R. C. et al. The role of HMGB1-RAGE axis in migration and invasion of hepatocellular carcinoma cell lines. Mol. Cell. Biochem. 390, 271–280, 10.1007/s11010-014-1978-6 (2014).
Chen, J., Liu, X., Zhang, J. & Zhao, Y. Targeting HMGB1 inhibits ovarian cancer growth and metastasis by lentivirus-mediated RNA interference. J. Cell. Physiol. 227, 3629–3638, 10.1002/jcp.24069 (2012).
Gnanasekar, M., Thirugnanam, S. & Ramaswamy, K. Short hairpin RNA (shRNA) constructs targeting high mobility group box-1 (HMGB1) expression leads to inhibition of prostate cancer cell survival and apoptosis. Int. J. Oncol. 34, 425–431 (2009).
Völp, K. et al. Increased expression of high mobility group box 1 (HMGB1) is associated with an elevated level of the antiapoptotic c-IAP2 protein in human colon carcinomas. Gut 55, 234–242 (2006).
Brezniceanu, M.-L. et al. HMGB1 inhibits cell death in yeast and mammalian cells and is abundantly expressed in human breast carcinoma. The FASEB journal 17, 1295–1297 (2003).
Tang, D., Kang, R., Zeh, H. J., 3rd & Lotze, M. T. High-mobility group box 1 and cancer. Biochim. Biophys. Acta 1799, 131–140, 10.1016/j.bbagrm.2009.11.014 (2010).
Naglova, H. & Bucova, M. HMGB1 and its physiological and pathological roles. Bratisl. Lek. Listy 113, 163–171 (2012).
Yan, W. et al. High-mobility group box 1 activates caspase-1 and promotes hepatocellular carcinoma invasiveness and metastases. Hepatology 55, 1863–1875, 10.1002/hep.25572 (2012).
Wang, C. et al. HMGB1 was a pivotal synergistic effecor for CpG oligonucleotide to enhance the progression of human lung cancer cells. Cancer Biol. Ther. 13, 727–736, 10.4161/cbt.20555 (2012).
Campbell, K. J., Rocha, S. & Perkins, N. D. Active repression of antiapoptotic gene expression by RelA(p65) NF-kappa B. Mol. Cell 13, 853–865 (2004).
Pacifico, F. & Leonardi, A. NF-kappaB in solid tumors. Biochem. Pharmacol. 72, 1142–1152, 10.1016/j.bcp.2006.07.032 (2006).
Hoesel, B. & Schmid, J. A. The complexity of NF-kappaB signaling in inflammation and cancer. Mol. Cancer 12, 86, 10.1186/1476-4598-12-86 (2013).
Su, J. L. et al. The role of the VEGF-C/VEGFR-3 axis in cancer progression. Br. J. Cancer 96, 541–545, 10.1038/sj.bjc.6603487 (2007).
Basseres, D. S. & Baldwin, A. S. Nuclear factor-kappaB and inhibitor of kappaB kinase pathways in oncogenic initiation and progression. Oncogene 25, 6817–6830, 10.1038/sj.onc.1209942 (2006).
Ferrara, N. & Davis-Smyth, T. The biology of vascular endothelial growth factor. Endocr. Rev. 18, 4–25, 10.1210/edrv.18.1.0287 (1997).
Musumeci, D., Roviello, G. N. & Montesarchio, D. An overview on HMGB1 inhibitors as potential therapeutic agents in HMGB1-related pathologies. Pharmacol. Ther. 141, 347–357, 10.1016/j.pharmthera.2013.11.001 (2014).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408, 10.1006/meth.2001.1262 (2001).
Chen, Q. et al. Apo2L/TRAIL and Bcl-2-related proteins regulate type I interferon-induced apoptosis in multiple myeloma. Blood 98, 2183–2192 (2001).
Huang, Z. et al. A chronic obstructive pulmonary disease negatively influences the prognosis of patients with bladder urothelial carcinoma via hypoxia inducible factor-1alpha. Int. J. Clin. Exp. Med. 7, 3344–3353 (2014).
Wu, D., Ding, Y., Wang, S., Zhang, Q. & Liu, L. Increased expression of high mobility group box 1 (HMGB1) is associated with progression and poor prognosis in human nasopharyngeal carcinoma. J. Pathol. 216, 167–175, 10.1002/path.2391 (2008).
Author information
Authors and Affiliations
Contributions
Z.H. and X.Z. conceived the idea and concept. Z.H., Z.Z. and L.Z. designed the experiments. Z.H., R.X., L.Z. and X.W. analysed the data. Z.H., Z.W. and S.H. performed the experiments. Z.H. and X.Z. wrote the manuscript and prepared the figures. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Huang, Z., Zhong, Z., Zhang, L. et al. Down-regulation of HMGB1 expression by shRNA constructs inhibits the bioactivity of urothelial carcinoma cell lines via the NF-κB pathway. Sci Rep 5, 12807 (2015). https://doi.org/10.1038/srep12807
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep12807
This article is cited by
-
Toll-like receptors 2 and 4 stress signaling and sodium-glucose cotransporter-2 in kidney disease
Molecular and Cellular Biochemistry (2023)
-
Autophagy-associated HMGB-1 as a novel potential circulating non-invasive diagnostic marker for detection of Urothelial Carcinoma of Bladder
Molecular and Cellular Biochemistry (2022)
-
Extracellular HMGB1 interacts with RAGE and promotes chemoresistance in acute leukemia cells
Cancer Cell International (2021)
-
miR-449a inhibits cell proliferation, migration, and inflammation by regulating high-mobility group box protein 1 and forms a mutual inhibition loop with Yin Yang 1 in rheumatoid arthritis fibroblast-like synoviocytes
Arthritis Research & Therapy (2019)
-
The evaluation of monocyte lymphocyte ratio as a preoperative predictor in urothelial malignancies: a pooled analysis based on comparative studies
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.