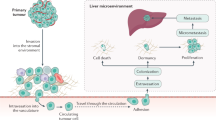
Abstract
We present a model of hepatic colorectal metastases which represents monoclonal cell lines double-labeled by luciferase and tdTomato. These cells form liver metastasis in varying numbers and patterns similar to those observed in patients. Using in vivo and ex vivo luminescent and fluorescent imaging we determine the growth kinetics and clonogenic frequency of tumor cells colonizing liver. Molecular profiling detected stable expressional differences between clones consistent with their phenotypes. The data indicate that clinically relevant phenotypes of liver metastases can be modeled in vivo.
Similar content being viewed by others
Introduction
Patients with colorectal cancer often present with liver metastases, which frequently results in a fatal outcome. Between 5 to 20% of patients with limited numbers of hepatic metastases and slow rates of progression/recurrence are successfully treated with local treatment approaches, with or without systemic therapy1,2. Little is known about liver metastases heterogeneity and animal models that reflect heterogeneity are still absent.
The tumor burden of hepatic metastases has been previously quantified by weight or volume of whole liver or the macroscopic findings of liver tumors3,4,5. Alternatively, models of experimental liver metastasis based on luminescently-labeled tumor cells have been previously reported6,7. However there are no reports about selection of tumor clones containing double luminescent and fluorescence tags and demonstrating different abilities to colonize and grow in the liver microenvironment. Successful development of such model can significantly assist in the understanding of mechanisms of development of liver metastases.
Here we present a new approach to model liver metastases, which includes generation of tumor clones with different liver colonization efficiencies and growth properties. This model represents monoclonal populations of tumor cells recapitulating oligometastatic, potentially curable disease and widespread metastatic disease. We employ dual labeling of clones with luciferase and tdTomato to provide different ex vivo and in vivo imaging modalities, including diffuse luminescent Imaging tomography (DLIT) for detection and resolution of individual tumor colonies during the time course of liver colonization. Our data indicate that observed phenotypes of liver metastases are based on the combination of two parameters- the efficiency of tumor colonization and doubling time (Td) following adaptation of tumor cells to the liver microenvironment.
Results
Generation of a panel of monoclonal cell lines with different metastatic ability
To generate a panel of labeled monoclonal HCT116 cell lines, HCT116 cell lines were double-labeled with Luc2 and tdTomato proteins8. tdTomato-positive cells were collected using flow cytometry and 16 monoclonal double-labeled HCT116 cell lines (HCT116-L2T) were generated. In vitro fluorescence and luminescence of HCT116-L2T were quantified using increasing amounts of cells and both fluorescence and luminescence were in good correlation with cell numbers (Pearson’s R = 0.99; p < 0.0001) (Fig. 1).
The correlation between the number of cells and luminescence or fluorescence in vitro.
(a) The luminescent image of different number of cells triplicated in 96-well plate. (b) The correlation between the number of cells and luminescence (R = 0.99, p < 0.0001). (c) The fluorescent image of different number of cells triplicated in 96-well plate. Intensities were acquired as radiant efficiency with 535 nm excitation and 580 nm emission. (d) The correlation between the number of cells and fluorescence (R = 0.99, p < 0.0001).
Sixteen individual clones were injected into spleen followed by splenectomy (Supplementary Video 1, 2, 3, 4, 5). Bioluminescence of mice and ex vivo fluorescence of harvested livers were quantified at 2 weeks after spleen injection (Fig. 2a,b). Quantification of these data demonstrated that clones had different abilities to colonize and grow in the liver as was determined by both luminescent measurements in vivo and fluorescent ex vivo imaging (see Fig. 2c,d). Clones #21 and #17 which showed high propensity to colonize liver had overall luminescence equal to 2.7 × 1010 ± 2.5 × 1010 and 2.0 × 1010 ± 7.7 × 108 p/sec/cm2/sr (steradian). Clones #12 and #18 with low colonization had overall luminescence equal to 2.4 × 107 ± 3.0 × 107 and 6.6 × 108 ± 4.1 × 108 p/sec/cm2/sr, respectively. We correlated in vivo luminescence and ex vivo fluorescence for each clone (Pearson’s R = 0.89; p < 0.0001) (Fig. 2e). We further selected clones #12 and #18 as “oligometastatic” clones, potentially recapitulating limited metastatic disease and designated them as O1 and O2, respectively. We also selected clones #21 and #17 as further model for widespread dissemination, also referred to as polymetastatic disease, in liver and named them P1 and P2, respectively (Fig. 2c,d).
Bioluminescence and ex vivo fluorescence of liver in 16 clones at 2 weeks after spleen injection.
(a) Representative bioluminescent images. (b) Representative ex vivo fluorescent images of harvested livers. Intensities were acquired as radiant efficiency with 535 nm excitation and 580 nm emission. (c) The distribution of bioluminescent intensities of each clone. (d) The distribution of ex vivo fluorescent intensities of each clone. (e) The correlation between in vivo bioluminescence and ex vivo fluorescence (R = 0.89, p < 0.0001).
To further characterize these metastatic phenotypes clones O1, O2, P1 and P2 were intrasplenically injected in corresponding groups of mice and bioluminescence was measured weekly. Ex vivo fluorescence of livers were quantified at 4 weeks after spleen injection. The bioluminescence was higher in P1 and P2 mice (6.7 × 106 ± 4.7 × 106 and 3.6 × 106 ± 2.5 × 106 p/sec/cm2/sr) than in O1 and O2 mice (2.0 × 105 ± 1.3 × 105 and 1.7 × 105 ± 9.1 × 104 p/sec/cm2/sr) 3 weeks after spleen injection (Fig. 3a,b). The ex vivo fluorescence in P1 and P2 was higher than in O1 and O2; this was consistent with macroscopic findings (Fig. 3c,d).
Bioluminescence and ex vivo fluorescence of liver in selected clones P1, P2, O1 and O2.
(a) Representative bioluminescent images, (b) Bioluminescence weekly measured from 1 to 3 weeks in P1 and P2, 1 to 4 weeks in O1 and O2. (c) Representative ex vivo fluorescent images of livers. Intensities were acquired as radiant efficiency with 535 nm excitation and 580 nm emission. (d) Macroscopic liver images (arrows indicate white small tumors). P1 and P2; polymetastatic clones, O1 and O2; oligometastatic clones.
Quantification of colonization ability and growth properties of selected clones
For further evaluation of metastatic colonization we measured macroscopically defined numbers and sizes of tumors and ex vivo fluorescence of individual colonies (Fig. 4). The total number of tumors was higher in P1 than P2, O1 and O2, whereas the size of tumors in P2 was larger than in P1, O1 and O2 (p < 0.001) (Fig. 5a,b). However total tumor volume in liver was larger in P1 and P2 than O1 and O2 (Fig. 5c). The colonizing ability of the tumor cells is expressed as the colonizing fraction (Fc) which is calculated as the number of tumors in liver divided by the number of cells injected. Assuming that the number of tumors in liver corresponds to the number of cells which have an ability to colonize liver, P1 had more ability to colonize liver than the other clones (Fc = 8.2 × 10−5 ± 1.1 × 10−5, 6.0 × 10−6 ± 2.3 × 10−6 in P1 and O1, respectively, p < 0.001) (Fig. 5d). P2 had similar ability to colonize the liver as clones O1 and O2, but the size of tumor was larger than the other clones. We further correlated total tumor volumes and in vivo luminescence or ex vivo fluorescence for each investigated clone. As is shown in Supplementary Fig. S1, these parameters were in good correlation with each other (Pearson’s R = 0.99; p < 0.0001).
Quantification of fluorescent intensities of individual liver colonies.
Left images indicate representative macroscopic findings and right images indicate fluorescent intensities in each clone. The fluorescent intensities were acquired as radiant efficiency with 535 nm excitation and 580 nm emission. P1 and P2; polymetastatic clones, O1 and O2; oligometastatic clones.
Quantitative estimation of colonizing ability and growth kinetics.
(a) The macroscopic total number of tumors, (b) macroscopic size of individual tumors, (c) calculated total tumor volume in liver, (d) Fc; colonizing fraction of tumor colony forming cells, (e) total cell number per colony, (f) doubling time and (g) number of cell divisions of each clone. P1 and P2; polymetastatic clones, O1 and O2; oligometastatic clones.
To characterize the basic properties of tumor colony forming cells, we calculated the total cell number per colony, doubling time (Td) and number of cell divisions by the calibration from correlation between fluorescent intensity and number of cells (Fig. 1 and Fig. 5e–g). P1 which had many small tumors showed larger colonizing fraction (Fc) and Td similar to O1 and O2 (p < 0.05). P2 which had large tumors showed similar Fc but shorter Td than O1 and O2 (p < 0.05). The data indicated that there are two parameters which define the characteristics of metastatic potential- colonizing fraction (Fc) and doubling time (Td) of tumor colony forming cells. Consistent with in vivo data, P2 showed faster growth as compared with the other clones in vitro (Supplementary Fig. S2).
Diffuse luminescent Imaging tomography (DLIT) provided the 3D distribution of metastatic tumors in liver (Supplementary Video 6). Using this technique it was possible to detect and measure the bioluminescence of individual hepatic metastatic colonies in the time course of metastases development therefore tracing spatial-temporal dynamic behavior of metastases (Fig. 6).
Representative images of quantification of bioluminescence of individual tumor using diffuse luminescent imaging tomography (DLIT) in P1- polymetastatic clone.
(a) 3 dimensional overhead view, (b) coronal section, (c) sagittal section, (d) transaxial section, (e) macroscopic image of liver, (f) ex vivo fluorescent image of liver.
Molecular profiling of selected monoclones
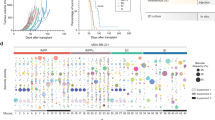
To detect the molecular differences between poly- and oligometastatic clones we used expressional profiling of corresponding cell lines in vitro with Illumina HT 12v4 bead arrays. We found 756 differentially expressed genes (DEGs) in P1 as compared to O1 and O2 and 461 differentially expressed genes in P2 as compared to O1 and O2 (Fig. 7a and Supplementary Table S1, S2). To estimate the functional significance of the detected DEGs we used Ingenuity Pathway Analysis (IPA) as previously described before9,10,11. As is shown in Fig. 7b, P1 is enriched by constitutively expressed genes associated with inflammatory responses. Interestingly, many of these genes are presented by an interferon-related gene signature (IRDS) previously described by us and others in the context of tumorigenicity, metastases development and radio-/chemo-resistance (Fig. 7b,c)9,12,13,14,15,16,17. P2 was enriched by genes, associated with regulation of growth and survival – a regulatory network of a subset of these genes is presented in Fig. 7d. Taken together, these data provide evidence that polymetastatic clones demonstrate molecular differences from oligometastatic clones. As well, two polymetastatic but phenotypically distinct clones (P1 and P2) express regulatory networks of genes which are consistent with their phenotypic properties.
Gene expression differences between polymetastatic and oligometastatic clones.
(a) Hierarchical clustering of 1,225 genes differentially expressed (fold-change ≥1.5 and FDR ≤5%) in P1 and P2 as compared to O clones. (b) Functional annotation of differentially expressed genes in P1 (left) and P2 (right) vs. O clones. Shown are the top 5 functions for each comparison. Values represent the -log10 p-value. (c, d) Ingenuity Network analysis of P1 (c) and P2 (d) specific gene patterns. Red color indicates over-expression and green color indicates suppression. Solid line indicates activation; dashed line indicates deactivation. P1 and P2; polymetastatic clones, O clones; O1 and O2, oligometastatic clones.
Discussion
Our data indicate at least three potential scenarios of colorectal metastases development in liver which can be recapitulated in our experimental system. The first scenario presented by clone P1 is determined by high ability of the clone to colonize liver (Fc = 8.2 × 10−5 ± 1.1 × 10−5) but is associated with relatively low growth rate (Td = 26.5 ± 0.2 hours). A combination of these parameters leads to increased number of small metastatic tumors. A second case is P2, which had lower Fc (1.1 × 10−5 ± 1.8 × 10−5) and shorter Td (22.7 ± 0.3 hours) resulted in relatively small number of large secondary tumors. And a third case is presented by clones O1 and O2, which had lower Fc and longer Td resulting in less number of small tumors, which might be clinically considered as oligometastatic liver disease (Fig. 8)18,19. Indeed, clinical observations are consistent with these models. Fong and co-authors provided a comprehensive review of more than 1000 cases of hepatic resections of colorectal metastases to liver1. Recently it was updated to include 1600 patients with maximal follow-up time 17.4 years20. In both reviews, based on comprehensive statistical analysis, authors refer to the size of liver metastases (parameter related to growth potential) and their numbers (parameter related to colonization ability) as to independent risk factors for distant metastasis, which is consistent with P2 and P1 properties. Our recent observations of patients, treated by stereotactic body radiotherapy (SBRT) or with surgically resected lung metastases points both to the number and rate of progression as key factors determining overall and disease-free outcome18,19. While these findings are an oversimplification of the problem, the complexity of the metastatic processes is far from a comprehensive description21,22,23. However, colonizing and growth abilities seem to be among the most critical factors determining the development of metastatic clones in distant sites24,25. Experimental models, which can capture differences of metastatic clones in these abilities, may provide useful tools both for detection of molecular properties discriminating oligo- and polymetastatic pathways (Fig. 7) and for validation of potential regulatory molecules identified in high-throughput clinical screenings11,18.
Three different scenarios of metastases development.
First, P1 had higher Fc, longer Td and lower nav resulted in increased number of small tumors. Second, P2 had lower Fc, shorter Td and higher nav resulted in limited number of larger tumors. Thirdly, O1 and O2 had lower Fc, longer Td and lower nav resulted in less number of small tumors. P1 and P2; polymetastatic clones, O1 and O2; oligometastatic clones. Fc; colonizing fraction of tumor forming cells, Td; doubling time (hours), nav; average number of cells per colony.
An increasing number of studies have demonstrated the optical imaging as a modality for preclinical in vivo examination of tumor detection, growth and response to different treatments26,27,28,29,30. Because of its efficiency, the clinical potential for the in vivo optical imaging is becoming apparent31,32. Compared to the previously reported models using cells single-labeled with luciferase6,7, our model demonstrates the advanced evaluation of tumor characterization using ex vivo optical imaging with fluorescence. Higher sensitivity of the bioluminescence provides the detection and quantification of small burden of metastases and enables quantitative longitudinal tumor growth without sacrifice of the mice. However, ex vivo fluorescent optical imaging enables more detailed quantification of tumor distribution, has higher resolution and allows visualizing interaction of the tumor cells with surrounding tissues. In addition, it provides a unique opportunity to quantify growth parameters of metastatic colonies on the cellular level without interference of stromal tissues and has the potential to quantitatively separate tumor cells from stromal cells for their separate investigation.
In summary, we describe a new technique to model limited and widespread liver metastatic disease and to quantify the tumor burden of hepatic metastases. This technique can also provide 3D imaging of metastases distribution in liver. With quantitative estimation of tumor burden, this technique may be helpful in further investigations of the biological mechanisms of hepatic tumors and potential approaches to their treatments.
Study approval
All studies performed on mice were approved by the IACUC of the University of Chicago.
Methods
Generation of clonal double-labeled HCT 116 cell line
The stably double-labeled HCT116 cell lines were generated with Luc2 and tdTomato genes using lentiviral-based gene delivery8. The Luc2-tdTomato plasmid and HCT116 cell line were obtained from Dr. Geoffrey Greene at the University of Chicago. The cell lines were maintained in DMEM (Life Technologies Corporation, NY, USA) with 10% fetal bovine serum,100 U/mL penicillin and 100 mg/mL streptomycin. All cell lines were maintained in culture with 5% CO2 at 37 degrees Celsius. The tdTomato-positive cells were collected using FACS Calibur Flow Cytometer (BD Immunocytometry Systems, CA, USA). Collected tdTomato-positive cells were diluted to 1 cell per 200 μl and plated at 1 cell per a well in 96-well plate. We generated 16 clones of HCT116 cell lines double-labeled with Luc2 and tdTomato (HCT116-L2T).
In vitro quantification of fluorescent and bioluminescent signals
The HCT116-L2T cells were plated at a density of 0, 103, 104, 2 × 104, 3 × 104, 5 × 104, 7 × 104, 9 × 104 and 105 cells per well in 96-well plates. Triplicates were performed in each density of cells. After 5 hours incubation, fluorescent and luminescent intensities were quantified with IVIS 200 (Xenogen, MA, USA) imaging system. Firefly D-luciferin potassium salt (GoldBio Technology, MO, USA, 150 μg/mL per well) was added just before luminescent assay
An animal model of hepatic colorectal metastases
All animal procedures were carried out in accordance with the approved guidelines. The procedures were approved by the Institutional Animal Care and Use Committee of the University of Chicago (Protocol # 72213-09).
Mice were anesthetized with 2% isoflurane in oxygen. Spleen was exposed through a 8 mm left flank incision (Supplementary Video 1). Six - eight weeks old female athymic nude mice (Harlan, WI, USA) were splenically injected with 2.0 × 106 cells per 100 μL phosphate buffered saline into liver (n = 3, in each group) (Supplementary Video 2). Five minutes post-injection, splenectomy was performed to avoid carcinomatous peritonitis and residual growth in spleen (Supplementary Video 3). Bioluminescent intensities were measured weekly using the IVIS 200 (Xenogen, MA, USA) imaging system after intra-peritoneal injection of 150 μg firefly D-luciferin potassium salt (GoldBio Technology, MO, USA). Data were analyzed using LivingImage 4.0 Software (Caliper Life Sciences, MA, USA). Two or 4 weeks after injections, livers were harvested and ex vivo fluorescent intensities of liver tumors were quantified as radiant efficiency. Sixteen clones were tested to select candidates for oligo- and polymetastatic phenotypes. Diffuse luminescent imaging tomography (DLIT) was performed for evaluation of tumor burden and distribution using real-time 3D reconstruction of bioluminescent signals which provide bioluminescent intensities of individual tumor.
Quantitative estimation of colonizing ability and growth kinetics
The numbers and sizes of liver tumors were macroscopically measured. The total tumor volumes were calculated by assuming it to be a sphere. The fluorescent intensities of 5 representative colonies in each liver were quantified using the IVIS 200 (Xenogen, MA, USA) imaging system. The fraction of tumor colony forming cells (Fc) was calculated as the number of tumors in liver divided by the number of cells splenically injected. The total cell number per colony, doubling time (Td) and number of cell divisions were calibrated from the correlation between fluorescent intensity and number of cells in vitro.
In vitro growth curve
HCT116-L2T clones were plated at a density of 3,000 cells per well in 96-well plates. Triplicates were performed per cell lines. Cell growth was evaluated at 0, 24, 48, 72 and 96 hours after plating. Cells were incubated with CellTiter-Blue (Promega, WI, USA) for 3 hours and fluorescent intensities were quantified per the manufacture’s instructions. Signals were normalized to the intensities at time zero.
Gene expression profiling and analysis
Selected HCT116 L2T clones (P1, P2, O1 and O2) were collected in cell lysis buffer and RNA was isolated using the TRIzol Reagent (Ambion, Austin, TX). 100 ng of RNA was labeled per manufacturer’s instructions and profiled in duplicate using the Illumina Human HT12v4 array (Illumina, San Diego CA). Background subtraction and quantile normalization was performed across arrays using Illumina Beadstudio software. Log-transformed gene expression was compared using Significance Analysis of Microarrays (SAM) for Excel (Stanford University, CA) with a False Discovery Rate (FDR) of 5% and a fold-change threshold of greater than or equal to 1.5 to identify differentially expressed genes33.
Ingenuity Pathway Analysis (IPA, Redwood City CA) was used to identify over-represented functions and pathways. Over-representation of gene sets in a canonical pathway was calculated using hypergeometric testing with an alpha value of 0.05. Significantly enriched pathways were manually distributed into specific functional groups.
Statistics
Data were analyzed using JMP 10 software (SAS Institute, NC, USA). Data were represented as the mean ± standard deviation for all figure panels in which error bars were shown. Pearson’s product-moment correlation coefficients were used to assess associations between parameters. The p values were assessed using 2-tailed Student t tests and p < 0.05 was considered statistically significant.
Additional Information
How to cite this article: Oshima, G. et al. Imaging of tumor clones with differential liver colonization. Sci. Rep. 5, 10946; doi: 10.1038/srep10946 (2015).
References
Fong, Y., Fortner, J., Sun, R. L., Brennan, M. F. & Blumgart, L. H. Clinical score for predicting recurrence after hepatic resection for metastatic colorectal cancer: analysis of 1001 consecutive cases. Ann. Surg. 230, 309–321 (1999).
Pawlik, T. M. et al. Effect of surgical margin status on survival and site of recurrence after hepatic resection for colorectal metastases. Ann. Surg. 241, 715-714 (2005).
Ishizu, K. et al. Development and characterization of a model of liver metastasis using human colon cancer HCT-116 cells. Biol. Pharm. Bull. 30, 1779–1783 (2007).
Kopper, L., Vanhanh, T. & Lapis, K. Experimental-Model for Liver Metastasis Formation Using Lewis Lung-Tumor. J. Cancer Res. Clin. Oncol. 103, 31–38 (1982).
Warren, R. S., Yuan, H., Matli, M. R., Gillett, N. A. & Ferrara, N. Regulation by vascular endothelial growth factor of human colon cancer tumorigenesis in a mouse model of experimental liver metastasis. J. Clin. Invest. 95, 1789–1797 (1995).
Smakman, N., Martens, A., Kranenburg, O. & Borel Rinkes, I. H. Validation of bioluminescence imaging of colorectal liver metastases in the mouse. J. Surg. Res. 122, 225–230 (2004).
Rajendran, S. et al. Murine bioluminescent hepatic tumour model. JoVE, 41, e1977 (2010).
Liu, H. et al. Cancer stem cells from human breast tumors are involved in spontaneous metastases in orthotopic mouse models. Proc. Natl. Acad. Sci. USA 107, 18115–18120 (2010).
Widau, R. C. et al. RIG-I-like receptor LGP2 protects tumor cells from ionizing radiation. Proc. Natl. Acad. Sci. USA 111, E484–E491 (2014).
Pitroda, S. P. et al. DNA repair pathway gene expression score correlates with repair proficiency and tumor sensitivity to chemotherapy. Sci. Transl. Med. 6, 229ra42 (2014).
Uppal, A. et al. 14q32-encoded microRNAs mediate an oligometastatic phenotype. Oncotarget 6, 3540–3552 (2015).
Khodarev, N. N. et al. Signal transducer and activator of transcription 1 regulates both cytotoxic and prosurvival functions in tumor cells. Cancer Res. 67, 9214–9220 (2007).
Weichselbaum, R. R. et al. An interferon-related gene signature for DNA damage resistance is a predictive marker for chemotherapy and radiation for breast cancer. Proc. Natl. Acad. Sci. USA 105, 18490–18495 (2008).
Khodarev, N. N. et al. STAT1 pathway mediates amplification of metastatic potential and resistance to therapy. PLoS One 4, e5821 (2009).
Cheon, H. et al. IFNbeta-dependent increases in STAT1, STAT2 and IRF9 mediate resistance to viruses and DNA damage. EMBO J. 32, 2751–2763 (2013).
Luszczek, W., Cheriyath, V., Mekhail, T. M. & Borden, E. C. Combinations of DNA methyltransferase and histone deacetylase inhibitors induce DNA damage in small cell lung cancer cells: correlation of resistance with IFN-stimulated gene expression. Mol. Cancer Ther. 9, 2309–2321 (2010).
Khodarev, N. N., Roizman, B. & Weichselbaum, R. R. Molecular pathways: interferon/stat1 pathway: role in the tumor resistance to genotoxic stress and aggressive growth. Clin. Cancer Res. 18, 3015–3021 (2012).
Lussier, Y. A. et al. MicroRNA expression characterizes oligometastasis(es). PLoS One 6, e28650 (2011).
Lussier, Y. A. et al. Oligo- and polymetastatic progression in lung metastasis(es) patients is associated with specific microRNAs. PLoS One 7, e50141 (2012).
House, M. G. et al. Survival after hepatic resection for metastatic colorectal cancer: trends in outcomes for 1,600 patients during two decades at a single institution. J. Am. Coll. Surg. 210, 744–755 (2010).
Chiang, A. C. & Massague, J. Molecular basis of metastasis. N. Engl. J. Med. 359, 2814–2823 (2008).
McAllister, S. S. & Weinberg, R. A. The tumour-induced systemic environment as a critical regulator of cancer progression and metastasis. Nat. Cell Biol. 16, 717–727 (2014).
Quail, D. F. & Joyce, J. A. Microenvironmental regulation of tumor progression and metastasis. Nat. Med. 19, 1423–1437 (2013).
Calon, A. et al. Dependency of colorectal cancer on a TGF-beta-driven program in stromal cells for metastasis initiation. Cancer Cell 22, 571–584 (2012).
Vanharanta, S. & Massague, J. Origins of metastatic traits. Cancer Cell 24, 410–421 (2013).
Cronin, M. et al. High resolution in vivo bioluminescent imaging for the study of bacterial tumour targeting. PLoS One 7, e30940 (2012).
Gutpell, K., McGirr, R. & Hoffman, L. Molecular imaging to target transplanted muscle progenitor cells. JoVE, 73, e50119 (2013).
Baban, C. K. et al. Bioluminescent bacterial imaging in vivo. JoVE, 69, e4318 (2012).
Fujii, T., Kamiya, M. & Urano, Y. In vivo imaging of intraperitoneally disseminated tumors in model mice by using activatable fluorescent small-molecular probes for activity of cathepsins. Bioconjug. Chem. 25, 1838–1846 (2014).
Sjollema, J. et al. The potential for bio-optical imaging of biomaterial-associated infection in vivo. Biomaterials 31, 1984–1995 (2010).
Bu, L., Shen, B. & Cheng, Z. Fluorescent imaging of cancerous tissues for targeted surgery. Adv. Drug Delivery Rev. 76, 21–38 (2014).
Byrne, W. L. et al. Use of optical imaging to progress novel therapeutics to the clinic. J. Control. Release 172, 523–534 (2013).
Tusher, V. G., Tibshirani, R. & Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl. Acad. Sci. USA 98, 5116–5121 (2001).
Acknowledgements
We thank Dr. Geoffrey L. Greene (University of Chicago) for the Luc2-tdTomato plasmid and HCT116 cell line, Mr. Ani Solanki (Animal Resource Center) for mice management and Dr. Lara Leoni for assistance with DLIT. This work was supported by the Virginia and D.K. Ludwig Fund for Cancer Research, Lung Cancer Research Foundation (LCRF) and Prostate Cancer Foundation (PCF). This work was supported in part by The Virginia and D.K. Ludwig Fund for Cancer Research, Lung Cancer Research Foundation (LCRF) and Prostate Cancer Foundation (PCF). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
N.K., R.W., S.H. and M.P. conceived the project and designed the experiments; G.O., S.W., A.U., M.S., S.P., J.O. and X.H. performed the experiments and analyzed the data; G.O., S.P., M.P., S.H., N.K. and R.W. interpreted data and wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Oshima, G., Wightman, S., Uppal, A. et al. Imaging of tumor clones with differential liver colonization. Sci Rep 5, 10946 (2015). https://doi.org/10.1038/srep10946
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep10946
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.