Abstract
Targeted drug delivery is important in cancer therapy to decrease the systemic toxicity resulting from nonspecific drug distribution and to enhance drug delivery efficiency. We have developed an aptamer-based DNA dendritic nanostructure as a multifunctional vehicle for targeted cancer cell imaging and drug delivery. The multifunctional DNA dendrimer is constructed from functional Y-shaped building blocks with predesigned base-pairing hybridization including fluorophores, targeting DNA aptamers and intercalated anticancer drugs. With controllable step-by-step self-assembly, the programmable DNA dendrimer has several appealing features, including facile modular design, excellent biostability and biocompatibility, high selectivity, strong binding affinity, good cell internalization efficiency and high drug loading capacity. Due to the unique structural features of DNA dendrimers, multiple copies of aptamers can be incorporated into each dendrimer, generating a multivalent aptamer-tethered nanostructure with enhanced binding affinity. A model chemotherapeutic anticancer drug, doxorubicin, was delivered via these aptamer-based DNA dendrimers and exerted a potent toxicity for target cancer cells (human T cell acute lymphoblastic leukemia cell line) with low side effects for the non-target cells (human Burkitt’s lymphoma cell line). This controllable aptamer-based DNA dendrimer is a promising candidate for biomedical applications.
Similar content being viewed by others
Introduction
Chemotherapy is widely used for cancer treatment using traditional small molecule drugs. However, the chemotherapeutic drugs may lead to serious toxic side effects and inefficient delivery to tumor tissues due to poor water solubility, nonspecific distribution and systemic toxicity1,2. Thus, development of desirable therapeutics which can penetrate biological barriers, distinguish normal and diseased tissues and intelligently respond to the tumor microenvironment for on-demand drug release, is an urgent need3. Nanotechnology applied in medicine, known as nanomedicine, has become a promising approach for efficient cancer therapeutics. This technology uses precisely engineered materials at the scale of 1–100 nm to develop novel therapeutic and diagnostic modalities4,5. For successful nanomedicine, nanoparticle sizes and surface properties must be controlled and targeting ligands must be incorporated for site-specific on-demand release of pharmacologically active agents at therapeutically optimal rates and dose regimens6.
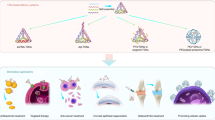
Various types of nanoparticles, including self-assembled polymers and metal nanoparticles, have been used as potential diagnostic and therapeutic agents, representing a promising breakthrough7,8,9. However, most inorganic and organic nanomaterials suffer from multiple drawbacks, such as limited biocompatibility and inability to engineer spatially addressable surfaces that can be utilized for multifunctional activities. Alternatively, DNA-based nanostructures are promising materials for biomedical applications10, because of their excellent biocompatibility, specific base pairing interactions, automated synthesis and programmability11. For example, uniform sized DNA tetrahedrons equipped with immune-stimulatory CpG oligonucleotides or small interface RNA have shown enhanced intracellular immunoregulation or gene delivery with excellent biostability and biocompatibility12,13 Likewise, triangular DNA origami loaded with the anticancer drug doxorubicin exhibited improved anti-tumor efficacy and lower systemic toxicity in vivo compared to anti-cancer drugs14.
Among the diverse DNA nanostructures, DNA dendrimers have attracted increasing interest in the past decade due to their monodispersity, excellent stability, globular shape and highly branched and porous structures15. Mintzer and coworkers used DNA dendrimers for delivery of functional molecules, such as the CpG motif16, into cells with excellent intracellular uptake via passive delivery. Tan and coworkers used Y-shaped monomers and DNA linkers to form DNA hydrogel for targeted gene therapy17. Although passive delivery is useful for cancers with leaky vasculatures, it is not suitable for other types of cancers, such as leukemia, that require specific targeting. In this regard, incorporation of a ligand which targets a particular cell receptor to facilitate receptor-mediated endocytosis could provide enhanced versatility for the treatment of a variety of diseases18.
Aptamers are single-stranded DNA or RNA oligonucleotides screened by a process called Systematic Evolution of Ligands by Exponential Enrichment (SELEX)19. Aptamers have excellent advantages as targeting ligands, such as high target affinity, excellent specificity and low immunogenicity. Aptamers can recognize a large range of targeting molecules, including organic and inorganic small molecules, proteins, cells and even tissues. Furthermore, the easy synthesis and functionalization of aptamers make it possible to design various aptamer chimeras, such as aptamer-dye, aptamer-drug, aptamer-biomolecule and aptamer-nanomaterial conjugates, to generate diversified molecular probes in sensing, imaging and targeted therapy20,21. Most importantly, aptamers can be easily designed and integrated into 3D nucleic acid structures without any need of chemical modification. Combined with the dendritic DNA structures and designed hybridization, it is possible to embed a variety of ligands and functional reagents to generate multifunctional nano-platforms.
In this work, we designed an aptamer-based DNA dendrimer as a multifunctional nanostructure for biomedical applications. In our proof-of-principle study, we have successfully incorporated functional domains, including aptamers, fluorophores and drug loading sites, into DNA dendrimers to achieve selective cancer cell recognition, bioimaging and targeted anticancer drug delivery. The aptamer used in this study, sgc8, which selectively recognizes cell membrane protein PTK722, was artificially designed to spontaneously hybridize with building blocks of the outer shell of the DNA dendrimer. PTK7-overexpressed cell lines, human T cell acute lymphoblastic leukemia cell line (CCRF-CEM) and cervical cancer HeLa cell line, were chosen as targets and human Burkitt’s lymphoma cell line (Ramos) with low expression of PTK7 was used as a control. By combining the advantages of DNA dendrimers and the sgc8 aptamer, these nanostructures can selectively distinguish and be internalized by the target cells. Because of their abundant double-stranded sequences, DNA dendrimers have high capacity to load intercalating therapeutic drugs. In this work, doxorubicin (Dox), a model chemotherapeutic anticancer drug, was loaded into DNA dendrimer for evaluating the targeted therapeutic effect. This aptamer-based self-assembled DNA dendrimer provides the advantages of facile modular design and assembly, high programmability and biocompatibility, as well as selective recognition. With controllable functional groups, these DNA dendrimers have remarkable potential for application in multifunctional bioimaging and drug delivery.
Results and Discussions
Design, preparation and characterization of multifunctional DNA Dendrimer
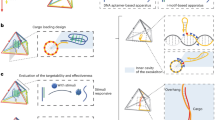
Multifunctional DNA dendrimers were prepared from three-armed Y-shaped DNA monomers using an enzyme-free, step-by-step base-pairing assembly strategy23. Y-shaped DNA monomers contained 13-base sticky-end segments, which further hybridized with other Y-DNAs as each generation was added. The Y-DNA called Y0 was assembled from the hybridization of three single strands, Y0a, Y0b and Y0c, by slowly cooling from 95 oC to 4 oC in 91 minutes. The others, Y1, Y2 and Y3, were prepared according to the same procedure from their three respective single-stranded sequences. The as prepared Y-DNAs were then characterized by native-PAGE gel electrophoresis (Fig. S1) and ready to use without purification. Different generations of DNA dendrimer (Gn) were prepared from Y-shaped building blocks by layer-by-layer assembly. The first generation G0 was the initial building block Y0. For Gn, it was synthesized by mixing Gn-1 and Yn with the ratio of 1: 3 × 2n−1 (n > 1) at room temperature for 1 hour. Based on the G3 structure, aptamers (sgc8) with sticky-end pairing with Y3 were added in the solution to form the aptamer-based DNA self-assembled nanostructure G3-sgc8, as shown in Fig. 1. Each generation was characterized by agarose gel electrophoresis and dynamic light scattering (Fig. 2) and directly used without purification. Only one major band for each generation was detected by gel electrophoresis suggesting the formation of highly pure DNA dendrimer. The band mobility decreased with increasing generation, suggesting the success of self-assembly (Fig. 2a). DLS measurement showed the average diameter of G1, G2, G3 and G3-sgc8 to be 13.7, 21.0, 24.3 and 43.8 nm, respectively (Fig. 2b). These data verified that higher generations of DNA dendrimer were heavier in mass and larger in size. To confirm the structure of DNA dendrimer, atomic force microscopy (AFM) was also used to characterize G3-sgc8. The measured diameter was correlated to the result of dynamic light scattering measurement. And it showed G3-sgc8 as a spherical structure which indicated the formative assembly was a DNA dendrimer (Fig. S2).
Working principle of aptamer-based dendritic DNA nanostructure. The final generation Dox-G3-sgc8 is designed with various functional groups, including fluorophores, targeting ligands and anticancer drugs to endow the DNA dendrimer with the capability of cancer cell recognition, imaging and drug delivery.
Selective Recognition of Target Cancer Cells by FITC labeled G3-sgc8
After confirming the successful formation of DNA dendrimers, the cancer cell recognition property of G3-sgc8 was investigated. With aptamer sgc8 coating on the surface, G3-sgc8 is expected to specifically recognize target cancer cell line CEM. In this study, an organic dye, FITC, was labeled on Y3 to stain G3-sgc8 with green fluorescence to investigate specific binding. Fig. 3a,b shows the flow cytometric comparison of target (CCRF-CEM) cells and control negative (Ramos) cells after incubation with random DNA library (lib), individual aptamer (sgc8), dendrimer decorated with random sequences (G3-rs) and aptamer-embedded dendrimer (G3-sgc8). Weak fluorescence was observed with random DNA and random DNA dendrimer for both CEM and Ramos cells, indicating low non-specific binding. There was a noticeable change in the fluorescence signal observed for CEM cells treated with free sgc8 because of the specific binding between aptamer and target cell line, while CEM cells treated with G3-sgc8 showed significantly higher fluorescence intensity than aptamer stained cells. No significant change in fluorescence intensity was observed for Ramos cells with either free sgc8 or G3-sgc8, further confirming the specific recognition of G3-sgc8 to target CEM cells. The binding affinities of free sgc8 and G3-sgc8 were determined by incubating CCRF-CEM cells with varying concentration of aptamer probes on ice for 30 min. As demonstrated in Fig. 3c, G3-sgc8 showed a significantly enhanced binding affinity (Kd = 0.07 ± 0.02 nM) compared to individual sgc8 aptamer (Kd = 2.95 ± 1.47 nM). G3-sgc8 showed at least a 40-fold enhancement in fluorescence intensity compared to the free aptamer with CCRF-CEM cells. This high binding affinity of G3-sgc8 is indicative of multivalent-mediated enhancement of binding affinity, because multiple aptamers on the outer-shell dendrimer can each recognize the receptors on the cell surface.
Selective cancer cell recognition by multifunctional dendrimer. Flow cytometry results demonstrated the selective recognition of G3-sgc8 (FITC-incorporated on G3) to target CCRF-CEM cells (a) but not to control Ramos cells (b). (c) Binding affinity of G3-sgc8 and fluorescein-labeled free sgc8 (inside figure) to CCRF-CEM cells. G3-sgc8 showed about 40-fold higher binding affinity than single aptamer.
Bioimaging is a visual method to investigate specific recognition and cellular trafficking of DNA assembled-nanoparticles. A strong green fluorescence signal was observed by the DeltaVision Elite cell imaging system after incubating G3-sgc8 with CEM at 37 °C for 2 h (Fig. 4). In contrast, a weak fluorescence signal was observed for G3-rs and Ramos cells. Previous work suggested that sgc8 aptamer enters the targeted cell line, such as CEM and HeLa, through receptor PTK7-mediated endocytosis. A colocalization assay was performed in this work to track the final destination of G3-sgc8 in live HeLa cells. Most of the green fluorescence signal from G3-sgc8 overlapped with the red fluorescence signal generated from a lysotracker (a lysosome marker, Fig. S3), indicating that aptamer-based DNA dendrimer can recognize and internalize into target cancer cells through receptor-mediated endocytosis instead of passive delivery, an important property for use as a multifunctional nano-platform for efficient delivery of imaging and therapeutic reagents into the cytoplasm.
Selective cytotoxicity of anticancer drug-loaded G3-sgc8
Benefitting from the large number of hybridized DNA base pairs, the DNA dendrimer is spatially well equipped for loading chemical anticancer drugs such as doxorubicin (Dox). Dox can preferentially insert between G-C pairs, resulting in the quenching of Dox fluorescence due to Förster resonance energy transfer. We then determined Dox intercalation by monitoring Dox fluorescence intensity changes to evaluate the amount of Dox loaded into the G3-sgc8. Dox fluorescence was dramatically quenched by G3-sgc8 with a molar ratio of 666/1, indicating a high loading capacity of 10 nM G3-sgc8 with ∼6.66 μM Dox, as shown in Fig. S6. We next investigated the release kinetics of Dox loaded into G3-sgc8 by detecting the intensity of Dox fluorescence. The G3-sgc8 (2 nM) with a drug payload of 666 nM released less than 7% of the DOX after 60 hours in a physiological environment (pH = 7.4 PBS buffer, Fig. S7). Therefore, the Dox payload was sufficiently stable to prevent drug leakage during the circulation in blood.
To investigate the drug transport into cell and drug release, the uptake and distribution of Dox-G3-sgc8-cy5 were studied with target HeLa cells using microscope imaging. We incubated Dox-G3-sgc8-cy5 with HeLa cells for 30 min (Fig. S4) and 2.5 hours (Fig. 5), then washed 3 times with PBS buffer, followed by staining with Hoechst and lysotracker. As shown in Fig. 5, G3-sgc8-cy5 was still predominantly localized in lysotracker-labeled acidic organelles after 2.5 hours, while most of the inserted Dox had escaped from the dendrimer and appeared colocalized with Hoechst in the nucleus. We believe that the transported Dox-G3-sgc8-cy5 first entered cells via receptor-mediated endocytosis and then escaped from the endosome, finally localizing in the lysosome. The acidic lysosome environment and enzyme catalysis facilitated the rapid release of the loaded anticancer drug, which traveled to the nucleus.
Subcellular distribution of Dox (red) loaded G3-sgc8-cy5 (cyan). Hoechest (blue) and Lysotracker Green (green) were used to stain the cell nuclei and acidic organelles. Cells were imaged using a 60x oil-immersion objective. The merged images were used to confirm that drug had released and escaped into the nucleus within 2.5 hours, but the nanostructure still stayed in the lysosome. Scar bar: 20 μm.
The in vitro cytotoxicity of Dox-G3-sgc8 and free Dox were evaluated by the 3-(4, 5-dimethylthiazol-2-yl)-5-(3-carboxymethoxyphenyl)-2-(4-sulfophenyl)-2H-tetrazolium (MTS) assay. In target CEM cells, Dox-G3-sgc8 showed the similar inhibition of cell proliferation as free Dox (Fig. 6a), while in non-target cell Ramos, Dox-G3-sgc8 showed significantly less inhibition of cell proliferation than free Dox. In contrast, for free drug, there was no drug selectivity in either target or non-target cancer cells. These results indicate that aptamer-induced targeted internalization enhanced site-specific drug delivery and suggest that our aptamer-based DNA dendrimer is an excellent vehicle for targeted cancer therapy.
MTS assay results showing selective cytotoxicity of Dox delivered by DNA dendrimer in target CEM cells (a) but much less in nontarget Ramos cells (b), in contrast to nonselective cytotoxicity of free Dox in both target cells and nontarget cells. The selective cytotoxicity of Dox delivered by DNA dendrimer indicates the capability of DNA dendrimers for targeted drug delivery.
Conclusions
We have demonstrated an aptamer-based DNA dendrimer as an anticancer drug carrier specific recognition behavior and selective cytotoxicity to target cancer cell lines. This designable DNA nanostructure can be equipped with different functional groups using oligonucleotide base pairing without complicated chemical modification. For biomedical applications, the DNA dendrimer has several remarkable features: (1) Facile design and preparation. All the DNA assembly mentioned in this paper occurs at room temperature by mixing different building blocks in a fixed ratio and the size can be controlled by adding more or fewer generations. (2) Multifunctionality. Different functional domains, including imaging dyes, targeting ligands and inserted anticancer drugs, are integrated in a single platform to fulfill diverse demands. (3) Good biocompatibility. Our DNA dendrimer shows very little toxicity without drug cargo (Fig. S7), while it is very toxic to target cells when carrying anticancer drugs. (4) Excellent stability. The stability of dendritic DNA nanostructure has been examined in many previous publications. For example, there was no structural change after direct treatment with endonuclease DNase I (1 U/mL, a considerably higher concentration than would be found in living cells)24. The DNA dendrimer also has excellent and stable loading capacity for anticancer drugs, as shown by a 60-hour test in physiology environment (Fig. S6). With these remarkable features, together with the attractive properties of cancer cell specific recognition, imaging and drug delivery, the multifunctional DNA dendrimer offers a promising new modality for selective multimodal cancer theranostics.
Methods
Materials
All oligonucleotides were purchased from Sangon (Shanghai, China) and used without further purification. Lysotracker and Hoechst were purchased from Life Technology (Beijing, China). RPMI-1640 and DMEM Medium were obtained from HyClone (Beijing, China). Fetal bovine serum (FBS) was obtained from Gibco through Life Technology (Beijing, China). Doxorubicin was purchased from Huafeng United Technology Co., Ltd. (Beijing, China). CellTiter 96® AQueous Non-Radioactive Cell Proliferation Assay (MTS) was purchased from Promega, Madison, WI, USA.
Cell culture
CCRF-CEM (CCL-119, T-cell line, human acute lymphoblastic leukemia), HeLa (Human cervical cancer cell line) and Ramos (CRL-1596, B-cell line, human Burkitt’s lymphoma) cells were obtained from American Type Culture Collection. CCRF-CEM and Ramos were cultured in RPMI 1640 and HeLa was cultured in DMEM, which contained 10% fetal bovine serum (FBS, 10%) and penicillin–streptomycin (100 IU/mL) at 37 oC in a humid atmosphere with 5% CO2.
Synthesis of DNA dendrimer
Y-shaped DNA was assembled according to the method as reported23. For preparation of Y-shaped DNA (e.g. Y0), three strands (Y0a, Y0b, Y0c) were mixed in the phosphate buffer (50 mM phosphate, 100 mM Na+, pH = 8.0) and the final concentration of each strand was 20 uM. Then, the mixture was heat to 95 oC for 2 min and cooled to 4 oC at a rate of 1 oC/min. To prepare DNA dendrimers (Gn), Y0 (G0) and Y1 were mixed at a 1:3 molar ratio and the mixture was kept at room temperature for 2 h and at 4 oC for 2 h to prepare G1. Gn was prepared using the same method by mixing Gn-1 and Yn in a ratio of 1:32n−1.
Characterization of DNA dendrimer
The Y-shaped DNAs used in the experiment were characterized by 10% native PAGE at 75 V in 1 × TBE buffer for 2 h on ice. They were all directly used for further assembly without purification. Gn was characterized by 1% agarose gel at 55 V in 1 × TAE buffer for 60 min on ice. In DLS experiments, described specifically for Gn, 2 μL solution of Gn (50 mM, pH 8.0 phosphate buffer with 100 mM NaCl) was diluted to 200 μL by the same phosphate buffer and characterized by DLS to give the radius of Gn. AFM imaging was performed on an Agilent 5500ILM SPM (Agilent Technologies, Inc.) equipped with a N9520A scanner with a scan size of 10 μm in x−y and 2.08 μm in z directions. Gold coated silicon probes (NT-MDT) with a nominal force constant of 0.01−0.08 N/m (CSG/11 Au) and 5.5−22.5 N/m (NSG/10) were used for force measurements and imaging, respectively. In a typical experiment, 2 μL sample solution was dropped onto the cleaved mica, left standing for 1 min and then removed by aspiration. Then 10 μL water was dropped onto the surface and removed using a stream of nitrogen.
Flow Cytometric Analysis
For the fluorescence analysis, Y3a and Y3b were labelled with FITC to generate a fluorescent dendrimer. To demonstrate the targeting capabilities of aptamer-conjugated dendrimer toward specific cells, fluorescence measurements were performed using a flow cytometer (BD Biosciences FACSVerseTM cytometer), according to the following procedure: Approximately 1 × 105 cells of each type were suspended with 200 μL BB buffer (1 × PBS, 0.55 mM MgCl2, pH = 7.4) in individual test tubes. To the cell samples, 2 μL of the G3-aptamer solution (Captamer = 200 nM) was added and the mixture was incubated at room temperature for 30 min. After incubation, the cells were washed twice by centrifugation with 0.5 mL buffer and resuspended in 0.2 mL buffer. The fluorescence was determined by counting 10,000 events. The fluorescein-labeled G3-rs was used as a negative control.
The binding affinities of free and dendrimer-conjugated aptamer probes were determined by incubating CCRF-CEM cells (1 × 105) at room temperature for 30 min in the dark with varying concentrations of aptamer probes in a 200 μL volume of buffer. Cells were then washed twice with 0.5 mL buffer, suspended in 0.2 mL buffer and subjected to flow cytometric analysis within 30 min. All of the experiments for the binding assay were repeated two times. The equilibrium dissociation constants (Kd) of the aptamer-cell interaction were obtained by fitting the dependence of fluorescence intensity of specific binding on the concentration of the aptamers to the equation Y = BmaxX/ (Kd + X), using SigmaPlot (Jandel, San Rafael, CA).
Cell Imaging
For cell imaging, the treatment steps for cell incubation were the same as described in Flow Cytometric Analysis. Ten microliters of cell suspension bound with G3-rs or G3-sgc8 was dropped on a thin glass slide placed above an objective on the confocal microscope. HeLa cell line was cultured in the confocal culture plate. G3-sgc8 was directly added to FBS-free medium and incubated for 2 h. Then washed twice with buffer and 200 μL PBS buffer was added to keep cells alive. For colocalization with lysosome, Lysotracker with a standard concentration was added and incubating in 37 oC for 30 min. The Lysotracker was removed by washing and ready-to-use Hoechst solution was added (1 ug/mL in PBS buffer), followed by incubation at room temperature for 5 min. A 60x oil-immersion objective on a Nikon inverted microscope linked to a DeltaVisionTM deconvolution-imaging system (Applied Precision, Seattle, Washington) and a Leica TCS SP5 microscope with 100x oil-immersion objective (Leica Microsystems CMS GmbH, Germany) were used for imaging.
Drug loading into G3-aptamer
To load Dox into G3-aptamer, Dox and G3-aptamer were mixed with special ratio and stayed at room temperature to allow saturation of drug loading. To evaluate the amount of Dox (doxorubicin) loaded into the G3-aptamer, we measured Dox intercalation by monitoring Dox fluorescence intensity changes. When a fixed concentration of Dox was incubated with an increasing molar ratio of the G3-aptamer, a sequential decrease was found in the fluorescence intensity of Dox (Fig. S2), due to Förster resonance energy transfer between Dox molecules when intercalated into the DNA duplex. According to Fig. S2, 1 μL G3-aptamer can tolerate 0.5 nmol Dox. To prevent leakage, we chose a Dox/G3-aptamer of 333 as a better concentration for drug loading.
Cytotoxicity test
The cytotoxicities of G3-aptamer, free drug, or drug-G3-aptamer complexes were evaluated using CellTiter 96® AQueous Non-Radioactive Cell Proliferation Assay (MTS). Cells (1 × 104 CEM or Ramos cells/well) were treated with G3-aptamer, free drug, or drug-G3-aptamer complexes in FBS-free medium. After incubation for 2 h in a cell culture incubator, supernatant medium was removed and fresh medium (10% FBS, 100 μL) was added for further cell growth (24 h). Then MTS reagent (20 μL) was added to each well and incubated for 1–2 h at 37 oC. The absorbance (490 nm) was recorded using a model 680 BioRad plate-reader (Bio-Rad, Hertfordshire, UK).
Additional Information
How to cite this article: Zhang, H. et al. A Controllable Aptamer-Based Self-Assembled DNA Dendrimer for High Affinity Targeting, Bioimaging and Drug Delivery. Sci. Rep. 5, 10099; doi: 10.1038/srep10099 (2015).
References
Peer, D. et al. Nanocarriers as an emerging platform for cancer therapy. Nat Nanotechnol. 2, 751–760, doi: DOI 10.1038/nnano.2007.387 (2007).
Li, X., Huang, J., Tibbits, G. F. & Li, P. C. Real-time monitoring of intracellular calcium dynamic mobilization of a single cardiomyocyte in a microfluidic chip pertaining to drug discovery. Electrophoresis 28, 4723–4733, doi: 10.1002/elps.200700312 (2007).
Sun, T. et al. Engineered Nanoparticles for Drug Delivery in Cancer Therapy. Angew. Chem. Int. Ed. Engl., 53, 12320–12364, doi: 10.1002/anie.201403036 (2014).
Farokhzad, O. C. & Langer, R. Nanomedicine: Developing smarter therapeutic and diagnostic modalities. Adv. Drug Deliver Rev. 58, 1456–1459, doi: 10.1016/j.addr.2006.09.011 (2006).
Tian, J. et al. Cell-specific and pH-activatable rubyrin-loaded nanoparticles for highly selective near-infrared photodynamic therapy against cancer. J. Am. Chem. Soc. 135, 18850–18858, doi:10.1021/ja408286k (2013).
Yezhelyev, M. V. et al. Emerging use of nanoparticles in diagnosis and treatment of breast cancer. Lancet. Oncol. 7, 657–667, doi: 10.1016/S1470-2045(06)70793-8 (2006).
Petros, R. A. & DeSimone, J. M. Strategies in the design of nanoparticles for therapeutic applications. Nat. Rev. Drug Discov. 9, 615–627, doi: 10.1038/nrd2591 (2010).
Davis, M. E., Chen, Z. G. & Shin, D. M. Nanoparticle therapeutics: an emerging treatment modality for cancer. Nat. Rev. Drug Discov. 7, 771–782, doi: 10.1038/nrd2614 (2008).
Tian, J. et al. A multifunctional nanomicelle for real-time targeted imaging and precise near-infrared cancer therapy. Angew. Chem. Int. Ed. Engl. 53, 9544–9549, doi: 10.1002/anie.201405490 (2014).
Pinheiro, A. V., Han, D., Shih, W. M. & Yan, H. Challenges and opportunities for structural DNA nanotechnology. Nat. Nanotechnol. 6, 763–772, doi: 10.1038/nnano.2011.187 (2011).
Chhabra, R., Sharma, J., Liu, Y., Rinker, S. & Yan, H. DNA self-assembly for nanomedicine. Adv. Drug Deliver Rev. 62, 617–625, doi: 10.1016/j.addr.2010.03.005 (2010).
Li, J. et al. Self-assembled multivalent DNA nanostructures for noninvasive intracellular delivery of immunostimulatory CpG oligonucleotides. Acs Nano 5, 8783–8789, doi: 10.1021/nn202774x (2011).
Lee, H. et al. Molecularly self-assembled nucleic acid nanoparticles for targeted in vivo siRNA delivery. Nat. Nanotechnol. 7, 389–393, doi: 10.1038/nnano.2012.73 (2012).
Zhang, Q. et al. DNA Origami as an In Vivo Drug Delivery Vehicle for Cancer Therapy. Acs Nano 8, 6633–6643, doi: 10.1021/Nn502058j (2014).
Mintzer, M. A. & Grinstaff, M. W. Biomedical applications of dendrimers: a tutorial. Chem. Soc. Rev. 40, 173–190, doi: 10.1039/B901839p (2011).
Rattanakiat, S., Nishikawa, M., Funabashi, H., Luo, D. & Takakura, Y. The assembly of a short linear natural cytosine-phosphate-guanine DNA into dendritic structures and its effect on immunostimulatory activity. Biomaterials 30, 5701–5706, doi: 10.1016/j.biomaterials.2009.06.053 (2009).
Li, J. et al. Self-assembly of DNA Nanohydrogels with Controllable Size and Stimuli-Responsive Property for Targeted Gene Regulation Therapy. J. Am. Chem. Soc. 137, 1412–1415, doi: 10.1021/ja512293f (2015).
Grabow, W. W. & Jaeger, L. siRNA DELIVERY Loaded-up microsponges. Nat. Mater. 11, 268–269, doi: 10.1038/Nmat3286 (2012).
Tuerk, C. & Gold, L. Systematic Evolution of Ligands by Exponential Enrichment - Rna Ligands to Bacteriophage-T4 DNA-Polymerase. Science 249, 505–510, doi: 10.1126/science.2200121 (1990).
Vinkenborg, J. L., Karnowski, N. & Famulok, M. Aptamers for allosteric regulation. Nat. Chem. Biol. 7, 519–527, doi: 10.1038/Nchembio.609 (2011).
Yang, L. et al. Aptamer-conjugated nanomaterials and their applications. Adv. Drug Deliver Rev. 63, 1361–1370, doi: 10.1016/j.addr.2011.10.002 (2011).
Shangguan, D. et al. Aptamers evolved from live cells as effective molecular probes for cancer study. P. Natl. Acad. Sci. USA 103, 11838–11843, doi: 10.1073/pnas.0602615103 (2006).
Zhou, T. et al. pH-Responsive Size-Tunable Self-Assembled DNA Dendrimers. Angew. Chem. Int. Ed. Engl. 51, 11271–11274, doi: 10.1002/anie.201205862 (2012).
Meng, H. M. et al. DNA dendrimer: an efficient nanocarrier of functional nucleic acids for intracellular molecular sensing. Acs Nano 8, 6171–6181, doi: 10.1021/nn5015962 (2014).
Acknowledgements
We thank the National Science Foundation for Distinguished Young Scholars of China (21325522), National Science Foundation for Excellent Youth Scholars of China (21422506), the National Science Foundation of China (91313302, 21205100, 21275122, 21435004) the National Basic Research Program of China (2013CB933703) and Innovative Research Team in University by the MOE of China (IRT13036) for their financial support.
Author information
Authors and Affiliations
Contributions
H.Z. designed, performed research and wrote the mauscript, Y.M., Y.X. prepared the Fig. 1–3, Y. A., Y. H. prepared the Fig. 4–5 and the supporting information, Z.Z. and C.J.Y. conceived the project, supervised the project and revised the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Zhang, H., Ma, Y., Xie, Y. et al. A Controllable Aptamer-Based Self-Assembled DNA Dendrimer for High Affinity Targeting, Bioimaging and Drug Delivery. Sci Rep 5, 10099 (2015). https://doi.org/10.1038/srep10099
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep10099
This article is cited by
-
Advancing Plant Resilience Against Microplastics and Metals Through Nanotechnology
BioNanoScience (2024)
-
Self-assembling biomolecules for biosensor applications
Biomaterials Research (2023)
-
Advanced cancer targeting using aptamer functionalized nanocarriers for site-specific cargo delivery
Biomaterials Research (2023)
-
Preparation, applications, and challenges of functional DNA nanomaterials
Nano Research (2023)
-
Aptamer-Functionalized DNA Nanostructures for Biological Applications
Topics in Current Chemistry (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.