Abstract
Reactivation of wild-type p53 (wt-p53) function is an attractive therapeutic approach to p53-defective cancers. An ideal p53-based gene therapy should restore wt-p53 production and reduces mutant p53 transcripts simultaneously. In this study, we described an alternative strategy named as trans-splicing that repaired mutant p53 transcripts in hepatocellular carcinoma (HCC) cells. The plasmids which encoded a pre-trans-splicing molecule (PTM) targeting intron 6 of p53 were constructed and then transfected into HCC cells carrying p53 mutation. Phenotypic changes of HCC cells induced by p53-PTM were analyzed through cell cycle, cell apoptosis and the expression of p53 downstream target genes. Spliceosome mediated RNA trans-splicing (SMaRT) reduced mutant p53 transcripts and produced functional wt-p53 protein after the delivery of p53-PTM plasmids, which resulted in phenotype correction of HCC cells. In tumor xenografts established by p53-mutated HCC cells, adenovirus encoding p53-PTM induced cell cycle arrest and apoptosis and then blocked the growth of tumors in mice. Collectively, our results demonstrated for the first time that mutant p53 transcripts were functionally corrected in p53-defective HCC cells and xenografts using trans-splicing, which indicated the feasibility of using trans-splicing to repair p53 mutation in p53-defective cancers.
Similar content being viewed by others
Introduction
Liver cancer in men is the fifth most frequently diagnosed cancer worldwide and the second leading cause of cancer death1. An estimated 748,300 new liver cancer cases and 695,900 cancer deaths occurred worldwide in 20081. Half of these cases and deaths were estimated to occur in China1. The incidence of liver cancer is increasing worldwide because of the dissemination of hepatitis B and C virus infection. There were an estimated 746,000 deaths from liver cancer in the world in 2012 (http://publications.cancerresearchuk.org/downloads/Product/CS_INFOG_WORLD_MORT.PDF). Epidemiologic and experimental studies have demonstrated that the risk of developing liver cancer was attributable to environmental and genetic factors2,3,4. A multistep process involving accumulation of multiple genetic alterations can occur in malignant transformation of the hepatocytes3,5. Tumor protein 53 (TP53, well-known as p53) responds to diverse cellular stresses to regulate the expression of target genes, thereby inducing cell cycle arrest, apoptosis, senescence, DNA repair, or changes in metabolism6,7,8. It is now widely acknowledged that p53 mutation is one of the most common genetic events in human cancer9. p53 mutations are present in >50% of all human tumors, including hepatocellular carcinoma (HCC)10. In HCC, p53 mutation is one of the most common genetic alternations5,11,12. Systematic review and meta-analysis revealed that alterations of p53 were associated with a poor outcome for HCC patients13. p53 mutations exhibit a variety of distinct local structural changes and alter thermal stability, which results in the loss of its ability to bind to p53 response elements and trans-activate downstream genes. In addition, mutant p53 proteins are endowed with oncogenic properties such as dominant negative activity and gain-of-function effects, which contributes to the promotion of tumorigenesis9,14,15.
Some strategies for targeting p53 have been developed due to the pivotal role of p53 in tumorigenesis16,17,18. Several gene therapeutic strategies have been employed in the attempt to restore p53 function in p53-defective tumors16,19,20. An ideal approach is the repair of mutant p53 into wt-p53 at molecular level. Firstly, it reduces the expression of mutant p53 which drives tumor progression through dominant negative activity and gain-of-function. Secondly, it induces wt-p53 production which restores wt-p53 activity to suppress the growth of cancer cells. Trans-splicing is a natural process which involves splicing between two separately transcribed mRNAs such that a composite transcript is produced21,22. Manipulation of this process offers a therapeutic candidate for genetic disorders caused by dominant mutations21,22,23. This property is exploited as an attractive RNA-modification therapy named as spliceosome mediated RNA trans-splicing (SMaRT)21,22,23. SMaRT-mediated repair is achieved by exon replacement and subsequent removal of the defective portion of the target pre-mRNA so that a functional gene product can be transcribed21,22,24. In theory, this strategy simultaneously reduces the production of deleterious mutant p53 protein and restores wt-p53 production. In addition, another advantage of SMaRT over conventional gene therapy is that the corrected genes can be maintained in their native sequence context and are regulated by their endogenous regulatory machinery.
In the context of these considerations, we attempted to correct mutant p53 transcripts in p53-mutated HCC cells using trans-splicing. A pre-trans-splicing molecule (PTM) consisted of trans-splicing domain and 3′ exon that encoded the correct p53 sequence was delivered into HCC cells with p53 mutation. We demonstrated p53-PTM corrected mutant p53 transcripts and restored wt-p53 activities in p53-defective HCC cells, which resulted in the trans-activation of p53-responsive genes and the suppression of the growth of HCC cells in vitro. Further study revealed that adenovirus vector expressing p53-PTM blocked the growth of tumor xenografts established by p53-mutated HCC cells.
Results
SMaRT-mediated repair of mutant p53 transcripts in transfected cells
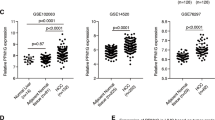
To test the feasibility of trans-splicing-mediated repair of mutant p53 transcripts in vitro, the plasmids encoding p53 trans-splicing expression cassette (Figure 1A) or the controls (Supplementary Figure S1) were delivered into PLC/PRF/5 cells which carried p53 mutation in Exon 7 of p53 cDNA. To confirm p53 trans-splicing in transfected cells, RT-PCR was performed on RNA isolated from the transfected cells using a trans-splicing specific primer set (Figure 1B). RT-PCR analysis detected an expected 893-bp fragments in PLC/PRF/5 cells transfected with p53-PTM (Figure 1C), which was identical to that of p53-cDNA. Moreover, DNA sequencing of the RT-PCR products (Figure 1D) revealed that correct p53 transcripts were generated from splicing events between endogenous target and trans-splicer transcripts, indicating that trans-splicing had occurred correctly with high fidelity in transfected cells. Preferably, the suitable antibodies should be applied to detect repaired p53 protein in transfected cell if possible. Although some antibodies differentiate mutant p53 protein (R175H) from wt-p5325; available antibodies do not discriminate mutant p53 protein (R249S) from wt-p53. Therefore, we evaluated the effect of p53-PTM on the expression of p53 protein using p53 antibody which recognizes both mutant forms and wild-type human p53. Western blot result showed that the level of p53 protein was not changed upon the treatment of p53-PTM (Supplementary Fig. S3). Thus, trans-splicing represents a potentially powerful approach for the correction of p53 mutation in HCC cells.
Schematic diagrams of the strategy for trans-splicing-mediated repair of mutant p53 transcripts and evaluation of trans-splicing-mediated repair of p53 mutation in transfected HCC cells.
(A). Schematic structure of trans-splicing cassettes and pre-RNA target. p53 pre-trans-splicing molecules contain a binding domain, branch point, polypyrimidine tract, splice site, a coding domain and Flag-tag. (B). Schematic illustration of trans-splicing-mediated repair of mutant p53 transcripts. Cis-splicing of the mutant p53 pre-mRNA yields mutant p53 transcripts in codon 249. Mature wt-p53 transcripts were generated by trans-splicing. Arrowheads indicate the PCR primers used for detection of trans-splicing-generated products. TS-FP: 5′-AGGGCAGCTACGGTT TCCGT-3′, TS-RP: 5′- TACTTGTCATCGTCGTCCTTG-3′. (C). Evaluation of trans-splicing-mediated repair of p53 mutation in PLC/PRF/5 cells by RT-PCR. Trans-spliced p53 RNA yields a reverse transcription-PCR product of 893 bp, no products were observed in PLC/PRF/5 cells transfected with pcDNA3.1(+) or pGFP-PTM. (D). Sequences of RT-PCR products generated from PLC/PRF/5 cells or PLC/PRF/5 cells transfected with p53-PTM. The observed sequences of trans-spliced p53 RNA was identical to that of wt-p53. TP53-S-P is the sequencing primer used for the detection of p53. TP53-S-P: 5′- ATGAGCCGCCTGAGGTTGG-3′. BD, Binding domain; BP, branchpoint; PPT, polypyrimidine tract; SS, splice site.
p53-PTM induced the expression of p53 downstream target genes in transfected cells
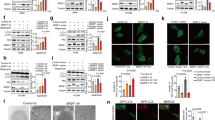
The above results confirmed trans-splicing-mediated repair of mutant p53 transcripts in HCC cells which carried p53 mutation in exon 7 of p53. As a transcription factor that both activates and represses a broad range of target genes, wt-p53 acts as a tumor suppressor by inhibiting cell cycle progression and inducing apoptosis7,15,26. To investigate whether trans-spliced p53 RNAs restore wt-p53 activities in HCC cells, cell apoptosis, cell cycle, cell proliferation and the expression of p53-dependent downstream genes were determined. Firstly, the apoptosis of transfected PLC/PRF/5 cells was evaluated using terminal deoxynucleotidyl transferase mediated dUTP nick end labeling (TUNEL) staining, annexin V/propidium iodide combined labeling flow cytometry and the expression of apoptosis-related genes. The result of TUNEL staining (Figure 2A) showed the proportion of TUNEL positive cells increased remarkably in PLC/PRF/5 cells transfected with p53-PTM compared with the controls (p < 0.05). As shown in Figure 2B, flow cytometry using Annexin-V/propidium iodide combined labeling also showed that an increase in the percentage of apoptotic cells was observed in PLC/PRF/5 cells upon the delivery of p53-PTM compared with the controls. PUMA, Bax, PARP-1, caspase-3 are important indicators of cell apoptosis. As shown in Figure 2 C and D, increased expression of caspase-3, PUMA and PARP was observed in PLC/PRF/5 cells transfected with p53PTM-E7-11. Moreover, p53-PTM suppressed the expression of the apoptosis-suppressing gene Bcl-2 and simultaneously stimulated the expression of Bax which encodes a dominant inhibitor of the Bcl-2 protein. Mdm2, a downstream molecule of p53, has been well characterized as a negative regulator. Mdm2 was also been induced in PLC/PRF/5 cells transfected with p53 PTM (Figure 2C and D).
SMaRT-mediated repair of mutant p53 transcripts resulted in the induction of the apoptosis in PLC/PRF/5 cells.
(A). SMaRT induced the apoptosis of PLC/PRF/5 cells determined by TUNEL Staining. (B). Cell apoptosis was detected by Annexin-V/propidium iodide combined labeling flow cytometry in PLC/PRF/5 cells. * p < 0.05. (C). mRNA relative expression of p53-responsive apoptotic genes was measured by SYBR Green qRT-PCR. * p < 0.05. (D). Representative Western blot and densitometric quantification of the proteins involved in apoptosis were shown.
Then, the effect of p53-PTM on cell cycle was evaluated through analyzing the distribution of cell cycle and the expression of regulatory genes for cell cycle. As shown in Figure 3A, flow cytometry analysis demonstrated an accumulation of cells in the G0-G1 phase following the delivery of the plasmid carrying p53-PTM into PLC/PRF/5 cells. Of the many genes involved in cell cycle control, cyclins control the progression of cells through the cell cycle by activating cyclin-dependent kinase (Cdk) enzymes. Partial repair of mutant p53 transcripts by SMaRT resulted in down-regulation of cyclins, which was demonstrated by quantitative RT-PCR analysis (Figure 3B) and immunoblot results (Fig 3C). p21, a regulator of cell cycle progression, is controlled by p53. Induction of p21 was observed in PLC/PRF/5 cells transfected with PTM-p53 compared with the controls (Figure 3B and C). Finally, proliferative activity was inhibited in PLC/PRF/5 cells after the transfection of p53-PTM compared with the controls (Figure 3D).
The effect of p53-PTM on the proliferation and cell cycle of PLC/PRF/5 cells.
(A). Flow cytometry combined with propidium iodide staining demonstrated that SMaRT induced cell cycle arrest in PLC/PRF/5 cells upon the treatment of p53-PTM. Numbers over each histogram indicated the percentage of the cells in G0-G1, S and G2-M phase. (B). mRNA relative expression of cyclins and p21 was measured by SYBR Green qRT-PCR. *p < 0.05. (C). Representative Western blot and densitometric quantification of the cyclins and p21 were shown. (D). Proliferation assay determined proliferative activity of PLC/PRF/5 cells after transfection of p53-PTM or the controls. * p < 0.05.
To further confirm whether p53-PTM induce p53-dependent downstream genes in cancer cells, the promoter activities of p53 downstream genes were determined using luciferase assay. As shown in Supplementary Figure S4, p53-PTM-E7-11 trans-activated the exogenously expressed luciferase gene driven by PUMA and p21 promoter. In addition, the results showed that the promoter activities of cyclin A, B, D and E and Bcl-2 were downregulated in PLC/PRF/5 cells treated with p53-PTM (Supplementary Figure S4).
Therefore, in vitro data demonstrated p53-PTM suppressed the growth of PLC/PRF/5 cells significantly compared with the negative control (pcDNA3.1), which resulted from an induction of wt-p53 production and a reduction in mutant p53 transcripts. Simultaneously, the above protocols revealed a weak induction of apoptosis, cell-cycle arrest in pGFPPTM-NC transfected cancer cells compared with the negative control (pcDNA3.1). It might be caused by a reduction in mutant p53 transcripts.
The effect of SMaRT on the growth of HCC xenograft tumors in nude mice
To evaluate trans-splicing-mediated repair of mutant p53 transcripts in vivo, adenovirus vectors carrying trans-splicing cassette of p53 were successfully prepared and their high infection efficiency was obtained in PLC/PRF/5 cells as indicated by eGFP expression (Figure 4A). Adenovirus vectors were administered directly into xenograft tumors developed by PLC/PRF/5 cells. 48 hours later, total RNA extracted from tumors was subjected to RT-PCR analysis for the detection of trans-spliced p53 RNA. As expected, an anticipated product of 893 bp was detected in RNA from tumors injected with adenovirus encoding p53-PTM and no amplification was detected from tumors injected with AD-GFP-PTM and AD-Null (Figure 4B). RT-PCR product was sequenced and the result showed point mutation of p53 transcripts was repaired by SMaRT (Figure 4B). Based on the crucial role of p53 in the pathogenesis of HCC, we further pursued the ability of SMaRT against p53 in suppressing the growth of xenograft tumors. Adenovirus vectors expressing p53-PTM and the controls were injected directly into the tumors and tumor size was monitored over time. Although the tumors grew progressively in all groups during the course of the experiment, tumors in p53 PTM-treated mice were significantly smaller than those in control mice reflected by the gross morphology (Figure 4C) and growth curves (Figure 4D). The growth curves of treated tumors became divergent on day 16 and the average fold increase of tumor volumes at the sacrifice with respect to the first measurements in AD-p53PTM-treated group was less than that in controls (Figure 4D).
Adenovirus vectors expressing p53-PTM inhibited the growth of xenograft tumors developed by PLC/PRF/5 cells in nude mice.
(A). The result showed efficient adenoviral transduction of PLC/PRF/5 cells as indicated by eGFP expression using a fluorescence microscope, 2 days following adenovirus vector administration. The black and white pictures showed the cells in the same field under normal white light. (B). Detection of trans-spliced p53 RNA in xenograft tumors. RT-PCR results showed that trans-spliced p53 RNAs were generated in PLC/PRF/5 cells infected with adenovirus vectors expressing p53-PTM (AD-p53PTM-E7-11). DNA sequencing of the RT-PCR products of trans-spliced p53 RNA confirmed trans-splicing-mediated repair of mutant p53 transcripts in the infected cells with high fidelity. (C). Effects of AD-p53PTM on the growth of pre-established PLC/PRF/5 xenografts at a gross morphology level. (D). Adenovirus vectors expressing p53-PTM inhibited the growth of xenograft tumors developed by PLC/PRF/5 cells in nude mice. The inhibitory effect of AD-p53PTM on the growth of xenograft tumors was revealed by the tumor growth curves (a) and the average volume fold increase of tumors at the sacrifice with respect to the first measurements (b). *p < 0.05. (E). Cell proliferation was assessed by immnofluorescence staining of Ki67. Representative immunofluorescence staining and the percentage of ki67-positive cells were shown. *p < 0.05. (F). The expression of the proteins involved in cell cycle and apoptosis was evaluated by immunohistochemical staining of the sections from the xenograft tumors. Representative immunohistochemical staining and percentages of cells staining positive for cyclins, Bax, Bcl-2, caspase3 and mdm2 were shown. Positive cells were counted in tumor tissues and presented as the mean ± SD (4 random fields per section and three sections per tumor). 1: AD-Null, 2: AD-GFP-PTM, 3: AD-p53PTM-E7-11. * p < 0.05.
The effect of p53-PTM on the proliferation, the expression of Cyclins, Bax, Bcl-2, caspase-3 and mdm2 in xenograft tumors
Proliferative phenotypes are fundamental components of malignant diseases. In contrast with AD-Null or AD-GFP-PTM, the tumor cells in AD-p53PTM-E7-11 group manifested a remarkable decrease in immunofluorescence of proliferative biomarker ki67 (Figure 4E). Bcl-2 family members, caspase-3, cyclins are important indicators of cell growth arrest and apoptosis. In tumor tissues examined by immunohistochemical staining, administration of AD-p53PTM-E7-11 resulted in remarkable up-regulation of caspase-3 and Bax, accompanied with decreased expression of Bcl-2 and cyclin A, B, D (Figure 4F). Simultaneously, tumors in mice treated with AD-p53PTM-E7-11 showed significant induction of p53-responsive molecule mdm2 compared with the controls. In addition, consistent with the effect of GFP-PTM on cell cycle and apoptosis in transfected cells in vitro, adenovirus enconding GFP-PTM showed week induction of cell-cycle arrest and cell apoptosis compared with the negative control (AD-Null) (Figure 4F).
Discussion
Rescuing the function of mutant p53 in cancer cells is an attractive cancer therapeutic strategy. In this study, a RNA-based strategy named as SMaRT has been designed to amend mutant p53 transcripts in HCC cells carrying mutant p53 transcripts. We demonstrated a plasmid carrying p53-PTM mediated the correction of mutant p53 transcripts in p53-defective HCC cells. The reactivation of wt-p53 in PLC/PRF/5 cells led to the transactivation of p53-dependent target genes and then induced cell cycle arrest and cell apoptosis in vitro. Human tumor xenografts have been widely used as predictive preclinical models for anticancer drug in humans. Therefore, HCC xenograft is used as a tumor model for evaluating the feasibility of trans-splicing. After intratumoral injection of adenovirus vector encoding p53-PTM, SMaRT reduced the growth of xenograft tumors developed by PLC/PRF/5 cells. Thus, it indicated that endogenous p53 mutant transcripts were partially corrected using SMaRT in vivo. Intra-tumor delivery is of limited value for most tumor types; however, local therapeutic approaches such as local ablation with radiofrequency or percutaneous ethanol injection, transcatheter arterial chemoembolization were also recommended in HCC clinical practice guideline. HCC xenografts and orthotopic HCC tumor model share similar characteristic for intratumor administration of anticancer drug. These indicate that intratumor delivery of vectors expressing p53-PTM is feasible for the treatment of p53-mutated HCC.
Mutation or functional inactivation of p53 is one of the most common genetic events in human cancer, including HCC. Most p53 mutations are missense point mutations that preferentially localized in the DNA binding domain9,15. A single missense mutation at codon 249 of the p53 gene has been identified as one of the “hotspot” mutation9,15. Therefore, PLC/PRF/5 cells containing mutation in codon 249 were selected as cell model for the detection of SMaRT-mediated correction of mutant p53 transcript. p53 mutations in the DNA binding domain result in the disruption of DNA binding and transcriptional trans-activation of p53 target genes, which abrogates p53 tumor-suppressive functions. In addition, mutant p53 protein gains additional oncogenic functions that endow cells with growth and survival advantages. A number of strategies based on the reactivation of wt-p53 function in p53-defective cancer cells have been developed18,27. Numerous studies have demonstrated that small synthetic molecules and peptides reversed phenotype of cancer cells in vitro and in vivo by reconstituting p53 tumor suppressor functions18,27. While, the chemicals had potential toxicity which might limit their application in humans. Their molecular mechanism underlying restoration of p53 function should be pursued and the application of the chemicals in clinical practice should be evaluated in further study. In theory, restoring mutant p53 function and/or enhancing wt-p53 by genetic means is a viable and attractive approach for developing cancer therapeutics. The efficacy of wt-p53 administration has been confirmed in preclinical and clinical models28,29,30. However, over-expression of p53 might be found in mice tissues after the administration of wt-p53 and increased p53 activity in mice model displayed the phenotype of accelerated and pronounced aging31,32,33. Mutant p53 proteins are unusually stable and accumulate to high levels in cancer cells, which exert gain-of-function in the promotion of carcinogenesis9,14,15. Therefore, the strategy of wt-p53 administration did not get a rid of detrimental mutant p53 transcripts. Alternative approach is knockdown of endogenous p53 mutant transcripts in cancer cells using siRNAs which inhibited tumor growth34,35,36. However, absence of wt-p53 expression in cancer cells contributed to the development of various tumor types32. Therefore, an ideal strategy for p53-mutated cancer cells is the repair of mutant p53 into wt-p53 in molecular level, which can simultaneously reduce mutant p53 expression and restore wt-p53 expression in cancer cells. Trans-splicing ribozymes had been utilized to repair mutant p53 transcripts with high fidelity and specificity in cancer cells37,38. Trans-splicing ribozymes-mediate repair of mutant p53 resulted in the production of functional p53 and the reduction of mutant p53 expression37,38. Unfortunately, ribozyme stability and efficiency are the main hurdles to applying the ribozyme to the clinic22,24,37,38. SMaRT can result in the replacement of defective target pre-mRNA via a correct version of the segment. This technology has been successfully applied in phenotypic correction of genetic defects, including cystic fibrosis (CF), haemophilia A and X-linked immunodeficiency39,40,41,42. Our studies have clearly demonstrated for the first time that partial correction of mutant p53 transcripts was achieved using SMaRT. All p53 isoforms share same structure in intron 6 and exon 743, which results in successful correction of all p53 isoforms containing p53 mutation in exon 7 using p53PTM-E7-11. The trans-spliced p53 transcripts induced apoptosis and cell cycle arrest in transfected cells and xenograft tumors developed by PLC/PRF/5 cells. As a RNA-based strategy, SMaRT has several advantages over conventional gene therapy. Trans splicing-mediated revision of mutant p53 transcripts can simultaneously reduce mutant p53 transcripts and restore wt-p53 production, which conventional gene replacement does not address. As the gene is repaired rather than introduced, the spatial and temporal expression of the gene should be controlled by endogenous regulation. Coordinated expression of the p53 is apparently critical for proper homeostasis of cells growth and incorrect p53 expression in mice model can lead to phenotypic aberrations33,44. The corrected p53 genes by SMaRT can be maintained in their native sequence context and are regulated by their endogenous regulatory machinery. In addition, the protein expression of p53 was not changed in HCC cells or normal hepatic cells expressing wt-p53 upon the treatment of p53-PTM (Supplementary Fig S3). Thus, trans-splicing-mediated repair of mutant p53 transcripts takes place in p53-defective cancer cells and undesirable effects of SMaRT on normal cells would be avoided.
PLC/PRF/5 cell line also contains other gene mutations such as CDKN2A (R112G). CDKN2A binds to mdm2 and inhibits the oncogenic action of mdm2 by blocking mdm2-induced degradation of p53 and enhancing p53-dependent transactivation and apoptosis45. However, Weber JD et al demonstrated that CDKN2A could also exert its tumor suppressive functions independently of the mdm2-p53 axis46. In addition, recent studies demonstrated that CDKN2A mutation did not occur frequently in human HCC tissues47,48. So, CDKN2A mutation may impair the therapeutic effects of p53-PTM on xenograft tumors created by PLC/PRF/5. However, our study focused on the repair of mutant p53 transcripts using trans-splicing in p53-mutated HCC cells. Therefore, the effects of p53-PTM on the function of other gene mutations were not investigated.
Of course, SMaRT has its limitations in the repair of p53 mutation. One of the major limitations is that a single PTM would not be able to address all mutations across a defective gene. Low efficiency would be a major potential disadvantage for the application of trans-splicing. Although p53-PTM suppressed the growth of p53-defective HCC cells in vitro and in vivo significantly, improvements in PTM design and delivery system which will increase their efficacy should be made in further study.
Mutations of p53 are one of the most frequent molecular events in the tumorigenesis of HCC and play an important role in the pathogenesis of HCC. In conclusion, our study demonstrated that SMaRT was successfully applied to correct mutant p53 transcripts in HCC cells carrying p53 mutation, which resulted in restoration of wt-p53 activity and phenotypic correction of HCC cells. These revealed that SMaRT would be developed as a good therapeutic candidate for p53-defective HCC.
Methods
See Supplementary Methods for detailed experimental methods
Plasmids construct and cell transfection
As shown in Fig 1A, a pre-trans-splicing molecule (PTM) contained one binding domain, the splice elements, a coding domain and Flag-tag. The PTM binding domain, complementary to intron 6 of human p53 gene, was generated by PCR on genomic DNA isolated from HepG2 cell. The splice elements consisted of the motifs necessary for a trans-splicing reaction included a spacer sequence, an extended polypyrimidine tract, a yeast branch-point consensus and a 3′ AG acceptor site. The coding domain encoded exon 7–11 of human p53 cDNA which was amplified by PCR using the plasmid pCMV-Neo-Bam wtp53. The plasmids pcDNA3.1(+), pGFPPTM-NC and p53-cDNA were used as the controls of p53PTM-E7-11. The schematic illustration of plasmids (pGFPPTM-NC and p53-cDNA) encoding GFP pre-trans-splicing molecule or p53 cDNA were shown in Supplementary Figure S1. Then, the plasmids were transfected into the p53-defective cells using Lipofectamine™2000 (Life Technologies, Gaithersburg, MD, USA). HCC cell line PLC/PRF/5 (mutant p53/R249S, confirmed in Supplementary Figure S2) was used as p53-defective cancer cells. 48 hours after transfection, total RNAs isolated from transfected cells were subjected to the analysis of RT-PCR and real-time RT-PCR. Western blot, cell proliferation and analysis of cell cycle and apoptosis were performed as we have described before49.
Production of recombinant adenovirus and animal experiment
Trans-splicing expression cassettes were cloned into shuttle plasmid of AdMax™ system with Cre-lox (Microbix Biosystems Inc., Ontario, Canada). The recombinant adenoviruses were produced and purified. Six to eight-week-old Nu/Nu mice obtained from Beijing HFK Bioscience Co. LTD (Beijing, China) were inoculated subcutaneously with PLC/PRF/5 cells to generate tumor xenografts and then adenovirus vectors were administered directly into the tumors. At the end of the experiment, the mice were sacrificed at day 36 post-inoculations and the tumors were collected. Immunohistochemistry and immunofluoresence were performed on tumor tissues as described previously50,51.
Ethic statement
The study was approved by the local research ethics committee at the Tongji Hospital of Huazhong University of Science and Technology. The methods used in this study were carried out in accordance with the approved guidelines.
Statistical analysis
All data are expressed as mean ± standard error from 3 separate experiments performed in triplicate except otherwise noted. The differences between groups were analyzed by Student's t test and P < 0.05 was considered to be statistically significant.
References
Jemal, A. et al. Global cancer statistics. CA: a cancer journal for clinicians 61, 69–90 (2011).
Bruix, J. & Sherman, M. Management of hepatocellular carcinoma. Hepatology 42, 1208–1236 (2005).
Dragani, T. A. Risk of HCC: genetic heterogeneity and complex genetics. Journal of hepatology 52, 252–257 (2010).
de Lope, C. R., Tremosini, S., Forner, A., Reig, M. & Bruix, J. Management of HCC. Journal of hepatology 56 Suppl 1S75–87 (2012).
Martin, J. & Dufour, J. F. Tumor suppressor and hepatocellular carcinoma. World J Gastroenterol 14, 1720–1733 (2008).
Lozano, G. Mouse models of p53 functions. Cold Spring Harb Perspect Biol 2, a001115 (2010).
Foulkes, W. D. p53-master and commander. N Engl J Med 357, 2539–2541 (2007).
Kruse, J. P. & Gu, W. Modes of p53 regulation. Cell 137, 609–622 (2009).
Muller, P. A. & Vousden, K. H. p53 mutations in cancer. Nature cell biology 15, 2–8 (2013).
Goh, A. M., Coffill, C. R. & Lane, D. P. The role of mutant p53 in human cancer. The Journal of pathology 223, 116–126 (2011).
Sell, S. Mouse models to study the interaction of risk factors for human liver cancer. Cancer Res 63, 7553–7562 (2003).
Aravalli, R. N., Steer, C. J. & Cressman, E. N. Molecular mechanisms of hepatocellular carcinoma. Hepatology 48, 2047–2063 (2008).
Liu, J. et al. Alterations of TP53 are associated with a poor outcome for patients with hepatocellular carcinoma: evidence from a systematic review and meta-analysis. Eur J Cancer 48, 2328–2338 (2012).
Joerger, A. C. & Fersht, A. R. Structure-function-rescue: the diverse nature of common p53 cancer mutants. Oncogene 26, 2226–2242 (2007).
Freed-Pastor, W. A. & Prives, C. Mutant p53: one name, many proteins. Genes & development 26, 1268–1286 (2012).
Bossi, G. & Sacchi, A. Restoration of wild-type p53 function in human cancer: relevance for tumor therapy. Head Neck 29, 272–284 (2007).
Selivanova, G. & Wiman, K. G. Reactivation of mutant p53: molecular mechanisms and therapeutic potential. Oncogene 26, 2243–2254 (2007).
Cheok, C. F., Verma, C. S., Baselga, J. & Lane, D. P. Translating p53 into the clinic. Nature reviews. Clinical oncology 8, 25–37 (2011).
Levesque, A. A. & Eastman, A. p53-based cancer therapies: Is defective p53 the Achilles heel of the tumor? Carcinogenesis 28, 13–20 (2007).
Suzuki, K. & Matsubara, H. Recent advances in p53 research and cancer treatment. J Biomed Biotechnol 2011, 978312 (2011).
Yang, Y. & Walsh, C. E. Spliceosome-mediated RNA trans-splicing. Mol Ther 12, 1006–1012 (2005).
Wood, M., Yin, H. & McClorey, G. Modulating the expression of disease genes with RNA-based therapy. PLoS Genet 3, e109 (2007).
Pergolizzi, R. G. & Crystal, R. G. Genetic medicine at the RNA level: modifications of the genetic repertoire for therapeutic purposes by pre-mRNA trans-splicing. C R Biol 327, 695–709 (2004).
O'Connor, T. P. & Crystal, R. G. Genetic medicines: treatment strategies for hereditary disorders. Nat Rev Genet 7, 261–276 (2006).
Yu, X., Vazquez, A., Levine, A. J. & Carpizo, D. R. Allele-specific p53 mutant reactivation. Cancer cell 21, 614–625 (2012).
Hainaut, P. & Wiman, K. G. 30 years and a long way into p53 research. Lancet Oncol 10, 913–919 (2009).
Lane, D. P., Cheok, C. F. & Lain, S. p53-based cancer therapy. Cold Spring Harb Perspect Biol 2, a001222 (2010).
Roth, J. A. et al. Retrovirus-mediated wild-type p53 gene transfer to tumors of patients with lung cancer. Nat Med 2, 985–991 (1996).
Guo, J. & Xin, H. Chinese gene therapy. Splicing out the West? Science 314, 1232–1235 (2006).
Shimada, H. et al. Phase I/II adenoviral p53 gene therapy for chemoradiation resistant advanced esophageal squamous cell carcinoma. Cancer science 97, 554–561 (2006).
Poyurovsky, M. V. & Prives, C. Unleashing the power of p53: lessons from mice and men. Genes & development 20, 125–131 (2006).
Donehower, L. A. & Lozano, G. 20 years studying p53 functions in genetically engineered mice. Nature reviews. Cancer 9, 831–841 (2009).
Tyner, S. D. et al. p53 mutant mice that display early ageing-associated phenotypes. Nature 415, 45–53 (2002).
Bossi, G. et al. Conditional RNA interference in vivo to study mutant p53 oncogenic gain of function on tumor malignancy. Cell Cycle 7, 1870–1879 (2008).
Zhu, H. B. et al. Silencing of mutant p53 by siRNA induces cell cycle arrest and apoptosis in human bladder cancer cells. World journal of surgical oncology 11, 22 (2013).
Bossi, G. et al. Mutant p53 gain of function: reduction of tumor malignancy of human cancer cell lines through abrogation of mutant p53 expression. Oncogene 25, 304–309 (2006).
Watanabe, T. & Sullenger, B. A. Induction of wild-type p53 activity in human cancer cells by ribozymes that repair mutant p53 transcripts. Proc Natl Acad Sci U S A 97, 8490–8494 (2000).
Shin, K. S., Sullenger, B. A. & Lee, S. W. Ribozyme-mediated induction of apoptosis in human cancer cells by targeted repair of mutant p53 RNA. Mol Ther 10, 365–372 (2004).
Mansfield, S. G. et al. Repair of CFTR mRNA by spliceosome-mediated RNA trans-splicing. Gene Ther 7, 1885–1895 (2000).
Liu, X. et al. Partial correction of endogenous DeltaF508 CFTR in human cystic fibrosis airway epithelia by spliceosome-mediated RNA trans-splicing. Nat Biotechnol 20, 47–52 (2002).
Chao, H. et al. Phenotype correction of hemophilia A mice by spliceosome-mediated RNA trans-splicing. Nat Med 9, 1015–1019 (2003).
Tahara, M. et al. Trans-splicing repair of CD40 ligand deficiency results in naturally regulated correction of a mouse model of hyper-IgM X-linked immunodeficiency. Nat Med 10, 835–841 (2004).
Khoury, M. P. & Bourdon, J. C. p53 Isoforms: An Intracellular Microprocessor? Genes & cancer 2, 453–465 (2011).
Garcia-Cao, I. et al. “Super p53” mice exhibit enhanced DNA damage response, are tumor resistant and age normally. The EMBO journal 21, 6225–6235 (2002).
Maggi, L. B., Jr et al. ARF tumor suppression in the nucleolus. Biochimica et biophysica acta 1842, 831–839 (2014).
Weber, J. D. et al. p53-independent functions of the p19(ARF) tumor suppressor. Genes & development 14, 2358–2365 (2000).
Ito, T. et al. Alteration of the p14(ARF) gene and p53 status in human hepatocellular carcinomas. Journal of gastroenterology 39, 355–361 (2004).
Taniguchi, K., Yamada, T., Sasaki, Y. & Kato, K. Genetic and epigenetic characteristics of human multiple hepatocellular carcinoma. BMC cancer 10, 530 (2010).
He, X. X. et al. MicroRNA-375 targets AEG-1 in hepatocellular carcinoma and suppresses liver cancer cell growth in vitro and in vivo. Oncogene 31, 3357–3369 (2012).
Wang, Z. et al. Embryonic liver fodrin involved in hepatic stellate cell activation and formation of regenerative nodule in liver cirrhosis. Journal of cellular and molecular medicine 16, 118–128 (2012).
Wang, Z. et al. beta-2 spectrin is involved in hepatocyte proliferation through the interaction of TGFbeta/Smad and PI3K/AKT signalling. Liver international 32, 1103–1111 (2012).
Acknowledgements
The work was supported by grants from the National Natural Science Foundation of China (No. 81072003, 81270506, 81472832) and by the Outstanding Youth Science Foundation of Tongji Hospital (No. YXQN005). We thank Pro. Bert Vogelstein (the Johns Hopkins University School of Medicine) for providing pCMV-Neo-Bam wtp53. We are grate to Prof Giulia Piaggio (Experimental Oncology Department, Istituto Regina Elena, Italy) for providing pcyclinB-prom-LUC.
Author information
Authors and Affiliations
Contributions
X.X.H., F.L. and Y.H.S. designed the research; X.X.H., F.L., J.J.Y., Y.N.Z. and J.W.Y. performed the research; H.T.S., Q.D. and Q.Z. contributed new reagents and analyzed the data; Y.H.S. and X.X.H. wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary data
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
He, X., Liu, F., Yan, J. et al. Trans-splicing repair of mutant p53 suppresses the growth of hepatocellular carcinoma cells in vitro and in vivo. Sci Rep 5, 8705 (2015). https://doi.org/10.1038/srep08705
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep08705
This article is cited by
-
Interferon-γ secreted by recruited Th1 cells in peritoneal cavity inhibits the formation of malignant ascites
Cell Death Discovery (2023)
-
Silencing E3 Ubiqutin ligase ITCH as a potential therapy to enhance chemotherapy efficacy in p53 mutant neuroblastoma cells
Scientific Reports (2020)
-
We skip to work: alternative splicing in normal and malignant myelopoiesis
Leukemia (2018)
-
β2 spectrin-mediated differentiation repressed the properties of liver cancer stem cells through β-catenin
Cell Death & Disease (2018)
-
NAT10 is upregulated in hepatocellular carcinoma and enhances mutant p53 activity
BMC Cancer (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.