Abstract
In primary fibroblasts from Leigh Syndrome (LS) patients, isolated mitochondrial complex I deficiency is associated with increased reactive oxygen species levels and mitochondrial morpho-functional changes. Empirical evidence suggests these aberrations constitute linked therapeutic targets for small chemical molecules. However, the latter generally induce multiple subtle effects, meaning that in vitro potency analysis or single-parameter high-throughput cell screening are of limited use to identify these molecules. We combine automated image quantification and artificial intelligence to discriminate between primary fibroblasts of a healthy individual and a LS patient based upon their mitochondrial morpho-functional phenotype. We then evaluate the effects of newly developed Trolox variants in LS patient cells. This revealed that Trolox ornithylamide hydrochloride best counterbalanced mitochondrial morpho-functional aberrations, effectively scavenged ROS and increased the maximal activity of mitochondrial complexes I, IV and citrate synthase. Our results suggest that Trolox-derived antioxidants are promising candidates in therapy development for human mitochondrial disorders.
Similar content being viewed by others
Introduction
Virtually every cell contains mitochondria, which are double membrane organelles that play multiple roles in cell metabolism, signal transduction and ATP generation. Mitochondrial function is sustained by the action of the oxidative phosphorylation (OXPHOS) system, consisting of four electron transport chain (ETC) complexes (CI-CIV) and the ATP-generating FoF1-ATPase (CV)1. ETC and CV action is linked via the highly negative potential (Δψ) across the mitochondrial inner membrane (MIM) through chemiosmotic coupling2. Malfunction of the OXPHOS system is observed in many human diseases including metabolic syndromes such as Leigh syndrome (LS)3,4,5. At the cellular level mitochondrial dysfunction and alterations in mitochondrial structure and Δψ are often paralleled by increased reactive oxygen species (ROS) levels6,7,8,9. This suggests that these aberrations might constitute linked therapeutic targets. In fibroblasts from LS patients with isolated CI deficiency (OMIM 253010), the α-tocopherol derivative Trolox (6-hydroxy-2, 5, 7, 8-tetramethylchroman-2-carboxylic acid) normalized increased levels of CM-H2DCF (5-(and-6)-chloromethyl-2′, 7′-dichlorodihydro-fluorescein) oxidizing ROS10. Trolox treatment also increased the amount/activity of the CI holo complex and improved Ca2+-stimulated mitochondrial ATP production in LS cells10,11. In fibroblasts from a healthy individual (CT) Trolox stimulated mitochondrial length and degree of branching12. Also studies in other models of mitochondrial disease highlighted beneficial effects of (mitochondria-targeted) antioxidants13,14,15,16. Although the above suggests that elevated ROS levels might play a pathophysiological role in mitochondrial dysfunction, antioxidant treatment of “real” mitochondrial disease patients appeared only (transiently) effective in a very limited number of cases17,18,19. These negative results raised doubts about the effectiveness of antioxidant treatment in ROS-related human pathologies20,21,22,23,24.
The discrepancy between results obtained in model systems and mitochondrial disease patients is striking and might be explained by the dual role of ROS as signaling and damaging molecules12,25,26. In addition, antioxidants can also display pro-oxidant properties depending on their concentration, physicochemical properties and reaction environment27,28,29. Importantly, therapeutic small molecules (including bioactive antioxidants) generally target multiple effectors thereby exerting subtle pleiotropic effects (both on- and off-target) at the cellular level30. This means that simply studying the in vitro potency and/or performing single-parameter high-throughput image-based cell screening is of limited use to identify potential drug-like molecules or understand their mode-of-action. Therefore the rate-of-success during lead selection and optimization in drug discovery benefits from multi-parameter phenotypic profiling31.
Here we present an integrated experimental and computational strategy that is broadly applicable for small molecule profiling and based on quantifying their phenotypical effects. This approach consists of: (i) automated calculation of phenotypic “fingerprints” reflecting mitochondrial morphology and Δψ (“morpho-function”) at the level of individual mitochondria in single living cells, (ii) using these morpho-functional fingerprints for supervised machine learning classification of CT and LS patient cells, (iii) automated evaluation of morpho-functional effects induced by small molecule oxidants in LS patient cells. In a proof-of-principle study, the above strategy was applied to evaluate the phenotypic effects of four newly developed Trolox variants in LS patient cells. This highlighted Trolox ornithylamide hydrochloride (KH003) as a therapeutically promising Trolox derivative, as confirmed by independent in vitro and in cellulo analysis. KH003 displayed cellular ROS scavenging, stimulated CI, CIV and citrate synthase activity. This suggests that Trolox-derivatives are promising candidates in therapy development for human mitochondrial disorders.
Results
Mitochondrial morpho-functional fingerprinting of human skin fibroblasts
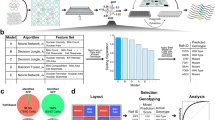
An automated approach was developed for phenotypic analysis of mitochondrial morphology and Δψ (“morpho-functional fingerprinting”) in primary human skin fibroblasts (Fig. 1). Cells were stained with TMRM (tetramethyl rhodamine methyl ester), a mitochondria-specific cation that accumulates in the mitochondrial matrix in a Δψ-dependent manner and visualized using epifluorescence microscopy (Fig. 1A; yellow boxes). Various quantitative parameters (Supplementary Table S1) describing mitochondrial morphology and TMRM intensity (“morpho-functional descriptors”) were extracted from the microscopy images by applying an automated image processing and analysis algorithm (Fig. 1A; blue boxes). This strategy was extensively validated previously in primary human skin fibroblasts (Supplementary Information). For every image, 31 descriptors were calculated for each mitochondrial object. Relative to typical control cells (CT5120), descriptor values in LS patient cells (P5175) were unaffected (8 descriptors), significantly increased (2 descriptors) or significantly decreased (21 descriptors; Supplementary Table S1).
Overall strategy for mitochondrial morpho-functional fingerprinting in primary human skin fibroblasts by combined automated image quantification and artificial intelligence techniques.
(A) Schematic depiction of the integrated experimental and computational strategy. Fibroblasts are stained with the mitochondria-specific cation TMRM and manually imaged by epifluorescence microscopy (yellow boxes: #1-#3). Next, the microscopy images are processed and the numerical values of 31 descriptors of mitochondrial morphology and membrane potential are extracted at the level of individual cells (blue boxes: #4-#13). The median value of each descriptor variable was calculated for each microscopy image (blue box: #14) and used for subsequent machine learning analysis (green boxes: #15-#23; for details see Results). (B) Typical images of a fibroblast from a healthy volunteer (CT5120) illustrating the various image processing steps in panel A (corresponding blue boxes and numbers).
Using mitochondrial morpho-functional fingerprints for supervised machine learning and cell classification
The dataset of 31 descriptors was used for training a machine learning algorithm to discriminate between CT5120 and P5175 cells. In total, 711 images were analyzed for the CT5120 cell line (containing 187465 mitochondrial objects) and 567 images were analyzed for the P5175 cell line (112615 objects). By calculating the median value of the 31 descriptors per image, a morpho-functional “fingerprint” of the P5175 cell line was obtained (Supplementary Table S1). Using the median value instead of the average value prevents that outlier values bias the analysis. Next, we used the multivariate fingerprints to determine whether various supervised machine learning algorithms were able to reliably discriminate between CT5120 and P5175 cells (Fig. 1A; green boxes). For this purpose, the data set for the CT5120 and P5175 cells was equally divided into a “training set” and a “test set”. The training set was used to “teach” the multivariate machine learning model, after which the test set was used to assess its classification (predictive) performance. Crucially, the test images were not used to train the machine learning model. To express the classification performance of the trained machine learning model, a score was used that reflects the probability that a given image in the test set is correctly classified as a CT5120 or P5175 cell. The performance of five popular machine learning algorithms was compared (Supplementary Information): Logistic Regression (LogReg), Linear Discriminant Analysis (LDA), Partial Least Squares Discriminant Analysis (PLSDA), Quadratic Determinant Analysis (QDA) and Support Vector Machines (SVMs). Best results were obtained with the LogReg and SVM model using the median per cell data-set (Supplementary Table S2). The LogReg model was used in the remainder of the study since it is computationally simpler than SVM and easier to interpret. The LogReg model condenses the total information contained in the 31 descriptors into a single parameter: the LogReg score. The latter represents the probability that a cell is classified as being CT5120 (LogReg value close to 1) or P5175 (LogReg value close to 0). The LogReg score is displayed against the first principal component (PC1), which captures the maximal variance in the overall data (Fig. 2A). This demonstrates that CT5120 and P5175 cells separate along the LogReg score, whereas no separation is observed along the PC1 axis, meaning that: (i) the difference in morpho-functional phenotype between CT5120 and P5175 cells is captured by the analysis (LogReg) and, (ii) non-supervised machine learning analysis is not appropriate to detect this difference (PC1). The LogReg model is designed in such a way that it takes a binary decision (i.e. any image scoring below 0.5 corresponds to P5175 cells whereas a score above 0.5 corresponds to CT5120 cells), leading to a correct classification rate (per individual image) of 78% on a blind test set. The corresponding area under the curve (AUC) of the receiver operator characteristic (ROC) curve (Fig. 2B), which has as maximal value of 1, equaled 0.85. This value reflects that the ROC-curve is above the line y = x, meaning that the LogReg model has predictive power. These results demonstrate that the two cell lines display observable differences and that the LogReg model is able to classify CT5120 and P5175 cells with 78% accuracy (based upon a single image). To further validate this model, two additional control cell lines from other individuals were analyzed (CT5118: n = 79 images, o = 16860 objects, N = 3 biological replicates; CT5119: n = 109, o = 31153, N = 4). The trained LogReg model also correctly classified these cell lines meaning that the mitochondrial morpho-functional fingerprint of all three control cell lines were classified as identical. Next, we estimated the relative importance of each descriptor in the model, as represented by the absolute value of the LogReg coefficients (Fig. 2C), using a backward elimination similar to the Recursive Feature Elimination (RFE) strategy32. In brief, the least informative descriptor (i.e. the one with the lowest LogReg coefficient) was removed from the dataset followed by recalculation of the LogReg model. This process was repeated until all descriptors were deleted from the model. RFE analysis revealed that at least 21 descriptors need to be included to conserve the predictive performance of the LogReg model (Fig. 2D). Three descriptors (“Perimeter Ratio”, “Margination” and “Area on Box”) ranked highest in the LogReg model (Fig. 2E).
Machine learning classification of control and patient cells.
(A) Visualization of the test samples according to the logistic regression scores vs. the main source of variance in the data (Principal component 1: PC1). Data from individual microscopy images of control cells (CT5120; green dots; n = 356 images; N = 13 days) and patient cells (P5175; red dots; n = 235 N = 13) separates along the horizontal axis. A cell image with a logistic regression (LogReg) score below 0.5 is classified as a P5175 cell. (B) Receiver Operator Curve (ROC) of the LogReg model demonstrating a correct single-image classification score of 78%. The area under the curve (AUC) of the ROC equaled 0.847. (C) Absolute value of the regression coefficient associated with the descriptors. This is a measure of the relative importance of each descriptor in the trained LogReg machine learning model. (D) Effect of Recursive Feature Elimination (RFE) on the classification performance of the LogReg model. Performance dropped when less than 21 descriptors are used (dotted line). (E) Background-corrected image of a TMRM-stained CT5120 fibroblast (top panel) and magnification of a region-of-interest (white box). The lower three panels show a “mitogram” depicting all mitochondrial objects in the top panel sorted according to the numerical value (indicated for typical objects) of the “Perimeter ratio”, “Margination” and “Area on Box” descriptor.
Morpho-functional fingerprinting and machine learning classification of cells treated with Trolox and newly developed Trolox variants
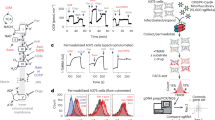
In cellular models of CI deficiency (i.e. LS cells) treatment with the antioxidant Trolox mitigated or fully reversed various cellular aberrations (see Introduction). As a proof-of-principle we applied the above strategy to quantify the effects of Trolox and 4 new Trolox derivatives (KH001-KH004) in P5175 cells. To this end we modified the Trolox sidechain and left the antioxidant part of the molecule (i.e. the HO-group on the chroman ring) intact (Fig. 3A). Cheminformatics analysis predicted that KH001-KH004 display similar physicochemical properties and that KH003 is chemically most similar to Trolox (Supplementary Figure S1 and Supplementary Table S3). Morpho-functional fingerprinting revealed that all five antioxidants reversed the patient phenotype to a variable extent (Fig. 3B and Supplementary Table S1). To condense this phenotypic information into a single parameter (LogReg score), images of antioxidant-treated P5175 cells were classified with the trained LogReg model. Only for KH003-treated patient cells the LogReg-score was larger than 0.5, meaning that they were classified as control cells.
Machine learning classification of the effect of Trolox and newly developed Trolox variants in control and LS patient cells.
(A) Chemical structure of the antioxidant Trolox and four novel Trolox derivatives (KH001-KH004). (B) Heat map representing the mitochondrial morpho-functional fingerprint of vehicle-treated patient cells (P5175), control cell (CT5120) and patient cells treated with the various antioxidants (500 μM, 72 h). The color coding in the heatmap represents significantly increased (red), significantly decreased (blue), or unchanged (white) descriptor values relative to CT5120. The overall effect of each antioxidant treatment is condensed into a single parameter (i.e. the LogReg score given by the machine learning model). The vertical boxes on the right indicate whether the mitochondrial morpho-functional fingerprint is classified as a P5175 cell (red) or as a CT5120 cell (green). For calculation of the heatmaps the following numbers of images were used: CT5120 (n = 711 images, o = 187465 objects, N = 35 days), P5175 (n = 467, o = 112615, N = 25), P5175 + Trolox (n = 260, o = 64638, N = 12), P5175 + KH001 (n = 43, o = 11484, N = 2), P5175 + KH002 (n = 74, o = 19158, N = 4), P5175 + KH003 (n = 74, o = 22364, N = 4), P5175 + KH004 (n = 44, o = 10927, N = 2).
In vitro and cellular antioxidant properties of Trolox and newly developed Trolox variants
As a measure of in vitro antioxidant capacity, we determined the oxygen radical absorbance capacity (ORAC) value. This parameter decreased in the order: KH003 > Trolox > KH001 > KH002 ≫ KH004. Analysis in LS-patient cells revealed that all antioxidants (500 μM, 72 h) reduced the increased levels of CM-H2DCF-oxidizing ROS (Fig. 4A). Trolox and KH004 did not affect the elevated levels of HEt (hydroethidium)-oxidizing ROS in LS-patient cells (Fig. 4A). These ROS levels were increased in KH001-treated cells and reduced by KH002 and KH003. None of the compound treatments affected cell shape or adherence (Fig. 4B; typical example of KH003-treated LS-patient cells).
Effect of Trolox and newly developed Trolox variants on ROS levels, ETC enzymatic activity and cell morphology in LS patient cells.
(A) Polar plot summarizing the in vitro peroxyl radical scavenging activity (ORAC), the levels of CM-H2DCF and HEt-oxidizing ROS and the activity of mitochondrial complex I (CI), complex IV (CIV) and citrate synthase (CS). CM-H2DCF, HEt and activity data were expressed as treated/vehicle condition (data taken from Supplementary Table S1). Vehicle-treated cells are indicated by a dotted line. (B) Typical microscopy images of LS cells (P5175) treated with KH003 and stained with the ROS-sensing reporter molecules 5-(and-6)-chloromethyl-2′, 7′-dichlorodihydro-fluorescein (CM-H2DCF) and hydroethidium (HEt). A magnification of the region within the white box (left panel) is shown on the right.
Effect of Trolox and newly developed Trolox variants on the maximal activity of complex I, complex IV and citrate synthase
We previously demonstrated in CT5120 cells that Trolox treatment (500 μM, 72 h) increases the maximal biochemical activity (Vmax) of key mitochondrial enzymes like CI, complex IV (CIV) and citrate synthase (CS)12. Here we observed that Trolox and its four variants displayed similar stimulatory effects in CT5120 cells (Fig 4A.). However in case of LS-patient cells, only Trolox and KH003 stimulated CI, CIV and CS activity. KH001 failed to stimulate CI activity, KH002 decreased CI, CIV and CS activity and KH004 failed to stimulate CIV and CS activity (Fig. 4A).
Discussion
The case study presented here illustrates the potential of phenotypic screening assisted by objective multivariate statistical methods. Machine learning techniques are particularly well suited to discriminate between phenotypes that are not easily described by a few parameters33 and were successfully applied previously for phenotypic profiling of drug effects34. The latter study applied a blind approach combining 11 probes to cover maximally the cellular biology and simultaneously testing complete dose response. This untargeted approach was developed in the context of non-specific drug screening.
For analysis of mitochondrial morphology unsupervised learning was used to define six morphological phenotypes in CHO cells and the effects thereupon of caspase inhibition35. Similarly, supervised learning was used to analyze mitochondrial morphology in BEAS-2B cells treated with CI and CII inhibitors36 and combined with fuzzy logic methods to investigate the link between mitochondrial morphology, Δψ and Bax activation37. These two last studies use a supervised strategy for mitochondrial morphology evaluation based on three predefined mitochondrial shapes (tubular, donut-shaped or swollen). This means that training was performed using a set of manually chosen images, which might introduce an undesirable selection bias. Our study differs from the above ones since it does not assume the existence of pre-defined mitochondrial morphology phenotypes. Moreover, as far as we are aware, supervised machine learning has not been previously used to evaluate the effects of small molecule treatment on mitochondrial morphology and Δψ in a human disease model (i.e. LS cells).
To provide numerical data for machine learning, microscopy images of cells stained with the Δψ-sensitive cation TMRM were manually acquired and automatically quantified using a well-established and extensively validated protocol. For every mitochondrial object in each microscopy image a set of 31 descriptors was calculated (“mitochondrial morpho-functional fingerprints”). Relative to a typical healthy control cell line (CT5120), typical LS patient cells (P5175) displayed alterations in 23 out of 31 descriptors. We evaluated various machine learning models for their ability to correctly classify CT5120 and P5175 cells. Although still supervised, the obtained LogReg model is based upon two objectively defined groups (i.e. cells were derived from a distinct healthy subject and an LS patient). A minimum of 21 descriptors was required to maintain the predictive power of the LogReg model. This means that the changes affecting the morpho-functional fingerprints can only be considered from a multivariate point of view. Within the model, three descriptors were of the highest relative importance: “Perimeter ratio” > “Margination” > “Area on box”. Remarkably, with respect to their average median value, the first two descriptors were not significantly different between CT5120 and P5175 cells (Supplementary Table S1). This demonstrates that parameters that appear uninformative using univariate statistics can be highly relevant in a multivariate analysis, which also considers the relationship between individual descriptors. Biologically, an increase in Perimeter ratio and Area on box indicates that mitochondrial objects and/or (sub)networks become smaller (Fig. 2E). Margination is an intensity-dependent parameter that reflects the homogeneity of the TMRM fluorescence between the center and edge of an object (Fig. 2E and Supplementary Table S1). This means that mitochondrial objects and/or (sub)networks with a high Margination value display inhomogeneous TMRM fluorescence, compatible with spatial Δψ inhomogeneities.
In a proof-of-principle study, the developed machine learning strategy was applied to phenotypically classify the effect of the reference compound Trolox and four newly developed Trolox derivatives (KH001-KH004) in P5175 cells. Trolox is a more water-soluble variant of the widely used antioxidant α-tocopherol38. Structurally, Trolox consists of a chroman headgroup and a short side chain (Fig. 3A). Both Trolox and α-tocopherol exert their antioxidant activity by reacting with various ROS and lipid hydroperoxyl (LOO•) radicals via the hydroxyl group at position 639,40. The resulting tocopheryl radical is resonance-stabilized and can be converted back to α-tocopherol by ascorbate leading to formation of an ascorbyl radical. The latter radical species is regenerated by the glutathione (GSH) system40. We here designed Trolox variants in such a way that only its side chain was modified whereas the hydroxyl and the methyl groups of the chroman ring were left unaltered. By doing so, we created Trolox variants with different lipophilicity (and thereby water/lipid solubility, cell partitioning and mitochondriotropic character), potentially affecting antioxidant potency, ROS specificity and biodistribution27,41,42,43,44. Visual inspection of the morpho-functional fingerprints reveals that Trolox and its four derivatives all display mitigating effects (Fig. 3B). However, only KH003-treated cells were classified as CT5120 cells by the trained machine learning model. Predictive cheminformatics analysis revealed that KH003 was chemically most similar to Trolox. In vitro ORAC measurements pointed at KH003 as a slightly more potent antioxidant towards peroxyl radicals than Trolox. Activity analysis showed that Trolox and KH003 stimulated the Vmax of key mitochondrial enzymes (CI, CIV and CS), whereas KH001, KH002, KH004 were less effective. At the cellular level, Trolox and KH001-KH004 reduced the elevated levels of CM-H2DCF oxidizing ROS in various LS patient fibroblasts. In the same cell lines, Trolox and KH004 did not reduce the increased level of HEt oxidizing ROS, whereas KH003 reduced HEt oxidation and KH001 displayed a pro-oxidant effect. These results demonstrate that modification of the Trolox side chain alters its ROS-scavenging properties. Moreover, they support the cheminformatics analysis and machine learning results by highlighting KH003 as displaying the most favorable properties at the cellular level. In summary, we conclude that supervised machine learning is a powerful method to evaluate small molecule effects on mitochondrial morphology and Δψ. Moreover our results suggest that Trolox-derived antioxidants are promising candidate molecules in therapy development for (ROS-related) mitochondrial disorders.
Methods
Cell lines, culture conditions and enzyme activity analysis
Primary human skin fibroblasts were obtained from healthy individuals (CT5119, CT5118, CT5120) and various Leigh syndrome (LS) patients (P) with isolated mitochondrial complex I (CI) deficiency (OMIM 252010) that were previously characterized at the genetic, biochemical and cellular level45. Cells were cultured under standardized conditions (see Supplementary Information for details). The activities of CI, complex IV (CIV) and citrate synthase (CS) were determined in mitochondria-enriched fractions as previously described46.
Live-cell microscopy
Mitochondrial morphology and membrane potential were analyzed in cells stained with the fluorescent cation TMRM (tetramethylrhodamine methyl ester). Cells were visualized using digital imaging microscopy as described in detail in the Supplementary Information. TMRM was used in non-quenching mode47. Reactive oxygen species levels were analyzed using the ROS-sensing reporter molecules 5-(and-6)-chloromethyl-2′, 7′-dichlorodihydro-fluorescein (CM-H2DCF) and hydroethidium (HEt), as described in the Supplementary Information.
ORAC assay
The ORAC assay measures the in vitro peroxyl radical scavenging potential of molecules. The antioxidant Trolox is used as a standard in this assay. For details see the Supplementary Information.
Synthesis of Trolox derivatives
Detailed information regarding the chemical synthesis strategy is provided in the patent application (Int. patent appl. no. PCT/NL2013/050528).
Cheminformatics, data handling, image quantification and machine learning
Physicochemical parameters were predicted and molecule structures were drawn using MarvinSketch 6.1.3 software (ChemAxon Ltd., Budapest, Hungary). Image processing and analysis were performed using Image Pro Premier software (Media Cybernetics, Inc., Bethesda, MD, USA). Data analysis, statistics and machine learning algorithms were programmed using custom scripts in MATLAB 6.1 (The Mathworks Inc., Natick, MA, USA). SPSS statistics 21 (IBM, Armonk, NY, USA) was used to perform hierarchical cluster analysis using average linkage between groups and the squared Euclidian distance. All data is presented as the mean ± SEM (standard error of the mean) unless stated otherwise and statistical significance was evaluated using a Mann-Whitney U test.
References
Smeitink, J., van den Heuvel, L. & DiMauro, S. The genetics and pathology of oxidative phosphorylation. Nat. Rev. Genet. 2, 342–352 (2001).
Mitchell, P. Coupling of phosphorylation to electron and hydrogen transfer by a chemi-osmotic type of mechanism. Nature 191, 144–148 (1961).
Koopman, W. J. H., Willems, P. H. G. M. & Smeitink, J. A. M. Monogenic Mitochondrial Disorders. N. Engl. J. Med. 366, 1132–1141 (2012).
Vafai, S. B. & Mootha, V. K. Mitochondrial disorders as windows into an ancient organelle. Nature 491, 374–383 (2012).
Koopman, W. J. H., Distelmaier, F., Smeitink, J. A. M. & Willems, P. H. G. M. OXPHOS mutations and neurodegeneration. EMBO J. 32, 9–29 (2013).
Iuso, A. et al. Dysfunctions of cellular oxidative metabolism in patients with mutations in the NDUFS1 and NDUFS4 genes of complex I. J. Biol. Chem. 281, 10374–10380 (2006).
Verkaart, S. et al. Superoxide production is inversely related to complex I activity in inherited complex I deficiency. Biochim. Biophys. Acta - Mol. Basis Dis. 1772, 373–381 (2007).
Guillery, O. et al. Modulation of mitochondrial morphology by bioenergetics defects in primary human fibroblasts. Neuromuscul. Disord. 18, 319–330 (2008).
Valsecchi, F. et al. Primary fibroblasts of NDUFS4−/− mice display increased ROS levels and aberrant mitochondrial morphology. Mitochondrion 13, 436–443 (2013).
Koopman, W. J. H. et al. Mitigation of NADH: Ubiquinone oxidoreductase deficiency by chronic Trolox treatment. Biochim. Biophys. Acta - Bioenerg. 1777, 853–859 (2008).
Distelmaier, F. et al. The antioxidant Trolox restores mitochondrial membrane potential and Ca2+ -stimulated ATP production in human complex I deficiency. J. Mol. Med. (Berl). 87, 515–22 (2009).
Distelmaier, F. et al. Trolox-sensitive reactive oxygen species regulate mitochondrial morphology, oxidative phosphorylation and cytosolic calcium handling in healthy cells. Antioxid. Redox Signal. 17, 1657–1669 (2012).
Calkins, M. J., Manczak, M., Mao, P., Shirendeb, U. & Reddy, P. H. Impaired mitochondrial biogenesis, defective axonal transport of mitochondria, abnormal mitochondrial dynamics and synaptic degeneration in a mouse model of Alzheimer's disease. Hum. Mol. Genet. 20, 4515–29 (2011).
Salmi, H., Leonard, J. V., Rahman, S. & Lapatto, R. Plasma thiol status is altered in children with mitochondrial diseases. Scand. J. Clin. Lab. Investig. 72, 152–157 (2012).
Mercer, J. R. et al. The mitochondria-targeted antioxidant MitoQ decreases features of the metabolic syndrome in ATM+/−/ApoE−/− mice. Free Radic. Biol. Med. 52, 841–849 (2012).
Goldschmidt, R. et al. Effects of cytoprotective antioxidants on lymphocytes from representative mitochondrial neurodegenerative diseases. Bioorg. Med. Chem. 21, 969–978 (2013).
Rötig, A., Sidi, D., Munnich, A. & Rustin, P. Molecular insights into Friedreich's ataxia and antioxidant-based therapies. Trends Mol. Med. 8, 221–224 (2002).
Hart, P. E. et al. Antioxidant treatment of patients with Friedreich ataxia: four-year follow-up. Arch. Neurol. 62, 621–626 (2005).
Martinelli, D. et al. EPI-743 reverses the progression of the pediatric mitochondrial disease-Genetically defined Leigh Syndrome. Mol. Genet. Metab. 107, 383–388 (2012).
Bjelakovic, G., Nikolova, D., Gluud, L. L., Simonetti, R. G. & Gluud, C. Mortality in randomized trials of antioxidant supplements for primary and secondary prevention: systematic review and meta-analysis. JAMA 297, 842–857 (2007).
Halliwell, B. Free radicals and antioxidants: updating a personal view. Nutr. Rev. 70, 257–65 (2012).
Pfeffer, G., Majamaa, K., Turnbull, D. M., Thorburn, D. & Chinnery, P. F. Treatment for mitochondrial disorders. Cochrane database Syst. Rev. 4, CD004426 (2012).
Heyland, D. et al. A randomized trial of glutamine and antioxidants in critically ill patients. N. Engl. J. Med. 368, 1489–97 (2013).
Kerr, D. S. Review of clinical trials for mitochondrial disorders: 1997–2012. Neurotherapeutics 10, 307–19 (2013).
Dickinson, B. C. & Chang, C. J. Chemistry and biology of reactive oxygen species in signaling or stress responses. Nat. Chem. Biol. 7, 504–511 (2011).
Sena, L. A. & Chandel, N. S. Physiological roles of mitochondrial reactive oxygen species. Mol. Cell 48, 158–166 (2012).
Lúcio, M. et al. Antioxidant Activity of Vitamin E and Trolox: Understanding of the Factors that Govern Lipid Peroxidation Studies In Vitro. Food Biophys. 4, 312–320 (2009).
Wattamwar, P., Hardas, S. S., Butterfield, D. A., Anderson, K. W. & Dziubla, T. Tuning of the Pro-oxidant and Antioxidant Activity of Trolox Through the Controlled Release from Biodegradable Poly(trolox ester) Polymers. Free Radic. Biol. Med. 49, S184 (2010).
Brigelius-Flohé, R. Vitamin E: The shrew waiting to be tamed. Free Radic. Biol. Med. 46, 543–554 (2009).
Bast, A. & Haenen, G. R. M. M. Ten misconceptions about antioxidants. Trends Pharmacol. Sci. 34, 430–436 (2013).
Feng, Y., Mitchison, T. J., Bender, A., Young, D. W. & Tallarico, J. A. Multi-parameter phenotypic profiling: using cellular effects to characterize small-molecule compounds. Nat. Rev. Drug Discov. 8, 567–578 (2009).
Guyon, I., Weston, J., Barnhill, S. & Vapnik, V. Gene selection for cancer classification using Support Vector Machines. Mach. Learn. 46, 389–422 (2002).
Sommer, C. & Gerlich, D. W. Machine learning in cell biology - teaching computers to recognize phenotypes. J. Cell Sci. 126, 1–11 (2013).
Perlman, Z. E. et al. Multidimensional drug profiling by automated microscopy. Science 306, 1194–1198 (2004).
Peng, J.-Y. et al. Automatic morphological subtyping reveals new roles of caspases in mitochondrial dynamics. PLoS Comput. Biol. 7, e1002212 (2011).
Ahmad, T. et al. Computational classification of mitochondrial shapes reflects stress and redox state. Cell Death Dis. 4, e461 (2013).
Reis, Y. et al. Multi-parametric analysis and modeling of relationships between mitochondrial morphology and apoptosis. PLoS One 7, e28694 (2012).
Kamal-Eldin, A. & Appelqvist, L. A. The chemistry and antioxidant properties of tocopherols and tocotrienols. Lipids 31, 671–701 (1996).
Nishikimi, M. & Machlin, L. J. Oxidation of alpha-tocopherol model compound by superoxide anion. Arch. Biochem. Biophys. 170, 684–9 (1975).
Traber, M. G. & Stevens, J. F. Vitamins C and E: Beneficial effects from a mechanistic perspective. Free Radic. Biol. Med. 51, 1000–1013 (2011).
Cort, W. M. et al. Antioxidant activity and stability of 6-hydroxy-2, 5, 7, 8-tetramethylchroman-2-carboxylic acid. J. Am. Oil Chem. Soc. 52, 174–178 (1975).
Wang, R., Fu, Y. & Lai, L. A New Atom-Additive Method for Calculating Partition Coefficients. J. Chem. Inf. Comput. Sci. 37, 615–621 (1997).
Kaempf-Rotzoll, D. E., Traber, M. G. & Arai, H. Vitamin E and transfer proteins. Curr. Opin. Lipidol. 14, 249–254 (2003).
Horobin, R. W., Trapp, S. & Weissig, V. Mitochondriotropics: A review of their mode of action and their applications for drug and DNA delivery to mammalian mitochondria. J. Control. Release 121, 125–136 (2007).
Distelmaier, F. et al. Mitochondrial complex I deficiency: from organelle dysfunction to clinical disease. Brain 132, 833–842 (2009).
Rodenburg, R. J. T. Biochemical diagnosis of mitochondrial disorders. J. Inherit. Metab. Dis. 34, 283–292 (2011).
Distelmaier, F. et al. Life cell quantification of mitochondrial membrane potential at the single organelle level. Cytometry. A 73, 129–38 (2008).
Author information
Authors and Affiliations
Contributions
L.B., J.S., S.v.E., C.V., M.P., L.M.C.B., J.B., P.W. and W.K. wrote the main manuscript. L.B., J.S., M.P., J.B., L.M.C.B., P.W. and W.K. designed the experiments. L.B., S.v.E., C.V., M.P., J.B. and W.K. analysed the data. L.B., S.v.E., C.V. and W.K. prepared the Figure 1. L.B., W.K. prepared the Figures 2 and 3. L.B., M.P. and W.K. prepared the Figure 4. A.J. and R.R. performed the measurements of maximal biochemical activity of Citrate Synthase and Complexes I and IV. L.B. and W.K. performed the data analysis and the prediction of the physicochemical properties of the compounds. S.v.E., C.V., M.P. and J.B. performed the microscopic and ORAC experiments.
Ethics declarations
Competing interests
This research was supported by the CSBR (Centres for Systems Biology Research) initiative from the Netherlands Organisation for Scientific Research (NWO; No: CSBR09/013V) and a PM-Rare (Priority Medicines Rare disorders and orphan diseases) grant from ZON-MW (Netherlands Organization for Health Research and Development-Medical Sciences, No: 40-41900-98-033), the Energy4All foundation (www.energy4all.eu), the TIM foundation (www.timfoundation.nl) and an equipment grant of NWO (Netherlands Organization for Scientific Research, No: 911-02-008). Part of this research was carried out at the SME Khondrion BV. LB, JS, SvE, CV, MP, JB, PW and WK are (fully or partly) employed by Khondrion BV. AJ, RR and Pr. LMCB declare no potential conflict of interest. The four newly generated antioxidants described in this study are covered by a patent (Int. patent appl. no. PCT/NL2013/050528).
Electronic supplementary material
Supplementary Information
Supplementary_Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Blanchet, L., Smeitink, J., van Emst - de Vries, S. et al. Quantifying small molecule phenotypic effects using mitochondrial morpho-functional fingerprinting and machine learning. Sci Rep 5, 8035 (2015). https://doi.org/10.1038/srep08035
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep08035
This article is cited by
-
Image-based profiling for drug discovery: due for a machine-learning upgrade?
Nature Reviews Drug Discovery (2021)
-
KH176 Safeguards Mitochondrial Diseased Cells from Redox Stress-Induced Cell Death by Interacting with the Thioredoxin System/Peroxiredoxin Enzyme Machinery
Scientific Reports (2018)
-
Deep Analysis of Mitochondria and Cell Health Using Machine Learning
Scientific Reports (2018)
-
Therapeutic effects of the mitochondrial ROS-redox modulator KH176 in a mammalian model of Leigh Disease
Scientific Reports (2017)
-
Computational imaging reveals mitochondrial morphology as a biomarker of cancer phenotype and drug response
Scientific Reports (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.