Abstract
The cyclin B-dependent protein kinase Cdk1 is a master regulator of mitosis and phosphorylates numerous proteins on the minimal consensus motif Ser/Thr-Pro (S/T-P). At least in several proteins, however, not well-defined motifs lacking a Pro in the +1 position, referred herein to as non-S/T-P motifs, have been shown to be phosphorylated by Cdk1. Here we show that non-S/T-P motifs in fact form consensus sequences for Cdk1 and probably play roles in mitotic regulation of physiologically important proteins. First, we show, by in vitro kinase assays, that previously identified non-S/T-P motifs all harbour one or more C-terminal Arg/Lys residues essential for their phosphorylation by Cdk1. Second, using Arg/Lys-scanning oriented peptide libraries, we demonstrate that Cdk1 phosphorylates a minimal sequence S/T-X-X-R/K and more favorable sequences (P)-X-S/T-X-[R/K]2–5 as its non-S/T-P consensus motifs. Third, on the basis of these results, we find that highly conserved linkers (typically, T-G-E-K-P) of C2H2 zinc finger proteins and a nuclear localization signal-containing sequence (matching P-X-S-X-[R/K]5) of the cytokinesis regulator Ect2 are inhibitorily phosphorylated by Cdk1, well accounting for the known mitotic regulation and function of the respective proteins. We suggest that non-S/T-P Cdk1 consensus motifs identified here may function to regulate many other proteins during mitosis.
Similar content being viewed by others
Introduction
In eukaryotic cells, the cyclin B-Cdk1 kinase complex is a master regulator of mitosis and plays roles in chromosome condensation, nuclear envelope breakdown and spindle formation1,2. Cyclin B-Cdk1, hereafter simply called Cdk1, phosphorylates hundreds of different proteins during mitosis, with its activity increasing rapidly as cells progress into prophase and prometaphase3,4. A number of studies, the first ones of which were begun at the end of 1980s, showed, employing various methods, that Cdk1 is a proline-directed kinase and phosphorylates substrates on the minimal consensus motif Ser/Thr-Pro (S/T-P), with the optimal sequence being S/T-P-X-R/K5,6,7,8,9,10. Such an identification of the Cdk1 consensus phosphorylation motif helped determine a number of Cdk1 substrates with S/T-P motifs (ref. 9 and references therein).
As early as in early 1990s, however, a few proteins, including vimentin, desmin and myosin II, were shown to be phosphorylated by Cdk1 on a Ser or Thr residue not followed by a Pro11,12,13,14. These phosphorylation sites lacking a Pro in the +1 position, which hereafter we call non-S/T-P motifs, were considered to be exceptional and regarded as “nonconsensus” Cdk1 sites6,9. Later, however, at least two other proteins, Wee1 and Boi1, were also shown to be phosphorylated by Cdk1 on non-S/T-P motifs15,16. More recently, we showed that the C-terminal tail of Emi2, an inhibitor of the anaphase-promoting complex/cyclosome (APC/C), is phosphorylated by Cdk1 on a non-S/T-P motif17. As these (not well-defined) non-S/T-P motifs are evolutionarily well-conserved at least in their respective protein species, they might have some structural similarity to each other, possibly forming a “consensus” sequence(s).
Besides these Cdk1 substrates having non-S/T-P motifs, a large number of human proteins have been shown to be mitotically phosphorylated apparently on non-S/T-P motifs that do not conform to even any known motifs for non-Cdk1 mitotic kinases18. Specifically, C2H2 zinc finger proteins, which include such proteins as Ikaros and YY1 and represent the largest family of transcription factors19, have been shown to be mitotically phosphorylated on their highly conserved linkers (containing a non-S/T-P motif) and, thereby, to be globally inactivated during mitosis20,21,22. In all of these cases, the kinase(s) responsible for the phosphorylation is not known and is rather thought not to be Cdk1 (because of the absence of a Pro in the phosphorylation sites)18,22. In contrast to these, Ect2, a central cytokinesis regulator23, has recently been shown to be phosphorylated by Cdk1 at prophase for its nuclear export and, hence, for mitotic cell rounding, but with Cdk1 phosphorylation of the conventional S/T-P motifs being insufficient for Ect2 nuclear export24, suggesting the occurrence of a relevant non-S/T-P motif(s) for Cdk1 in Ect2.
In this study, using both in vitro kinase assays of the known Cdk1 substrates with non-S/T-P motifs and Arg/Lys-scanning oriented peptide libraries, we show that Cdk1 phosphorylates a minimal S/T-X-X-R/K sequence and more favorable (P)-X-S/T-X-[R/K]2–5 sequences as its non-S/T-P consensus motifs. Furthermore, based on these results, we find that Cdk1 phosphorylates a non-S/T-P minimal consensus sequence in the linkers of C2H2 zinc finger proteins and also a non-S/T-P optimal consensus sequence overlapping a nuclear localization signal (rich in Arg/Lys) in Ect2 to inhibit Ect2-Importin binding, well accounting for the mitotic regulation and function of the respective proteins. Given these results, Cdk1 could also regulate many other physiologically important proteins by phosphorylating their non-S/T-P consensus motifs.
Results
Importance of a C-terminal Arg/Lys residue(s) for Cdk1 phosphorylation of the known non-S/T-P motifs
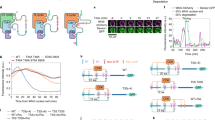
More than two decades ago, three intermediate filament proteins (vimentin, desmin and GFAP) and myosin II (regulatory light chain) were shown to be phosphorylated by Cdk1 on their N-terminal non-S/T-P motifs11,12,13,14,25. More recently, we showed that Emi2, an inhibitor of the APC/C, is phosphorylated on a C-terminal non-S/T-P motif to be inactivated in Xenopus eggs17. To determine whether the non-S/T-P motifs in these proteins (Fig. 1a) make any consensus sequence for phosphorylation by Cdk1, we performed in vitro Cdk1 kinase assays of the respective non-S/T-P motif-spanning peptides (WT or Ala-mutants, fused to glutathione S-transferase or GST) in the presence of [Υ-32P]ATP. Initially, we confirmed that all of the phosphoacceptor Ser or Thr residues in the non-S/T-P motifs tested can be phosphorylated by Cdk1, albeit to different degrees from each other (Fig. 1b). We then examined the requirement of the individual Ser/Thr-surrounding residues of the respective non-S/T-P motifs for their phosphorylation by Cdk1. Notably, an Arg in the +3 position in both vimentin (Fig. 1c) and desmin (Fig. 1d), three Arg/Lys residues in the +2, +3 and +4 positions in myosin II (Fig. 1e) and two Arg/Lys residues in the +4 and +5 positions in Emi2 (Fig. 1g) were all more important for the phosphorylation than the other tested Ser/Thr-surrounding residues of the respective non-S/T-P motifs, while, in GFAP, only two Arg residues in the +3 and +4 positions were important for the phosphorylation (Fig. 1f). Thus, in all the proteins tested, an Arg or Lys residue(s) C-terminal to the phosphoacceptor Ser/Thr is important for Cdk1 phosphorylation of the non-S/T-P motif, although, in most cases, the majority of the other Ser/Thr-surrounding residues tested are also required to some extent. In a previous study26, an Arg in the +3 position of the synthetic vimentin non-S/T-P motif peptide, although followed by two artificially introduced Arg residues, was also shown to be important for its phosphorylation by Cdk1.
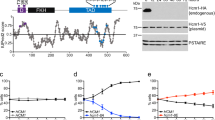
In vitro Cdk1 kinase assays of known non-S/T-P motif sequences.
(a) Non-S/T-P motif sequences of known Cdk1 substrate proteins. For each substrate protein, the phosphoacceptor Ser or Thr in its non-S/T-P motif is shown in parentheses and printed in blue in the sequence, while the Arg or Lys residue in red is a residue important for Cdk1 phosphorylation of the non-S/T-P motif, as revealed below in (c-g). Desmin, myosin II and Emi2 are from Xenopus, while vimentin and GFAP are from human. (b) GST-fused peptides (WT or a Ser/Thr → Ala mutant; for the residues of the peptides, see Methods) of the indicated substrate proteins were incubated with cyclin B1-Cdk1 and [Υ-32P]ATP, subjected to SDS-PAGE and stained with Coomassie brilliant blue (CBB) or autoradiographed (32P). For each GST-fused peptide of each substrate protein, relative intensity of the radioactive signal (WT = 100) is shown at the bottom. (c)–(g) GST-fused peptides (WT or the indicated Ala-mutants) of vimentin (c), desmin (d), myosin II (e), GFAP (f) and Emi2 (g) were subjected to in vitro Cdk1 kinase assays and analyzed as in (b). Full scans of all autoradiographies are included in Supplementary Fig. S4.
Cdk1 phosphorylation of the non-S/T-P motif sequences using Arg/Lys-scanning oriented peptide libraries
The above results indicated that one or more (consecutive) Arg/Lys residues in the +2 to +5 positions are important for Cdk1 phosphorylation of the known non-S/T-P motifs. To test more systematically for the position and number of C-terminal (consecutive) Arg/Lys residues that would be important for such phosphorylation, we used such Arg/Lys-scanning oriented peptide libraries as summarized in Fig. 2a. Each library contained an N-terminal biotin tag, a central Ser or Thr as the phosphoacceptor and a second fixed amino acid(s) (Ala, Pro, or an equimolar mixture (R′) of Arg and Lys) in an appropriate position(s), particularly after the phosphoacceptor site; in all other positions (within the 13-amino acid window) was present a degenerate equimolar mixture of 17 (X0), 16 (X1), 15 (X2) or 14 (X3) amino acids, which are defined respectively in Fig. 2a. The actual (truncated) sequences of the peptide libraries used are shown in Fig. 2b, in which R corresponds to R′ (a mixture of Arg and Lys) in Fig. 2a, while X is either X0, X1, X2 or X3 in Fig. 2a, depending on its corresponding positions in Fig. 2a. These peptide libraries were incubated individually with cyclin B-Cdk1 and [Υ-32P] ATP in solution, transferred to an avidin-coated membrane, washed and imaged with a phosphoimager.
In vitro phosphorylation of non-S/T-P and S/T-P motif sequences using Arg/Lys-scanning oriented peptide libraries.
(a) Schematic representation of Arg/Lys-scanning oriented peptide libraries. See the lower part of the panel for explanation of X0, X1, X2, X3 and R′. Slashes indicate “or”. (b) Each peptide library used (shown only for the -2 to the +6 positions) was incubated with [Υ-32P]ATP and either ERK2 or Cdk1 and analyzed as described in the text and Methods. The radioactive signal of each library (32P) was quantitated and its value is shown just above the signal and in the bar diagram, with the value of XXSPXXXXX being set at 1.0 (mean ± SD, n = 3 for ERK2 and N = 4 for Cdk1). R corresponds to R′ (a mixture of Arg and Lys) in (a), while X is either X0, X1, X2 or X3 in (a), depending on its corresponding positions in (a).
We first analyzed phosphorylation of the peptide libraries by ERK2-MAPK kinase, another proline-directed kinase27. ERK2 phosphorylated the conventional S/T-P and S/T-P-X-R/K motifs for Cdk1 to the same extent, but not detectably other (Arg/Lys-containing) non-S/T-P motifs (Fig. 2b). Importantly, the presence of a Pro in the −2 position considerably (~4-fold) enhanced ERK2 phosphorylation of the S/T-P and S/T-P-X-R/K motifs (Fig. 2b), consistent with previous reports28,29. Furthermore, Ser was a slightly better ERK2 substrate than Thr as the phosphoacceptor (Fig. 2b), also consistent with previous reports29,30. Thus, these results confirm the previously identified consensus sequence for ERK2 as P-X-S/T-P28, showing the validity of using the present peptide libraries and method for examining the phosphorylation motifs of Cdk1. So we next analyzed phosphorylation of the peptide libraries by Cdk1. In addition to the conventional S/T-P and S/T-P-X-R/K motifs, Cdk1 was able to phosphorylate various Arg/Lys-containing non-S/T-P motifs to different degrees (see below for a detailed description of these results) (Fig. 2b). Thus, obviously, Cdk1 differs from ERK2 in that it can phosphorylate even the non-S/T-P motifs.
Identification of non-S/T-P motif consensus sequences for Cdk1
From the data in Fig. 2b, we compared Cdk1 phosphorylations of the various Arg/Lys-containing non-S/T-P motifs by classifying them into several categories. When a single Arg/Lys was present in either of the five positions after the phosphoacceptor Ser, S-X-X-R/K, containing an Arg/Lys in the +3 position, was significantly most strongly phosphorylated, albeit to a much lesser extent than the conventional minimal consensus sequence S-P (Fig. 3a), suggesting that S-X-X-R/K may be the non-S/T-P minimal consensus sequence for Cdk1. As for two and three (consecutive) Arg/Lys residues, S-X-[R/K]2 and S-X-[R/K]3 were significantly much more strongly phosphorylated than the other [R/K]2- and [R/K]3-containing sequences, respectively, albeit still to a lesser extent than the conventional minimal consensus S-P (Fig. 3b, c), but much more strongly than S-X-X-R/K (see Figs 2b and 3a). These results indicate that, when the +1 position is not an Arg/Lys, a +2 Arg/Lys, but not a +4 or +5 Arg/Lys, greatly enhances the phosphorylation based on the +3 Arg/Lys. Furthermore, as for four and five consecutive Arg/Lys residues, S-X-[R/K]4 was a much better substrate than S-[R/K]4, as expected and was somewhat better than even the conventional minimal sequence S-P (Fig. 3d), while S-X-[R/K]5 was much better than both S-[R/K]5 and S-P, being phosphorylated to the same extent as the conventional S/T-P optimal sequence S-P-X-R/K (Fig. 3e). These results and the data summarized in Fig. 3f clearly show that, when present consecutively (on a +2/+3 Arg/Lys-background), either of the Arg/Lys residues in the +4, +5 and +6 positions has a positive and additive effect (about 3-fold each, under the present experimental conditions) on the +2/+3 Arg/Lys-based phosphorylation, whereas an Arg/Lys in the +1 position has a negative (less than 0.5-fold) effect (compare SXRR with SRRR, SXRRR with SRRRR and SXRRRR with SRRRRR in Fig. 3f).
Identification of non-S/T-P consensus sequences for Cdk1.
Data are taken from Fig. 2b (mean ± SD, n = 4) for comparison of the effects of the position of a single Arg/Lys residue (a) or two to five consecutive Arg/Lys residues (b)-(e), the number of (consecutive) Arg/Lys residues (f), the presence or absence of a -2 Pro (g) and the identity of the phosphoacceptor as Ser or Thr (h), on Cdk1 phosphorylation of the non-S/T-P sequences. In (g) and (h), two non-S/T-P or S/T-P sequences to be compared directly are vertically lined. In each panel, the phosphorylation level of XXSPXXXXX is also shown for comparison and is set at 1.0; note, however, that its scales are different from each other between (a), (b) and (c), (d)–(f) and (g) and (h). *P < 0.05; **P < 0.01.
As a Pro in the -2 position has been shown to positively influence the phosphorylation of the conventional S/T-P motif by Cdk1 (ref. 10; see also Fig. 2b), as for phosphorylation by ERK2 (ref. 28; see also Fig. 2b), we also compared Cdk1 phosphorylations of the non-S/T-P motifs with and without a -2 Pro. Interestingly, the presence of a -2 Pro increased the phosphorylation of all S-X-[R/K]2–5 sequences each about 2-fold and, notably, P-X-S-X-[R/K]5 was a 2-fold better substrate than even the conventional S/T-P optimal sequence S-P-X-R/K (Fig. 3g). Finally, Ser was a ~2-fold better substrate than Thr as the phosphoacceptor in both the non-S/T-P and S/T-P motifs (Fig. 3h), as shown recently for the S/T-P motif30. Thus, together with the results of in vitro Cdk1 kinase assays of previously identified non-S/T-P motifs (Fig. 1), these results show that Cdk1 phosphorylates a minimal sequence S/T-X-X-R/K and (much) more favorable sequences (P)-X-S/T-X-[R/K]2–5 as its non-S/T-P consensus motifs.
Cdk1 phosphorylates highly conserved linkers (T-G-E-K-P) of C2H2 zinc finger proteins
Having identified non-S/T-P consensus sequences for Cdk1, we next addressed whether the minimal consensus sequence S/T-X-X-R/K would occur in any physiologically important proteins. While such a sequence can be phosphorylated by Cdk1 at least in several proteins, including Wee1 (ref. 15) and Boi1 (ref. 16) as well as vimentin and desmin (Fig. 1), a large number of similar sequences (S-A/G-X-R/K) have been shown to be phosphorylated by an unknown kinase(s) in human mitotic cells18. Specifically, short, highly-conserved linkers (matching or resembling T-G-E-K-P) of C2H2 zinc finger proteins, which include such proteins as Ikaros, Sp1 and YY1 and represent the largest family of transcription factors19, have been shown to be mitotically phosphorylated by an unknown kinase(s), thereby causing inactivation of the zinc fingers during mitosis20,21,22. We therefore investigated whether the (multiple) linkers of Ikaros, Sp1 and YY1 (Fig. 4a) would be phosphorylated by Cdk1. For this, first we performed in vitro kinase assays of the respective linker-spanning peptides (WTs or Ser/Thr → Ala mutants, fused to GST), using [Υ-32P] ATP and either Cdk1 or other mitotic kinases, Plk1 and Aurora B, which are localized at kinetochores (potentially with C2H2 transcription factors) in prometaphase and metaphase2,10. Importantly, all of the phosphoacceptor Ser or Thr residues (seven in total) in the linkers of Ikaros/Sp1/YY1, except Thr378 in YY1, were phosphorylated by Cdk1, mostly if not very strongly, whereas none of them and only Ser196 in Ikaros were phosphorylated by Plk1 and Aurora B, respectively (Fig. 4b). Somewhat interestingly, Plk1 was able to (weakly) phosphorylate most of the Ikaros/Sp1/YY1 peptides at some other sites than their linker phosphoacceptors (Fig. 4b), although usually its activity requires priming phosphorylation of the substrates31. Thus, among the mitotic kinases tested, only Cdk1 can phosphorylate nearly all linkers of Ikaros/Sp1/YY1. Then we also asked whether an Arg/Lys in the +3 position of the respective linkers would be important for their phosphorylation by Cdk1. All the (seven) Arg/Lys residues in the linkers of Ikaros/Sp1/YY1, except Arg381 in YY1, were required for Cdk1 phosphorylation of the respective linkers (Fig. 4c–e), similar to and nearly to the same extent as, their respective phosphoacceptor Ser/Thr residues (Fig. 4b–e). Furthermore, in the linker-spanning peptide (containing S196/K199) of Ikaros, Lys199 was most important, among the seven Ser196-surrounding residues tested, for Ser196 phosphorylation by Cdk1 (Supplementary Fig. S1a). Thus, in nearly all cases, the S/T-X-X-R/K motif (typically, T-G-E-K) in the linker can apparently serve as a Cdk1 phosphorylation site in vitro.
Cdk1 phosphorylation of the linkers of C2H2 zinc finger proteins.
(a) Linker sequences of (human) Ikaros, Sp1 and YY1. For each linker, the phosphoacceptor Ser or Thr is shown in parentheses and printed in blue in the sequence, while the Arg/Lys in the +3 position in red. (b) GST-fused linker-spanning peptides (WT or a Ser/Thr → Ala mutant; for the residues of the peptides, see Methods) of the indicated zinc finger proteins were subjected to in vitro kinase assays by using [Υ-32P]ATP and either Cdk1, Plk1 or Aurora B. For each linker peptide of each protein, relative intensity of the radioactive signal (WT = 100) is shown at the bottom. (c)–(e) In vitro Cdk1 kinase assays of the linker-spanning peptides (WT or the indicated Ala-mutants) of Ikaros (c), Sp1 (d) and YY1 (e). (f) Flag/His6-tagged YY1 protein (WT or T348A) was incubated with either interphase (I) or M phase (M) extracts (from Xenopus eggs) in the presence or absence of roscovitine (Ros; 300 μM), BI2536 (BI; 15 μM) or ZM447439 (ZM; 30 μM) and then subjected to Flag-immunoprecipitation followed by immunoblotting with the indicated antibodies. In input, the arrowhead shows Flag/His6-tagged YY1(WT). Full scans of all autoradiographies and immunoblots are included in Supplementary Fig. S5.
We then asked whether mitotic phosphorylation of the linkers would require Cdk1 activity. To this end, we utilized anti-HpTGEKP antibody (α-HpTGEKP), which can recognize a mitotically- or Cdk1-phosphorylated Thr348 in the linker of YY1 (ref. 22 and Supplementary Fig. S1b). When incubated with M phase but not interphase extracts from Xenopus eggs and then immunoprecipitated for immunoblotting with α-HpTGEKP, recombinant full-length YY1 protein (tagged with Flag) was found to be phosphorylated at Thr348; importantly, however, this phosphorylation was strongly inhibited by treating the M phase extracts with the Cdk1 inhibitor roscovitine, but not with the Plk1 inhibitor BI2536 or the Aurora B (and A) inhibitor ZM447439 (Fig. 4f). Treating the M phase extracts with another (and more specific) Cdk1 inhibitor RO-3306 (ref. 32) also strongly inhibited Thr348 phosphorylation of YY1 (Supplementary Fig. S1c). Thus, Cdk1 is likely to be specifically required for mitotic phosphorylation of the linker (Thr348) in YY1. In these experiments, we also observed that α-HpTGEKP recognized numerous endogenous proteins (of diverse sizes), perhaps mostly C2H2 zinc finger proteins22, in M phase extracts in either a roscovitine- or an RO-3306-sensitive manner (see input of Fig. 4f and Supplementary Fig. S1c). These results, together with the results of in vitro kinase assays (Fig. 4b–e), suggest that Cdk1 is the principal kinase that phosphorylates the linkers (containing the non-S/T-P minimal consensus sequence S/T-X-X-R/K) of C2H2 zinc finger proteins, thereby perhaps globally inactivating the C2H2 family during mitosis20,22.
Cdk1 phosphorylates and inhibits a nuclear localization signal-containing sequence (P-X-S-X-[R/K]5) in Ect2
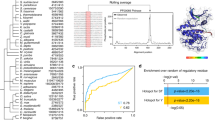
We also examined whether Cdk1 would phosphorylate more favorable non-S/T-P consensus sequences (P)-X-S/T-X-[R/K]2–5 in any physiologically important proteins (as for myosin II; Fig. 1). According to the phosphoproteomic analysis of human cells18, mitotic phosphorylation of such preferred non-S/T-P sequences, although mediated by an unknown kinase(s), occurs in numerous proteins, of which about 20 sequences (or proteins) have four or more consecutive Arg/Lys residues after the Ser/Thr + 1 position. In at least five of these 20 sequences, which include Ect2 and Cullin 4B, the four or more consecutive Arg/Lys residues are suggested to function as a (classical) nuclear localization signal (NLS, typically consisting of four Arg/Lys residues; ref. 33)34,35. Interestingly, the Cdk1 substrate Ect2, a key cytokinesis regulator23, has recently been shown to be exported from the nucleus at prophase in a Cdk1-dependent manner and, thereby, to play an essential role in mitotic cell rounding24. As phosphorylation of a Ser/Thr residue(s) near an NLS can inhibit NLS activity in certain proteins36, we suspected that Cdk1 phosphorylation of the well-conserved, NLS-containing non-S/T-P optimal sequence P-X-S-X-[R/K]5 in Ect2 (Figs 3g and 5a) might inhibit its NLS activity, thereby perhaps causing (or allowing) nuclear export of Ect2 at prophase for mitotic rounding. First, we confirmed, by in vitro kinase assays, that Ser345 in the P-X-S-X-[R/K]5 sequence of (human) Ect2 can be readily phosphorylated by Cdk1 (Fig. 5b). We also confirmed that the five consecutive Arg/Lys residues (or NLS) in the sequence are required for Cdk1 phosphorylation of Ser345 (Fig. 5b). Thus, clearly, Cdk1 can recognize and phosphorylate the NLS-containing non-S/T-P (optimal) consensus sequence P-X-S-X-[R/K]5 in Ect2. It should be noted, however, that, in these experiments, Thr342 followed by a Pro and preceding Ser345 (see Fig. 5a) was mutated to Ala, because, reportedly, it can also be mitotically phosphorylated, perhaps by Cdk1 (ref. 18; see also below).
Cdk1 phosphorylation and inhibition of the NLS-containing non-S/T-P consensus sequence of Ect2.
(a) Conservation of the NLS-containing non-S/T-P consensus sequence in Ect2 proteins from various species. (b) GST-fused peptides (WT or the indicated Ala-mutants; residues 334–363) of human Ect2 were subjected to in vitro Cdk1 kinase assays. In all constructs in this experiment, Thr342 preceding Ser345 (a) was replaced by Ala, because, reportedly, it can also be mitotically phosphorylated18, perhaps by Cdk1. RKRRR/5A, an NLS mutant. (c) Flag/His6-tagged Ect2-F constructs (WT or the indicated Ala-mutants; residues 328–388) were incubated with either interphase (I) or M phase (M) extracts (from Xenopus eggs) and then subjected to Flag-immunoprecipitation followed by λ phosphatase treatment and immunoblotting with the indicated antibodies. In the bar diagram, the levels of coimmunoprecipitated Importin β were quantitated and normalized to Ect2-F constructs; the relative values to Importin β coprecipitated with Ect2-F(WT) in interphase extracts are shown (mean ± SD, n = 4). KRR/3A, an NLS mutant. (d) Flag/His6-tagged Ect2-F constructs (WT or the indicated Asp-, Glu- or Ala-mutants) were incubated with interphase (I) extracts and analyzed as in (c) (mean ± SD, n = 4). (e) Flag/His6-tagged Ect2-F constructs (WT or the indicated Ala-mutant) were incubated with M phase extracts in the presence or absence of roscovitine (Ros; 300 μM) or RO-3306 (RO; 300 μM) and analyzed as in (c); in the bar diagram, the level of Importin β coimmunoprecipitated with Ect2-F(T342A/S345A) is set at 1.0 (mean ± SD, n = 3). Full scans of all autoradiographies and immunoblots are included in Supplementary Fig. S6.
We then asked whether Cdk1 phosphorylation of Ser345 would indeed inhibit the NLS activity of Ect2, by testing whether Ser345 phosphorylation can inhibit the binding of Ect2 to Importin, a complex of Importin α and β that binds an NLS and imports the NLS-containing protein to the nucleus33. For this, first we incubated Flag-tagged Ect2 fragments (residues 328-388: Ect2-F) with either interphase or M phase extracts from Xenopus eggs and subjected them to Flag-immunoprecipitation followed by immunoblotting for (endogenous) Importin β. Ect2-F(WT) was complexed with Importin β in interphase but not M phase extracts, whereas NLS-mutated Ect2-F(KRR/3A) showed no Importin β binding even in interphase extracts (Fig. 5c), suggesting that, in M phase extracts, the NLS activity of Ect2-F(WT) was inhibited possibly by phosphorylation of Ser345. Consistently, when Ser345 was mutated to non-phosphorylatable Ala, Ect2-F(S345A) was able to bind Importin β even in M phase extracts, albeit to a somewhat lesser (~60%) extent than was Ect2-F(WT) in interphase extracts (Fig. 5c). This partial restoration of Ect2-F(S345A)-Importin β binding in M phase was, however, most likely owing to yet another phosphorylation at Thr342 preceding Ser345 (see above), because Ect2-F(T342A) and Ect2-F(T342A/S345A) were able to restore their bindings to Importin β partially and completely, respectively (Fig. 5c). Furthermore and importantly, when Thr342 and Ser345 were replaced by phosphomimetic Glu and Asp, respectively, the binding of Ect2-F(T342E/S345D), but not necessarily Ect2-F(T342E) or Ect2-F(S345D), to Importin β was nearly completely (>95%) inhibited in interphase extracts (Fig. 5d). Finally, as was expected, treating the M phase extracts with the Cdk1 inhibitors roscovitine or RO-3306 was able to restore Ect2-F(WT)-Importin β binding almost to the level of Ect2-F(T342A/S345A)-Importin β binding in untreated M phase extracts (Fig. 5e). Thus, these results do suggest that Cdk1 phosphorylation of the P-X-S345-X-[R/K]5 sequence (and Thr342) inhibits Ect2 from binding Importin, probably accounting for the Cdk1-dependent nuclear export of Ect2 at prophase for mitotic rounding24. We also observed that Importin β binding of Cullin 4B (containing a P-X-S-X-[R/K]4 sequence), a scaffold protein of cullin-RING ubiquitin ligase complexes37, can be inhibited by a similar mechanism (although the biological significance for this is not known) (Supplementary Fig. S2). Thus, these results would seem to suggest a fairly general role for Cdk1 in regulating the NLS activity of NLS-containing non-S/T-P consensus motifs.
Discussion
Using both known cyclin B-Cdk1 substrates having non-S/T-P motifs and Arg/Lys-scanning oriented peptide libraries, here we have identified non-S/T-P consensus sequences for Cdk1 as (P)-X-S/T-X-[R/K]2–5 with the minimal sequence S/T-X-X-R/K. In these experiments, we simply measured the extents or levels, not the rates or Km, of Cdk1 phosphorylation of the substrate peptides, yet these assays did give highly significant differences in phosphorylation among the different substrates. Notably, S-X-[R/K]4 or 5, but not S-X-[R/K]2 or 3, can be phosphorylated by Cdk1 more strongly than the conventional minimal consensus sequence S/T-P, while P-X-S-X-[R/K]5 is a better substrate for Cdk1 than even the conventional optimal sequence S/T-P-X-R/K. Thus, the increasing numbers of consecutive Arg/Lys residues after the +1 position and, to a somewhat lesser extent, a Pro in the -2 position have a profound positive effect on Cdk1 phosphorylation of the non-S/T-P motifs (we have not tested for motifs having more than five consecutive Arg/Lys residues, which are absent in the human mitotic phosphoproteome18). Of course, the non-S/T-P minimal consensus sequence S/T-X-X-R/K is much less strongly phosphorylated by Cdk1 than the conventional S/T-P motifs as well as the (P)-X-S/T-X-[R/K]2–5 sequences (Figs 2 and 3), but it seems to become a moderate substrate for Cdk1, depending on the other phosphoacceptor Ser/Thr-surrounding residues (Fig. 1b-d and Supplementary Fig. S1a). Given these results, both of the two motifs, S-A/G-X-R/K and P-X-S-X-(R/K)-R/K-(R), that were shown to be very frequently phosphorylated in human mitotic cells18, could primarily be a substrate for Cdk1.
In our additional experiments, we have also observed that the presence of consecutive Arg/Lys residues, as well as a Pro in the −2 position, can substantially increase Cdk1 phosphorylation of even the conventional S/T-P motif (Supplementary Fig. S3). This result is consistent, at least in part, with the recent proposal that Cdk1 phosphorylates a more expanded sequence P/C/X-X-S/T-P-X-R/K-K as its S/T-P optimal phosphorylation motif10. Structurally, the consecutive basic Arg/Lys residues in the non-S/T-P consensus motifs might interact with acidic residues in cyclin B, as suggested for a +4 Lys in the expanded S/T-P consensus sequence10. Such potential interactions with cyclin B and perhaps also some other interactions with Cdk1, as for the interaction of a +3 Arg/Lys in the S/T-P motif with Cdk2 (ref. 38), could facilitate Cdk1 phosphorylation of the phosphoacceptor Ser/Thr even in the absence of a +1 Pro, which likely interacts indirectly with the activation loop of Cdk1 (refs 10, 38). Regardless, given their very close structural (and in vitro substrate-specificity) similarities to Cdk1 (refs 29, 39), some other cyclin-dependent kinases, particularly Cdk2, could also phosphorylate, at least in part, such non-S/T-P consensus motifs as identified here, although primarily in different substrates and in different stages of the cell cycle than Cdk1.
On the basis of our defined non-S/T-P consensus sequences for Cdk1, we found that both the highly conserved linkers (T-G-E-K-P or its analogs) of C2H2 zinc finger proteins, including Ikaros, Sp1 and YY1 and the NLS-containing sequence (matching P-X-S/T-X-[R/K]5) of Ect2 are phosphorylated by Cdk1. Phosphorylation at such sites is most probably indispensable for global inactivation of the C2H2 family during mitosis20,22,40 and is likely responsible for the Cdk1-dependent nuclear export of Ect2 for mitotic cell rounding24, which is supported by our findings that Cdk1 phosphorylation of the site can inhibit Ect2-Importin binding. It is possible that, besides cyclin B-Cdk1, cyclin A-Cdk2, whose activity peaks at the onset of mitosis4, also contributes to the phosphorylation of zinc finger proteins and Ect2; however, this contribution would be rather small if at all, given the very large inhibitory effect of the fairly Cdk1-specific inhibitor RO-3306 on the phosphorylation (Supplementary Fig. S1c and Fig. 5e). Thus, it seems likely that Cdk1 phosphorylation of the non-S/T-P consensus sequences can play considerable roles in mitotic regulation of physiologically important proteins. Indeed, given the occurrence of a large number of mitotically phosphorylated non-S/T-P Cdk1 consensus sequences in human cells18 (see also the above discussion), many proteins of physiological importance could be phosphorylated by Cdk1 at such non-S/T-P consensus sequences for their mitotic regulations. In this context, we note the recent report showing that Ser251 required for mitotic activation of FoxM1B lies in a sequence similar to S-X-[R/K]3, is mitotically phosphorylated and can be phosphorylated by Cdk1 in vitro (although it is not considered to be a direct substrate for Cdk1)41.
Why are there two different (although partially overlapping) types of phosphorylation motifs for Cdk1, namely, the conventional S/T-P motif and the non-S/T-P motif, which includes the currently defined Arg/Lys-containing motif and perhaps also an Arg/Lys-lacking motif, which can be seen in certain proteins15,42? Most simply, the S/T-P and the non-S/T-P motifs may be to function as the primary and the secondary Cdk1 substrates, respectively, as suggested by the (generally) stronger Cdk1 phosphorylation of the former motif than the latter (Fig. 2 and Supplementary Fig. S3). In this case, such factors as a cyclin-docking motif (RXL) in protein substrates and a Cdk1 accessory protein Cks1 could differently regulate Cdk1 phosphorylations of the two motifs (ref. 43 and references therein). Furthermore and interestingly, multiple S/T-P motifs are often clustered in the disordered regions of mitotic proteins44 and, in some cases, their Cdk1 phosphorylation is required for Cdk1 phosphorylation of the non-S/T-P motifs in the same protein15,16,17; notably, in the case of Emi2, the N-terminal S/T-P motifs (phosphorylated by Cdk1) bind cyclin B-Cdk1 itself, which in turn phosphorylates a C-terminal non-S/T-P motif17. Thus, in these cases, the S/T-P motifs could function to locally concentrate Cdk1 for phosphorylation of the non-S/T-P motifs. Besides these likely possibilities, it is also possible that the S/T-P motif and the non-S/T-P motif occur for them (or their functions) to be differently regulated on the level of dephosphorylation, as exemplified by their different sensitivities to PP2A in the case of Emi2 (ref. 17).
Finally, our definition of the non-S/T-P consensus sequences for Cdk1 could help identify many new Cdk1 substrates and their phosphorylation sites (as partly discussed above). In fact, in many previously identified Cdk1 substrates with conventional S/T-P motifs, such non-S/T-P consensus sequences as we identified here could have been overlooked because of their lacking a “canonical” Pro in the +1 position of the phosphoacceptor Ser/Thr. Thus, it seems to be worth reconsidering Cdk1 phosphorylation sites in so far examined/identified Cdk1 substrates.
Methods
Xenopus eggs and egg extracts
All experiments using Xenopus were approved by the Institutional Animal Care and Use Committee of Kyushu University (Fukuoka, Japan) and carried out in accordance with the guidelines set by this committee. CSF extracts, referred to as M phase extracts in the text, were prepared from unfertilized Xenopus eggs, as we described previously45, while interphase extracts were prepared from CSF extracts by the addition of 600 μM CaCl2 and 100 μg/ml cycloheximide, followed by incubation for 15 min at 23°C.
Biochemical reagents
Sources of major biochemical reagents were as follows: roscovitine (Calbiochem); RO-3306 (Calbiochem); ZM-447439 (Focus Biomolecules); BI2536 (JS Research Chemicals Trading); human recombinant cyclin B1-Cdk1 complex (Upstate); human recombinant ERK2 protein (abcam); human recombinant Aurora B-INCENP complex (Carna Biosciences); human recombinant Plk1 protein (Carna Biosciences); GST-Accept (Nacalai); His-Accept (Nacalai).
cDNAs and mutagenesis
The cDNAs encoding Xenopus desmin (residues 1–21), myosin II (3–32) and Emi2 peptides (629–651) were isolated from a Xenopus ovarian cDNA library, while the cDNAs encoding human vimentin (residues 34–54), GFAP (2–26), Ikaros (128–152, 156–180 and 184–208), Sp1b (635–659 and 662–686), YY1 (336–360 and 366–390) and Ect2 peptides (334–363) were isolated from a HEK293 cell cDNA library; these cDNAs were subcloned into the pGEX-3X plasmid vector encoding GST (GE Helthcare). The cDNAs encoding human Ect2 fragments (residues 328–388), Cullin 4B fragments (1–50) and full-length YY1 were isolated from a HEK293 cell cDNA library; these cDNAs were subcloned into the Flag/His6-tag-encoding pCold plasmid vector (Takara). Point mutations and deletions were generated by site-directed mutagenesis using appropriate primers (shown in Supplementary Table S1), a KOD-Plus-Neo kit (Toyobo) and a DpnI enzyme (NEB).
Recombinant proteins
GST-fused peptides and Flag/His6-tagged proteins, encoded by the above-described cDNAs, were bacterially expressed and purified by using GST-Accept (Nacalai) and His-Accept (Nacalai), respectively, according to the manufacture's instructions.
In vitro kinase assays
For phosphorylation of GST-fused peptides, 2 μg each of them was mixed with either 100 ng of cyclin B1-Cdk1 protein, 50 ng of Plk protein, or 50 ng of Aurora B(-INCENP) protein in 20 μl of a kinase buffer (10 mM HEPES-KOH, pH 7.5, 100 mM KCl, 20 mM MgCl2) supplemented with 50 μM cold ATP and 5 μCi of [Υ-32P]ATP, incubated for 20 min at 30°C and then subjected to SDS-PAGE.32P-labeled GST-fused peptides were analyzed by using the FLA-5000 (FUJI FILM) and quantified with the Image Gauge (FUJI FILM). GST-fused YY1 peptide (0.5 μg) was also mixed with 100 ng of cyclin B1-Cdk1 protein in 20 μl of the above-described kinase buffer supplemented with 1 mM cold ATP, incubated for 30 min at 30°C and analyzed by immunoblotting with appropriate antibodies.
In vitro phosphorylation of oriented peptide libraries
The oriented peptide libraries (shown in Fig. 2 and Supplementary Fig. S3) were made to our order by Sigma. For phosphorylation of each peptide library, 31.25 μM peptide library was mixed with 80 ng of either cyclin B1-Cdk1 protein or ERK2 protein in 16 μl of a kinase buffer (50 mM Tris-HCl, pH 7.5, 40 mM NaCl, 10 mM MgCl2, 1 mM DTT, 0.1% Tween 20) supplemented with 100 μM cold ATP and 25 μCi of [Υ-32P]ATP and then incubated for 4 hours at 30°C, essentially as described previously10. After the addition of 200 mM cold ATP, 4 μl of each reaction was transferred to a streptavidin-coated membrane (Promega), which was then washed three times with a buffer containing 10 mM Tris-HCl (pH 7.5), 140 mM NaCl and 1.0% SDS, three times with 2 M NaCl, twice with 2 M NaCl containing 1% H3PO4 and once with distilled water. Each32P-labeled peptide library was analyzed by using the FLA-5000 (FUJI FILM) and quantified with the Image Gauge (FUJI FILM).
Immunoprecipitation
Either M phase extracts or interphase extracts from Xenopus eggs, routinely 60 μl, were incubated with 0.8 μg of Flag/His6-tagged Ect2 or Cullin 4B fragments or with 2.1 μg of Flag/His6-tagged full-length YY1 protein for 10 min at 23°C and then subjected to immunoprecipitation with anti-Flag M2 affinity gels (Sigma) for 20 min at 4°C with constant mixing. The gels were washed three times with 100 μl of a buffer (20 mM Tris-HCl, pH 7.5, 100 mM NaCl, 5 mM 6DAP, 10 mM EDTA, 5 mM NaF, 1 mM Na3VO4, 1 μM okadaic acid, 20 μM leupeptin, 2 μM pepstatin, 10 μg/ml aprotinin, 400 μM PMSF, 1 mM DTT, 0.1% Tween) at 4°C. The immunoprecipitates were treated or not treated with λ phosphatase before being subjected to immunoblotting45.
Antibodies and immunoblotting
Immunoblotting was performed by using anti-Flag M2 antibody (Sigma; F1804), anti-Xenopus cyclin B1 antibody (gift from J. Maller), anti-Importin β antibody (abcam; ab36775), or anti-HpTGEKP motif antibody (Millipore; ABE319), essentially as we described previously45; dilutions of the respective antibodies are presented in Supplementary Table S2.
Statistical analysis
Data are expressed as mean ± standard deviation (SD). P-values were calculated using the Student's t-test with MS Excel software. The results were considered significantly different at P < 0.05.
References
Nurse, P. Universal control mechanism regulating onset of M-phase. Nature 344, 503–508 (1990).
Nigg, E. A. Mitotic kinases as regulators of cell division and its checkpoints. Nat. Rev. Mol. Cell Biol. 2, 21–32 (2001).
Ubersax, J. A. et al. Targets of the cyclin-dependent kinase Cdk1. Nature 425, 859–864 (2003).
Pines, J. & Rieder, C. L. Re-staging mitosis: a contemporary view of mitotic progression. Nat. Cell Biol. 3, E3–6 (2001).
Moreno, S. & Nurse, P. Substrates for p34cdc2: in vivo veritas? Cell 61, 549–551 (1990).
Nigg, E. A. Cellular substrates of p34(cdc2) and its companion cyclin-dependent kinases. Trends Cell Biol. 3, 296–301 (1993).
Songyang, Z. et al. Use of an oriented peptide library to determine the optimal substrates of protein kinases. Curr. Biol. 4, 973–982 (1994).
Holmes, J. K. & Solomon, M. J. A predictive scale for evaluating cyclin-dependent kinase substrates. A comparison of p34cdc2 and p33cdk2. J. Biol. Chem. 271, 25240–25246 (1996).
Errico, A., Deshmukh, K., Tanaka, Y., Pozniakovsky, A. & Hunt, T. Identification of substrates for cyclin dependent kinases. Adv. Enzyme Regul. 50, 375–399 (2010).
Alexander, J. et al. Spatial exclusivity combined with positive and negative selection of phosphorylation motifs is the basis for context-dependent mitotic signaling. Sci. Signal. 4, ra42; 10.1126/scisignal.2001796 (2011).
Pines, J. & Hunter, T. p34cdc2: the S and M kinase? New Biol. 2, 389–401 (1990).
Chou, Y. H., Ngai, K. L. & Goldman, R. The regulation of intermediate filament reorganization in mitosis. p34cdc2 phosphorylates vimentin at a unique N-terminal site. J. Biol. Chem. 266, 7325–7328 (1991).
Kusubata, M. et al. Cdc2 kinase phosphorylation of desmin at three serine/threonine residues in the amino-terminal head domain. Biochem. Biophys. Res. Commun. 190, 927–934 (1993).
Satterwhite, L. L. et al. Phosphorylation of myosin-II regulatory light chain by cyclin-p34cdc2: A mechanism for the timing of cytokinesis. J. Cell Biol. 118, 595–605 (1992).
Harvey, S. L., Charlet, A., Haas, W., Gygi, S. P. & Kellogg, D. R. Cdk1-dependent regulation of the mitotic inhibitor Wee1. Cell 122, 407–420 (2005).
McCusker, D. et al. Cdk1 coordinates cell-surface growth with the cell cycle. Nat. Cell Biol. 9, 506–515 (2007).
Isoda, M. et al. Dynamic regulation of Emi2 by Emi2-bound Cdk1/Plk1/CK1 and PP2A-B56 in meiotic arrest of Xenopus eggs. Dev. Cell 21, 506–519 (2011).
Dephoure, N. et al. A quantitative atlas of mitotic phosphorylation. Proc. Natl Acad. Sci. USA 105, 10762–10767 (2008).
Wolfe, S. A., Nekludova, L. & Pabo, C. O. DNA recognition by Cys2His2 zinc finger proteins. Annu. Rev. Biophys. Biomol. Struct. 29, 183–212 (2000).
Dovat, S. et al. A common mechanism for mitotic inactivation of C2H2 zinc finger DNA-binding domains. Genes Dev. 16, 2985–2990 (2002).
Rizkallah, R. & Hurt, M. M. Regulation of the transcription factor YY1 in mitosis through phosphorylation of its DNA-binding domain. Mol. Biol. Cell 20, 4766–4776 (2009).
Rizkallah, R., Alexander, K. E. & Hurt, M. M. Global mitotic phosphorylation of C2H2 zinc finger protein linker peptides. Cell Cycle 10, 3327–3336 (2011).
Tatsumoto, T., Xie, X., Blumenthal, R., Okamoto, I. & Miki, T. Human ECT2 is an exchange factor for Rho GTPases, phosphorylated in G2/M phases and involved in cytokinesis. J. Cell Biol. 147, 921–928 (1999).
Matthews, H. K. et al. Changes in Ect2 localization couple actomyosin-dependent cell shape changes to mitotic progression. Dev. Cell 23, 371–383 (2012).
Matsuoka, Y. et al. Two different protein kinases act on a different time schedule as glial filament kinases during mitosis. EMBO J. 11, 2895–2902 (1992).
Ando, S. et al. Phosphorylation of synthetic vimentin peptides by cdc2 kinase. Biochem. Biophys. Res. Commun. 195, 837–843 (1993).
Roskoski, R. Jr. ERK1/2 MAP kinases: structure, function and regulation. Pharmacol. Res. 66, 105–143 (2012).
Gonzalez, F. A., Raden, D. L. & Davis, R. J. Identification of substrate recognition determinants for human ERK1 and ERK2 protein kinases. J. Biol. Chem. 266, 22159–22163 (1991).
Miller, M. L. et al. Linear motif atlas for phosphorylation-dependent signaling. Sci. Signal. 1, ra2; 10.1126/scisignal.1159433 (2008).
Chen, C. et al. Identification of a major determinant for serine-threonine kinase phosphoacceptor specificity. Mol. Cell 53, 140–147 (2014).
Elia, A. E., Cantley, L. C. & Yaffe, M. B. Proteomic screen finds pSer/pThr-binding domain localizing Plk1 to mitotic substrates. Science 299, 1228–1231 (2003).
Vassilev, L. T. et al. Selective small-molecule inhibitor reveals critical mitotic functions of human CDK1. Proc. Natl. Acad. Sci. USA. 103, 10660–10665 (2006).
Stewart, M. Molecular mechanism of the nuclear protein import cycle. Nat. Rev. Mol. Cell Biol. 8, 195–208 (2007).
Saito, S. et al. Deregulation and mislocalization of the cytokinesis regulator ECT2 activate the Rho signaling pathways leading to malignant transformation. J. Biol. Chem. 279, 7169–7179 (2004).
Zou, Y. et al. Characterization of nuclear localization signal in the N terminus of CUL4B and its essential role in cyclin E degradation and cell cycle progression. J. Biol. Chem. 284, 33320–33332 (2009).
Jans, D. A. The regulation of protein transport to the nucleus by phosphorylation. Biochem. J. 311, 705–716 (1995).
Jackson, S. & Xiong, Y. CRL4s: the CUL4-RING E3 ubiquitin ligases. Trends Biochem. Sci. 34, 562–570 (2009).
Brown, N. R., Noble, M. E., Endicott, J. A. & Johnson, L. N. The structural basis for specificity of substrate and recruitment peptides for cyclin-dependent kinases. Nat. Cell Biol. 1, 438–443 (1999).
Malumbres, M. & Barbacid, M. Mammalian cyclin-dependent kinases. Trends Biochem. Sci. 30, 630–641 (2005).
Jantz, D. & Berg, J. M. Reduction in DNA-binding affinity of Cys2His2 zinc finger proteins by linker phosphorylation. Proc. Natl. Acad. Sci. USA 101, 7589–7593 (2004).
Chen, Y. J. et al. A conserved phosphorylation site within the forkhead domain of FoxM1B is required for its activation by cyclin-CDK1. J. Biol. Chem. 284, 30695–30707 (2009).
Kraft, C. et al. Mitotic regulation of the human anaphase-promoting complex by phosphorylation. EMBO J. 22, 6598–6609 (2003).
Kõivomägi, M. et al. Multisite phosphorylation networks as signal processors for Cdk1. Nat. Struct. Mol. Biol. 20, 1415–1424 (2013).
Holt, L. J. et al. Global analysis of Cdk1 substrate phosphorylation sites provides insights into evolution. Science 325, 1682–1686 (2009).
Sako, K. et al. Emi2 mediates meiotic MII arrest by competitively inhibiting the binding of Ube2S to the APC/C. Nat. Commun. 5, 3667; 10.1038/ncomms4667 (2014).
Acknowledgements
We thank members of the Sagata laboratory for discussions. This work was supported by grants from the MEXT, Japan, to N.S.
Author information
Authors and Affiliations
Contributions
N.S. conceived the project; N.S. and K. Suzuki designed the experiments; K. Suzuki, K. Sako, K.A., M.I., C.S. and N.N. performed the experiments; K. Suzuki, K. Sako and N.S. analyzed the data; N.S. and K. Suzuki wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Suzuki, K., Sako, K., Akiyama, K. et al. Identification of non-Ser/Thr-Pro consensus motifs for Cdk1 and their roles in mitotic regulation of C2H2 zinc finger proteins and Ect2. Sci Rep 5, 7929 (2015). https://doi.org/10.1038/srep07929
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07929
This article is cited by
-
Qualitative rather than quantitative phosphoregulation shapes the end of meiosis I in budding yeast
The EMBO Journal (2024)
-
Function of SYDE C2-RhoGAP family as signaling hubs for neuronal development deduced by computational analysis
Scientific Reports (2022)
-
CDK1–cyclin-B1-induced kindlin degradation drives focal adhesion disassembly at mitotic entry
Nature Cell Biology (2022)
-
Kinase inhibition profiles as a tool to identify kinases for specific phosphorylation sites
Nature Communications (2020)
-
Multisite phosphorylation code of CDK
Nature Structural & Molecular Biology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.