Abstract
Sequence-specific recognition of DNA by small turn-on fluorescence probes is a promising tool for bioimaging, bioanalytical and biomedical applications. Here, the authors report a novel cell-permeable and red fluorescent hemicyanine-based thiazole coumarin (TC) probe for DNA recognition, nuclear staining and cell cycle analysis. TC exhibited strong fluorescence enhancement in the presence of DNA containing AT-base pairs, but did not fluoresce with GC sequences, single-stranded DNA, RNA and proteins. The fluorescence staining of HeLa S3 and HEK 293 cells by TC followed by DNase and RNase digestion studies depicted the selective staining of DNA in the nucleus over the cytoplasmic region. Fluorescence-activated cell sorting (FACS) analysis by flow cytometry demonstrated the potential application of TC in cell cycle analysis in HEK 293 cells. Metaphase chromosome and malaria parasite DNA imaging studies further confirmed the in vivo diagnostic and therapeutic applications of probe TC. Probe TC may find multiple applications in fluorescence spectroscopy, diagnostics, bioimaging and molecular and cell biology.
Similar content being viewed by others
Introduction
Small organic dyes capable of exhibiting turn-on fluorescence through sequence-specific interaction with nucleic acids play an integral role in fluorescence spectroscopy, diagnostics, imaging and biomedical applications1,2. Selective targeting of double-stranded (ds) DNA using organic dyes offers powerful strategies to develop i) probes for molecular biology and immunohistochemistry, flow cytometry and DNA quantitation, ii) genome-specific binders of potential theranostic interest in conjugation with predesigned oligonucleotides and iii) diagnostic therapy of gene-related human diseases especially cancer and parasitic and viral infections3,4,5. In this regard, various small sequence-specific fluorophores have been developed for biological assays, including cell imaging and DNA-quantitation in cells6.
The discovery of DNA as a genetic material and its double helical structure led to numerous studies directed at understanding DNA-small molecule interactions7,8,9,10.Typically, DNA-small molecule interaction has two prominent modes viz., intercalation and minor groove binding. The research efforts of Dervan11,12 and Lown13 have set forth the foundation for developing a series of small molecules with affinity for the AT-rich minor groove of B-DNA. 4′,6-Diamidino-2-phenylindole (DAPI)14 and bisbenzimides (Hoechst dyes)15 are some of the well-studied nuclear staining agents that bind with the minor groove. Unfortunately, these blue-emitting DNA binders require ultraviolet (UV) light for excitation which is known to cause cellular DNA damage through free radical generation, eventually causing cell death16. Moreover, Hoechst dyes, despite being faithful to the AT-rich minor groove of DNA, are nonetheless prone to fasten onto GC-rich sites, too and also show binding affinity towards single-stranded (ss) DNA and RNA17,18. Similarly, DAPI suffers from the disadvantages of non-specific binding to RNA19 and requirement of high concentration for effective imaging, thereby limiting its use only to fixed cells20. Apart from minor groove binders, ethidium bromide (EtBr) and propidium iodide (PI) have been used for cell staining and gel-imaging studies. However, impermeable nature of these dyes limited their application to fixed and dead cells only15. Recently, cyan and green fluorescence DNA-selective probes such as BENA43521 and C6122 have been reported. However, these probes, upon binding to DNA, exhibit fluorescence enhancement with low quantum yields. Thomas et al. recently reported a dinuclear ruthenium(II) polypyridyl complex as a DNA-staining probe, which nonetheless required high concentration for cellular imaging23.
Apart from these molecules, a large number of cyanine dyes have been extensively used in DNA gel staining, microchip-based DNA sensing and fluorescence staining of DNA in cells24,25,26,27. Among the cyanine family dyes, thiazole orange (TO) and yellow orange (YO) are two important classes of fluorescence probes that display significant fluorescence enhancement upon binding with DNA. Further, the homodimeric forms of TOTO-1 and YOYO-1 are found to be highly sensitive for DNA detection28,29,30,31. However, these cyanine-based probes also exhibit significant fluorescence enhancement in the presence of RNA and ssDNA32,33,34. Recently, two other classes of cyanine dyes, SYBR green I and PicoGreen I, have been developed and successfully used for DNA staining in the picogram scale, although they also show fluorescence enhancement in the presence of ssDNA18,35.
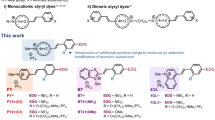
The limitations of existing probes discussed above necessitates the development of highly specific DNA-selective probes with i) long-wavelength excitation/emission, ii) strong turn-on fluorescence, iii) increased cell permeability, iv) non-toxicity to live cells, v) fidelity to dsDNA and vi) sensitivity at low concentrations. In the present study, we report a turn-on red fluorescence hemicyanine probe TC as an effective cell-permeable, base-pair specific dsDNA recognition and nuclear DNA staining probe (Fig. 1). Inspired by the basic core structure of cyanine probes, we have designed three hemicyanine-based molecular probes, namely thiazole-coumarin (TC), coumarin-lepidine (CL) and thiazole-pyrene (TP) with the objective of finding a superior DNA staining reagent. To elucidate the role of positively charged quaternary amine group in benzothiazole-based probes we have replaced the benzothiazole group with a quinoline moiety in CL. Similarly, coumarin group has been substituted with the hydrophobic pyrene in TP to understand the role of heterocyclic fluorophore moieties (coumarin/quinoline) in the benzothiazole-based probes.
The choice of coumarin chromophore in the probes is owing to its excellent fluorescence properties in the visible region36. Further, these probes are likely to display excitation and emission in the longer wavelength of the visible region owing to extended conjugation, an essential prerequisite to avoid autofluorescence and DNA photo-damage during cellular imaging. One of the main characteristic optical properties of a dye to qualify as a potential DNA binding and staining reagent is that it must be non-fluorescent or weakly fluorescent in the unbound state and display highly enhanced fluorescence in the longer wavelength of the visible region (red) upon interaction with DNA. Interestingly, all three hemicyanine probes are almost non-fluorescent in buffer solution (100 mM Tris-HCl, pH = 7.4) with very low quantum yields (Table S1), a property that partially satisfies one of the primary requirements of a suitable turn-on fluorescence DNA binding molecular probe. Intramolecular twisting of unsymmetrical cyanine dyes has been shown to be responsible for efficient quenching of fluorescence in unconstrained environments while their binding to compact packets of DNA results in enhanced fluorescence due to restriction of intramolecular twisting37,38. Similar molecular transformations are anticipated from our hemicyanine probes upon interaction with DNA.
Results and Discussion
Photophysical properties of probe TC
We first studied molecular interactions of TC, CL and TP in the absence and presence of DNA through photophysical (absorption and emission) measurements in 100 mM Tris-HCl buffer (pH = 7.4) solution. TC, CL and TP showed absorption in the visible region with absorption maxima at 528, 519 and 460 nm, respectively (supplementary Fig. S2a). Interestingly, the absorption spectrum of TC in the presence of dsDNA (AT)10 showed a significant bathochromic shift (Δλmax = 14 nm) along with hypochromicity (supplementary Fig. S2b). Emission spectra of TC, CL and TP exhibited very weak fluorescence with maxima at 641, 689 and 623 nm, respectively and large Stoke's shifts (Fig. 2a). Next, we recorded the emission spectra of probes in the presence of DNA duplexes [(AT)10 and (GC)10]. TC (10 µM) showed strong fluorescence enhancement in the presence of (AT)10, which further increased with DNA concentration (0 to 12 µM) with overall ~ 8-fold enhancement accompanied by hypsochromic shift (Δλmax = 26 nm) in the emission maxima at 615 nm. Interestingly, TC did not display such fluorescence enhancement in the presence of (GC)10. Emission spectra of CL (10 µM) showed ~ 30 and ~ 6-fold fluorescence enhancement in the presence of (AT)10 and (GC)10, respectively, although with appreciably lower fluorescence intensity (QY: 0.040 and 0.007, respectively). We did not observe a detectable change in the emission spectra of TP (10 µM) in the presence of (AT)10 and (GC)10 (Fig. 2a).
Selective AT-base pair fluorescence probe.
(a), Fluorescence spectra of all the probes TC, CL and TP (10 µM) in presence of (AT)10 and (GC)10 in Tris-HCl (100 mM, pH = 7.4) buffer solution. (b), Photographs of TC and TC+ (AT)10 samples illuminated under UV light (365 nm). (c), Fluorescence spectra of probe TC (10 µM) in presence of various dsDNAs. (d), Fluorescence response of probe TC (10 µM) with increasing concentration of (AT)20, (AT)10, (GC)10, (D1)mix and (D2)mix upon excitation at 521 nm. (e), Shows the side and top view of intercalation mode of probe TC in presence of 2AT- and 2GC-base pairs.
These fluorescence emission studies of TC, CL and TP in the presence of (AT)10 and (GC)10 clearly indicate TC as an effective and selective turn-on fluorescence probe for DNA containing AT-base pairs. The proficiency of probe TC as compared to CL and TP may be explained by the positively charged benzothiazole-conjugated N,N-diethylamino-coumarin fluorophore that balanced amphiphilicity and the possibility achieving maximum π-electron overlap through molecular planarity (benzothiazole and coumarin) in a constrained environment.
In addition to this, the remarkable turn-on fluorescence enhancement of TC only in the presence of dsDNA containing AT-base pairs indicates that the AT-region provides favorable constrained environment to TC by restricting its intramolecular twisting. This was further verified by recording the emission spectra of TC with increasing glycerol percentage in Tris-HCl buffer solution. The emission spectra showed an increase in fluorescence intensity with the percentage of glycerol content (supplementary Fig. S4). Addition of glycerol increases the viscosity of the buffer solution. This viscous nature of glycerol aids in freezing the intramolecular rotation of probe TC to attain the planar molecular structure. The planar structure of TC leads to maximum electron delocalization which is further reflected in the enhanced fluorescence of probe TC. A similar effect i.e., ‘freezing of intramolecular twisting’ is assumed to occur when TC binds to dsDNA. These results confirm that the origin of the fluorescence enhancement of probe TC in the presence of dsDNA containing AT-base pairs is due to the restriction of intramolecular twisting. Further, we recorded the fluorescence spectra of TC in the presence of oligonucleotides (dA10, dT10, dG10 and dC10). TC did not show significant changes in its emission behavior in the presence of ssDNA (supplementary Fig. S3). Therefore, the selectivity of TC for AT-region of dsDNA over ssDNA is a highly useful property because most of the commonly employed staining dyes such as ethidium bromide (EtBr), SYTO, Picogreen I and SYBR green I show weak fluorescence enhancement in the presence of ssDNA sequences39.
Selective fluorescence enhancement in the presence of dsDNA containing AT-base pairs
Highly selective turn-on fluorescence enhancement of TC towards (AT)10 prompted us to explore its sequence-selective binding affinity towards various other dsDNA strands. We selected three different dsDNA (AT)20, self-complementary mixed sequences (D1)mix and (D2)mix composed of a variable number of consecutive AT-base pairs, i.e., 20, 4 (two sets) and 2 (two sets), respectively (supplementary Fig. S1). TC (10 µM) displayed ~ 16-fold fluorescence enhancement with 20 consecutive AT-base pairs in (AT)20, which is two-fold higher than (AT)10. Next, we recorded the emission spectra of self-complementary AT-rich d(AATT)5 sequence40. The fluorescence measurements of probe TC showed ~16-fold enhancement in the presence of d(AATT)5 (supplementary Fig. S5). The mixed dsDNA (D1)mix and (D2)mix with 4 (two sets) and 2 (two sets) consecutive AT-base pairs exhibited fluorescence enhancement of ~6 and 1.5-folds, respectively (Fig. 2c). These data were further validated by recording the emission spectra of TC (10 µM) with increasing concentration (0 µM to 12 µM) of dsDNA (AT)20, (AT)10, (D1)mix and (D2)mix that displayed overall ~16, ~8, ~6 and ~1.5-fold fluorescence enhancement, respectively, with hypsochromic shift in the fluorescence maxima (Fig. 2d). Further, we determined the quantum yield of probe TC in buffer solution. In the absence of dsDNA, TC showed very low quantum yield (ΦF = 0.03). However, the quantum yield increased significantly (ΦF = 0.36) upon binding with (AT)20. In general, above-discussed studies revealed significant red fluorescence enhancement with increasing number of AT-base pairs in dsDNA and increased quantum yield specifically in the presence of AT-rich regions, thereby suggesting the preferential binding affinity of TC towards AT-sites rather than GC-sites.
Subsequently, the concentration-dependent fluorescence response of TC against fixed concentration of dsDNA (12 µM) containing AT-base pairs was investigated (Fig. 3a). In the absence of dsDNA, increasing concentrations of TC (0 to 20 µM) showed very weak fluorescence emission. In the presence of (AT)20, increasing concentration of TC from 0 to 18 µM showed regular enhancement in fluorescence intensity. Saturation was achieved beyond 18 µM (18 to 22 µM). Interestingly, for concentrations of TC> 24 µM a decrease in fluorescence intensity accompanied by bathochromic shift in the emission maxima was observed. Similar changes in the fluorescence emission were observed with increasing concentration of TC in the presence of (D1)mix and (D2)mix (supplementary Fig. S6). The observed fluorescence emission behavior of TC suggests that, at lower concentrations, TC can readily occupy the available binding sites (AT-region) of dsDNA, resulting in strong fluorescence enhancement. However, at higher concentrations of TC (18–22 µM) the fluorescence emission reaches a peak due to saturation of binding sites. After the saturation of binding sites (for > 24 µM of TC) in dsDNA (12 µM), competitive binding interactions occur between the unbound and bound TC molecules leading to displacement of some of the bound molecules from dsDNA41. This explains the linear increase in fluorescence at lower concentrations < 18 µM, saturation at 18–22 µM and a dramatic decrease in the fluorescence intensity at > 24 µM of TC.
Gel –electrophoresis and FRET between TC and Hoechst.
(a), Plot of fluorescence intensity of increased concentration of TC (0–20 µM) in presence of fixed concentrations (12 µM) of (AT)20, (AT)10,(D1)mix and (D2)mix. (b–c), Agarose gel-electrophoresis of dsDNAs followed by staining the bands with probe TC and ethidium bromide respectively. Lane 1–4: (AT)20, (D1)mix, (D2)mix and (GC)10 respectively. Inset ring (red) in (b) shows non staining of (GC)10 band by TC. Inset ring (red) in (c) shows staining of (GC)10 band by ethidium bromide. (d), Normalized densitometric signal intensity data of TC stained agarose-gel. (e), Fluorescence spectra of preformed Hoechst (4 µM)+(AT)20 complex with increasing concentration of TC from 0 µM to 40 µM. Inset: Overlap spectra of emission of preformed Hoechst (4 µM)+(AT)20 complex and absorption spectra of TC (10 µM)+(AT)20 complex. (f), Schematic representation of FRET between Hoechst 33258 (donor) and probe TC (acceptor) in presence of dsDNA upon excitation at 350 nm. Also shown is the direct emission from TC upon excitation at 521 nm.
The turn-on red fluorescence behavior of probe TC in the presence of AT-rich DNA inspired us to explore the use of TC as a staining agent for dsDNA in gel electrophoresis. The dsDNA samples (0.6 µg) of (AT)20, (D1)mix, (D2)mix and (GC)10 were subjected to agarose gel electrophoresis and the gel was visualized under UV-illumination after staining with TC. UV-illuminated gel image showed strong fluorescence intensity for (AT)20, (D1)mix and (D2)mix, respectively while the band for (GC)10 was not stained by TC at all (Fig. 3b). Besides, densitometric signal intensity data of TC-stained agarose gel exhibited relative fluorescence intensities in the following order: (AT)20>(D1)mix>(D2)mix (Fig. 3d). Next, the agarose gel was stained with the well-known gel staining agent ethidium bromide (EtBr). Ethidium bromide stained bands of (D1)mix, (D2)mix and (GC)10 on the gel. While the staining was very weak for (AT)20, it was strongest in the case of (GC)10 as expected from its weaker affinity towards AT-region as compared to GC-region of dsDNA (Fig. 3c). From the gel-electrophoresis staining studies, it is clear that TC probe selectively discriminates dsDNA containing AT-base pairs from DNA containing only GC-base pairs. In addition, the comparative fluorescence response studies among TC, ethidium bromide (supplementary Fig. S7a) and propidium iodide (PI) (supplementary Fig. S7b) in the presence of (AT)20 revealed TC as an excellent fluorescence marker for AT-rich DNAs while ethidium bromide and propidium iodide did not display appreciable fluorescence changes.
To gain insights into preferential binding of TC to AT-base pair regions in dsDNA, competitive binding studies with Hoechst were conducted. Hoechst dyes are known to interact with the minor groove of dsDNA. First, we recorded the emission spectrum of preformed (AT)20-Hoechst 33258 complex upon excitation at 350 nm (corresponding to Hoechst), which showed fluorescence enhancement at 450 nm. Next, we recorded the emission spectra of preformed (AT)20-Hoechst 33258 complex with increasing concentration of TC (0 to 40 µM) upon excitation at 350 nm (Fig. 3e). A marked decrease in emission intensity of Hoechst at 450 nm was observed with increasing concentration of TC. Simultaneously, there was a gradual increase in emission at 610 nm of TC from 0 to 20 µM, above which emission intensity reached saturation (supplementary Fig. S8a). Interestingly, excitation at TC absorption (521 nm) showed strong fluorescence enhancement at 610 nm with increasing concentration of the probe from 0 to 20 µM, which is specific of TC upon binding to dsDNA. This phenomenon of decrease in fluorescence intensity at 450 nm (Hoechst) and the corresponding increase in emission at 610 nm (TC) upon excitation at 350 nm (Hoechst) is due to efficient FRET from Hoechst to TC (Fig. 3f). Further increase in concentration of TC (20 to 40 µM) led to a decrease in fluorescence, a characteristic phenomenon where unbound excess probe molecules (TC) assist the displacement of bound ones resulting in partial quenching of fluorescence (supplementary Fig. S8b)41.
To understand this unprecedented fluorescence behavior observed in competitive study, circular dichroism (CD) studies of the (AT)20-Hoechst complex were conducted in the presence of TC. CD-spectrum of (AT)20 displayed an intense positive and a negative Cotton signal in the 200–300 nm region. Hoechst alone has no characteristic CD signal, but in the presence of (AT)20 it showed an induced positive Cotton signal in the 300–400 nm region (supplementary Fig. S9). It is known that the minor groove of dsDNA comprises of chiral environment and, upon binding, induces its chirality to the achiral Hoechst dye molecules. However, increasing concentration of TC (0 to 40 µM) did not depict any detectable change in the CD signal of (AT)20-Hoechst complex. These data revealed that TC is not competing with Hoechst for binding site in the minor groove of dsDNA. Rather, it bound to the adjacent region containing AT-base pairs. This is evident by the effective FRET from the Hoechst (donor) to the TC (acceptor). Normally, the efficient energy transfer in FRET mechanism depends on the distance between donor-acceptor pair and proper overlap of emission of donor and absorption of acceptor42. The effective spectral overlap of emission of Hoechst and excitation of TC indicated a perfectly matched donor-acceptor pair for the FRET process in AT-rich DNAs (Fig. 3einset). Thus, the observed FRET between the two probes clearly suggests that the binding site for TC must be in close proximity to that of Hoechst in the AT-rich minor groove of dsDNA. Similarly, FRET was also observed between DAPI (donor) and TC (acceptor) in the presence of dsDNA containing AT-base pairs.
To understand the thermal stability of dsDNA-TC complexes, we carried out temperature-dependent UV-vis absorption studies. The UV-melting data of TC bound (AT)20, (AT)10, (D1)mix and (D2)mix showed increasing melting temperatures (Tm) with ΔTm = 3.8, 2.6, 1.9 and 1.6°C, respectively as compared to their unbound forms (supplementary Table S2). This moderate thermal stabilization of TC-bound AT-containing dsDNA is indicative of favorable interaction without any structural deformation in the duplex DNA structure. In contrast, melting study of (GC)10 with the addition of TC showed an appreciable decrease in Tm (ΔTm = −1.7°C), which depicts the destabilization of duplex DNA structure in the presence of the probe. Overall, a significant increase in Tm of TC-bound dsDNA containing AT-base pairs provided substantial evidence for preferential binding of TC to AT-base pairs region of DNA.
Simulation studies of probe TC binding to dsDNA
To gain greater insights into the binding mode and preferential fluorescence enhancement in AT-rich over GC-rich regions of dsDNA we carried out ab initio Density Functional Theory (DFT) calculations of probe TC in the absence and presence of dsDNA. Each molecule was optimized with the help of DFT and their photophysical properties were calculated using time-dependent DFT (TD-DFT) as implemented in Gaussian 09 package. For the optimization, ωb97XD exchange correlation functional was used to include empirical dispersion while for TD-DFT calculations B3LYP exchange and correlation functional was employed. We used 6-31g(d) basis set for all the atoms (see supplementary information). The DNA phosphate backbone was neutralized by protonating one of the oxygen atoms of the phosphate groups, which does not alter any property of the DNA duplex structure43,44,45. The DFT and TD-DFT calculations were performed with surrounding water, using a Polarized Continuum Model (PCM) to include the solvent (water) effect. To predict the interaction mode and relative binding affinity of probe TC, we estimated the stabilization energies of TC in presence of AT- and GC-base pairs in all possible modes. The optimized structures showed that intercalation was the most preferred binding mode for TC (Fig. 2e). Studies also revealed that TC was stabilized more strongly by AT-base pairs (37.65 kcal mol−1) than by GC-base pairs (34.98 kcal mol−1). These energy values support the observed moderately-increased melting temperatures of AT-rich DNAs over GC-rich DNAs in the presence of probe TC.
To predict the preferential fluorescence enhancement of probe TC in AT-base pair regions over the GC-base pairs, we calculated the absorbance and emission spectra using optimized ground state (S0) geometry and optimized first excited state (S1) geometry, respectively. The TD-DFT calculation provided excitation energies and oscillator strengths to the lowest singlet states (supplementary Table. S3). It was found that for all the possible modes, the lowest energy transitions occur from Highest Occupied Molecular Orbital (HOMO) to Lowest Unoccupied Molecular Orbital (LUMO). For both TC and TC@2AT complexes, the HOMOs and LUMOs were localized on the TC moiety for both absorption and emission. It is interesting to note that, in case of the TC@2GC complex for absorption, the corresponding MOs (HOMO and LUMO) were localized on the TC moiety, but for emission, the transition occurred from the TC moiety to the guanine moiety of TC@2GC (Fig. 4), which quenched the fluorescence emission. In fact, in the latter case, the fluorescence quenched because of charge transfer.
Cellular uptake and selective DNA-imaging properties of probe TC
The selective fluorescence enhancement of TC in the presence of AT-rich dsDNA further encouraged us to study its cellular uptake and application in nuclear DNA staining. The HeLa S3 cells were incubated with probe TC and then imaged using confocal fluorescence microscopy. HeLa S3 cells were excited at 521 nm and the emission was measured at 540–750 nm, corresponding to the excitation of the probe TC for selective binding of nuclear DNA. Fluorescence imaging of HeLa S3 cell lines with probe TC (5 µM) and co-staining with commercially available Hoechst 33258 nuclear stain showed selective targeting of cell nucleus by the probe TC (supplementary Fig. S10). Further, we carried out nuclear staining studies in HEK 293 cells with cytoplasmic α-tubulin marker to demonstrate the specificity of TC to nuclei and did not stain the cytoplasm. It was revealed that the probe TC did not colocalize with the cytoplasmic α- tubulin marker, but did so with the properly placed nucleus (Fig. 5 a–h). Collectively, these results suggest that the probe TC is cell membrane-permeable and highly selective towards the nucleus.
Cellular uptake properties of probe TC.
(a–h), Confocal fluorescence microscope images of HEK 293T cells incubated (at 37°C for 30 min) with a, TC (5 µM). b, anti α-tubulin antibody. c, Hoechst 33258. d, differential interference contrast (DIC, bright field image) of HEK 293T cells respectively. Overlay fluorescence images, e: a and c. f: a and b. g: a, b and c. h: a and d. Scale bar 5 µm. i, Metaphase chromosomal staining with probe TC. Metaphase chromosomal plate was prepared using HEK 293 cell line and stained with probe TC (2 μM). Inset: Arrow shows the maximum centromere localization of probe TC.
To further validate the nuclear DNA binding of TC, we performed staining of the metaphase chromosome of HEK 293 cell line. The results clearly illustrated that the probe TC (at 2 and 5 μM concentration) stained the chromosomes efficiently (Fig. 5 i). The careful observation of these images revealed that some areas of chromosomes (for example, the AT-rich centromeric region) were more densely stained (shown with arrows in Fig. 5i, inset). However, at higher concentration we observed uniform chromosomal staining compared to the lower concentration of probe TC, which showed remarkably greater staining of centromere region (supplementary Fig. S11).
Fidelity of Probe TC to dsDNA
One of the foremost problems with most DNA staining dyes is non-specific binding with cellular ribonucleic acids (RNA), proteins and other biomacromolecules46,47. To explore the possibilities of such non-specific interactions, we recorded the emission spectrum of TC in the presence of fetal bovine serum albumin (FBA), which did not show any detectable change in basal fluorescence of the probe (supplementary Fig. S12a). Further, we recorded the fluorescence spectra of preformed [TC + d(AATT)5] with the addition of excess FBA and vice versa. Interestingly, we did not observe any detectable change in the strong fluorescence intensity of probe TC. While, upon addition of d(AATT)5 to TC + FBA mixture, ~70% fluorescence of probe TC was achieved compare to [TC + d(AATT)5] complex (supplementary Fig. S12b). In order to check the effect of other parameter like salt and pH we have measured the fluorescence of [TC + d(AATT)5] complex in presence of NaCl (100 mM) and with the variation pH from 5 to 8 (supplementary Fig. S13). In presence of 100 mM NaCl, [TC + d(AATT)5] did not show appreciable change in the strong fluorescence intensity. Similarly, variation of pH from 5 to 8 had no effect on the fluorescence of [TC + d(AATT)5] complex. Overall, these results suggests probe TC is selective towards dsDNA and retains its fluorescence even in complex samples containing proteins and in presence of variable salt and pH conditions. To demonstrate the selectivity of TC for dsDNA over RNA, we performed deoxyribonuclease I (DNase I) and ribonuclease (RNase) digestion studies in HEK 293 cell lines. DNase I is known to degrade nuclear DNA while RNase degrades RNA by catalyzing phosphodiester bond cleavage. After DNase I digestion of cells the fluorescence of probe TC (red) and control Hoechst 33258 (blue) almost completely diminished from the nuclear region compared to untreated cells. However, as predicted, the RNase treatment of HEK 293 cells did not affect the fluorescence staining of TC and Hoechst 33258 in the nuclear region (Fig. 6a).
DNA-specific fluorescence probe.
(a), The DNase and RNase studies in HEK 293 cell lines. Immunofluorescence staining of control and cells treated with DNase I (100 µg/mL) and RNase (40 µg/mL) with TC (red). DNA was counterstained with Hoechst (blue). Scale bar, 5 µm. (b–c), Cell cycle analysis by staining with PI and probe TC respectively. HEK 293 cells stained with 6 µg of PI and probe TC for 30 mins. FACS analysis was done by FACS aria instrument.
Application of probe TC in Cell Cycle Analysis
The FACS analysis using flow cytometry is a versatile tool for cell cycle analysis with the aid of DNA-specific binders. To quantitate the DNA content in the cells, we employed probe TC for the cell cycle analysis of HEK293 cells using flow cytometry. For FACS analysis, HEK293 cells were separately stained with PI (10 μg/mL) and probe TC (5 μM) in the presence and absence of RNase. FACS data showed a similar pattern of each cell cycle phase with almost similar percentage of populations of G1, S and G2-M (Fig. 6b–c). These results suggest that the probe TC binds to DNA only and may be applied in FACS analysis to quantify DNA and cell cycle estimation without employing RNase. It was also observed that the intensity of probe TC was more as compared to the same concentration of PI used in the cell cycle analysis at FL2-A scale.
Cytotoxicity Studies of Probe TC
Another significant aspect is that of cytotoxicity, which is a major constraint with several DNA-binding probes including PI. To understand whether probe TC displayed any cytotoxicity towards HEK293 cells, MTT assay was performed. In a dose-dependent experiment, three different concentrations of probe TC were taken. It was observed that TC was non-toxic to HEK293 cells for treatment time intervals (30 min) used in the present experiments (supplementary Fig. S14a). Time-dependent experiments were also performed for 6, 12, 18 and 24 h by taking 5 µM of TC. These experiments revealed a gradual decrease in cell viability, but the percentage of dead cells was infinitesimal. As many as 64% of cells were viable after 24 h of treatment (supplementary Fig. S14b).
Preferential fluorescence staining of malaria parasites
The strong and specific binding of probe TC towards the AT-rich DNA raises the possibility whether this probe can be used for purposes other than mammalian nuclear staining and cell sorting. To test this possibility we selected Plasmodium falciparum as a model system which has AT rich genome (~80%). The parasites were incubated with low concentrations of probe TC (1 and 2 µM) with trophozoite stage parasites growing in erythrocytes. Live fluorescence imaging of trophozoite stage parasites depicted that probe TC could specifically bind to the parasite cells within the red blood cells at 1 and 2 µM concentrations. Vehicle control (water) did not show any detectable fluorescence under similar experimental conditions (Fig. 7). Incubation of the HepG2 cells (human liver cancer cells) with 2 µM of probe TC did not show any detectable fluorescence (supplementary Fig. S15). These results clearly indicate that probe TC preferentially bound to the AT-rich genome of the parasite at very low concentrations. Collectively, these results suggest the selectivity of probe TC for AT-base pairs in dsDNA, fidelity to DNA over RNA and protein, non-toxicity and preferential staining of malarial parasites in human erythrocytes, indicating that probe TC is a useful diagnostic and possibly therapeutic, red-fluorescence biomarker for AT-rich genomes48,49.
Binding of probe TC to the AT rich genome of Plasmodium falciparum.
Live cell fluorescence imaging showed probe TC is accumulated specifically in the nucleus of the parasitized red blood cell at 1 and 2 µM. Vehicle control (water) did not show any detectable fluorescence under the same experimental conditions. DAPI stains the nuclei and all merged panel shows phase images of the parasites along with DAPI and TC fluorescence signal. Scale bar: 2 µm.
Conclusion
In summary, we have developed red fluorescence hemicyanine-based thiazole-coumarin (TC) probe for base pair-specific recognition of dsDNA. Fluorescence spectroscopy and gel-electrophoresis studies of TC showed strong fluorescence enhancement in the presence of DNA containing AT-base pairs, but non-fluorescence with DNA containing only GC-base pairs, single-stranded DNA, RNA and proteins. The increase in fluorescence of TC as a function of consecutive AT-base pairs in dsDNA also suggests AT-rich regions are the preferable binding sites. The FRET studies demonstrated that minor-groove binding Hoechst and probe TC act as an efficient donor-acceptor pair. The fluorescence spectroscopy studies, fluorescence-imaging of HeLaS3, HEK 293, digestion by nuclease enzymes and flow cytometry studies of TC revealed AT-base pair-specific fluorescence enhancement, effective cell-permeability, non-toxicity, selective nuclear DNA staining over cytoplasmic region, cell cycle analysis and DNA quantitation (See Table S4 for the advantages of probe TC over other commercial probes).
The tendency of TC to localize in a particular region of chromosomes and preferential affinity (at low concentrations) towards the malarial parasite DNA in human erythrocyte authenticate the diagnostic and therapeutic potential of probe TC. We anticipate that the AT-selective red fluorescence DNA probe TC could be a promising and useful reagent in molecular biology, cell biology and possibly other DNA-based techniques such as fluorescence spectroscopy, flow cytometry, DNA quantitation and also for the development of genome-specific binders of theranostic interest in conjugation with designed oligonucleotides.
Methods
General Information
All the chemicals, reagents, oligos (dA20, dT20, dA10, dT10, dG10, dC10 and d(AATT)5), control probes (Hoechst 33258, ethidium bromide and propidium iodide), primary antibody, Alexa A488-coupled secondary antibody, DNase I and RNase were purchased from Sigma-Aldrich. 1H and 13C-NMR spectra were recorded on a Bruker AV-400 MHz spectrometer with chemical shifts reported as parts per million (ppm)(in CDCl3/DMSO-d6, tetramethylsilane as an internal standard) at 20 °C. UV-vis absorption and emission spectra were measured in quartz cuvettes of 1 cm path length. The absolute fluorescence quantum yields were determined using an integrating sphere for the samples on FLSP920 spectrometer (Edinburgh Instruments). High resolution Mass spectra (HRMS) were obtained on Agilent Technologies 6538 UHD Accurate-Mass Q-TOF LC/MS spectrometer.
Synthetic procedure for compounds TC, CL and TP
To a stirred solution of N-methylated benzothiazole (1) or lepidine (2) (1.2 eq.) in ethanol, piperidine (0.2 eq.) was added and the reaction mixture was allowed to stir for 10 min. 7-(Diethylamino)-2-oxo-2H-chromene-3-carbaldehyde50 (1 eq.) in ethanol solution was added dropwise to the above reaction mixture, leading to immediate yellow to purple color change in the solution after which the reaction mixture was allowed to stir for 4 h. The completion of the reaction was monitored with thin layer chromatography (TLC). After completion of the reaction, the solvent was evaporated under vacuum. The crude product was purified using column chromatography on silica gel using CHCl3/MeOH as an eluent to afford TC or CL in good yields.
To a stirred solution of N-methylated benzothiazole (1) (164 mg, 5.65 mmol) in ethanol piperidine (60 µL, 5.65 mmol) was added and allowed to stir for 10 min. Pyrene-1-carboxaldehyde (100 mg, 4.347 mmol) in ethanol solution was added dropwise to the reaction mixture with pressure equalizer and stirred under reflux conditions for 7 h. The completion of the reaction was monitored with TLC. The solvent was vacuum-evaporated after completion of the reaction. The crude product was purified using column chromatography on silica gel using DCM/MeOH (98/2) as eluent to afford probe TP in good yield.
Characterization data for TC
Brown color powder, yield 50%51. 1H-NMR (DMSO-d6, 400 MHz) δ 8.59 (s, 1H), 8.37 (dd, J = 0.8 Hz, J = 8Hz, 1H), 8.21 (d, J = 8.4 Hz, 1H), 8.02 (dd, J = 8 Hz, J = 15.6 Hz, 2H), 7.83 (m, 1H), 7.74 (td, J = 1.2 Hz, J = 7.6 Hz, 1H), 7.57 (d, J = 9.2 Hz, 1H), 6.87 (dd, J = 2.4 Hz, J = 9.2 Hz, 1H), 6.67 (d, J = 2.4 Hz, 1H), 4.22 (s, 3H), 3.53 (q, J = 7.2 Hz, 4H), 1.17 (t, J = 7.2 Hz, 6H). 13C-NMR (DMSO-d6, 100 MHz) δ 171.6, 159.5, 157.1, 153.3, 148.3, 144.3, 142.0, 131.8, 129.2, 128.0, 127.4, 124.1, 116.5, 112.0, 111.3, 110.9, 108.9, 96.4, 44.6, 35.8, 12.4. HRMS (ESI-MS): calcd for C23H23N2O2SI [M-I] + m/z = 391.1475, found 391.1456.
CL
Violet color powder, yield 60%. 1H-NMR (DMSO-d6, 400 MHz) δ 9.25 (d, J = 6.4 Hz, 1H), 8.72 (d, J = 8 Hz, 1H), 8.43 (m, 4H), 8.24 (m, 1H), 8.02 (m, 2H), 7.54 (d, J = 9.2 Hz, 1H), 6.82 (dd, J = 2.4 Hz, J = 9.2 Hz, 1H), 6.61 (d, J = 2.4 Hz, 1H), 4.46 (s, 3H), 3.50 (q, J = 7.2 Hz, 4H), 1.16 (t, J = 7.2 Hz, 6H). 13C-NMR (DMSO-d6, 100 MHz) δ 160.0, 158.2, 157.8, 152.4, 152.1, 147.7, 145.1, 138.7, 138.2, 134.8, 130.9, 129.3, 126.0, 125.6, 119.4, 118.4, 115.3, 114.8, 113.9, 110.2, 108.6, 96.3, 44.4, 12.4. HRMS (ESI-MS): calcd for C25H25IN2O2 [M-I]+ m/z = 385.1911, found 385.1902.
TP
Brown color powder, yield 78%. 1H-NMR (DMSO-d6, 400 MHz) δ 9.19 (d, J = 15.6 Hz, 1H), 8.97 (d, J = 8.4 Hz, 1H), 8.88 (d, J = 9.6 Hz, 1H), 8.39 (m, 9H), 8.18 (t, J = 7.6 Hz, 1H), 7.92 (m, 1H), 7.84 (td, J = 0.8 Hz, J = 7.6 Hz, 1H), 4.46 (s, 3H). 13C-NMR (DMSO-d6, 100 MHz) δ 171.5, 144.1, 142.0, 133.7, 130.8, 130.3, 130.1, 129.9, 129.5, 128.5, 128.1, 127.4, 127.3, 127.0, 126.7, 125.7, 125.5, 124.2, 124.0, 123.5, 122.6, 116.9, 115.4, 36.5. HRMS (ESI-MS): calcd for C26H18INS [M-I]+ m/z = 376.1154, found 376.1156.
Metaphase chromosome preparation and Probe TC staining
HEK293 cells were cultured in a 60 mm dish, harvested with 0.25% EDTA-Trypsin and collected by centrifugation at 2000 rpm for 3 min. Supernatant was discarded and washed twice with 1XPBS. Cell pellet was resuspended in 10 mL hypotonic solution (0.56% KCl) and incubated for 30 min at 37°C. Recentrifugation was conducted and the pellet was again suspended in 8 mL fixative (glacial acetic acid: ethanol, 1:3). Then 2–3 drops of cell suspension were dropped at some distance on slide. Slides were either air dried or flame dried. Hoechst (1 μg/mL) and probe TC (5 μM) were poured on the slide and kept for 10 min for DNA staining and mounted with 70% glycerol. Images were taken at 100× by Carl Zeiss Laser Scanning Microscope (LSM510 META).
Plasmodium falciparum culture and probe TC treatment
The growth and maintenance of parasite culture have been described elsewhere52. In brief, P. falciparum 3D7 strain was grown at 4–10% parasitemia in RPMI 1640 medium supplemented with 0.5% albumax (Invitrogen), 5% NaHCO3, 50 µg/mL gentamicin and 50 µg/mL ampicillin at 37°C. The culture was synchronized by incubating ring-staged parasites for 5 minutes with 5% sorbitol. Probe TC (resuspended in Mili-Q water) was added to parasites (30 ± 5 hpi) at a final concentration of 1 and 2 µM and the parasites were incubated for 30 minutes at 37°C. Milli-Q water was used as a vehicle control. Fluorescence images were captured using Carl Zeiss AXIO Imager Z1 and the software used for image capturing was AxioVision Rel. 4.8.
References
Armitage, B. A. Cyanine Dye–DNA Interactions: Intercalation, Groove Binding and Aggregation. Top. Curr. Chem. 253, 55–76 (2005).
Ranasinghe, R. T. & Brown, T. Fluorescence based strategies for genetic analysis. Chem. Commun. 2005, 5487–5502 (2005).
Waring, M. J. DNA Modification and Cancer. Annu. Rev. Biochem. 50, 159–192 (1981).
Neidle, S. DNA minor-groove recognition by small molecules. Nat. Prod. Rep. 18, 291–309 (2001).
Hurley, L. H. DNA and its associated processes as targets for cancer therapy. Nat. Rev. Cancer. 2, 188–200 (2002).
Blackburn, G. M., Gait, M. J., Loakes, D. & Williams, D. M. Nucleic Acids in Chemistry and Biology. (RSC Publishing, Cambridge, UK, 2006).
Lerman, L. S. Structural considerations in the interactions of deoxyribonucleic acid and acridines. J. Mol. Biol. 3, 18–30 (1961).
Waring, M. J. Drugs which affect the Structure and Function of DNA. Nature 219, 1320–1325 (1968).
Zimmer, C. & Wähnert, U. Nonintercalating DNA-binding ligands: specificity of the interaction and their use as tools in biophysical, biochemical and biological investigations of the genetic material. Prog. Biophys. Mol. Biol. 47, 31–112 (1986).
Neidle, S. & Abraham, Z. Structural and sequence-dependent aspects of drug intercalation into nucleic acids. CRC Crit. Rev. Biochem. 17, 73–121 (1984).
White, S., Szewczyk, J. W., Turner, J. M., Baird, E. E. & Dervan, P. B. Recognition of the four Watson-Crick base pairs in the DNA minor groove by synthetic ligands. Nature 391, 468–471 (1998).
Dervan, P. B. Molecular recognition of DNA by small molecules. Bioorg. Med. Chem. 9, 2215–2235 (2001).
Reddy, B. S. P., Sondhi, S. M. & Lown, J. W. Synthetic DNA minor groove-binding drugs Pharmacol. Ther. 84, 1–111 (1999).
Kubista, M., Akerman, B. & Norden, B. Characterization of interaction between DNA and 4′,6-diamidino-2-phenylindole by optical spectroscopy. Biochemistry 26, 4545–4553 (1987).
Crissman, H. A. & Hirons, G. T. Staining of DNA in live and fixed cells. Methods Cell Biol. 41, 195–209 (1994).
Pfeifer, G. P., You, Y.-H. & Besaratinia, A. Mutations induced by ultraviolet light. Mutat. Res. 571, 19–31 (2005).
Loontiens, F. G., Regenfuss, P., Zechel, A., Dumortier, L. & Clegg, R. M. Binding characteristics of Hoechst 33258 with calf thymus DNA, poly[d(A-T)] and d(CCGGAATTCCGG): multiple stoichiometries and determination of tight binding with a wide spectrum of site affinities. Biochemistry 29, 9029–9039 (1990).
Singer, V. L., Jones, L. J., Yue, S. T. & Haugland, R. P. Characterization of PicoGreen reagent and development of a fluorescence-based solution assay for double-stranded DNA quantitation. Anal. Biochem. 249, 228–238 (1997).
Tanious, F. A., Veal, J. M., Buczak, H., Ratmeyer, L. S. & Wilson, W. D. DAPI (4′,6-diamidino-2-phenylindole) binds differently to DNA and RNA: minor-groove binding at AT sites and intercalation at AU sites. Biochemistry 31, 3103–3112 (1992).
Zink, D., Sadoni, N. & Stelzer, E. Visualizing chromatin and chromosomes in living cells. Methods 29, 42–50 (2003).
Erve, A. et al. BENA435, a new cell-permeant photoactivated green fluorescent DNA probe. Nucleic Acids Res. 34, e43 (2006).
Feng, S., Kim, Y. K., Yang, S. & Chang, Y. T. Discovery of a green DNA probe for live-cell imaging. Chem. Commun. 46, 436–438 (2010).
Gill, M. R. et al. Ruthenium(II) polypyridyl complex for direct imaging of DNA structure in living cells. Nat. Chem. 1, 662–667 (2009).
Haugland, R. P. Handbook of Fluorescent Probes and Research Products; Molecular Probes. (Inc.: Eugene, OR, 2002).
Joo, C., Balci, H., Ishitsuka, Y., Buranachai, C. & Ha, T. Advances in single-molecule fluorescence methods for molecular biology. Annu. Rev. Biochem. 77, 51–76 (2008).
vanMameren, J., Peterman, E. J. G. & Wuite, G. J. L. See me, feel me: methods to concurrently visualize and manipulate single DNA molecules and associated proteins. Nucleic Acids Res. 36, 4381–4389 (2008).
Peng, X. et al. An effective minor groove binder as a red fluorescent marker for live-cell DNA imaging and quantification. Angew. Chem. Int. Ed. 50, 4180–4183 (2011).
Selvin, P. Science innovation '92: the San Francisco sequel. Science 257, 885–886 (1992).
Glazer, A. N. & Rye, H. S. Stable dye-DNA intercalation complexes as reagents for high-sensitivity fluorescence detection. Nature 359, 859–861 (1992).
Rye, H. S. et al. Stable fluorescent complexes of double-stranded DNA with bis-intercalating asymmetric cyanine dyes: properties and applications. Nucleic Acids Res. 20, 2803–2812 (1992).
Rye, H. S. et al. Picogram detection of stable dye-DNA intercalation complexes with two-color laser-excited confocal fluorescence gel scanner. Methods Enzymol. 217, 414–431 (1993).
Ogura, M. & Mitsuhashi, M. Fluorometric method for the measurement of nuclease activity on plastic plates. BioTechniques. 18, 231–233 (1995).
Ogura, M. & Mitsuhashi, M. Use of the fluorescent dye YOYO-1 to quantify oligonucleotides immobilized on plastic plates. BioTechniques. 16, 1032–1034 (1994).
Rye, H. S. & Glazer, A. N. Interaction of dimeric intercalating dyes with single-stranded DNA. Nucleic Acids Res. 23, 1215–1222 (1995).
Zipper, H., Brunner, H., Bernhagen, J. & Vitzthum, F. Investigations on DNA intercalation and surface binding by SYBR Green I, its structure determination and methodological implications. Nucleic Acids Res. 32, e103 (2004).
Yuan, L., Lin, W., Zheng, K. & Zhu, S. FRET-based small-molecule fluorescent probes: rational design and bioimaging applications. Acc. Chem. Res. 46, 1462–1473 (2013).
Furstenberg, A. et al. Ultrafast excited-state dynamics of DNA fluorescent intercalators: new insight into the fluorescence enhancement mechanism. J. Am. Chem. Soc. 128, 7661–7669 (2006).
Silva, G. L., Ediz, V., Yaron, D. & Armitage, B. A. Experimental and computational investigation of unsymmetrical cyanine dyes: understanding torsionally responsive fluorogenic dyes. J. Am. Chem. Soc. 129, 5710–5718 (2007).
Cosa, G., Focsaneanu, K.-S., McLean, J. R. N., McNamee, J. P. & Scaiano, J. C. Photophysical properties of fluorescent DNA-dyes bound to single- and double-stranded DNA in aqueous buffered solution. Photochem. Photobiol. 73, 585–599 (2001).
Nguyen, B., Neidle, S. & Wilson, W. D. A role for water molecules in DNA-ligand minor groove recognition. Acc. Chem. Res. 42, 11–21 (2009).
Dragan, A. I. et al. Characterization of PicoGreen interaction with dsDNA and the origin of its fluorescence enhancement upon binding. Biophys. J. 99, 3010–3019 (2010).
Stender, A. S. et al. Single cell optical imaging and spectroscopy. Chem. Rev. 113, 2469–2527 (2013).
Samanta, P. K., Manna, A. K. & Pati, S. K. Structural, electronic and optical properties of metallo base pairs in duplex DNA: a theoretical insight. Chem. Asian. J. 7, 2718–2728 (2012).
Samanta, P. K. & Pati, S. K. Structural and magnetic properties of a variety of transition metal incorporated DNA double helices. Chem. Eur. J. 20, 1760–1764 (2014).
Churchill, C. D. M. & Wetmore, S. D. Developing a computational model that accurately reproduces the structural features of a dinucleoside monophosphate unit within B-DNA. Phys. Chem. Chem. Phys. 13, 16373–16383 (2011).
Martin, R. M., Leonhardt, H. & Cardoso, M. C. DNA labeling in living cells. Cytometry A. 67, 45–52 (2005).
vanZandvoort, M. A. et al. Discrimination of DNA and RNA in cells by a vital fluorescent probe: lifetime imaging of SYTO13 in healthy and apoptotic cells. Cytometry 47, 226–235 (2002).
Roset, R., Subirana, J. A. & Messeguer, X. MREPATT: detection and analysis of exact consecutive repeats in genomic sequences. Bioinformatics 19, 2475–2476 (2003).
Smale, S. T. & Kadonaga, J. T. The RNA polymerase II core promoter. Annu. Rev. Biochem. 72, 449–479 (2003).
Maity, D. & Govindaraju, T. Highly selective colorimetric chemosensor for Co2+. Inorg. Chem. 50, 11282–11284 (2011).
Sun, Y.-Q., Liu, J., Zhang, J., Yang, T. & Guo, W. Fluorescent probe for biological gas SO2 derivatives bisulfite and sulfite. Chem. Commun. 49, 2637–2639 (2013).
Deshmukh, A. S. et al. The role of N-terminus of Plasmodium falciparum ORC1 in telomeric localization and var gene silencing. Nucleic Acids Res. 40, 5313–5331 (2012).
Acknowledgements
Prof. C. N. R. Rao, FRS for his constant support and encouragement, JNCASR, Council of Scientific and Industrial Research (CSIR) New Delhi, [grant No. 02/(0128)/13/EMR-II], Innovative Young Biotechnologist Award (IYBA) and Department of Biotechnology (DBT), India (BT/03/IYBA/2010)for financial support, Alexander von Humboldt Foundation, Germany for special equipment donation to T.G, CSIR for SRF fellowship to N.N. P.K.S. and R.S. T.K.K and S.K.P are supported by Sir J C Bose National Fellowship, India, T.K.K by DBT (grant No. BT/01/CEIB/10/III/01), SKD by Centre of Excellence (CoE) in Parasitology (JNU), Builder programme in Chemical Biology (JNU), National Biosciences Award (all funded DBT). The authors acknowledge Prof. R. K. Tyagi (SCMM, JNU) for providing HepG2 cells.
Author information
Authors and Affiliations
Contributions
N.N. and T.G. designed the project. N.N. undertook the synthesis, photophysical studies and in vitro studies of the probes. T.K.K. and S.K.D. designed in vivo studies. M.K., S.D. and R.S. performed immuno-fluorescence imaging and malaria parasite staining studies. P.K.S. and S.K.P. carried out the theoretical calculations. All authors contributed to writing the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Narayanaswamy, N., Kumar, M., Das, S. et al. A Thiazole Coumarin (TC) Turn-On Fluorescence Probe for AT-Base Pair Detection and Multipurpose Applications in Different Biological Systems. Sci Rep 4, 6476 (2014). https://doi.org/10.1038/srep06476
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep06476
This article is cited by
-
Benzocoumarin-Styryl Hybrids: Aggregation and Viscosity Induced Emission Enhancement
Journal of Fluorescence (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.