Abstract
Regulated promoters are an important basic genetic tool allowing, for example, gene-dosage and gene depletion studies. We have previously described a cumate-inducible promoter (PQ5) that is functional in diverse Alphaproteobacteria. This promoter has been engineered by combining a synthetic minimal promoter, Psyn2 and operator sites and the repressor of the Pseudomonas putida F1 cym/cmt system. In the present study, we engineered a vanillate-regulated promoter using Psyn2 and the regulatory elements of the Caulobacter crescentusvanR-vanAB system. We show that the resulting promoter, which we called PV10, responds rapidly to the inducer vanillate with an induction ratio of about two orders of magnitude in Sphingomonas melonis Fr1. In contrast to the switch-like behavior of PQ5, PV10 shows a linear dose-response curve at intermediate vanillate concentrations, allowing graded gene expression. PV10 is functionally compatible with and independent of PQ5 and cumate and viceversa, suggesting that both systems can be used simultaneously.
Similar content being viewed by others
Introduction
Regulated promoters are an essential genetic tool for studying bacterial physiology as well as for synthetic biology and industrial applications. For example, they provide means to study essential gene function by depletion analysis and to conditionally express toxic genes. Although multiple of such systems are usually available for a particular model organism, they are often underdeveloped for many non-model organisms and even constitutive (minimal) promoters are not always available. We have previously described a synthetic approach that allowed us to develop a cumate-inducible expression system that is functional in diverse Alphaproteobacteria, including several sphingomonads, Methylobacterium extorquens and Caulobacter crescentus1. In this approach, we first identified a minimal promoter consensus based on alignment of several Sphingomonas melonis Fr1 housekeeping gene promoters, then screened for mutations in non-conserved positions in the −10 element of this minimal promoter for increased expression and finally combined this mutant promoter (termed Psyn2) with operator sequences and the repressor of the heterologous cym/cmt system, which naturally controls cumate and cymene catabolism in Pseudomonas putida F12,3. This engineered promoter (called PQ5) was cumate-regulated and resulted in induction ratios of two- to three orders of magnitude in the different organisms tested. For sphingomonads, a group of bacteria with great potential in bioremediation, industrial biotechnology and plant protection4,5,6,7,8,9, this was the first dedicated inducible gene expression system described to date.
Because certain applications call for more than one inducible promoter, we wondered whether it would be possible to combine Psyn2 with yet other heterologous operator sequences and repressors so it would be regulated by a stimulus other than cumate. Here we describe such a promoter, termed PV10, that combines Psyn2 with the vanO operator sequences of the vanAB operon and the vanillate-responsive repressor VanR naturally involved in vanillate degradation in the freshwater bacterium Caulobacter crescentus10. Our experiments demonstrate that PV10 is vanillate-inducible and shows a high dynamic range of gene expression in S. melonis Fr1 and that PV10 and PQ5 are orthogonal, with each promoter only responding to its designated stimulus, vanillate and cumate, respectively.
Results
Design of PV10
A scheme of the organization of vanR and PV10 is shown in Fig. 1a. The design rationale is described in the following. In Caulobacter crescentus, vanAB is divergently transcribed from vanR, encoding the GntR-type transcriptional repressor of the vanAB operon. The vanAB promoter (vanABp) has been mapped and shows −35 (TTGACG) and −10 (AAGATT) boxes indicative of a housekeeping, σ70-dependent promoter10. Two perfect inverted repeats (ATTGGATCCAAT), each comprising one operator site (vanO) to which VanR is thought to bind, are present in the vanR-vanAB intergenic region, one immediately upstream of the vanABp −35 box and the second overlapping the −10 box (positions −9 to −7) and the +1 transcriptional start site10. The strong synthetic promoter Psyn2 harbors −35 (TTGACG) and −10 (TAACTGC) boxes characteristic for σ70-dependent promoters, with positions −12, −11, −7 and −6 (underlined) in the −10 box highly conserved in numerous S. melonis Fr1 housekeeping promoters1. To render Psyn2 regulated by vanillate, we combined Psyn2 with vanO sites in essentially the same configuration observed in the C. crescentusvanAB promoter and we termed the resulting promoter PV10. This required changing the −10 box from TAACTGC to TAAATTG (Fig. 1b), so that the downstream vanO sequence overlaps the −10 box (the modified core promoter is referred to as Psyn2′); although this might lead to a weakened promoter (see below), we reasoned that, at the same time, this configuration would allow tight repression. In fact, Psyn2 is very strong1 and such strong expression is probably not needed in most cases where expression levels in the physiological range are desired. vanR was placed under control of the constitutive promoter Pbla-mut1T, which we have used before to drive expression of the PQ5 repressor CymR*1. The basic plasmid for vanillate-regulated gene expression is pVH, a derivative of the broad-host-range plasmid pCM6211, in which downstream of PV10 a multiple cloning site (Fig. 1c) is present that is flanked 5′ by a sequence encoding a hemagglutinin (HA) tag and 3′ by a sequence coding for a triple FLAG (3XFLAG) tag and a putative rho-independent transcriptional terminator (Fig. 1a).
Genetic organization and nucleotide sequence of PV10.
(a) Organization of vanR, PV10 and the multiple cloning site (MCS) on plasmid pVH. Bended arrows denote promoters and dark grey boxes indicate ribosome bind sites (RBS). Start and stop codons of the open reading frame encoding HA and 3XFLAG tags (white boxes) up- and downstream of the MCS (black box) are indicated by ATG and TGA, respectively. T193* denotes a putative transcriptional terminator. Unique restriction sites in pVH are shown. (b) Nucleotide sequence of PV10. The 3′-truncated Psyn2 core promoter (Psyn2′; see the main text) containing −35 and −10 boxes (highlighted in light gray) and PV10 are indicated by dashed arrows above the nucleotide sequence. Palindromic vanO operator sites are indicated by inverted solid arrows and are highlighted in italics in the nucleotide sequence. The RBS and start codon are in bold. (c) Nucleotide sequence, translation and restriction sites of the MCS.
Characterization of PV10
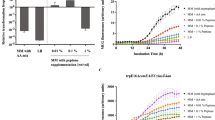
In order to characterize PV10-dependent gene expression in S. melonis Fr1, PV10 was transcriptionally fused to E. colilacZ (plasmid pVH-lacZ) and PV10-lacZ+ activity was followed in strain JVZ857/pVH-lacZ grown with different vanillate concentrations using β-galactosidase assays. As shown in Fig. 2a, PV10-lacZ+ activity was dependent on the inducer concentration, showing a low basal activity (48 +/− 8 Miller units) without vanillate and high activity (3600 +/− 280 Miller units) at the highest vanillate concentration tested (250 μM). This represents a maximal induction ratio of 74-fold. In the range of 6.5 to 74 μM vanillate, the dose-response curve was essentially linear (R2 = 0.994), indicating that PV10 allows graded gene expression, rather than showing switch-like behavior. To follow induction dynamics, JVZ857/pVH-lacZ was grown to mid-exponential phase, PV10 was induced by addition of 250 μM vanillate and PV10-lacZ+ activity was repeatedly measured over 4.5 h. As seen from Fig. 2b, the response is rapid and sustained. To see whether PV10 could be used simultaneously with the previously characterized cumate-inducible promoter PQ5, we tested both promoters for their response to vanillate and/or cumate. Like for PV10, PQ5 activity was followed using strain JVZ857 harboring a plasmid-borne PQ5-lacZ+ transcriptional fusion described previously1. As shown in Fig. 2c, PV10 did not respond to cumate and cumate had no effect on the induction by vanillate. Similarly, PQ5 showed no response to vanillate and vanillate did not affect the capacity of PQ5 to respond to cumate.
Characterization of PV10.
(a) Dose-response curve of PV10 as measured with β-galactosidase (β-gal.) assays on S. melonis Fr1 carrying the PV10-lacZ+ transcriptional fusion (JVZ857/pVH-lacZ). Vanillate concentrations tested ranged from 6.5 to 250 μM, in 1.5-fold increments and no vanillate as control. The inset is a log-log representation of the same data to illustrate induction at low vanillate concentrations. (b) Induction kinetics of PV10 were determined by addition of 250 μM vanillate to a non-induced culture of JVZ857/pVH-lacZ grown to mid-exponential phase and following β-gal. activity over time. (c) Strains JVZ857/pVH-lacZ (white bars) and JVZ857/pQF-lacZ carrying the PQ5-lacZ+ transcriptional fusion (grey bars) were grown in the presence (+) or absence (−) of 250 μM vanillate and/or 25 μM cumate as indicated and β-gal. activities were determined. All data represent the mean +/− SD of three independent biological replicates.
In summary, our result demonstrate that PV10 is a rapidly responding, vanillate-inducible promoter in S. melonis Fr1. Furthermore, they suggest that PV10 is orthogonal to the previously described cumate-inducible promoter PQ5 and both promoters can be used simultaneously without interference.
Other vanillate-regulated expression plasmids
In addition to the basic vanillate-inducible expression plasmid pVH, we have also constructed two destination plasmids for Gateway cloning, pVHD and pVYD, that allow C-terminal fusions to the HA tag and SYFP2, respectively and three plasmids for N-terminal fusions to mCherry, SYFP2 and mTq2 (pVCC, pVCY and pVCTq, respectively). Plasmids will be made available from Addgene (www.addgene.org).
Discussion
Altogether, our results demonstrate that PV10 allows tuning gene expression over a high dynamic range in S. melonis Fr1 and possibly other Alphaproteobacteria. This graded response to vanillate is in contrast to the more switch-like behavior of the previously described cumate-inducible promoter PQ51, at least in S. melonis Fr1 and might make PV10 the better choice when gene-dosage should be reproducibly and carefully regulated. In contrast, compared to PV10, PQ5 shows both higher absolute expression and relative induction upon inducer addition1, making PQ5 more suitable when very high gene expression levels are desired, e.g. for overexpression studies. Thus, the two promoters are complementary and one or the other might be better suited depending on the biological question. Importantly, because there is no crosstalk between the CymR*/PQ5 and VanR/PV10 systems, i.e. they are orthogonal, both can be used simultaneously, allowing more sophisticated genetic studies of bacterial physiology.
Methods
Strains and growth conditions
Escherichia coli TOP10 (Invitrogen) or “ccdB survival” (Invitrogen) were used for cloning and routinely grown in LB-Lennox at 37°C. S. melonis Fr1 wild-type strain JVZ85712 was grown in LB-Lennox at 28°C. Plasmids were transformed in S. melonis by electroporation as previously described13. When appropriate, antibiotics were added at the following concentrations: tetracycline (10 μg/ml) and chloramphenicol (34 μg/ml). Vanillate (4-hydroxy-3-methoxybezoic acid) was purchased from Sigma-Aldrich (Cat. No. W398802-25G) and dissolved in ethanol to give 1000× stock solutions for final concentrations indicated in the figure legends. Cumate stocks were prepared as described previously1. For “no vanillate” and “no cumate” controls, cultures were mock treated with 0.1% (vol/vol) ethanol.
Plasmid construction
Standard molecular biology protocols were followed14. Phusion DNA polymerase for PCR and restriction enzymes were from Thermo Scientific and T4 DNA ligase was from New England Biolabs. pVH was constructed in two steps. First, vanR was amplified from plasmid pRVYFPC-210 using primers VanR_PciI_F (5′-ATT TAC ATG TTT TCA GTC GGC GCG AAT GC-3′) and VanR_NheI_R (5′-ATT TTG CTA GCA GGG AGA GAC CCC GAA TGG ACA TGC CGC GCA TAA-3′) and cloned in pQH1 via PciI/NheI, replacing cymR*. Then, a synthetic fragment (Eurofins, MWG Operon, Germany) containing Pbla-mut1T1 for vanR expression and PV10 for vanillate-regulated expression was amplified using primers PV10_F (5′-ATT TGC TAG CAT CAG GGT TAT TG-3′) and PV10_R (5′- ATT TAA GCT TCC TCT ACT AGT ATT G-3′) and cloned between NheI/HindIII. To construct pVH-lacZ, lacZ was excised from pAK127lacZ(MCS)1 using XbaI/EcoRI and cloned in pVH via SpeI/EcoRI. pVHD was constructed by amplification of the Gateway cassette from pDEST-565 (Addgene plasmid 11520) using primers oJVZ739 (5′-ATT TGG TAC CTC TAG CTA GCG ATA TCA CC-3′) and oJVZ740 (5′-ATT TTC TAG AGA CAA GTT TGT ACA AAA AAG C-3′) and cloning in pVH via XbaI/Acc65I. pVYD was obtained by subcloning a PsiI/SpeI fragment of pVH containing vanR and PV10 in between the same sites of pQYD1, replacing cymR* and PQ5. Plasmids pVCY, pVCC and pVCTq were constructed by amplifying the genes encoding fluorescent proteins with primers mTq2C_F (5′-ATT TGG TAC CGA GCT CCA ATT GGG GCG GCG GCA GCG GCG GCG GCA GCG TGA GCA AGG GCG AGG AGC-3′) and mTq2C_R (5′-ATT TTG AAT TCT CAC TTG TAC AGC TCG TCC ATG CC-3′), digestion of the PCR product with Acc65I/EcoRI and cloning in pVH via Acc65I/MunI. Templates for PCR were: pQY1 for “super” yellow fluorescent protein 2 (SYFP2); pQR1 for mCherry; pmTurquoise2-C115 for mTurqouise2 (mTq2).
Reporter assays
Promoter activities were measured essentially as described previously1. For dose-response curves and cross-induction experiments, S. melonis Fr1 carrying pVH-lacZ or pQF-lacZ1 were grown in LB-Lennox containing different concentrations of vanillate and/or cumate overnight to mid-exponential phase and β-galactosidase activity was measured according to Miller16. To follow induction kinetics, S. melonis carrying pVH-lacZ was grown to mid-exponential phase and induced by the addition of 250 μM vanillate and β-galactosidase was measured at different time points. All results are presented as mean +/− SD of three biological replicates. Linear regression analysis to evaluate linearity of the dose-response curve was performed in GraphPad Prism 5 (version 5.04, Graphpad Software Inc., USA).
References
Kaczmarczyk, A., Vorholt, J. A. & Francez-Charlot, A. Cumate-inducible gene expression system for sphingomonads and other Alphaproteobacteria. Appl. Environ. Microbiol. 79, 6795–6802 (2013).
Eaton, R. W. p-Cumate catabolic pathway in Pseudomonas putida Fl: cloning and characterization of DNA carrying the cmt operon. J. Bacteriol. 178, 1351–1362 (1996).
Eaton, R. W. p-Cymene catabolic pathway in Pseudomonas putida F1: Cloning and characterization of DNA encoding conversion of p-cymene to p-cumate. J. Bacteriol. 179, 3171–3180 (1997).
Balkwill, D. L., Fredrickson, J. K. & Romine, M. F. in The Prokaryotes: A Handbook on the Biology of Bacteria, Vol.7, Proteobacteria, Delta and Epsilon Subclasses. Deeply Rooting Bacteria. 605–629 (Springer-SBM, New York, 2006).
Fialho, A. M. et al. Occurrence, production and applications of gellan: current state and perspectives. Appl. Microbiol. Biotechnol. 79, 889–900 (2008).
Innerebner, G., Knief, C. & Vorholt, J. A. Protection of Arabidopsis thaliana against leaf-pathogenic Pseudomonas syringae by Sphingomonas strains in a controlled model system. Appl. Environ. Microbiol. 77, 3202–3210 (2011).
Takeuchi, M., Hamana, K. & Hiraishi, A. Proposal of the genus Sphingomonassensu stricto and three new genera, Sphingobium, Novosphingobium and Sphingopyxis, on the basis of phylogenetic and chemotaxonomic analyses. Int. J. Syst. Evol. Microbiol. 51, 1405–1417 (2001).
Vogel, C., Innerebner, G., Zingg, J., Guder, J. & Vorholt, J. A. Forward genetic in planta screen for identification of plant-protective traits of Sphingomonas sp. strain Fr1 against Pseudomonas syringae DC3000. Appl. Environ. Microbiol. 78, 5529–5535 (2012).
White, D. C., Sutton, S. D. & Ringelberg, D. B. The genus Sphingomonas: physiology and ecology. Curr. Opin. Biotechnol. 7, 301–306 (1996).
Thanbichler, M., Iniesta, A. A. & Shapiro, L. A comprehensive set of plasmids for vanillate- and xylose-inducible gene expression in Caulobacter crescentus. Nucleic Acids Res. 35, e137 (2007).
Marx, C. J. & Lidstrom, M. E. Development of improved versatile broad-host-range vectors for use in methylotrophs and other Gram-negative bacteria. Microbiology 147, 2065–2075 (2001).
Kaczmarczyk, A. et al. Role of Sphingomonas sp. strain Fr1 PhyR-NepR-σEcfG cascade in general stress response and identification of a negative regulator of PhyR. J. Bacteriol. 193, 6629–6638 (2011).
Kaczmarczyk, A., Vorholt, J. A. & Francez-Charlot, A. Markerless gene deletion system for sphingomonads. Appl. Environ. Microbiol. 78, 3774–3777 (2012).
Sambrook, J. & Russel, D. Molecular Cloning: A Laboratory Manual. Third edn, (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, 2001).
Goedhart, J. et al. Structure-guided evolution of cyan fluorescent proteins towards a quantum yield of 93%. Nature communications 3, 751 (2012).
Miller, J. H. Experiments in Molecular Genetics. 352–355 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, 1972).
Acknowledgements
We thank Martin Thanbichler, Dominic Esposito, Joachim Goedhart and Theodorus W. J. Gadella for plasmids. This work was supported by Swiss National Science Foundation (SNF) grant 31003B-152835.
Author information
Authors and Affiliations
Contributions
A.K. designed and performed experiments. A.K., J.A.V. and A.F.-C. conceived the project and wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Kaczmarczyk, A., Vorholt, J. & Francez-Charlot, A. Synthetic vanillate-regulated promoter for graded gene expression in Sphingomonas. Sci Rep 4, 6453 (2014). https://doi.org/10.1038/srep06453
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep06453
This article is cited by
-
Endotoxin-free gram-negative bacterium as a system for production and secretion of recombinant proteins
Applied Microbiology and Biotechnology (2023)
-
Complex general stress response regulation in Sphingomonas melonis Fr1 revealed by transcriptional analyses
Scientific Reports (2019)
-
Heterologous production of a new lasso peptide brevunsin in Sphingomonas subterranea
Journal of Industrial Microbiology and Biotechnology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.