Abstract
Both cytosine-Ag-cytosine interactions and cytosine modifications in a DNA duplex have attracted great interest for research. Cytosine (C) modifications such as methylcytosine (mC) and hydroxymethylcytosine (hmC) are associated with tumorigenesis. However, a method for directly discriminating C, mC and hmC bases without labeling, modification and amplification is still missing. Additionally, the nature of coordination of Ag+ with cytosine-cytosine (C-C) mismatches is not clearly understood. Utilizing the alpha-hemolysin nanopore, we show that in the presence of Ag+, duplex stability is most increased for the cytosine-cytosine (C-C) pair, followed by the cytosine-methylcytosine (C-mC) pair and the cytosine-hydroxymethylcytosine (C-hmC) pair, which has no observable Ag+ induced stabilization. Molecular dynamics simulations reveal that the hydrogen-bond-mediated paring of a C-C mismatch results in a binding site for Ag+. Cytosine modifications (such as mC and hmC) disrupted the hydrogen bond, resulting in disruption of the Ag+ binding site. Our experimental method provides a novel platform to study the metal ion-DNA interactions and could also serve as a direct detection method for nucleobase modifications.
Similar content being viewed by others
Introduction
In DNA duplexes, silver ions specifically interact with C-C mismatches1,2,3,4, while mercury ions specifically interact with T-T mismatches5,6,7,8. These interactions that strongly stabilize DNA duplexes have been extensively studied recently9, but the nature of coordination of Ag+ with C-C mismatches is not clearly understood4,10,11,12. Considering that cytosine (C) modifications such as 5-methylcytosine (mC) and 5-hydroxymethylcytosine (hmC) are important epigenetic markers associated with gene expression and tumorigenesis13,14,15, we were motivated to explore the interactions of Ag+ with a DNA duplex containing a single C-C, C-mC or C-hmC mismatch in the alpha-hemolysin nanopore (α-HL). The α-HL has a nanocavity (2.6 nm opening with a 1.4 nm constriction site) that can capture and hold the DNA duplex (Supplementary Figure S1), providing an ideal platform for studying both the C-Ag-C interaction and how cytosine modifications change this interaction. In a nanopore experiment, an electric field drives charged molecules through a nanometer-scale pore that spans an insulating membrane, which separates two aqueous solutions. The baseline ionic current through the pore is transiently blocked by larger macromolecules (such as DNA) that enter the pore. The ion current through a nanopore is sensitive to target molecules that interact with the pore, therefore different molecular states can be electrically clarified from characteristic changes in the nanopore current. The α-hemolysin nanopore has been studied for DNA sequencing16,17,18, various single-molecule detections19,20,21 and biomolecular interactions22,23,24,25.
In previous nanopore studies26,27,28,29, researchers have found that C, mC or hmC can be recognized by immobilizing the DNA with streptavidin28, or by chemical modifications26 in α-HL. When in a solid-state nanopore, it was found that DNA duplexes containing mC and hmC can be discriminated29 and by using methylated CpG binding proteins, C and mC themselves could also be discriminated27. Several other methods can be used to distinguish hmC, mC and C bases with chemical modifications via sequencing30,31,32. In this report, we described a unique nanopore sensor that can directly discriminate cytosine and cytosine modifications simultaneously (evidenced by ionic current signals such as dwell times (toff, Supplementary Figure S1) and residual currents (Supplementary Figure S1) without modifications. The key principle of this novel method for cytosine modifications determination is the fact that Ag+ stabilizes a C-C containing DNA duplex, which was confirmed in the nanopore for the first time. By molecular dynamics (MD) simulations, we found that cytosine modifications such as mC and hmC disrupted both the hydrogen bonds and Ag+ interactions, which subsequently affected DNA-Ag+ stability (in the term of rate of dissociation).
Results
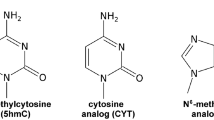
The study involved three 16-nt AT rich ssDNAs as the targets, which contain a cytosine (TC), 5′-methylcytosine (TmC) and 5′-hydromethylcytosine (ThmC) at the 10th nucleotide (5′ → 3′), respectively (Table 1). Their common probe, P, contains a cytosine at the corresponding position, such that when P is hybridized with the three targets, their hybrids P·TC, P·TmC and P·ThmC, form a C-C, C-mC and C-hmC mismatched base-pair respectively. Since Ag+ was tested in the experiments, we could not use KCl buffer due to AgCl precipitation. Therefore, we first tested how the single-stranded DNA (ssDNA) P (Figure 1) interacts with the nanopore in KNO3 solution. Short (<1 ms) and long events in the range of 1–10 ms were easily identified (Figure 1a,b). The residual current also has a wide distribution, with a peak at 17.4 ± 0.84 pA (Figure 1c). Others have previously noted that KNO3 has unknown effects on DNA translocation and some extraordinary long events were seen, with about 10-fold lower occurrence rate constant (Kon) of ssDNA in KNO3 than in the KCl buffer8, as well as in certain cations such as Li+33 and ion liquid34. In order to ensure the ssDNA interactions were excluded, we only considered events longer than 10 ms as the DNA duplexes interact with the nanopore. A control experiment demonstrated that Ag+ itself does not affect the open pore current (Figure 1d.e). The positively charged Ag+ is driven away from the nanopore by the applied voltage.
The ssDNA P interacts with the nanopore.
(a) The representative current trace recorded at 150 mV. Two types of events were identified: a1: spike-like current profile which last about 200 us and a2: rectangular-like current profile which last about 1 to 10 ms. (b) The histogram of the dwell time in Log form. The long events (>100 = 1 ms) were easily identified. (c) The histogram of residual currents when the ssDNA P interacts with the nanopore. The nanoporecurrent traces of the empty pore (d) and with the addition of 50 uM AgNO3 (e) recorded at 150 mV in 1 M KNO3.
Ag+ stabilizes a DNA duplex with C-C mismatches
The addition of Ag+ increases the stability of dsDNA containing a C-C mismatch, which leads to an increase in the complex's dwell time within the nanopore (Figure 2a,b). We can see that ssDNAs (dwell time <10 ms) and dsDNAs (dwell time >10 ms) were well separated (Figure 2c). For details on the probe screening process, please refer to the supplementary information (Supplementary Note S1). The events with an ending spike35,36,37,38 were identified (Figure 2a,b enlarged single current traces), indicating the DNA duplex capturing and dissociation (See Supplementary Note S2 for detailed description). The difference in dwell time provides a key differentiator between C-C and C-Ag-C. In detail, P·TC hybrid (C-C) yielded the dwell time distribution with a peak at 59 ± 5 ms (Figure 2c, blue), while C-Ag-C yielded a dwell time distribution with the first peak at 51 ± 6 ms and the second peak at 384 ± 12 ms (Figure 2c, red). Molecular dynamics (MD) simulations indicate that hydrogen bonds are alternatively formed between N4A-N3B and N3A-N4B atoms (simulations described in details below) and there is a 2.6-fold difference in binding energy bewteen these two conformations. This difference in binding energy could be the reason that we observed two dwell time distributions peaks. This second peak demonstrates dwell times with C-Ag-C that are 6.5-fold longer than those with C-C (Figure 2c). We interpret that the prolonged blocking events are due to the binding of Ag+ to the C-C mismatch in the P·TC hybrid. As reported previously, the binding of Ag+ forms a C-Ag-C bridge base pair that stabilizes the P·TC complex1,2,3,4, resulting in an extended dwell time under the same holding potential. The Ag+ effect is equivalent to an increase in dsDNA hybridization energy, which was calculated to be 3.8 ± 0.5 kJ·mol−1 using ΔE = RTln(τ+Ag/τ−Ag), where τ−Ag and τ+Ag are block durations before and after the addition of Ag+. We also found a decrease in residual current after the addition of Ag+. They are 41.5 ± 0.4 pA (without Ag+) and 36.8 ± 0.2 pA (with Ag+), respectively (Figure 2d). The change is 4.7 ± 0.45 pA (by error propaganda equation). The hydrated radius of Ag+ is 0.34 nm39 and as a result, the substantial radius of Ag+ in complex with the DNA blocks more current flow. Thus it is reasonable to see a deeper current blockage for DNA with Ag+.
Ag+ stabilizes DNA duplex containing C-C mismatches.
(a) The capturing of C-C duplex (ssDNA TC hybridized with P) in the nanopore. (b) The capturing of C-C duplex with the addition of Ag+. (c) The histogram of the dwell time in Log form. The C-C generated a single peak of 59 ± 5 ms (blue). The C-Ag-C generated two peaks of 51 ± 6 ms and 384 ± 12 ms (red), which increased the dwell time by 6.5 fold compared to the C-C duplex. (d) The histogram of residual currents. The C-C generated a single peak of 41.5 ± 0.4 pA (blue); The C-Ag-C generated a peak of 36.8 ± 0.2 pA. The difference was 4.7 ± 0.45 pA between C-C and C-Ag-C. The red circles indicate the capturing of DNA duplexes. The enlarged single traces in a and b demonstrated the DNA duplex dissociation signature with an ending spike. Recordings were made at 150 mV.
We further compared the equilibrium dissociation constant (Kd) for P·TC in the absence and in the presence of Ag+. We have derived an expression to obtain Kd from the block frequency (See Supplementary S1: nanopore measurement of double-stranded DNA equilibrium constant). The expression is Kd = (fss/kon)2/2([ssDNA]0-fss/kon), where kon is the average ssDNA (P or TC) capture rate in the nanopore and fss is the total frequency of blocks generated by unhybridized ssDNA (P and TC) in the mixture. We found that Ag+ can decreases fss from 6.52 ± 0.38 s−1 to 4.10 ± 0.19 s−1. This decrease of fss is also confirmed by an increase of the ton (See Supplementary Figure S1 for definition) for 1.6-fold (Supplementary Figure S2). We found a decrease of Kd from 0.12 ± 0.01 μM−1 to 0.04 ± 0.004 μM−1 (Table 2), suggesting that the stabilization of P·TC by Ag+ shifts the equilibrium of the reaction P + TC↔P·TC toward the product P·TC. The decrease of Kd is expected to increase the melting temperature (Tm). Indeed, the UV measurement shows that Tm for the mixture of P/TC in 1 M KNO3 increased from 28.5 ± 0.6°C (without Ag+) to 43.5 ± 0.6°C (with addition of Ag+), confirming the equilibrium shift toward the duplex formation due to the Ag+ stabilization of dsDNA. Overall, the C-Ag-C bridge-pair functions as an interstrand lock, or SilverLock, that greatly stabilizes dsDNA hybridization. The resulting nanopore signature for SilverLock can identify a single C-C mismatch in a dsDNA.
Weak interaction of Ag+ with a DNA duplex containing mC-C mismatches
The addition of Ag+ also increases the stability of dsDNA containing an mC-C mismatch (probe P is hybridized with the target TmC, their hybrid P·TmC forms a single C-mC mismatch), though the increase in dwell time is less than those for C-C (Figure 3a,b). We found that P·TmC yielded a dwell time distribution peaked at 69 ± 6 ms (Figure 3c, blue), while P·TmC with Ag+ yielded a peak at 92 ± 10 ms (Figure 3c, red), which represents a 1.3-fold increase in dwell time, corresponding to a 0.53 ± 0.07 kJ·mol−1 increase of the energy for dsDNA dehybridization. This energy increase is lower than the 3.8 kJ·mol−1 for dsDNA containing a C-C mismatched base pair bound with Ag+, suggesting that the effect of Ag+ on stabilization of dsDNA with a C-mC mismatch is much weaker than that with a C-C mismatch.
Weak interaction of Ag+ with a DNA duplex containing mC-C mismatches.
The representative current traces of mC-C (a) and mC-Ag-C (b) capturing. (c) The histogram of the dwell time in Log form. The mC-C generated a single peak of 69 ± 6 ms (blue). The mC-Ag-C generated a single peak of 92 ± 10 ms (red), which increased the dwell time by 1.3 fold. (d) The histogram of residual currents. The mC-C generated a single peak of 37.4 ± 0.7 pA (blue). The mC-Ag-C generated two peaks of 33.9 ± 0.8 pA and 38.1 ± 0.8 pA (red). The difference was 3.5 ± 1.1 pA between mC-C and the first peak of mC-Ag-C. The red circles indicate the capturing of DNA duplexes. Recordings were made at 150 mV.
For residual currents, P·TmC yielded a peak at 37.4 ± 0.7 pA and P·TmC with Ag+ yielded two residual current peaks at 33.9 ± 0.8 pA and 38.1 ± 0.8 pA (Figure 2d). The difference was about 3.5 ± 1.1 pA between the peak of mC-C and the first peak of mC-Ag-C (Figure 2d). This suggests that the interaction between mC-C and Ag+ was weaker than that between C-C and Ag+ (See Supplementary Note S3 for discussion).
No observable interaction of Ag+ with a DNA duplex containing hmC-C mismatches
We also measured the effect of Ag+ on the dsDNA containing a C-hmC mismatched base pair (probe P is hybridized with the target ThmC, their hybrid P·ThmC forms a single C-hmC mismatch). The addition of Ag+ does not appear to affect the stability of dsDNA containing an hmC-C mismatch, though dwell time is lower than those for C-C and mC-C mismatches (Figure 4). We found that P·ThmC yielded a dwell time distribution which is very similar to that of P·ThmC with Ag+ (Figure 4a,b,c). The hmC-C yielded a dwell time distribution peaked at 19.6 ± 1 ms (Figure 4c, blue), while hmC-Ag-C yielded a peak at 17.3 ± 1 ms (Figure 4c, red). For residual current, P·ThmC yielded a peak at 36.3 ± 0.95 pA and P·ThmC with Ag+ yielded a similar peak at 36.2 ± 0.71 pA (Figure 4d). The difference was 0.1 ± 1.19 pA. Overall, these data demonstrate that hmC-C mismatches are less stable than mC-C or C-C mismatches. Therefore, the presence of Ag+ seems to have little effect on the C-hmC mismatch.
No observable interaction of Ag+ with a DNA duplex containing hmC-C mismatches.
The representative current traces of hmC-C (a) and hmC-Ag-C (b) capturing. (c) The histogram of the dwell time in Log form. The hmC-C generated a single peak of 19.6 ± 1 ms (blue). The hmC-Ag-C generated a single peak of 17.3 ± 1 ms (red). (d) The histogram of residual currents. The hmC-C generated a single peak of 36.3 ± 0.95 pA (blue); The hmC-Ag-C generated a single peak of 36.2 ± 0.71 pA (red). The difference was 0.1 ± 1.19 pA. The red circles indicate the capturing of DNA duplexes. Recordings were made at 150 mV.
Besides the dwell time, the addition of Ag+ decreased the residual current at different degrees for the tested DNA duplexes, which provides the second key differentiator to discriminate C,mC and hmC (Supplementary Figure S3). We also found that Ag+ does not interact with ssDNAs TC, TmC or ThmC (Supplementary Figure S4).
Molecular dynamics (MD) simulations
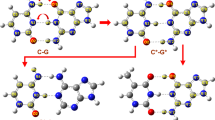
Molecular dynamics (MD) simulations of DNA duplexes containing these mismatches reveal how Ag+ may bind to the mismatches, as well as different coordination configurations between the mismatched bases (Supplementary Note S4 for simulation description). As shown in Figure 5a, a DNA duplex, with the same sequence as that in experiment was solvated in an electrolyte. The C-C base pairing was formed by the hydrogen bond between the N3 atom of one cytosine base (in the strand A) and the N4 atom of the other cytosine base (in the strand B) (Figure 5b). Besides the conformation shown in Figure 5b, another possible paring was formed by the hydrogen bond between N4A and N3B atoms (Supplementary Movie S1). The distances between N3 and N4 atoms of different bases, as shown in Figure 5d, indicate that hydrogen bonds are alternatively formed between N4A and N3B atoms and between N3A and N4B atoms. This type of pairing results in the formation of a binding site for a cation (Figure 5b). During the simulation, K+ ions were found in the binding site and the mean residence time for K+ was about 10 ns (Supplementary Movie S2). As confirmed in an independent MD simulation (Supplementary Figure s5, Movie S3), Ag+ can also enter the binding site and further stabilize the paring between mismatched C-C bases. Correspondingly, these simulations also indicate that the dwell time of the duplex with a Ag+ is longer (Figure 2c) due to the enhanced stability.
Molecular dynamics simulations of DNA duplex containing C-C, mC-C and hmC-C mismatches.
(a) (LBQ is the creator of figure 5a). Side-view of the simulation system. The DNA duplex is in the “stick” presentation and two backbones are illustrated as yellow and green belts respectively. Potassium ions that neutralize the entire simulation system are shown as tan balls. Water in a cubic box (78.5 × 78.5 × 78.5 Å3) is shown transparently. (b) A snap-shot of pairing between two cytosine bases. The dashed circle highlights the binding site for a cation. (c) A snap-shot of hmC-C pairing before the pairing was broken. (d–f) Time-dependent distances between the N3 atom of one base and the N4 atom of the other base, in C-C(d), mC-C(e) and hmC-C(f) mismatches.
The simulations also reflect our experimental results for the differences in stability between the complexes. Figure 5e shows that, because of the switching between the two states of N4A-N3B and N3A-N4B (Figure 5b), the hydrogen bonds were formed and broken more frequently in mC-C compared to the C-C mismatch (Supplementary Movie S4). Additionally, the probability for having longer bond lengths was higher for the mC-C than for the C-C mismatch (Supplementary Figure S6). Therefore, these results suggest that the cation binding site in the mC-C duplex was less stable than in the C-C duplex, consistent with the experimental results that the dwell time of C-Ag-C was longer than mC-Ag-C duplex (Figure 2c, Figure 3c). Interestingly, for the duplex with the hmC-C, the base pairing was broken at about 25 ns during the simulation (Figure 5f, Supplementary Movie S5). Right before the breakage, Figure 5c shows that, because of the hydrogen bond between the hydroxyl group in the hmC base and the phosphate group, the hmC base rotated towards the backbone of the duplex. Such interaction could also be mediated by a water molecule (Supplementary Figure S7). Meanwhile, base pairing was formed between the O2 atom in the hmC base and the N4 atom of the C base. After the breakage, the hmC and C bases can temporarily form inter-strand base-stacking, which causes the breakage of a neighboring base-pair. Because the binding site falls apart in the duplex with the hmC-C mismatch, the effect of Ag+ on the dwell time should be negligible, as also demonstrated in nanopore experiments with hmC-C (Figure 4). Overall, this shows tight agreement between the theoretical and experimental results.
Discussion
Studies have shown that Ag+ forms dinuclear complexes with cytosine and the complexes have been observed by X-ray diffraction. This study suggests that each of the methylcytosine residues doubly cross-linked by two Ag+ at the base binding sites N3 and O211. Thermodynamic properties of C-Ag-C complexes were studied by isothermal titration calorimetry (ITC) and circular dichroism (CD) and the results suggest that the specific binding between the Ag+ and the single C-C mismatch was mainly driven by the positive dehydration entropy change of Ag+ and the negative binding enthalpy change from the bond formation between the Ag+ and the N3 positions of the two cytosine bases4,10. However, our MD simulation of C-Ag-C shows that Ag+ is dynamically coordinated between N3A and O2B, or N3B and O2A (Figure 5b, Supplementary Figure S5). This finding suggests that the coordination of Ag+ in C-Ag-C complexes may have a different mechanism.
Different binding affinities for Ag+ ions with DNA duplexes containing C-C, mC-C or hmC-C could be explained in several ways. Firstly, the Tm measurement demonstrates that Ag+ coordination raises the melting temperature through the stabilization effect of Ag+ on the C-C containing duplexes. Secondly, previous MD simulations found that H2O molecules have the highest affinity for hmC when compared to C and mC, which increases the rotation probability29. Our MD simulation revealed that the water molecule can mediate or directly interact with the phosphate group and the hydroxyl group in hmC. These results suggest a mechanism behind the lower stability of the base-pairing in hmC-C mismatches. Thirdly, using atomic force microscopy (AFM), studies have found that the persistence length follows the trend mC > C > hmC29, suggesting that hmC-containing DNA has the largest flexibility and least structural stability. Finally, the –OH group in hmC can chelate with the phosphate group40 which may prevent a stable hmC-Ag-C complex formation.
Conclusion
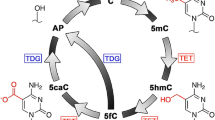
Overall, we have demonstrated that chemical interactions between Ag+ and cytosine and its modifications could be applied to study C, mC and hmC differences. Without Ag+, the residual current follows C-C > mC-C > hmC-C (Figure 2,3,4d, blue; Figure S3a) and the dwell time follows mC-C > C-C > hmC-C (Figure 2,3,4c, blue). The residual current differences with the addition of Ag+ are C-C > mC-C > hmC-C (Figure 2,3,4d and Figure S3a,b). The dwell time differences (ratios) with the addition of Ag+ are also C-C > mC-C > hmC-C (Figure 2,3,4c). With these two key differentiators, we can discriminate C, mC and hmC bases. It is therefore concluded that the C-Ag-C mismatch is the most stable and the hmC-Ag-C is the least stable. This direct discrimination was successfully demonstrated without modification and amplification of target DNA. We also demonstrated that it is a dynamic coordination between Ag+ and C-C mismatches, which indicates a new binding mechanism. By utilizing the chemical interactions with metal ions, this approach might be extended to study other cytosine modifications, such as 5-formylcytosine (5fC) and 5-carboxylcytosine and to investigate metallo-pair interactions41,42, including copper ion-stabilizing pyridine-2,6-dicarboxylate-pyridine mismatches and silver/mercury interacting with modified uracil pairs. Finally, it is also possible that a target fragment of a genomic sample could be obtained by a suite of restriction endonucleases. The target fragments can then be purified and segregated for nanopore research.
Methods
Electrophysiology and single channel recording
The electrophysiology setup and nanopore experimental methods have been well-documented43. Briefly, the recording apparatus was composed of two chambers (cis and trans) that were partitioned with a Teflon film. The planar lipid bilayer of 1,2-diphytanoyl-sn-glycerophosphatidylcholine (Avanti Polar Lipids) was formed spanning a 100–150 μm hole in the center of the partition. The α-hemolysin (αHL) protein monomers (Sigma, St. Louis, MO) can be self-assembled in the bilayer to form molecular pores, which can last for hours during electrical recordings. Both cis and trans chambers were filled with symmetrical 1 M salt solutions (KNO3) buffered with 10 mM 3-(N-morpholino)propanesulfonic acid (Mops)2 and titrated to pH 7.02. All solutions were filtered before use. DNA oligonucleotides (Table 1) were synthesized and electrophoresis purified by Integrated DNA Technologies (IDT), IA. Before testing, the mixtures of DNA and probes were heated to 90°C for 5 minutes and then slowly cooled to room temperature. Single-channel currents were recorded with an Axopatch 200A patch-clamp amplifier (Molecular Device Inc., former Axon Inc.), filtered with a built-in 4-pole low-pass Bessel Filter at 5 kHz and acquired with Clampex 9.0 software (Molecular Device Inc.) through a Digidata 1332 A/D converter (Molecular Device Inc.) at a sampling rate of 20 kHz·s−1. DNAs were presented in the solution on cis side of the pore (grounded) and a holding potential was applied from the trans side to produce an ion current across the pore. Data was based on at least four separate experiments and obtained by single channel search. The histograms were fitted by exponential log probability (dwell time histogram distribution) or Gaussian function (residual current histogram distribution). The red circles in each figure represent the capturing of DNA duplex in the nanopore. The electrophysiology experiments were conducted at 22 ± 1°C. Data was presented as AVE ± SD (average ± standard deviation).
The ratio of Ag+ to DNA duplex was set to 100:1 in all the experiments. Varying the concentration of Ag+ (50X, 500X) does not change the number of DNA duplex capturing events significantly. This was similar to the previous findings that the melting temperature reached a plateau when the Ag+ concentration was 1.5 fold higher than the DNA2. By isothermal titration calorimetry (ITC) and electrospray ionization mass spectrometry measurement, the binding of Ag+ to a DNA duplex containing a single C-C mismatch was identified at a 1:1 molar ratio4,10. The lines under each current trace mark the 0 current.
Melting temperature measurement
The melting temperatures of duplexes containing C-C, mC-C, or hmC-C mismatches were determined by monitoring the increase in absorbance at 260 nm as a function of temperature (Cary 100 Bio UV-Visible spectrophotometer). The temperature was increased from 4°C to 50°C (for samples without Ag+ ion), or from 10°C to 60°C (for samples with Ag+), at a rate of 0.5°C/min. P/TC (2/2 μM) and 2 μM Ag+ ions were used in the experiment, because previous studies found that the melting temperature reached a plateau when the silver(I) ion concentration was 1.5 fold higher than the DNA2. The melting temperature was calculated from the collected data using the Cary WinUV Thermal software. Each sample was repeated at least three times.
Molecular dynamics simulation
The software NAMD44 was used to perform all-atom MD simulation on the IBM bluegene supercomputer. Force fields used in simulations were the CHARMM2745 for DNA, the TIP3P46 model for water molecules and the standard one47 for ions. Long-range coulomb interactions were computed using the particle-mesh Ewald (PME) method. A smooth (10–12 Å) cutoff was used to compute the van der Waals interaction. After each simulation system was equilibrated at 1 bar, following simulations were carried out in the NVT (T = 300 K) ensemble. The temperature of a simulated system was kept constant by applying the Langevin dynamics on Oxygen atoms of water molecules.
Change history
10 July 2015
A correction has been published and is appended to both the HTML and PDF versions of this paper. The error has been fixed in the paper.
References
Lin, Y. H. & Tseng, W. L. Highly sensitive and selective detection of silver ions and silver nanoparticles in aqueous solution using an oligonucleotide-based fluorogenic probe. Chem Commun 43, 6619–21 (2009).
Ono, A. et al. Specific interactions between silver(I) ions and cytosine-cytosine pairs in DNA duplexes. Chem Commun 39, 4825–7 (2008).
Wen, Y. et al. A graphene-based fluorescent nanoprobe for silver(I) ions detection by using graphene oxide and a silver-specific oligonucleotide. Chem Commun 46, 2596–8 (2010).
Torigoe, H. et al. Thermodynamic and structural properties of the specific binding between Ag(+) ion and C:C mismatched base pair in duplex DNA to form C-Ag-C metal-mediated base pair. Biochimie 94, 2431–40 (2012).
Miyake, Y. et al. MercuryII-mediated formation of thymine-HgII-thymine base pairs in DNA duplexes. J Am Chem Soc 128, 2172–3 (2006).
Tanaka, Y. et al. 15N-15N J-coupling across Hg(II): direct observation of Hg(II)-mediated T-T base pairs in a DNA duplex. J Am Chem Soc 129, 244–5 (2007).
Torigoe, H., Ono, A. & Kozasa, T. Hg(II) ion specifically binds with T:T mismatched base pair in duplex DNA. Chemistry 16, 13218–25 (2010).
Wen, S. et al. Highly sensitive and selective DNA-based detection of mercury(II) with alpha-hemolysin nanopore. J Am Chem Soc 133, 18312–7 (2011).
Ono, A., Torigoe, H., Tanaka, Y. & Okamoto, I. Binding of metal ions by pyrimidine base pairs in DNA duplexes. Chem Soc Rev 40, 5855–66 (2011).
Torigoe, H., Miyakawa, Y., Ono, A. & Kozasa, T. Thermodynamic properties of the specific binding between Ag+ ions and C:C mismatched base pairs in duplex DNA. Nucleosides Nucleotides Nucleic Acids 30, 149–67 (2011).
Kistenmacher, T. J., Rossi, M. & Marzilli, L. G. Crystal and molecular structure of (nitrato)(1-methylcytosine)silver(I): An unusual cross-linked polymer containing a heavy metal and a modified nucleic acid constituent. Inorganic Chemistry 18, 240–244 (1979).
Urata, H., Yamaguchi, E., Nakamura, Y. & Wada, S. Pyrimidine-pyrimidine base pairs stabilized by silver(I) ions. Chem Commun 47, 941–3 (2011).
Brena, R. M., Huang, T. H. & Plass, C. Quantitative assessment of DNA methylation: Potential applications for disease diagnosis, classification and prognosis in clinical settings. J Mol Med (Berl) 84, 365–77 (2006).
Kriaucionis, S. & Heintz, N. The nuclear DNA base 5-hydroxymethylcytosine is present in Purkinje neurons and the brain. Science 324, 929–30 (2009).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–22 (2009).
Clarke, J. et al. Continuous base identification for single-molecule nanopore DNA sequencing. Nat Nanotechnol 4, 265–70 (2009).
Branton, D. et al. The potential and challenges of nanopore sequencing. Nat Biotechnol 26, 1146–53 (2008).
Bayley, H. Sequencing single molecules of DNA. Curr Opin Chem Biol 10, 628–37 (2006).
Bayley, H. & Cremer, P. S. Stochastic sensors inspired by biology. Nature 413, 226–30 (2001).
Gu, L. Q. & Shim, J. W. Single molecule sensing by nanopores and nanopore devices. Analyst 135, 441–51 (2010).
Howorka, S. & Siwy, Z. Nanopore analytics: sensing of single molecules. Chem Soc Rev 38, 2360–84 (2009).
Deamer, D. Nanopore analysis of nucleic acids bound to exonucleases and polymerases. Annu Rev Biophys 39, 79–90 (2010).
Olasagasti, F. et al. Replication of individual DNA molecules under electronic control using a protein nanopore. Nat Nanotechnol 5, 798–806 (2010).
Chu, J., Gonzalez-Lopez, M., Cockroft, S. L., Amorin, M. & Ghadiri, M. R. Real-time monitoring of DNA polymerase function and stepwise single-nucleotide DNA strand translocation through a protein nanopore. Angew Chem Int Ed Engl 49, 10106–9 (2010).
Benner, S. et al. Sequence-specific detection of individual DNA polymerase complexes in real time using a nanopore. Nat Nanotechnol 2, 718–24 (2007).
Li, W. W., Gong, L. & Bayley, H. Single-molecule detection of 5-hydroxymethylcytosine in DNA through chemical modification and nanopore analysis. Angew Chem Int Ed Engl 52, 4350–5 (2013).
Shim, J. et al. Detection and quantification of methylation in DNA using solid-state nanopores. Scientific Reports 3 (2013).
Wallace, E. V. et al. Identification of epigenetic DNA modifications with a protein nanopore. Chem Commun 46, 8195–7 (2010).
Wanunu, M. et al. Discrimination of methylcytosine from hydroxymethylcytosine in DNA molecules. J Am Chem Soc 133, 486–92 (2011).
Yu, M. et al. Tet-assisted bisulfite sequencing of 5-hydroxymethylcytosine. Nat Protoc 7, 2159–70 (2012).
Yu, M. et al. Base-resolution analysis of 5-hydroxymethylcytosine in the mammalian genome. Cell 149, 1368–80 (2012).
Booth, M. J. et al. Quantitative sequencing of 5-methylcytosine and 5-hydroxymethylcytosine at single-base resolution. Science 336, 934–7 (2012).
Kowalczyk, S. W., Wells, D. B., Aksimentiev, A. & Dekker, C. Slowing down DNA translocation through a nanopore in lithium chloride. Nano letters 12, 1038–1044 (2012).
de Zoysa, R. S. S. et al. Slowing DNA translocation through nanopores using a solution containing organic salts. The Journal of Physical Chemistry B 113, 13332–13336 (2009).
Kang, I. et al. Designing DNA interstrand lock for locus-specific methylation detection in a nanopore. Scientific reports 3 (2013).
Wang, Y., Zheng, D., Tan, Q., Wang, M. X. & Gu, L. Q. Nanopore-based detection of circulating microRNAs in lung cancer patients. Nat Nanotechnol 6, 668–74 (2011).
Tian, K., He, Z., Wang, Y., Chen, S.-J. & Gu, L.-Q. Designing a Polycationic Probe for Simultaneous Enrichment and Detection of MicroRNAs in a Nanopore. ACS nano 7, 3962–3969 (2013).
Zhang, X., Wang, Y., Fricke, B. L. & Gu, L.-Q. Programming Nanopore Ion Flow for Encoded Multiplex MicroRNA Detection. ACS nano (2014).
Nightingale, E. R., Jr Phenomenological theory of ion solvation. Effective radii of hydrated ions. Journal of Physical Chemistry 63, 1381–1387 (1959).
Gunther, L, E. et al. The Effect of Metal Ions on the Structure of Nucleic Acids. in Bioinorganic Chemistry Vol. 100 135–154 (AMERICAN CHEMICAL SOCIETY, 1971).
Atwell, S., Meggers, E., Spraggon, G. & Schultz, P. G. Structure of a copper-mediated base pair in DNA. Journal of the American Chemical Society 123, 12364–12367 (2001).
Okamoto, I., Iwamoto, K., Watanabe, Y., Miyake, Y. & Ono, A. Metal-ion selectivity of chemically modified uracil pairs in DNA duplexes. Angew Chem Int Ed Engl 48, 1648–51 (2009).
Shim, J. W., Tan, Q. & Gu, L. Q. Single-molecule detection of folding and unfolding of the G-quadruplex aptamer in a nanopore nanocavity. Nucleic Acids Res 37, 972–82 (2009).
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J Comput Chem 26, 1781–802 (2005).
MacKerell, A. D., Jr et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. Journal of Physical Chemistry B 102, 3586–3616 (1998).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. The Journal of Chemical Physics 79, 926–935 (1983).
Beglov, D. & Roux, B. Finite representation of an infinite bulk system: Solvent boundary potential for computer simulations. The Journal of Chemical Physics 100, 9050–9063 (1994).
Acknowledgements
We thank Dr Edward H. Blaine for invaluable discussions on the experiment design and data analysis. This work was supported by grants from NSF CAREER 0546165 (LQG), NIH GM079613 (LQG).
Author information
Authors and Affiliations
Contributions
Y.W. conceived the principal idea and designed the experiments. Y.W. performed the nanopore experiments, collected and analyzed the nanopore data. B.-Q.L. designed and performed the molecular dynamics simulation, collected and analyzed the data. Z.-Y.Y. and K.G. designed and performed the melting temperature measurements, collected and analyzed the data. Y.W., B.-Q.L. and L.-Q.G. wrote the manuscript. All the authors, Y.W., B.-Q.L., Z.-Y.Y., K.G., X.-Y.Z., B.R. and L.-Q.G. discussed the results and commented on the manuscript and co-wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Wang, Y., Luan, BQ., Yang, Z. et al. Single Molecule Investigation of Ag+ Interactions with Single Cytosine-, Methylcytosine- and Hydroxymethylcytosine-Cytosine Mismatches in a Nanopore. Sci Rep 4, 5883 (2014). https://doi.org/10.1038/srep05883
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep05883
This article is cited by
-
Role of pH in the stability of cytosine-cytosine mismatch and canonical AT and GC base pairs mediated with silver ion: a DFT study
Structural Chemistry (2022)
-
Nanopore electric snapshots of an RNA tertiary folding pathway
Nature Communications (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.