Abstract
Super resolution microscopy such as STORM and (F)PALM is now a well known method for biological studies at the nanometer scale. However, conventional imaging schemes based on sparse activation of photo-switchable fluorescent probes have inherently slow temporal resolution which is a serious limitation when investigating live-cell dynamics. Here, we present an algorithm for high-density super-resolution microscopy which combines a sparsity-promoting formulation with a Taylor series approximation of the PSF. Our algorithm is designed to provide unbiased localization on continuous space and high recall rates for high-density imaging and to have orders-of-magnitude shorter run times compared to previous high-density algorithms. We validated our algorithm on both simulated and experimental data and demonstrated live-cell imaging with temporal resolution of 2.5 seconds by recovering fast ER dynamics.
Similar content being viewed by others
Introduction
Single-molecule localization microscopy methods, such as STORM1 and (F)PALM2,3, utilize sparse activation of photo-switchable fluorescent probes in both temporal and spatial domains. Each activated probe can be assimilated to an ideal point source so that the acquired images consist of isolated replicates of the point spread function of the microscope (PSF). This allows one to achieve sub-pixel accuracy on the order of tens of nanometers for the estimated location of each probe1,2,3,4,5,6. In general, reconstruction of sub-cellular structures relies on numerous localized probes and the required acquisition time of these methods is therefore relatively long, i.e. on the order of minutes. This is a serious limitation when investigating live-cell dynamics.
One possible approach for overcoming this limitation is high-density imaging7. By increasing the density of activated probes, shorter acquisition times for a single super-resolution image can be achieved. However this complicates the localization task due to overlapping PSFs. DAOSTORM7, for example, fits multiple overlapping PSFs in an iterative manner by analyzing pixel clusters in the residual image. The positions of the probes are determined by minimizing a least-squares criterion. CSSTORM8 (Compressed sensing STORM) and deconSTORM9 (deconvolution STORM) impose sparsity priors on the distribution of probes. In CSSTORM algorithm, the localization task is formulated as a convex optimization problem and solved by means of linear programming, while deconSTORM uses a modified Lucy-Richardson deconvolution algorithm by exploiting temporal correlation of activated probes. In general, these sparsity promoting methods provide increased recall rates compared to multi-emitter fitting at the expense of higher computational complexity. In a different approach, super resolution optical fluctuations imaging (SOFI)10 and 3B analysis11 utilize stochastic photon-emission processes such as photo-bleaching and blinking to reconstruct high-density data. For example, 3B analysis based on realistic models of photo-bleaching and blinking processes reconstructs the high-density data using a Bayesian approach. The Current implementation of 3B analysis has a relatively long reconstruction time but it can be made faster by using a computationally efficient Bayesian algorithm or parallel computing12.
In addition, all of these sparsity-promoting methods are based on similar discrete formulations. They reconstruct a high resolution image on a pre-defined sub-pixel grid, e.g., with a pixel size of 20 nm. Such formulations, however, have three inherent limitations. First, discrete-domain formulations can account for only pre-defined locations, not all possible probe locations over a continuum. Therefore, these methods need to extract the localization information from their reconstructed high resolution images. A partial solution to this problem is to compute local centers of mass in the reconstructed image8, but this tends to result in a biased estimation of the probe locations. Second, using a finer sub-pixel grid increases the computational load, especially with the linear-programming approach used in CSSTORM. The final issue is the underestimation of photon counts. Indeed, many sparse solvers are based on iterative thresholding, which often truncates signals toward zero, resulting in a spatial bias of the recovered locations of the probes.
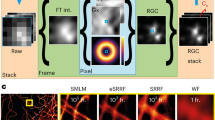
To address these limitations of current high-density localization algorithms, we introduce a FAst Localization algorithm based on a CONtinuous-space formulation (FALCON) for high density super-resolution microscopy data. In particular, to obtain a grid-free reconstruction, our approach combines a sparsity-promoting formulation with a Taylor approximation of the PSF. It consists of three main stages: deconvolution with a sparsity prior, deconvolution with a fixed spatial support and continuous-domain refinement (Fig. 1 and 2 and Supplementary Fig. 1, Supplementary note). The first step generates a high-resolution sparse image similarly to CSSTORM. The second deconvolution step corrects underestimated pixel values. The last stage uses a Taylor series approximation of the PSF for refining the position of each probe over a continuum. It also refines the photon count of each probe. For the Taylor expansions of the PSF with respect to the spatial offset from the grid location, we only use linear terms to make the optimization procedure faster. Importantly, this approximation is not limited to Gaussian functions. The final output of the algorithm are positions and photon counts of the localized probes. All these steps are implemented in a computationally efficient way by utilizing ADMM (Alternating Direction Method of Multipliers)13 for fast reconstruction of the high resolution sparse images and by using alternating refinements.
Schematic illustration for FALCON.
For a detailed description, see Supplementary note.
Taylor approximation of the shifted PSF for continuous refinement of FALCON.
(a) First-order Taylor approximation of the shifted PSF: the shifted PSF is approximated by the PSF centered on the sub-pixel and the derivative of the PSF. (b) Approximation error of Gaussian PSF of 360 nm FWHM. The error of the approximation does not exceed 2% up to 30 nm displacement.
Results
We validated the performance of the proposed algorithm using both simulated and real STORM/PALM data. First, we confirmed that the linear approximation of the PSF provides a good approximation for a PSF that is shifted by a few nanometers from the sub-pixel grid. Specifically, the relative approximation error is less than 2% for a displacement of up to 30 nm for a Gaussian function with a FWHM (Full Width at Half Maximum of 360 nm (Fig. 2b). Next, we investigated the localization performance of our algorithm for a single probe. Starting from a uniform probability distribution over an area of a single sub-pixel of 33.3 nm (Fig. 3a–c), we localized 50, 000 individual probes using two methods: center-of-mass on a deconvolved image8 and our continuous refinement by Taylor series approximation. The former method resulted in estimations that were strongly biased towards the sub-pixel grid points, while our approach produced a much more uniform distribution of the estimated particle locations. Similar results were also observed for probes that were uniformly distributed along a diagonal line (Fig. 3d–f). These simulations were performed with high photon-emission rates (5, 000 photons on average, standard deviation of 2, 000).
Numerical analysis of the continuous refinement on 50,000 simulated images.
In each image, a single molecule is randomly placed with uniform probability distribution within a single sub-pixel area (a–c) or along a diagonal line (d–f). Histograms of ground-truth positions (a,d), histograms of the centroid fit that is applied to the high resolution images reconstructed by a sparsity-based deconvolution (l1 minimization) (b,e) and histograms of FALCON (c,f).
We also compared our algorithm with a least-square fitting method, DAOSTORM and CSSTORM over a wide range of imaging densities. We simulated two different photon-emission rates: high (5, 000 photons on average, standard deviation of 2, 000) and low (350 photons on average, standard deviation of 70). The FWHM of the PSF was fixed at 385 nm. For the high photon-emission rate simulations (Fig. 4a and b), FALCON and CSSTORM achieved two times higher recall rates than DAOSTORM did for densities larger than 6 μm−2. In terms of localization accuracy, the proposed algorithm outperforms all the other algorithms in high imaging densities (4–9.3 μm−2) and our method is compatible with the least-square fitting showing only 5% difference in the lowest imaging density. For the low photon-emission rate simulations (Fig. 4c and d), the improvement is even more distinct in terms of recall rate and FALCON demonstrates better localization accuracy over the whole range of imaging densities (0.05–6.5 μm−2).
Performances of FALCON in comparison with Least-square fitting, DAOSTORM and CSSTORM over a wide range of imaging densities.
Simulation on the random distribution of molecules over a wide range of imaging densities with high-photon emission rates: recall rates (a), localization accuracy (b) and low-photon emission rates: recall rates (c), localization accuracy (d). The error bars represent standard deviations.
The performance dependency on the size of PSF was also investigated. Fixing the imaging density to 6 μm−2, we investigated the performance of FALCON, DAOSTORM and CSSTORM for various FWHMs in the range of 250–430 nm (Fig. 5). As the FWHM becomes smaller, the performance of all algorithms improves, leading to comparable results. Nevertheless, as the FWHM becomes larger, FALCON performs better than CSSTORM, leading to better accuracy and higher recall rates with a notable improvement in the low photon-emission cases (Fig. 5c and d). DAOSTORM always shows minimum recall rates and the differences with the others are very clear with the broader PSFs.
Performances of FALCON in comparison with DAOSTORM and CSSTORM over a wide range of PSF widths from 250 nm to 430 nm.
Simulated data with high-photon emission rates: (a) molecular recall rates, (b) localization accuracy and low-photon emission rates: (c) molecular recall rates, (d) localization accuracy. The error bars indicate standard deviations.
In order to quantify the performance of our algorithm on specific geometric structures, we simulated circular structures whose radii varied between 150 nm and 400 nm (Fig. 6). In every frame, 4 activated probes are randomly placed on the circle, resulting in overlapping PSFs even at the largest radius. We estimated the radius of the reconstructed circle by averaging the distances between all localized probes from 300 simulated images and the center of the circle. The results of the simulation demonstrate that our algorithm performs better for extracting this geometrically relevant measure, over the entire range of simulations. In contrast, the values by CSSTORM were underestimated by up to 18 nm as a result of underestimated photons; CSSTORM also shows spurious peaks as artifacts in the histogram (Fig. 6b) because of the discrete formulation as shown in (Fig. 3). These artifacts were not created by FALCON and DAOSTORM. In terms of the estimation bias, DAOSTORM was comparable to FALCON, but it had smaller recall rates and larger standard deviation of localized locations (Fig. 6c).
Performance analysis for synthetic “ring” phantoms with various radii.
(a) Rendered images of the algorithms overlaid by a cyan circle with 150 nm radius as a ground truth. (b) Histogram of the distances from localized molecules by FALCON, DAOSTORM and CSSTORM to the center of the ring in (a). (c) The differences between the true radius and the distances on average along various radii from 150 nm to 400 nm with 4 fixed molecules on the circle in every simulated image. The error bars represent standard deviations.
Our ADMM implementation of the two deconvolution stages exhibits fast convergence and the refinement stage is computationally efficient thanks to the closed-form expression of each update step. For example, on an Intel i7 3.4 GHz CPU, our stand-alone Matlab implementation of FALCON took 5 minutes to reconstruct 100 high-density (6 μm−2) STORM images of 100 μm2 (100 × 100 pixels); on a GPU (Nvidia GTX Titan) it took FALCON 40 seconds to reconstruct the same data. Since the run time of our method mainly depends on the raw camera size and over-sampling factor M for the reconstruction grid, the computation cost will be further reduced by using a coarser sub-pixel grid for the deconvolution steps. In FALCON, the role of the deconvolution steps is mainly to estimate the number of sources and initial locations and these initial values can be refined. We demonstrated that FALCON using coarser grid of 50 nm provides compatible results at the small expense of recall rates in comparison with FALCON using the finer grid of 33.3 nm in spite of only using reduced number of variables to estimate as shown in Supplementary Fig. 5.
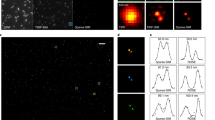
We demonstrate the performance of FALCON on a real STORM data of fixed microtubules (MTs). FALCON reconstructed alpha tubulin subunits of microtubules labeled with Alexa 647 in Cos-7 cells, by using 500 frames imaged at 25 Hz (Fig. 7a). For the quantitative analysis, as increasing a number of raw frames to reconstruct a image, we measured resolutions of the reconstructed images by means of Fourier ring correlation (FRC) criteria14. For the FRC analysis, each half of localized locations were binned into an image of 5nm pitch and cross correlation between the two images was calculated, then converted as a FRC resolution. FALCON shows a resolution improvement of up to 20% over the other high-density algorithms (Fig. 7b). In other words, FALCON can achieve the same spatial resolution using 100 times fewer frames. The superiority of FALCON is especially visible at the intersection of MTs. FALCON offers an unbiased estimation of distances between MTs, showing fewer spurious and noisy peaks than other high density algorithms (Fig. 7c). This is in accordance with our results concerning the radii of simulated circles. FALCON also provides a reliable distribution of photon counts which are well matched to results of another low-density MT data, acquired under identical preparation protocols15, whereas DAOSTORM (green line) loses many molecules with photon counts <5,000 in comparison and CSSTORM yields many false-positive molecules with photon counts <500 (Supplementary Fig. 12). Furthermore, the proposed method showed improvements even in low-density data; especially around regions of complex MT structures or inserted fiducial beads, in comparison with Least-square fitting method (Supplementary Fig. 13).
FALCON performance on fixed microtubules data.
(a) FALCON reconstructed α-tubulin subunits of microtubules (MTs) labeled with Alexa 647 in Cos-7 cells; 500 raw frames were used to reconstruct a super-resolution image. White box highlights the intersection of MTs reconstructed by FALCON. (b) Fourier ring correlation(FRC) analysis on the reconstructed images by DAOSTORM, CSSTORM and FALCON. For the FRC analysis, fiducial beads indicated by yellow circles are discarded. The error bars represent standard deviations. Cross-sectional profiles across region in white box (yellow line) were measured by DAOSTORM, CSSTORM and FALCON and plotted in (c). Scale bars are 1 μm in (a).
We also applied FALCON to reconstruct live, PALM data. In particular, the dynamics of the endoplasmic reticulum (ER) were captured in U2OS cells labeled with tdEos fused to reticulon-4; reconstructed movies were made, with each frame being a single PALM image acquired over 2.5 seconds (160 frames of raw data). Over the course of 20 seconds, we observed several changing features of the ER network. In particular, expanding and shrinking tubules were visualized along with the dynamic motion of tubule junction points (Fig. 8f–j, white arrow). We also captured distinct features of this organelle reshaping itself: the disappearance of tubule junctions (Fig. 8f–j, dotted circle) and emergence of tubules (Fig. 8f–j, blue arrow) to form new junction points are just a few examples of this. Importantly, we measured an average tubule thickness of approximately 60 nm (Fig. 8g), which is in line with previously reported diameter measurements16,17. Furthermore, we compared super-resolution images reconstructed by DAOSTORM, CSSTORM and FALCON by increasing accumulation time scales (1–3 s) in Supplementary Fig. 14. FALCON shows better reconstruction of complex ER structures than the others and detected more molecules, up to 50% in comparison with DAOSTORM and CSSTORM, which can improve temporal resolution.
FALCON performance on live ER data.
Live imaging of the endoplasmic reticulum protein, reticulon-4 fused to tdEos imaged over 20 seconds in a U2OS cell. Conventional (a–e) and super-resolution (f–j) snapshots are shown with a 2.5 seconds temporal resolution. The average size of tubules measured from the reconstruction is approximately 60 nm (at FWHM). Yellow markers in (g) highlight a representative tubule width. The dynamic motions of these structures are highlighted in (g–j) indicated by white and blue arrows and dashed, yellow circles. All scale bars are 1 μm.
Discussion
We presented a fast localization algorithm for high-density data which incorporates a sparsity promoting formulation together with a Taylor series approximation of the PSF for continuous localization. The robustness of the proposed method has been extensively studied by simulated and experimental data in comparison with previous high-density algorithms based on multi-emitter fitting (DAOSTORM) or deconvolution using sparsity-promoting priors (CSSTORM and deconSTORM). Localization performances have been quantitatively analyzed not only in terms of recall rates and localization accuracy, but also in terms of localization bias. Importantly, even though localization bias can be easily observed in case of high-density imaging and is important for quantitative biological analyses, it has not been properly investigated before7,8,9.
We have shown that a localization bias can be induced by several factors. Low recall rates is one factor, which can occur by counting multiple closely-spaced probes as one and then localizing the centroid position. In general, a deconvolution based approaches utilizing a sparsity promoting priors can achieve higher recall rates than multi-emitter fitting based methods by effectively resolving closely placed probes. However, this approach can introduce an additional bias for location estimation mainly originating from its use of the discrete sparsity-promoting formulation. Our experiments have illustrated grid-pattern artifacts and spatial distortions appearing in the final image reconstructed when using the discrete sparsity-promoting formulation. In particular, spatial distortions can be problematic when imaging intracellular structures. For instance, quantifying the sizes of mitochondria, microtubules or the tubules of the ER would be done with less confidence, since structures would be falsely constricted. On the other hand, the proposed method based on the continuous formulation minimized these bias errors and has shown almost bias-free localization results.
In live imaging, the variance of the localization accuracy is composed of both the algorithmic localization variance and the variance of the fluctuations. Indeed, the motion artifacts due to dynamic structures that move on time-scales shorter than our exposure time can be problematic, often resulting in artificially larger or smaller structures- as previously reported18. In the ER experiment, we sought to probe the dynamic motion of ER tubules. Previous studies19,20, with similar exposure times demonstrated that motion of this organelle is dynamic on longer timescales than the one used here.
Our method is currently using a two dimensional PSF model which is appropriate to a relatively flat sample region having a thickness below 1 μm. However, it must be noted that our continuous sparsity-promoting formulation is not limited to a two-dimensional Gaussian PSF model. It can be easily extended to non-Gaussian models including three dimensional PSF models. For example, multi-plane imaging systems with three-dimensional PSF engineering techniques can be accommodated with our formulation.
The proposed formulation with a Gaussian noise assumption can be further improved by considering more realistic noise statistics. Since many state-of-the-art cameras for localization microscopy, such as an EMCCD (Electron multiplying charge-coupled device) or sCMOS(scientific complementary metal–oxide–semiconductor) camera, have very small read-out noise21, Poissonian shot noise becomes a dominant noise source to resolve. Basically, exploiting Poisson statistics22 would be more advantageous especially for live-imaging data, which is likely to be achieved at low SNR due to low emission photons and high background. However, even with our current implementation based on a Gaussian noise model, at the SNR levels from our experimental conditions, the proposed algorithm is robust under Poisson noise statistics.
Concerning a modeling of readout noise for our simulations, we assumed that a CCD-type camera was used, where readout noise is known to be well approximated by the normal distribution. The well-established high-density algorithms such as DAOSTORM and CSSTORM also assumed Gaussian readout noise of small variance for their simulations7,8. Although sCMOS cameras have readout noise with pixel-to-pixel variations resulting in a non-Gaussian distribution such as the log-normal distribution23, our simulation results indicate that the proposed algorithm is not sensitive to the choice of the noise model (Supplementary Fig. 9).
In general, it is difficult to make a fair comparison between algorithms in terms of run time due to the difference in implementation environment. Since the speed of the algorithm is determined by various factors including optimization, programming language, operating system and usage of parallel implementation, the implementation environment must also be thoroughly evaluated to maximize calculation efficiency. For example, a computationally optimized CSSTORM implementation24 is faster by two orders of magnitude than the initial MATLAB-based implementation8 which takes approximately one day to reconstruct a 100 μm2 region of 100 raw frames of STORM data with high-density activation(6 μm−2) on an Intel i7, 3.4 GHz CPU.
Although the speed of our approach is also dependent on the implementation environment, a generally fast algorithm speed can be achieved since the algorithm starts from a coarser grid which inherently reduces the problem complexity. Specifically, the proposed method would be very beneficial for three dimensional localization problems with high-density data, where calculation time reduction is critical since discrete formulation requires numerous finer voxels to be estimated.
In conclusion, Taylor approximation of the PSF can be jointly utilized with sparsity-promoting formulation for high-density imaging to yield continuous localization with reduced complexity (up to several orders of magnitude). Furthermore, we have also investigated how a spatial bias is introduced in the localization estimation of high-density imaging. We also have showed that the proposed continuous formulation with specially designed sparsity priors substantially reduces the bias and provides better localization accuracy and higher recall rates than other currently available methods over a wide variety of experimental conditions. Moreover, our method is efficient in terms of implementation, which reduces computation time by orders of magnitude in comparison with the previous high density algorithms. These results were well confirmed through experimental data obtained using fixed microtubules sample. We also successfully reconstructed fast dynamic motions of ER and measured its cross-sectional profiles reliably.
Methods
Simulation & Evaluation
Photon-emission statistics:
We assume that the total number of photons emitted from a probe is a random variable that follows a log-normal distribution8. We considered two photon-emission rates: high and low. The former setting was primarily meant to simulate STORM data while the latter was closer to PALM data. The log-normal parameters for the high photon-emission case are (mean, standard deviation) = (5,000, 2,000) and for the low photon-emission case, they are (350, 70). Constant background fluorescence photons of 100 or 10 are added to every camera pixel for high and low emission cases, respectively.
PSF model:
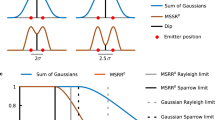
When reconstructing real STORM data, we used low-density STORM images for fitting the measured PSF. Our PSF model is a sum of two Gaussian functions8. Low-density data is used as a reference of the PSF model, in order to optimize their widths and weights. For our simulated data, we chose to have two Gaussian functions that have a standard deviation of σ and 2σ and that are linearly combined with a ratio of 4:1.
Noise statistics:
For simulating noise, we assumed that a high-performance CCD camera was used for data acquisition and considered two type of noise contribution, shot noise and a small readout noise21,23. The shot noise follows Poisson statistics. Specifically, the mean of the detected photons is determined by the expected number of incoming photons and by the background fluorescence level at each pixel. We also added CCD-type readout noise which follows a normal distribution with zero mean and variance of 2. In Supplementary Fig. 9, we also considered a CMOS-type camera whose readout noise has intrinsic pixel to pixel variation, resulting in non-Gaussian distribution. Specifically, CMOS-type readout noise follows a log-normal distribution with unit mean and variance of 2.
Probe distribution and performance analysis:
For the experiment of Fig. 4 and 5, we randomly distributed N probes in the central region of the image; the size of the region is 100 × 100 pixels. The size of the image is slightly larger, 108 × 108 pixels. The camera pixel size is 100 nm. In Fig. 4, the imaging densities vary between 0.05 μm−2 and 9.3 μm−2 for the high photon-emission rates (Fig. 4a and b) and between 0.05 μm−2 and 6.5 μm−2 for the low photon-emission rates (Fig. 4c and d). The FWHM of the PSF was set to 385 nm. We used 50 realizations for every density. Fig. 5 depicts additional analysis with FWHM PSF between 250 nm and 430 nm. The imaging density is 6 μm−2 and we used 30 realizations for each FWHM.
The performance analysis was carried out by matching each localized probe with the closest ground-truth probe. Localization errors larger than 300 nm were excluded from the analysis. The localization accuracy is expressed in terms of standard deviation or the FWHM of the histograms of localization errors. The recall rate is defined by the ratio between the number of matched probes and N.
Simulation of geometric structures:
We generated STORM images by distributing particles randomly on a circle of radius r. The radius varies between 150 nm and 400 nm. The probes are activated at different time instances, covering 300 frames. The number of probes at each frame is N = 4 (Fig. 6). Supplementary Fig. 4 depicts estimation results for various number of molecules N between 1–6 with fixed r = 200 nm. For every localized probe, we calculated the distance from the probe to the center of the circle and estimated the radius by averaging all distances. Then, we plot the differences between the true radii and the estimated values. Here, high photon-emission rates are used.
Image and signal processing
We used Gaussian rendering to generate super-resolution images. For a rendered image, every localized molecule is convolved with a Gaussian kernel and then scaled by its estimated photon count. In Fig. 8, normalized Gaussian kernels were used without photon scaling. For the experimental STORM data of fixed microtubule (Fig. 7), stage drifts are corrected by using fiducial beads. To correct the drift of the sample in Supplementary Fig. 13, the mean vertical (resp. horizontal) position of a straight horizontal(resp. vertical) segment of microtubules was measured as a function of frame number and the polynomial fit of this function was subtracted from the vertical (resp. horizontal) coordinates of all the peaks.
In order to determine the achievable temporal resolution of our live data, we used the Nyquist criteria ( ) as commonly applied to super-resolution imaging25. We counted the number of molecules on our structure of interest and obtained an estimate of its effective area by thresholding the super-resolution image; from this we had an estimate of our localization density. With a 2.5 s reconstructed super-resolution image, our localization density corresponds to R = 40 nm.
) as commonly applied to super-resolution imaging25. We counted the number of molecules on our structure of interest and obtained an estimate of its effective area by thresholding the super-resolution image; from this we had an estimate of our localization density. With a 2.5 s reconstructed super-resolution image, our localization density corresponds to R = 40 nm.
Sample preparation & Data acquisition
Cell culture:
COS-7 and U2OS cells were cultured in DMEM supplemented with 10% FBS (Sigma Aldrich) in a cell culture incubator (37°C and 5% CO2) and plated at low confluency on cleaned 25 mm size 1 cover-glass (Menzell).
Sample preparation for microtubule imaging in fixed cells:
Prior to fixation, all solutions were pre-warmed at 37°C: 24 hours after plating, Cos-7 cells were pre-extracted for 10 s in 0.5% Triton X-100 (Triton) in BRB80 (80 mM PIPES, 1 mM MgCl2, 1 mM EGTA, adjusted to pH 6.8 with KOH) supplemented with 4 mM EGTA, washed in PBS, fixed for 10 min in - 20°C-Methanol (Sigma) and washed again in PBS. The samples were then blocked 30 minutes in 5% BSA, before being incubated for 1.5 hour at room temperature with 1:1000 mouse α-tubulin antibodies (Sigma, T5168) in PBS - 1% BSA - 0.2% Triton (PBST), followed by 3 washes with PBS-0.2% Triton and then incubated for 45 min in PBST with 1:1000 goat anti-mouse Alexa-647 F(ab)2 secondary antibody fragments (Life Technologies, A-21237).
Sample preparation for live-cell imaging of the endoplasmic reticulum (ER):
After letting cells grow to approximately 60% confluency, U2OS cells were transfected with 2 μg of the reticulon 4 (Rtn4)-tdEos plasmid using FuGENE 6 transfection reagent (Roche). Cells were washed 24 hours post transfection and imaged in Leibovitz.
Live and fixed super-resolution imaging:
Imaging of the live and fixed samples was performed on a modified Olympus IX71 inverted microscope. In the case of Alexa-647 labeled α-tubulin, a laser at 641 nm (Coherent, CUBE 640-100C) was reflected by a multi-band dichroic (89100 bs, Chroma) onto the back aperture of a 100 × 1.3 NA oil objective (Olympus, UplanFL) mounted on a piezo objective scanner (P-725 PIFOC, Physicinstrument). The collected fluorescence from the sample was filtered by a band-pass emission filter (ET700/75, Chroma) and imaged onto an EMCCD camera (IxonEM+, Andor) with a 100 nm pixel size and using the conventional CCD amplifier at a frame rate of 25 fps. Laser intensity on the sample was ≈ 1-2 kW. cm−2. A 405 nm laser was added to maintain a high peak density. Blinking was induced according to15. Briefly, 30 μL of a mixture of 20% Vectashield, 80% Glycerol was pipeted on top of the sample and a cover-glass was added on top of the buffer to spread it evenly on the sample. For live-cell imaging, transfected cells were identified using a 488 nm laser (Sapphire 488, Coherent) at an intensity <1 kW cm−2 to prevent activation of tdEos. Subsequently, cells were imaged using a 561 nm laser (Sapphire 561, Coherent) with an intensity of approximately 3 kW cm−2. A 405 nm laser was used to convert tdEos molecules from their green to red form and was left on throughout acquisition. Fluorescence was collected after passing through a ET605/70 (Chroma) emission filter. Imaging was performed at a frame rate of 64 fps.
References
Rust, M. J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy STORM. Nat. Methods 3, 793–796 (2006).
Hess, S. T., Girirajan, T. P. & Mason, M. D. Ultra-high resolution imaging by fluorescence photoactivation localization microscopy. Biophys. J. 91, 4258 (2006).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Smith, C. S., Joseph, N., Rieger, B. & Lidke, K. A. Fast, single-molecule localization that achieves theoretically minimum uncertainty. Nat. Methods 7, 373–375 (2010).
Henriques, R. et al. Quickpalm: 3d real-time photoactivation nanoscopy image processing in imagej. Nat. Methods 7, 339–340 (2010).
Parthasarathy, R. Rapid, accurate particle tracking by calculation of radial symmetry centers. Nat. Methods 9, 724–726 (2012).
Holden, S. J., Uphoff, S. & Kapanidis, A. N. DAOSTORM: an algorithm for high-density super-resolution microscopy. Nat. Methods 8, 279–280 (2011).
Zhu, L., Zhang, W., Elnatan, D. & Huang, B. Faster storm using compressed sensing. Nat. Methods 9, 721–723 (2012).
Mukamel, E. A., Babcock, H. & Zhuang, X. Statistical deconvolution for superresolution fluorescence microscopy. Biophys. J. 102, 2391–2400 (2012).
Dertinger, T., Colyer, R., Iyer, G., Weiss, S. & Enderlein, J. Fast, background-free, 3D super-resolution optical fluctuation imaging (SOFI). Proc. Natl. Acad. Sci. U.S.A. 106, 22287 (2009).
Cox, S. et al. Bayesian localization microscopy reveals nanoscale podosome dynamics. Nat. Methods 9, 195–200 (2011).
Hu, Y. S., Nan, X., Sengupta, P., Lippincott-Schwartz, J. & Cang, H. Accelerating 3b single-molecule super-resolution microscopy with cloud computing. Nat. Methods 10, 96–97 (2013).
Afonso, M. V., Bioucas-Dias, J. M. & Figueiredo, M. A. Fast image recovery using variable splitting and constrained optimization. IEEE Trans. Image Process 19, 2345–2356 (2010).
Nieuwenhuizen, R. P. et al. Measuring image resolution in optical nanoscopy. Nat. Methods 10, 557–562 (2013).
Olivier, N., Keller, D., Rajan, V. S., Gönczy, P. & Manley, S. Simple buffers for 3d storm microscopy. Biomed. Opt. 4, 885–899 (2013).
Hu, J., Prinz, W. A. & Rapoport, T. A. Weaving the web of er tubules. Cell 147, 1226–1231 (2011).
Snapp, E. L. ER biogenesis: proliferation and differentiation. The Biogenesis of Cellular Organelles (New York: Landes Bioscience and KluwerAcademic/Plenum Publishers, 2004).
Frost, N. A., Lu, H. E. & Blanpied, T. A. Optimization of cell morphology measurement via single-molecule tracking palm. PloS one 7, e36751 (2012).
Shim, S.-H. et al. Super-resolution fluorescence imaging of organelles in live cells with photoswitchable membrane probes. Proc. Natl. Acad. Sci. U.S.A. 109, 13978–13983 (2012).
York, A. G. et al. Instant super-resolution imaging in live cells and embryos via analog image processing. Nat. Methods 10, 1122–1126 (2013).
Beier, H. T. & Ibey, B. L. Experimental comparison of the high-speed imaging performance of an em-ccd and scmos camera in a dynamic live-cell imaging test case. PloS one 9, e84614 (2014).
Kim, K. et al. Fast maximum likelihood high-density low-snr super-resolution localization microscopy. 10th International Conference on Sampling Theory and Applications: Invited Session VII: Sampling in Bio Imaging, Jacobs University, Bremen, Gemerny (2013 July 3).
Fowler, B., McGrath, D. & Bartkovjak, P. Read noise distribution modeling for cmos image sensors. 2013 INTERNATIONAL IMAGE SENSOR WORKSHOP, Snowbird Resort, Utah, USA (2013 June 13).
Babcock, H. P., Moffitt, J. R., Cao, Y. & Zhuang, X. Fast compressed sensing analysis for super-resolution imaging using l1-homotopy. Opt. Express 21, 28583–28596 (2013).
Shroff, H., Galbraith, C. G., Galbraith, J. A. & Betzig, E. Live-cell photoactivated localization microscopy of nanoscale adhesion dynamics. Nat. Methods 5, 417–423 (2008).
Acknowledgements
This research was supported by the Korea Science and Engineering Foundation (KOSEF) grant funded by the Korea government (MEST) (No.2009-0081089) and the Swiss National Science foundation under grant (No.200020-144355).
Author information
Authors and Affiliations
Contributions
M.U. supervised the project. M.U., J.M., C.V., H.K. and J.Y. designed the algorithm. J.M. implemented the algorithm and ran the simulated experiments. J.M., C.V. and H.K. designed and analysed the simulated data. N.O. prepared the sample of COS-7 cell and acquired STORM data of microtubule. L.C. prepared the sample of U2OS cell and J.M. and L.C. acquired live PALM data of ER. J.M., L.C., N.O. and S.M. performed analysis on the experimental data of the microtubule and ER. S.H. performed the DAOSTORM analysis on simulated and experimental STORM data. J.M. generated the figures for all simulations and J.M., L.C. and N.O. generated the figures of the experimental data. All authors wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Note and Figures
Supplementary Information
Super-resolution movie of live-ER data
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported license. The images in this article are included in the article's Creative Commons license, unless indicated otherwise in the image credit; if the image is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the image. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Min, J., Vonesch, C., Kirshner, H. et al. FALCON: fast and unbiased reconstruction of high-density super-resolution microscopy data. Sci Rep 4, 4577 (2014). https://doi.org/10.1038/srep04577
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep04577
This article is cited by
-
Quantitatively mapping local quality of super-resolution microscopy by rolling Fourier ring correlation
Light: Science & Applications (2023)
-
Super-resolution SRS microscopy with A-PoD
Nature Methods (2023)
-
Bayesian posterior density estimation reveals degeneracy in three-dimensional multiple emitter localization
Scientific Reports (2023)
-
ReCSAI: recursive compressed sensing artificial intelligence for confocal lifetime localization microscopy
BMC Bioinformatics (2022)
-
Smoothness correction for better SOFI imaging
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.