Abstract
Cancer is a genomic disease associated with a plethora of gene mutations resulting in a loss of control over vital cellular functions. Among these mutated genes, driver genes are defined as being causally linked to oncogenesis, while passenger genes are thought to be irrelevant for cancer development. With increasing numbers of large-scale genomic datasets available, integrating these genomic data to identify driver genes from aberration regions of cancer genomes becomes an important goal of cancer genome analysis and investigations into mechanisms responsible for cancer development. A computational method, MAXDRIVER, is proposed here to identify potential driver genes on the basis of copy number aberration (CNA) regions of cancer genomes, by integrating publicly available human genomic data. MAXDRIVER employs several optimization strategies to construct a heterogeneous network, by means of combining a fused gene functional similarity network, gene-disease associations and a disease phenotypic similarity network. MAXDRIVER was validated to effectively recall known associations among genes and cancers. Previously identified as well as novel driver genes were detected by scanning CNAs of breast cancer, melanoma and liver carcinoma. Three predicted driver genes (CDKN2A, AKT1, RNF139) were found common in these three cancers by comparative analysis.
Similar content being viewed by others
Introduction
Wide genomic aberration is a hallmark of the genomes of all cancer types. Deep sequencing technology1,2 has recently characterized the geographic and functional spectrum of cancer genomic aberrations and revealed insights into the mutational mechanisms3,4,5,6. These somatic mutations in cancer genomes may encompass several distinct classes of DNA sequence variations, including point mutations, copy number aberrations (CNA) and genomic rearrangements7. CNAs are deletions or additions of large segments of a genome and usually include one to tens of genes. Although these somatically acquired changes have been observed in cancer cell genomes, it does not necessarily mean that all of the abnormal genes are also involved in the development of cancers. Indeed, some genes are likely to make no contribution to cancer progress at all. In order to draw a distinction between them, these mutated genes have been coined driver and passenger genes7,8. A driver gene is causally implicated in the process of oncogenesis, while a passenger gene makes no contribution to cancer development itself, but is simply a by-product of the genomic instability observed in cancer genomes. Distinguishing driver genes from passenger genes has thus been considered an important goal of cancer genome analysis, especially in the field of personalized medicine and therapy9,10.
Driver and passenger genes can be differentiated by the functional roles they play in cells. Different genomic data that measure gene functions at different dimensions would be highly informative to separate potential driver from passenger genes. Recently, several methods have been proposed to identify potential driver genes based on systematic integration of genome scale data of CNA and gene expression profiles and applied to melanoma8, gingivobuccal cancer11 and liver carcinoma12. Apart from using gene expression data, integrating other types of genomic datasets such as those for protein-protein interaction13,14,15, epigenetic16, metabolism pathways17,18, sequence similarity19 and Gene Ontology20 should greatly increase the predictive power for driver genes and thus enable researchers to systematically investigate the mechanisms underling a great variety of cancers.
To this aim, we developed a computational method, MAXDRIVER, for the identification of driver genes from aberrant regions throughout cancer genomes by integrating multiple omics data. Several computational strategies are used to optimize gene similarities, filter noise and search maximal information flow among a query disease and candidate genes through a heterogeneous network. Large-scale validation results suggest MAXDRIVER is a useful method for genomic data integration and the discovery of cancer driver genes from aberrant regions and their flanks. By comparative analysis of breast cancer, melanoma and liver carcinoma, common potential drivers and their associated pathways are proposed. The present work highlights the importance of systematic integration and optimization of multiple omics data to investigate the mechanisms that underlie cancer development.
Results
Overview of MAXDRIVER
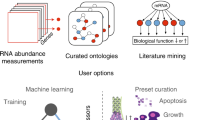
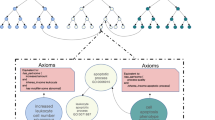
MAXDRIVER mainly performed three steps to integrate multiple data sources for the identification of driver genes for a given cancer (Fig. 1). In the first step, we adopted a multiple regression model to construct a fused gene functional similarity network, in which edge weights were derived from four data sources, including protein-protein interactions (PPI), gene co-expression patterns (GCE), gene sequence similarities (GSS) and pathway co-occurrence relationships (PCO). For this purpose, we calculated a gene similarity profile using each of these data sources and derived a gene functional similarity profile using the GO function (GO). The above 5 gene similarity profiles were used to calculate a functional similarity between a gene pair of the gene functional similarity network through a multiple regression model. With parameters of the model estimated, we further used the trained model to calculate a score for every pair of genes, obtaining edge weights of the fused gene functional similarity network. In the above procedure, we adopted a heuristic filtration strategy to filter out noises that indicated low confidence relationships for gene pairs. In the second step, we combined the fused gene functional similarity network with a disease phenotypic similarity network and gene-disease associations to construct a heterogeneous network. In the third step, we applied an information flow method to the heterogeneous network to trace the relationships among cancers and candidate genes located in CNA regions. In this procedure, we measured the strength of association between a cancer and a gene as the maximum value of the information flowing from the cancer to the gene (Fig. 1a). With this method established, we used genes located in CNA regions as candidates and ranked them according to their maximal information flow values (Fig. 1b).
Workflow of MAXDRIVER.
(a) A heterogeneous network was constructed by two-step integrations. First, a fused gene functional similarity network was calculated by fusing 4 datasets. To any gene pair gene gi and gj, the gene network was weighted by fused values  ,
,  were optimized parameters. The values were further filtered by setting parameter β = 0.25. Second, a heterogeneous network was constructed by merging the fused gene functional similarity network, gene-disease associations and a disease phenotypic similarity network that was calculated by text mining. The disease phenotypic similarities were further filtered by setting parameter γ = 0.19. Edges from each candidate gene to an additional sink are added for calculating the maximal information flow from a query cancer to the sink. The information flow through each candidate gene is then calculated as the strength of association between the candidate gene and the query disease. (b) CNA regions were estimated from deep-sequenced data of common or personal cancer genomes. Genes located in CNA regions were ranked by the maximal information flow values calculated from cancer node to candidate gene nodes in the heterogeneous network.
were optimized parameters. The values were further filtered by setting parameter β = 0.25. Second, a heterogeneous network was constructed by merging the fused gene functional similarity network, gene-disease associations and a disease phenotypic similarity network that was calculated by text mining. The disease phenotypic similarities were further filtered by setting parameter γ = 0.19. Edges from each candidate gene to an additional sink are added for calculating the maximal information flow from a query cancer to the sink. The information flow through each candidate gene is then calculated as the strength of association between the candidate gene and the query disease. (b) CNA regions were estimated from deep-sequenced data of common or personal cancer genomes. Genes located in CNA regions were ranked by the maximal information flow values calculated from cancer node to candidate gene nodes in the heterogeneous network.
Performance of MAXDRIVER
Identification of cancer driver genes is usually done by performing biological experiments, however only a few driver gene sets are available to date. Therefore, there are no large gene lists that can be used to validate the performance of MAXDRIVER. Alternatively, here we used known disease genes as simulations for leave-one out large-scale validations and test if MAXDRIVER was able to find known drivers of cancers. First, we selected previously identified disease genes from the OMIM database as positive controls and then tested if they can be recalled from artificially constructed control sets, including linkage intervals and random controls. On linkage interval control gene set, a known disease gene was simulated as a driver gene and its neighbour genes in 10 M distance as passengers, considering that 10 M was much larger than most CNAs were. Cross validations for recalling the known cancer genes from interval control gene sets indicated that MAXDRIVER can achieve top one ranked precision (TOP) as high as 64.06%, with parameters β = 0.25 and γ = 0.19 (Fig. 2). The mean rank ratio (MRR) of all 2,496 test cases was 7.19%, suggesting that known disease genes were ranked highly. We calculated the area under the rank receiver operating characteristic curve (ROC), named AUC and achieved an AUC value of 93.78% (see Methods for detailed definitions of TOP, MRR and AUC). We also performed a validation on random control sets, whose 99 genes were randomly selected on a genome-wide scale. This test allowed for simulation of cases where no prior gene knowledge was available and also avoided a bias towards better-characterized regions. Results for this validation showed that MAXDRIVER can successfully rank 1,558 known disease genes at top 1, yielding a TOP precision of 62.42% (Fig. 2). It also achieved a MRR of 14.37% and an AUC of 86.61%. These results indicate that MAXDRIVER achieves high precision on both control sets. Permutation experiments were performed by shuffling edges in the gene network while fixing the degree (i.e., the number of neighbours of each node). For two control sets, the AUC scores were both reduced by approximately 50%, but the result for the linkage interval (55.11%) was slightly higher (Fig. 2). These permutation validations suggest that the ability of MAXDRIVER to prioritize known disease genes is mainly due to its ability to exploit the informative connections in the heterogeneous network.
Access of individual genomic data sources
To test the contribution of all genomic data sets, we validated the performances of each individual data on random control gene sets. At each case, the edge weights of the gene functional similarity network were only calculated from individual gene genomic data. Among the four data sources, PPI obtained the highest TOP of 53.41%, a MRR of 16.7% and an AUC value of 84.3%, while the GCE was as low as 47.22% (TOP), 18.21% (MRR) and 80.15% (AUC). When all four data sources were fused without noise filtering, the performances of MAXDRIVER were improved to 56.54% (TOP), 16.58% (MRR) and 84.38% (AUC) (Supplementary Fig. S1). These results suggest that the PPI information is more useful than the other three genomic data sets.
We further performed a noise filtering process on disease phenotypic similarities and gene functional similarities, to filter out low confidence values to improve predicting precisions. The histograms of the disease similarities were plotted and fitted to Weibull distribution by using MATLAB Fitting ToolBox (p-value 1.34E-16; Kolmogorov-Smirnov test, two-sided) with a scale of 0.161 (Std. Err 9.43E-05) and a shape of 1.65 (Std. Err 1.17E-03). The edge weights were fitted to Gamma distribution (p-value 2.79E-13; Kolmogorov-Smirnov test, two-sided) with a scale of 0.014 (Std. Err 1.33E-05) and a shape of 0.633 (Std. Err 4.11E-04). Both probability densities exhibited a clear positive skewness, indicating that most small scores presumably were noise background (Supplementary Fig. S2). We first used the thresholds as the mean of gene similarity (β = 0.23) and disease similarity (γ = 0.14), respectively and achieved 62.63% (TOP), 15.58% (MRR) and 85.09% (AUC) on a random control. We also optimized parameters β = 0.25 and γ = 0.19 by using a grid search. In this case, MAXDRIVER achieved additional improvements of 1.21% (MRR) and 1.52% (AUC), but only a little reduction in TOP (0.21%). These results indicate that these two processes of noise filtering are both useful and improved the precision in biological data integration.
Robustness of MAXDRIVER
Threshold parameters β, γ play important functions in filtering the noise and in making the heterogeneous network sparse. Here we changed them with their respective change ratios to test their effects on MAXDRIVER. For parameter x, the parameter changed ratio is defined as (x − x0)/x0 and the related changed ratios are defined as (TOP(x) − TOP(x0))/TOP(x0), (AUC(x) − AUC(x0))/AUC(x0) and (MRR(x) − MRR(x0))/MRR(x0), where x0 represents the optimized parameter value. When the β changed ratios were set from -1 to 1, the changed ratios of TOP, MRR and AUC were all less than 0.14. When the γ changed ratios were set from −1 to 1, the changed ratios of TOP, MRR and AUC were all less than 0.2 (Fig. 3). In comparison, the change ratios of TOP were more than MRR and AUC, suggesting that these two parameters affected many of the top one ranked genes. More specifically, TOP reached its optimum at the optimal point of β (Fig. 3a, b), but TOP was optimal when γ was 0 (Fig. 3c, d). These results indicate that disease phenotypic similarity is useful in achieving higher AUC and MRR, albeit at the expense of a slightly reduced TOP score. In summary, the parameters β and γ exhibit varying effects on the different criteria without significantly affecting the robustness of the overall performance of MAXDRIVER.
Robustness of MAXDRIVER according to parameters changes.
When β and γ changed ratios were ranged from −1 to 1 respectively, TOP, MRR and AUC changed ratios were calculated on random control gene sets (a, c) and linkage interval genes sets (b, d). The results were calculated by only changing β, while γ was kept as the optimized value 0.19 (a, b). The results were calculated by only changing γ, while β was kept as the optimized value 0.25 (c, d).
Predicted driver genes of breast cancer, melanoma and liver carcinoma
To test the capability of MAXDRIVER in detecting novel driver genes, we employed it to scan CNAs of breast cancer (OMIM 114480), melanoma (OMIM 155600) and liver carcinoma (OMIM 114550). Recently, cancer genes and mutational processes in breast cancer had been systemically reported21,22,23,24. Within 218 breast cancer samples, 78 CNAs were detected by using GISTIC, a method that was used for CNAs analysis from deep sequencing data24,25. By ranking the genes located in the regions of CNAs, we detected 70 top one ranked genes as potential driver genes (Supplementary Table S1). Five genes, namely AKT1, PIKSCA, TP53, AKT2 and MAP3K were successfully predicted amongst breast cancer genes that had been verified experimentally in previous reports (e.g. AKT1, BRCA1, CDH1, GATA3, PIK3CA, PTEN, RB1, TP53, AKT2, ARID1B, CASP8, CDKN1B, MAP3K1, MAP3K13, NCOR1, SMARCD1 and TBX3)22. Besides the 5 known driver genes, a total of 64 genes had been annotated before to be associated with diverse diseases by genetic association database26 or OMIM database27 (see more detailed information in Supplementary Table S1). Melanoma is a well-studied type of tumor and four genes (MITF, KLF6, TBC1D16 and RAB27A) have been reported previously to act as driver genes8. From two early genomic variation studies1,28, a total of 91 CNAs had been identified. Using MAXDRIVER, we predicted 63 potential genes to be driver genes (Supplementary Table S2), with one of the four genes, MITF, detected successfully. Whilst RAB27 was not identified itself, one of its interacting protein partners, MYO5A, was also ranked top one position. Furthermore, a total of 58 genes are annotated as disease related (Supplementary Table S2). We also tested MAXDRIVER on liver carcinoma, the third leading cause of cancer-related death worldwide. Thirty frequently altered genes had been described before, including TP53, CTNNB1, ARIDI1, ARID2, AXIN1, PRS6KA3, VCAM1, CDK14, TERT, MLL4, CCNE1, CDK2A, AXIN1, PTEN, PIK3CA, KRAS, NRAS, MYC, MET, CCND1, CDKN2A, RB1, ARID1A, ARID1B, DRID2, IRF2, NFE2L2, ERRFI1, RPSKA3 and MLL329,30. We reported 57 potential driver genes, including TP53 and CDKN2A (Supplementary Table S3), detected by scanning for 57 CNAs12. Except for gene PAK2, all other 56 genes had been annotated previously as disease related (Supplementary Table S3). These predicted driver genes exhibit diverse associations not only to cancers, but also to various human diseases, providing novel candidates for further experimental validations.
Common driver genes and biological processes of breast cancer, melanoma and liver carcinoma
Early research had revealed similar genomic aberrations and mechanisms underlying tumorigenesis in various cancers31. Here, we performed a comparative analysis of 70, 63 and 57 predicted driver genes of breast cancer, melanoma and liver carcinoma respectively. Specifically, 9 genes were found to be common in liver carcinoma and melanoma, 7 genes in liver carcinoma and breast cancer and 11 genes in melanoma and breast cancer. Three genes (CDKN2A, AKT1, RNF139) were predicted to be driver genes in all three cancer types (Fig. 4a). Amongst these three genes, dysfunction of CDKN2A and AKT1 had been identified previously to cause disease. For example, AKT1 was reported to drive development of breast cancer32 and proteus syndrome33,34. CDKN2A (p16) acted as a tumor suppressor gene and its inactivation by deletion was frequently found in cancers such as, bladder carcinoma35 and lung adeno-carcinomas36. Based on our analysis, we propose that RNF139 is a novel cancer driver gene. RNF139 encoded an endoplasmic reticulum-resident E3 ubiquitin ligase and it contained a sterol-sensing domain and a RING finger motif37,38. This gene located in a CNA region in melanoma (121,760,777–128,860,000), a CNA region in liver carcinoma (120,876,000–134,690,000) and near a CNA region in breast cancer (128,816,653–128,993,129). Although it was not reported early as a driver gene, MAXDRIVER predicted it as top one from 15, 19 and 23 genes that were located in the CNA regions of breast cancer, melanoma and liver carcinoma, respectively, suggesting it may be a novel potential driver gene.
Common driver genes and functional enrichment analysis of predicted driver genes of breast cancer, melanoma and liver carcinoma.
(a) Venn graph showing common driver genes of predicted driver genes of three cancers. (b) Functional enrichment of 164 potential driver genes. The sizes of circles are correlated to the gene numbers. The colours used indicate enrichments p-values. The graph was analyzed and printed by using BINGO software67.
We next performed a functional enrichment analysis on all 164 unique predicted genes, a union of 70, 63 and 57 genes of breast cancer, melanoma and liver carcinoma respectively. These 164 genes were almost uniformly distributed across the human chromosomes, except for the X and Y chromosomes, with only one gene located on X and none on Y (Supplementary Fig. S3). These genes were significantly related to several cancer pathways, such as melanoma (hsa05218), endometrial cancer (hsa05213), non-small cell lung cancer (hsa05223), renal cell carcinoma (hsa05211), glioma (hsa05214), bladder cancer (hsa05219) and chronic myeloid leukemia (hsa05220) (all p-values < 1E-5, Supplementary Table S4). Furthermore, 164 genes were found to be enriched in a number of biological processes related to cell fate, cell survival and genome maintenance (Fig. 4b, p-value < 1E-5, Supplementary Table S5). Within the category ‘regulation of biological processes’, negative regulation of cell death, programmed cell death and apoptosis were most enriched (Fig. 4b-b1). Comparatively, genes with functions involved in positive regulation of cellular processes were found to be enriched even more highly (Fig. 4b-b2). In addition, genes involved in the response to chemical stimuli were also found highly enriched (Fig. 4b-b3), as well as genes involved in developmental processes (Fig. 4b-b4), indicating that dysfunctions of these genes could be important to cancer development.
The 164 genes identified by MAXDRIVER showed not only a tight functional relationship among them, but also with other well-established oncogenes. From 163 genes that were reported previously as being involved in cancer pathways (KEGG, hsa05200)39, 23 genes were successfully predicted as driver genes by our method (Supplementary Fig. S4), achieving a p-value as low as 2.2E-16 (Fisher exact test, one-sided). In total, 558 interactions and 717 regulatory connections were found to exist among 163 cancer genes and 164 predicted driver genes (Fig. 5, Supplementary Table S6), suggesting these genes were tightly correlated. The functional relationships among the 164 predicted driver genes were also supported by other functionally related evidences such as activation, inhibition, binding, post-translation or co-expression, all of which were analyzed by using STRING database34 (Supplementary Fig. S5). The identification of these driver genes is consistent with earlier analyses of cancer genomes and should reveal new potential oncogenes involved in cancer onset as well as progress.
Correlations among known cancer genes and predicted driver genes.
Total 558 interactions and 717 regulatory connections existed among 163 known cancer genes and 164 predicted driver genes were extracted from the DAVID database66. The graph was drawn by using Cytoscape software68. Among 163 known cancer genes, 23 genes are predicted as driver genes (red).
Detecting driver genes in CNAs flanks
MAXDRIVER can detect gene drivers located not only in regions of CNAs, but also accompanying flanking regions. This is important for identifying real driver mutations (or genes), since genome aberrations may be not located in protein-coding regions but in intronic, intergenic or gene regulatory regions and could affect genes at great distance within the chromosome40. One example is a single nucleotide polymorphism (SNP), rs6983267, which interacts physically with the MYC proto-oncogene, which is located some 330 kilobases away on the chromosome41,42. Here, we show that MAXDRIVER has the ability to detect the genes CDKN2A, AKT1 and RNF139 as driver genes, even though they not all are located in regions with CNAs. AKT1 is located in a CNA region of breast cancer on chromosome 14, while it is located at the 5′ side of a liver carcinoma CNA region and at the 3′ side of a melanoma CNA region. CDKN2A is located in two CNA regions of liver carcinoma and breast cancer, respectively, while it is located at the 5′ side of a melanoma-associated CNA region. RNF139 is located in two liver carcinoma and melanoma CNAs, but at the 5′ side of a breast cancer CNA region (Fig. 6). MAXDRIVER can rank these three genes at top one position by scanning all extended CNA regions with flanks of three cancers. Detailed analysis of RNF139 revealed that the nearest breast cancer CNA region to RNF139 is 3,315,974 bp far away and 43,347 bp in length. Intriguingly, this CNA region does not harbour any protein coding sequences. Setting this CNA at the centre, we extended it from 123,838,327 bp to 133,838,327 bp to include 14 genes. MAXDRIVER successfully ranked RNF139 at top one position of these 14 genes. Similar results on two other CNAs of liver carcinoma and melanoma are also obtained by MAXDRIVER. In summary, these results suggest that MAXDRIVER is a powerful tool for identifying driver genes that are located even at large distances from mutated regions within cancer genomes.
Discussion
Next-generation sequencing technology has significantly contributed to cancer genomics research and has revealed a plethora of genomic changes that accumulate in cancer cells. However, methods that can perform deep analysis of such large genomic data sets and effectively utilize the redundance and complementarity of multiple data, are largely absent. To bridge this gap, we have developed MAXDRIVER, a method shown here to be efficient and reliable in its ability to predict novel cancer driver genes. It can be used not only to scan common CNAs collected from multiple cancer genomes for the identification of driver genes that are likely to be common in cancers, but also to scan CNAs of personal genomes, in order to identify personalized driver genes and such knowledge can then contribute to the development of personalized medicine as well as therapies43,44. Its superior performances benefits from two key approaches, namely its optimizing strategy of genomic data integration using a noise filtering function and the utilization of a maximal information flow method to maximize the relationship between queried cancer and its candidate genes.
Using MAXDRIVER, we were not only able to predict many novel potential driver genes, but it also allowed us to perform a comparative view of liver carcinoma, breast cancer and melanoma cancer. We predicted 164 potential driver genes, many of which had been reported previously as oncogenes (or tumor suppressors) and which displayed a tight correlation to 163 genes reported in the cancer pathway (hsa05200) suggesting that MAXDRIVER's predictions are reliable. Interestingly, we found that the three genes CDKN2A, AKT1 and RNF139 were predicted in all three cancers. CDKN2A and AKT1 are well characterized as driver genes that are crucially involved in a diverse range of cancers. Although further experiments are needed to investigate the exact function of RN139, particularily in cancer progress, our analysis strongly indicates that dysfunction of RNF139 may be involved in the development of a multitude of cancers.
Although genomic aberrations identified by deep sequencing technologies can now be scrutinized by deep analysis utilizing MAXDRIVER, the integration of additional genomic data should allow for even greater acceleration of this process. For example, time-series genomic measurements of different stages of cancer development will allow for dynamical descriptions of genomic aberrations43,45. Meanwhile, genome-wide patterns of epigenetic modification is another useful information, since the importance of epigenetic markers in maintaining normal cellular functions is reflected by the observation that many diseases develop upon spatial and temporal dysregulation of these histone and DNA-associated modifications16. A miRNA regulatory network could also be useful for driver gene prediction, as dysfunction of miRNA and their targeted genes had been reported in diverse cancers46,47. The genes that are regulated by one or more miRNAs can be viewed as a functional linkage and can then be embedded into our heterogeneous network. Considering that cancer is mainly a disease of signaling problems, we can also add a gene signaling network into our heterogeneous network. At present, the largest curated human signaling network is available for 6,309 proteins, including 62,737 signaling actions (released on Oct. 26, 2013, http://www.bri.nrc.ca/wang/)48,49,50,51. The human signaling network is not only useful for detecting cancer-related genes, but more importantly for investigating how differential signaling processes occur throughout cancer development. Compared to our constructed non-directed gene network, the gene signaling network is directed and includes three types of signaling actions, i.e. activation, inhibition and physical interaction, which allow for presenting more precise relationships among gene pairs. Therefore, a more precise modelling is desired to efficiently use this signaling network information for detailed investigation of cancer driver genes and cancer-related signaling processes.
An interesting and quantitative set of data is gene differential expression patterns calculated from cancer and normal cells. In our study, we have defined functional relationships between genes from the correlations of their expression vectors that are measured in 79 human tissues. This definition is useful for describing generic relationships among genes, but may not be adequate in providing sufficiently precise information for a special type of cancer, since different cancers usually have different expression profiles52,53. By utilizing differential expression patterns of genes specific for certain type of cancer, we can present a more precise prediction for the cancer. Technically, the utilization of differential expression patterns can be taken in two ways. First, we can further filter the predicted driver genes by analyzing their differential expression patterns. Genes exhibit highly differential expression patterns may possess a higher potential to be involved in the cancer development. Second, it might be also possible to embed differential expression patterns of genes into the heterogeneous network by using degrees of differential expression as capacities of gene pairs. Although this strategy of using differential expression patterns is desired in achieving higher prediction power for well-studied cancers, such a strategy could only be used case-by-case and thus is not useful for rarely studied cancers (because of the limited availability of gene expression data). In summary, MAXDRIVER can be used to scan CNAs of personal genomes, in order to identify molecular attributes of individual patients and it is easily extended to utilize more types of omics data that can sever as a useful tool for integrative network analysis in cancer systems biology54,55,56. By predicting and analyzing driver genes triggering different stages of tumorigenesis, we can track a rippling effect along the cancer differentiation path and track how cancer development is triggered and then amplified under the involvement of these genes. We are confident that the power and resolution offered by MAXDRIVER will allow biologists to better distinguish the different functional roles of genes underlying cancer development.
Methods
MAXDRIVER is designed to identify driver genes by integrating multiple omics data. It first integrates 5 gene datasets to build a fused gene functional similarity network. A heterogeneous network is then constructed by integrating a fused gene functional similarity network, a disease phenotypic similarity network and gene-disease associations. A maximum information flow strategy is then employed to prioritize candidate genes that are located in CNA regions and the top one-ranked genes were considered to be driver genes, whilst others were considered passenger genes. In the following, we describe, in details, the calculation of gene similarity, construction of heterogeneous network and relationships among a query disease and candidate genes by calculating the maximal information flow.
Calculation of gene similarity profiles
To build the fused gene functional similarity network, we describe the calculations of gene similarity from each of five datasets. We extracted protein-protein interactions from version 9.0 (released on Apr 13, 2011) of the HPRD database57 composed of 9,515 genes and 37,364 interactions. Given a pair of genes i and j, we defined their similarity  as the unit weight 1, if these two genes were linked in the network, otherwise we defined them as 0. Finally, we obtained a 9,515 by 9,515 binary matrix as a gene interacting profile.
as the unit weight 1, if these two genes were linked in the network, otherwise we defined them as 0. Finally, we obtained a 9,515 by 9,515 binary matrix as a gene interacting profile.
We downloaded from the NCBI Refseq database (released Mar, 2012) the protein sequences of all the aforementioned 9,515 proteins. We then aligned these sequences against each other using the NCBI BLASTP program58. We then calculated the sequence similarity between a pair of gene i, j as

where eij was the e-value outputted by BLASTP program with defaulted parameters. By doing this, we obtained a 9,515 by 9,515 numeric matrix as a gene sequence similarity profile.
We downloaded the gene expression data from http://biogps.gnf.org/downloads/ (GEO code: GSE1133), in which whole genome gene expressions were measured for 79 human tissues59. We then represented each of the 9,515 genes as a 79-dimensional expression vector, with each dimension corresponding to the expression level of the gene in a particular tissue. We then calculated the co-expression pattern between a pair of genes i, j as the absolute Pearson's correlation coefficient of their gene expression vectors, calculated as

where ei and ej were the expression vectors of genes i and j, respectively. By doing so, we obtained a 9,515 by 9,515 numeric matrix that served as a gene expression similarity profile.
We downloaded a total of 200 human pathways from the KEGG database (released Mar, 2012)39. We then represented a gene i as a 200 dimensional binary vector by assigning 1 to a dimension if the gene was present in the corresponding pathway and assigned 0 otherwise. We then calculated the gene co-occurrence relationship of two genes i and j as the cosine of the angle between their vectors, calculated by

where pi and pj were the vectors corresponding to the genes i and j, respectively. By doing so, we obtained a 9,515 by 9,515 numeric matrix as a gene pathway similarity profile.
For each of the above profiles, similarity values are in the range of [0, 1], where 1 means the highest similarity and 0 the lowest similarity. In the case that the relationship between a pair of genes is absent from a data source, the lowest similarity 0 is assigned.
Fusion of gene similarities
Although each of the above gene similarity profiles reflects functional similarity of genes from a certain perspective, none of them provides a thorough explanation about relationships between gene properties and gene functions. We therefore integrated these similarity profiles and generated a single fused gene similarity profile that can link gene properties to gene functions in a more comprehensive way. For this purpose, we downloaded the biological process domain of the Gene Ontology (GO)20 and the corresponding annotations for human genes (both released on Jan, 2012). Focusing on genes annotated with at least fifteen informative GO terms that appeared at or below the fifth level of the GO hierarchy and had at least five genes annotated, we obtained a total of 3,842 well-annotated genes. We then calculated the functional similarities between every pair of these genes, using the method of Resnik, as implemented in the software package GOSemSim60,61, obtaining a 3,842 by 3,842 numeric matrix serving as similarities with high confidence. On the basis of this method the coverage of this profile was low, we therefore adopted the following regression model to build a more comprehensive gene functional similarity profile.
Given the high confidence functional similarity between a pair of genes i and j, denoted by Rij, we used the four gene similarity scores  to explain Rij as
to explain Rij as

Where αk, k = 1, …, 5 were regression coefficients. Considering the huge number of i-j pairs in the high confidence gene functional similarity profile, we adopted a sampling strategy to estimate the parameters. Briefly, in each sampling procedure, we selected at random 1,000 gene pairs to train a model, obtaining estimates of αk for k = 1, …, 5. By repeating the sampling procedure 1,000 times and averaging over the resulting estimates, we obtained the mean estimates αk for k = 1, …, 5. According to this strategy, we calculated  for the regression intercept,
for the regression intercept,  for the gene sequence similarity,
for the gene sequence similarity,  for the gene coexpression pattern,
for the gene coexpression pattern,  for the pathway co-occurrence relationship and
for the pathway co-occurrence relationship and  for protein-protein interactions.
for protein-protein interactions.
With the parameters estimated, we further predicted functional similarity of each gene pairs, obtaining a 9,515 by 9,515 numeric matrix S of gene functional similarities.
Construction of a heterogeneous network
With gene functional similarities inferred from multiple data sources, we constructed a heterogeneous network that is composed of a fused gene functional similarity network, a disease phenotypical similarity network and known associations among diseases and genes.
The fused gene functional similarity network, Gg = {Vg, Eg}, was constructed by introducing a parameter β, to filter small similarity values in the predicted similarity matrix S. More precisely, with the cut-off value given by β, we set all sij = 0 if sij < β and kept sij unchanged otherwise. We noticed that such a noise filtering process not only helped in filtering out low confidence similarities and thus improving the performance of our method, but also resulted in a sparse network to speed up subsequent calculations. We used two methods to determine the optimum value for β. First, we used the mean of the fused similarities and obtained β = 0.23. With this parameter value, the gene functional similarity network contains 9,515 nodes and 1,756,050 edges. Second, we used a grid search with step 0.01 to find the optimal β that could yield the best performance in a cross-validation experiment and obtained β = 0.25. With this parameter value, the gene functional similarity network contained 9,515 nodes and 579,620 edges. Among these two methods, the later was more time consuming, but typically resulted in higher prediction accuracy in cross-validation experiments.
We constructed the disease phenotypic similarity network, Gd = {Vd, Ed}, as follows. First, we obtained from the literature62 a phenotype similarity profile that was represented as a matrix of similarity scores among 5,080 human disease phenotypes. Then, we introduced a parameter γ to filter out small similarity values in this matrix. Similar to our strategy for building the gene functional similarity network, we set all similarity scores to 0 if the score was less than the threshold value given by γ. To determine an appropriate value of γ, we also used the two methods mentioned above for determining β. First, we used the mean of the phenotypic similarities and obtained γ = 0.14. With this parameter value, the disease phenotypic similarity network contained 1,609 nodes and 1,013,622 edges. Second, we used a grid search with step 0.01 to find the optimal γ that could yield the best performance in a cross-validation experiment and obtained γ = 0.19. With this parameter value, the disease phenotypical similarity network contained 1,609 nodes and 600,090 edges.
Focusing on genes in the functional similarity network and diseases in the phenotype similarity network, we further obtained 2,496 associations among 1,609 human diseases and 1,460 genes using the BioMart tool63. We denote these associations as a set Ea.
With these components prepared, we constructed a heterogeneous network, G = {V, E}. Briefly, we defined the set of vertices V as the union of all diseases in the disease phenotypical similarity network and all genes in the gene functional similarity network, as  . In a similar way, we defined the set of edges E as the union of all edges in the disease phenotypical similarity network, all edges in the gene functional similarity network and all known associations among diseases and genes, as
. In a similar way, we defined the set of edges E as the union of all edges in the disease phenotypical similarity network, all edges in the gene functional similarity network and all known associations among diseases and genes, as  .
.
Prioritization of candidate genes by maximizing information flow in the heterogeneous network
We formulated the problem of prioritization of candidate genes as searching for the maximum information flow in the heterogeneous network and solved this problem as follows. First, we converted the undirected heterogeneous network to a directed graph by treating each undirected edge as two distinct directed edges of opposite directions. Second, we assigned a capacity value to each directed edge to measure the maximal connection transduction through this edge. As an edge of a disease pair, we defined the phenotypic similarity of that pair as their capacity. As an edge of a gene pair, we assigned the functional similarity between these two genes as their capacity. For the association component, we assigned positive infinites to all edges. Third, given a query disease and a set of candidate genes, we introduced a sink node and linked the candidate genes (nodes) to the sink node by edges of positive infinite capacities (Fig. 1a). Fourth, the strength of association from the query disease to each candidate gene is calculated. Formally, for the heterogeneous network G = (V, E, C) where V, E, C > 0 representing the node, edge and nonnegative capacity on each edge respectively, the maximal information flow from the query node to the sink through all candidate genes is calculated as follows.

Where the information flow f(v, w) is defined as the flow value transmitted from node v to node w and cap(v, w) is the capacity of the edge linked nodes v and w. we resorted to the HR_PR algorithm64 to search for the maximum flow from the query disease (node) to the sink node. The flow value pushed from the node corresponding to a candidate gene to the sink node is calculated to indicate the strength of association between the candidate gene and the query disease. Finally, we ranked candidate genes according to their scores to obtain a ranking list.
Validation methods
We conducted two cross-validation experiments to assess the performance of our method and further applied this method to three real CNA data sets for breast cancer, melanoma and liver carcinoma.
In the first cross-validation experiment, we ranked each gene (positive control) known as associated with a disease against a set of genes located within 10 Mb around the positive control (negative controls). We call this experiment validation against a linkage interval to simulate the genes in CNAs. In the second cross-validation experiment, we ranked each gene known as associated with a disease (a positive control) against a set of 99 genes selected at random from all 9,519 genes in the functional similarity profile (negative controls). We called this experiment ‘validation against random controls’.
We used three criteria to evaluate the performance of our method in a cross-validation experiment. First, we calculated the proportion of positive cases ranked first to obtain a criterion called the top one precision (TOP). Second, we divided the rank of a positive case by the number of corresponding negative controls to obtain the relative rank of the positive case and we averaged relative ranks of all positive cases to obtain a criterion called the mean rank ratio (MRR). Third, given a rank threshold, we calculated the proportion of positive cases ranked above the threshold as sensitivity and the proportion of negative controls ranked below the threshold as specificity. We then plotted the rank receiver operating characteristic curve (ROC) and calculated the area under this curve to obtain a criterion called the AUC score. Thus, the larger the TOP and AUC and the smaller the MRR, the higher the performance of the prioritization method.
For real case studies, we collected 78 CNA regions from breast cancer data sets24,25, 91 CNA regions from melanoma1,28 and 57 CNA regions from liver carcinoma data sets12. For each of the three cancers, we scanned each of their CNA regions to obtain a set of candidate genes and we applied our method to rank these candidate genes. Finally, we proposed genes ranked at the top of ranking lists as potential driver genes. Gene functional enrichment analysis was performed by using the DAVID database65,66, the STRING database34 and the KEGG database39.
References
Wang, J. et al. CREST maps somatic structural variation in cancer genomes with base-pair resolution. Nat Methods 8, 652–4 (2011).
Stratton, M. Genome resequencing and genetic variation. Nat Biotechnol 26, 65–6 (2008).
Koboldt, D. C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res 22, 568–76 (2012).
Kidd, J. M. et al. A human genome structural variation sequencing resource reveals insights into mutational mechanisms. Cell 143, 837–47 (2010).
Beroukhim, R. et al. The landscape of somatic copy-number alteration across human cancers. Nature 463, 899–905 (2010).
Abecasis, G. R. et al. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
Stratton, M. R., Campbell, P. J. & Futreal, P. A. The cancer genome. Nature 458, 719–24 (2009).
Akavia, U. D. et al. An integrated approach to uncover drivers of cancer. Cell 143, 1005–17 (2010).
Santarius, T., Shipley, J., Brewer, D., Stratton, M. R. & Cooper, C. S. A census of amplified and overexpressed human cancer genes. Nat Rev Cancer 10, 59–64 (2010).
Futreal, P. A. et al. A census of human cancer genes. Nat Rev Cancer 4, 177–83 (2004).
Ambatipudi, S. et al. Genome-wide expression and copy number analysis identifies driver genes in gingivobuccal cancers. Genes Chromosomes Cancer 51, 161–73 (2012).
Woo, H. G. et al. Identification of potential driver genes in human liver carcinoma by genomewide screening. Cancer Res 69, 4059–66 (2009).
Chen, Y., Jiang, T. & Jiang, R. Uncover disease genes by maximizing information flow in the phenome-interactome network. Bioinformatics 27, i167–76 (2011).
Jirawatnotai, S. et al. A function for cyclin D1 in DNA repair uncovered by protein interactome analyses in human cancers. Nature 474, 230–4 (2011).
Zhang, Q. C. et al. Structure-based prediction of protein-protein interactions on a genome-wide scale. Nature 490, 556–60 (2012).
Portela, A. & Esteller, M. Epigenetic modifications and human disease. Nat Biotechnol 28, 1057–68 (2010).
Dutta, B. et al. A network-based, integrative study to identify core biological pathways that drive breast cancer clinical subtypes. Br J Cancer 106, 1107–16 (2012).
Lango Allen, H. et al. Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 467, 832–8 (2010).
Mushegian, A. R., Bassett, D. E., Jr, Boguski, M. S., Bork, P. & Koonin, E. V. Positionally cloned human disease genes: patterns of evolutionary conservation and functional motifs. Proc Natl Acad Sci U S A 94, 5831–6 (1997).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25, 25–9 (2000).
Shah, S. P. et al. The clonal and mutational evolution spectrum of primary triple-negative breast cancers. Nature 486, 395–9 (2012).
Stephens, P. J. et al. The landscape of cancer genes and mutational processes in breast cancer. Nature 486, 400–4 (2012).
Ellis, M. J. et al. Whole-genome analysis informs breast cancer response to aromatase inhibition. Nature 486, 353–60 (2012).
Staaf, J., Jonsson, G., Ringner, M., Baldetorp, B. & Borg, A. Landscape of somatic allelic imbalances and copy number alterations in HER2-amplified breast cancer. Breast Cancer Res 13, R129 (2011).
Mermel, C. H. et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol 12, R41 (2011).
Becker, K. G., Barnes, K. C., Bright, T. J. & Wang, S. A. The genetic association database. Nat Genet 36, 431–2 (2004).
Hamosh, A. et al. Online Mendelian Inheritance in Man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res 30, 52–5 (2002).
Lin, W. M. et al. Modeling genomic diversity and tumor dependency in malignant melanoma. Cancer Res 68, 664–73 (2008).
Zhang, Z. Genomic landscape of liver cancer. Nat Genet 44, 1075–7 (2012).
Nakagawa, H. & Shibata, T. Comprehensive genome sequencing of the liver cancer genome. Cancer Lett 340, 234–40 (2012).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–58 (2013).
Spears, M. et al. Proximity ligation assays for isoform-specific Akt activation in breast cancer identify activated Akt1 as a driver of progression. J Pathol 227, 481–9 (2012).
Lindhurst, M. J. et al. A mosaic activating mutation in AKT1 associated with the Proteus syndrome. N Engl J Med 365, 611–9 (2011).
Szklarczyk, D. et al. The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored. Nucleic Acids Res 39, D561–8 (2011).
Rebouissou, S. et al. CDKN2A homozygous deletion is associated with muscle invasion in FGFR3-mutated urothelial bladder carcinoma. J Pathol 227, 315–24 (2012).
Suzuki, Y., Oonishi, T., Kudo, T. & Doi, H. LKB1, TP16, EGFR and KRAS somatic mutations in lung adenocarcinomas from a Chiba Prefecture, Japan cohort. Drug Discov Ther 6, 24–30 (2012).
Poland, K. S. et al. A constitutional balanced t(3;8)(p14;q24.1) translocation results in disruption of the TRC8 gene and predisposition to clear cell renal cell carcinoma. Genes Chromosomes Cancer 46, 805–12 (2007).
Gimelli, S. et al. The tumor suppressor gene TRC8/RNF139 is disrupted by a constitutional balanced translocation t(8;22)(q24.13;q11.21) in a young girl with dysgerminoma. Mol Cancer 8, 52 (2009).
Klukas, C. & Schreiber, F. Dynamic exploration and editing of KEGG pathway diagrams. Bioinformatics 23, 344–50 (2007).
Trynka, G. et al. Dense genotyping identifies and localizes multiple common and rare variant association signals in celiac disease. Nat Genet 43, 1193–201 (2011).
Tuupanen, S. et al. The common colorectal cancer predisposition SNP rs6983267 at chromosome 8q24 confers potential to enhanced Wnt signaling. Nat Genet 41, 885–90 (2009).
Pomerantz, M. M. et al. The 8q24 cancer risk variant rs6983267 shows long-range interaction with MYC in colorectal cancer. Nat Genet 41, 882–4 (2009).
Ziogas, D. & Roukos, D. H. Genetics and personal genomics for personalized breast cancer surgery: progress and challenges in research and clinical practice. Ann Surg Oncol 16, 1771–82 (2009).
Wistuba, I. I, Gelovani, J. G., Jacoby, J. J., Davis, S. E. & Herbst, R. S. Methodological and practical challenges for personalized cancer therapies. Nat Rev Clin Oncol 8, 135–41 (2011).
Chen, R. et al. Personal omics profiling reveals dynamic molecular and medical phenotypes. Cell 148, 1293–307 (2012).
Iorio, M. V. et al. MicroRNA gene expression deregulation in human breast cancer. Cancer Res 65, 7065–70 (2005).
Calin, G. A. & Croce, C. M. MicroRNA signatures in human cancers. Nat Rev Cancer 6, 857–66 (2006).
Cui, Q. et al. A map of human cancer signaling. Mol Syst Biol 3, 152 (2007).
Awan, A. et al. Regulatory network motifs and hotspots of cancer genes in a mammalian cellular signalling network. IET Syst Biol 1, 292–7 (2007).
Li, L. et al. The human phosphotyrosine signaling network: evolution and hotspots of hijacking in cancer. Genome Res 22, 1222–30 (2012).
Newman, R. H. et al. Construction of human activity-based phosphorylation networks. Mol Syst Biol 9, 655 (2013).
Zaman, N. et al. Signaling network assessment of mutations and copy number variations predict breast cancer subtype-specific drug targets. Cell Rep 5, 216–23 (2013).
Liang, P. & Pardee, A. B. Analysing differential gene expression in cancer. Nat Rev Cancer 3, 869–76 (2003).
Wang, E. Understanding genomic alterations in cancer genomes using an integrative network approach. Cancer Lett 340, 261–9 (2012).
Wang, E. et al. Cancer systems biology in the genome sequencing era: part 1, dissecting and modeling of tumor clones and their networks. Semin Cancer Biol 23, 279–85 (2013).
Wang, E. et al. Cancer systems biology in the genome sequencing era: part 2, evolutionary dynamics of tumor clonal networks and drug resistance. Semin Cancer Biol 23, 286–92 (2013).
Peri, S. et al. Development of human protein reference database as an initial platform for approaching systems biology in humans. Genome Res 13, 2363–71 (2003).
Shiryev, S. A., Papadopoulos, J. S., Schaffer, A. A. & Agarwala, R. Improved BLAST searches using longer words for protein seeding. Bioinformatics 23, 2949–51 (2007).
Su, A. I. et al. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc Natl Acad Sci U S A 101, 6062–7 (2004).
Pihur, V. & Datta, S. RankAggreg, an R package for weighted rank aggregation. BMC Bioinformatics 10, 62 (2009).
Wang, J. Z., Du, Z., Payattakool, R., Yu, P. S. & Chen, C. F. A new method to measure the semantic similarity of GO terms. Bioinformatics 23, 1274–81 (2007).
van Driel, M. A., Bruggeman, J., Vriend, G., Brunner, H. G. & Leunissen, J. A. A text-mining analysis of the human phenome. Eur J Hum Genet 14, 535–42 (2006).
Smedley, D. et al. BioMart--biological queries made easy. BMC Genomics 10, 22 (2009).
Andrew, V. & Goldberg, S. R. beyond the flow decomposition barrier. Journal of the ACM 45, 783–797 (1998).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4, 44–57 (2009).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 37, 1–13 (2009).
Maere, S., Heymans, K. & Kuiper, M. BiNGO: a Cytoscape plugin to assess overrepresentation of gene ontology categories in biological networks. Bioinformatics 21, 3448–9 (2005).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13, 2498–504 (2003).
Acknowledgements
This work was partly supported by the National Basic Research Program of China (2012CB316504), the National High Technology Research and Development Program of China (2012AA020401), the National Science Foundation (61175002, 60928007 and 61273228). The authors would like to thank Dr. Torsten Juelich for critical reading of this manuscript and helpful suggestions.
Author information
Authors and Affiliations
Contributions
R.J., T.J. and X.Z. provided guidance and planning for the project. Y.C. and R.J. produced the program and wrote the manuscript, particularly producing the results section. J.H., W.J. and T.H. contributed in preparing data and analysis of the results.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Supplementary Information
Supplementary Data
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Chen, Y., Hao, J., Jiang, W. et al. Identifying potential cancer driver genes by genomic data integration. Sci Rep 3, 3538 (2013). https://doi.org/10.1038/srep03538
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep03538
This article is cited by
-
Novel ratio-metric features enable the identification of new driver genes across cancer types
Scientific Reports (2022)
-
DriveWays: a method for identifying possibly overlapping driver pathways in cancer
Scientific Reports (2020)
-
SV-HotSpot: detection and visualization of hotspots targeted by structural variants associated with gene expression
Scientific Reports (2020)
-
Ontology-based prediction of cancer driver genes
Scientific Reports (2019)
-
TOPDRIVER: the novel identifier of cancer driver genes in Gastric cancer and Melanoma
Applied Network Science (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.