Abstract
Soil salinity problems are widespread around the globe with increased risk of spreading over the years. The fungus Piriformospora indica, identified in Indian Thar desert, colonizes the roots of monocotyledon plants and provides resistance towards biotic as well as abiotic stress conditions. We have identified a cyclophilin A-like protein from P. indica (PiCypA), which shows higher expression levels during salinity stress. The transgenic tobacco plants overexpressing PiCypA develop osmotic tolerance and exhibit normal growth under osmotic stress conditions. The crystal structure and NMR spectroscopy of PiCypA show a canonical cyclophilin like fold exhibiting a novel RNA binding activity. The RNA binding activity of the protein and identification of the key residues involved in the RNA recognition is unique for this class of protein. Here, we demonstrate for the first time a direct evidence of countering osmotic stress tolerance in plant by genetic modification using a P. indica gene.
Similar content being viewed by others
Introduction
Abiotic stress leads to a series of morphological, physiological, biochemical and molecular changes that adversely affect plant growth and productivity1,2. High salinity is one of the major threats to plant growth and productivity; it causes both hyperionic and hyperosmotic stress and can lead to plant death3. Plants basically counteract these negative effects of salinity stress by activation of various biochemical and metabolic pathways4 that include accumulation of osmolytes like proline, fructose, myo-inositol, betaine, sucrose etc.5, maintaining the intracellular ion homeostasis and scavenging of reactive oxygen species (ROS)6,7,8. The overexpression of various plant genes has been shown to provide salinity stress tolerance to plants3,9,10.
P. indica, a plant-root-colonizing basidiomycete fungus, was discovered in the Indian Thar desert and has been shown to provide strong growth-promoting activity during its symbiosis with a broad spectrum of plants11. Symbiotic relationship of P. indica with plants results in better plant growth, higher seed yield, early flowering and biotic and abiotic stress tolerance responses12,13. It has been also reported that mutual nutrients like phosphate transport take place by P. indica14. Salt-stress studies have shown positive effects of P. indica on barley12. P. indica also provides resistance towards fungal diseases and tolerance to salt stress in monocotyledonous plants15. However, genes from this beneficial fungus, P. indica, in providing salinity stress tolerance to plants have not been exploited yet.
Cyclophilins (Cyps) are highly conserved, ubiquitous in nature and catalyze the interconversion of peptidyl prolyl imide bonds in peptide and protein substrates16. In eukaryotic cells, Cyps are found in all cellular compartments with a variety of functions being ascribed to them including cell division, transcriptional regulation, protein trafficking, cell signaling, pre-mRNA splicing, molecular chaperone mechanism and stress tolerance16. A striking feature of plant Cyp genes is that their expression is induced in response to various stresses as well as abscisic acid (ABA). Further, wide distribution and ubiquitous nature of Cyps signifies their fundamental importance in plant survival. Although diverse functions of Cyps have been suggested in plants, their physiological relevance and the molecular basis of stress-responsive expression are still largely unexplored17,18,19,20. The role of plant Cyp in conferring salt-stress tolerance in rice has been reported21. Recently, we have identified 28 rice, 35 Arabidopsis and 8 yeast Cyps genes from their respective genomes through a genome-wide analysis16. Earlier, we have reported sequence-specific 1H, 13C and 15N resonance assignments for a Cyclophilin A-like protein from Piriformospora indica (PiCypA) which was found to be upregulated during osmotic stress conditions22.
Though Cyps are known to play role in pre-mRNA splicing, their RNA-binding activity has not been well described. RNA-binding proteins (RBPs) play major role in RNA metabolism, pre-mRNA processing, transport, stability and translation23,24. RBPs are involved in posttranscriptional regulatory machinery that further allows plants to respond against environmental stresses25. Many RBPs have been reported to be regulated during abiotic stresses in plants like MA16, a glycine rich RBP which is induced by dehydration stress and ABA in maize embryos26, Tudor-SN (TSN) protein which is highly expressed during drought stress in Arabidopsis27. DEAD-box RNA helicase (ZmDRH1) in Zea mays28 and glycine rich RBP from Sorghum bicolor29 have been reported to be upregukated under abiotic stress. Here we report the crystal structure of PiCypA protein, the role of PiCypA in imparting osmotic stress tolerance to plant and the unique property of PiCypA to bind RNA. Some of the interacting partners of PiCypA protein were also upregulated under salt stress in the transgenic plants. Furthermore this tolerance is observed in T2 generation of transgenic plants and the possible mechanism of the tolerance is by maintaining the antioxidant machinery of plants under stress conditions.
Results
Identification of PiCypA and its overexpression in tobacco plants
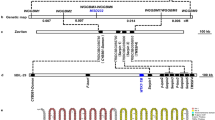
P. indica was grown at different salt concentrations and good growth of P. indica was observed at 200 and 400 mM NaCl, while there was moderate growth at 600 mM NaCl and no growth at 800 mM NaCl. To identify the genes involved in the high salt tolerance, the functional screening of salinity tolerant genes was performed. For this purpose E. coli cells were transformed with phagemids derived from a cDNA library (made from RNA isolated from P. indica grown at 600 mM NaCl) as described in materials and methods. A total of over one million recombinant bacterial cells (E. coli SOLR cells) were plated on LB agar medium supplemented with 600 mM NaCl and without salt as control. A lawn of colonies on the control and only 36 colonies on the plate containing 600 mM NaCl were observed. Out of 36 salt tolerant colonies we selected one for further analysis, which was found to be a gene encoding cyclophylin A-like protein (PiCypA; accession number GQ214003). Functional significance of the PiCypA gene was established by overexpressing the complete open reading frame (ORF; 495 bp) in tobacco plants using Agrobacterium-mediated transformation with the gene construct pCAMBIA1301-PiCypA ( Fig. 1a ).
Analysis of PiCypA overexpressing transgenic tobacco plants (T1 generation) under osmotic stress.
(a) Structure of T-DNA region of pCAMBIA1301 containing PiCypA gene (495 bp) in XhoI and EcoRI restriction enzyme sites of MCS region with CaMV35S promoter and poly A terminator (pCAMBIA1301-PiCypA). (b) Visualization of β-glucuronidase (GUS) activity in the leaf tissue of WT and vector control (VC) controls and the transgenic lines (L12.1, L12.2 and L12.3) by using X-gluc. The hygromycin resistant putative tobacco transformants were used for GUS assay. (c) PCR analysis of the transgenic lines by the use of PiCypA gene specific forward and reverse primers. Lane 1 (M) is DNA marker, lane 2 is WT, lane 3 is VC and rests of lanes (4–6) are transgenic lines showing the required amplification (495 bp). (d) The PiCypA transgenic lines and control plants (WT and VC) were analyzed by the Southern blot hybridization to show the integration of the PiCypA gene along with CaMV35S promoter and poly A terminator (1.1 kb) in the transgenic lines. The genomic DNAs were digested with HindIII and probed with radiolabelled ORF of PiCypA gene. The gel is cropped for clarity. (e) Relative mRNA expression in all the three transgenic lines checked by Real Time PCR. (f) Western blot analysis of the WT and VC control plants and T1 transgenic lines (12.1, 12.2 and 12.3) using PiCypA polyclonal antibodies. The gel is cropped for clarity. Full gel figures for (d) and (f) are shown in supplementary figure S4.
The same vector not containing the PiCypA gene called empty vector control (VC), pCAMBIA1301, was also transformed to plants treated as an additional control along with the wild type (WT) plants. Seeds from the T0 plants, when plated onto hygromycin-containing medium, segregated in the expected 3:1 ratio of Hygr/Hygs ( Table 1 ).
Analysis for the PiCypA transgene integration was carried out on the T1 generation of transgenic plants. We selected 16 independently transformed T1 transgenic plants and grew them to maturity; however, only 3 different lines (L12.1, L12.2 and L12.3) were randomly selected for subsequent analysis. The hygromycin resistant putative tobacco transformants and VC plants were confirmed by β-glucuronidase (GUS) assay and found to be positive whereas WT was GUS negative ( Fig. 1b ). Integration of PiCypA gene (495 bp) was confirmed in the GUS positive transgenic lines by PCR using gene specific primers, whereas WT and VC plants were PCR negative ( Fig. 1c ). The morphological and growth characteristics of T1 plants were similar to the WT and VC control plants. The stability of integration of the transgene was further confirmed by probing of HindIII digested genomic DNA with an equal amount of PiCypA cDNA. The expected band pattern per insertion was observed at 1.1 kb position (containing the entire cauliflower mosaic virus promoter and full ORF of PiCypA cDNA) ( Fig. 1d , lanes 2–4). No band was detected in the WT plants ( Fig. 1d , lane 1). Real time PCR of all the three lines showed the expression of transcript in all of the transgenic lines ( Fig. 1e ). Western blot of the T1 transgenics by PiCypA polyclonal antibodies identified an 18 kDa band corresponding to the purified recombinant protein and its intensity varied between the different transgenic lines ( Fig. 1f , lanes 3–5). A very faint protein band of the same size was also observed in the WT and VC plants ( Fig. 1f , lanes 1 and 2). This could be possibly due to the cross-reactivity of the polyclonal antibodies with native tobacco protein. All these findings confirmed the stable integration of the PiCypA gene into the transgenic plants.
PiCypA provides tolerance to excess osmotic stress
To test for osmotic stress tolerance, leaf disks from T1 transgenic, control tobacco plants (WT and VC) were floated separately on 200 mM NaCl and H2O for 72 h. Salinity-induced loss of chlorophyll was lower in PiCypA overexpressing lines (L12.1, L12.2 and L12.3) compared with those from the WT and VC plants ( Fig. 2a ). The damage caused by stress was reflected in the degree of bleaching observed in the leaf tissue after 72 h. It was evident that transgenic lines had a better ability to tolerate osmotic stress. The measurement of the chlorophyll content of the leaf disks from different transgenic lines stressed with 200 mM NaCl provided further support for a positive relationship between the overexpression of PiCypA and tolerance towards osmotic stress ( Fig. 2b ). To assess the tolerance, we germinated the seeds of all three transgenic lines along with WT and VC in 200 mM NaCl and in water as control. All the above seeds germinated well in water ( Fig. 2c , first row), while a better and high germination rate of transgenic lines (12.1, 12.2 and 12.3) was observed as compared to WT and VC ( Fig. 2c , bottom row). The T1 seedlings were morphologically similar when grown with or without NaCl. To assess the effect of high salt (200 mM NaCl) on the growth and development of PiCypA overexpressing lines, WT and VC plants, the three week old seedlings (after growing in water) were further grown in the soil continuously irrigated by 200 mM NaCl. In the presence of NaCl, the WT and VC plants showed growth retardation, whereas transgenic plants did not develop any sign of stress ( Fig. 2d , bottom row). It was observed that without salt all plants had similar growth ( Fig. 2d , upper row). The T1 transgenic plants grew normally and set viable seeds under continuous osmotic stress, whereas the WT and VC plants could not survive till maturity under continuous osmotic stress ( Fig. 2e ). Overall, the results showed that the PiCypA overexpressing T1 transgenic lines have shown better ability to tolerate osmotic stress. This is the first direct evidence to show osmotic stress tolerance in plant by a P. indica fungus gene.
Analysis of osmotic stress tolerance in the T1 transgenic lines.
(a) Leaf disk senescence assay for osmotic tolerance in transgenic tobacco plants (L12.1, L12.2 and L12.3) in 200 mM NaCl. Transgenic lines showing high osmotic stress tolerance as compared to controls (WT and VC). (b) Remaining chlorophyll content from leaf disks of WT, VC plants and the transgenic lines after incubation 200 mM NaCl solution (Clh A and Clh B are shown in dark red and light green bar respectively). Disks floated in water served as the experimental control (yellow bar represents total chlorophyll content in the control condition). (c) Germination and seedling growth of WT, VC and T1 transgenic lines (L12.1, L12.2 and L12.3) under water condition (for control) and 200 mM NaCl (for experiment). (d) Phenotypic comparison of growth of T1 transgenic tobacco plants overexpressing PiCypA gene (L12.1, L12.2 and L12.3) with controls plants (WT and VC) in the presence of water and 200 mM NaCl. Seeds were germinated for 7 days and plantlets grown on water (upper row) and on 200 mM NaCl (lower row). (e) WT, VC and transgenic plants (L12.1, L12.2 and L12.3) in soil pots supplied with 200 mM NaCl solution. Note that the WT and VC plants could not sustain growth under osmotic stress. (f) Phenotypic comparison of growth of T2 transgenic tobacco plants overexpressing PiCypA gene (L12.1.1, L12.2.1 and L12.3.1) with control plants (WT and VC) in the presence of water and 200 mM NaCl.
Growth parameters and viable seeds under osmotic stress in PiCypA-transgenic plants
The percent seedlings survival of transgenic lines grown in 200 mM NaCl did not significantly change as compared to water grown WT and VC controls ( Table 1 ). Several critical growth parameters, such as plant height, fresh weight of leaves, total protein content, time required for flowering and seed weight were scored as an indicator of osmotic stress to test tolerance in T1 transgenic lines ( Table 1 ) as compared with the WT plants grown in H2O. It should be noted here that similar data for WT/VC plants grown under osmotic stress could not be obtained because these plants failed to sustain growth in the presence of 200 mM NaCl ( Fig. 2e ). Overall performance or total seed yield of the transgenic plants was unaffected in the presence of 200 mM NaCl. The flowering time in osmotic-stressed plants was delayed by a week when compared with plants grown without stress, but there was no change in the flower morphology and yield ( Table 1 ). Significantly, the pod and seed size, seed number and seed weight of salt grown transgenic lines were statistically similar to those from WT and VC plants grown without osmotic stress ( Table 1 ).
PiCypA overexpressing tobacco lines confers tolerance to oxidative stress
In response to 200 mM NaCl, PiCypA transgenic lines exhibited lesser accumulation of malondialdehyde (MDA) (lipid peroxidation product), H2O2 and ion leakage as compared to controls seedlings (WT and VC), suggesting more tolerance to oxidative stress in transgenic tobacco under the osmotic stress ( Supplementary fig. S1 a–c ). We observed higher increase in the activities of Catalase (CAT), ascorbate peroxidases (APX) and glutathione reductase (GR) in transgenic lines as compared to control plants (WT and VC) under osmotic stress ( Supplementary fig. S1 d, e and f ). Moreover, the proline accumulation is another common adaptive mechanism for providing osmotolerance to plants against abiotic stress42. In this study we observed that the proline accumulation and relative water content were also maintained in transgenic lines as compared to control plants (WT and VC) ( Supplementary fig. S1 g and h ).
T2 transgenic tobacco plants grow normally and set viable seeds under osmotic stress
The hygromycin resistant T1 transgenic seeds were grown in soil under normal conditions. These plants (T2 generation) were morphologically similar to that of WT and tested for osmotic stress tolerance. Osmotic stress-induced loss of chlorophyll in leaf disk assay was lower in PiCypA overexpressing T2 lines (L12.1.1, L12.2.1 and L12.3.1) compared with those from the WT and VC plants ( Supplementary fig. S2 a and b ). PiCypA overexpressing T2 lines under osmotic (200 mM NaCl) stress accumulated less H2O2 in comparison with WT and VC as tested by DAB staining method and its quantitation ( Supplementary fig. S2 c and d ). The concentration of proline, an important osmoregulator, was higher in PiCypA overexpressing T2 lines as compared to WT and VC ( Supplementary fig. S2e ). The growth retardation was also reported in three weeks old WT and VC plants whereas T2 transgenic lines grew happily under continuous salt irrigation ( Fig. 2f , lower row). It was observed that without salt all the plants showed similar growth ( Fig. 2f , upper row).
Molecular architecture and dynamics of PiCypA
To understand the molecular basis of PiCypA's role in providing salt stress tolerance, we performed structural and dynamics studies. Crystal structure of PiCypA was solved at 1.97 Å resolutions by molecular replacement method. Crystallization and data collection statistics have been described previously30. The model includes the entire polypeptide chain and was refined to Rwork and Rfree values of 0.159 and 0.183, respectively ( Supplementary Table S1 ). Crystal arrangement has shown three molecules per asymmetric unit arranged in a side-by-side orientation. PiCypA is a monomeric single domain protein and forms a canonical cyclophilin fold comprising two α-helices and eight β-strands ( Fig. 3a ). Eight β-strands form a flattened cylinder with β-barrel like fold where two helices sit on top and bottom of the barrel. Six phenylalanines (F11, F25, F39, F56, F115 and F132) and one tyrosine (Y51) arranged in edge-to-face orientation form a tight hydrophobic core. Around this core, other hydrophobic aliphatic residues (V9, I13, I23, L27, L42, I59, L65, L101, M103, I117 and V135) form a closed cage structure ( Fig. 3b ). Eighty five residues comprising ~52% of a full polypeptide chain (N45–F56, G68-P98, A104-Q114 and T119-V131) are present in five loop regions that have similar B-factor compared to average B-factor of 29.5 Å2 for the entire polypeptide chain. The rigid nature of loops in solution was also confirmed from the NMR dynamics data. The residues forming these loops have similar order parameter like those forming α-helices and β-strands that confirms that the loops are rigid. We calculated order parameter by analyzing NMR relaxation parameters R1, R2 and 15N-{1H} nOe. Complete sequence-specific NMR resonance assignments of the protein have been reported previously22. We had observed that chemical shifts of amide protons of L34 (10.67 ppm), D88 (11.36 ppm) and H95 (10.74 ppm) and amide nitrogen of E89 (132.8 ppm) are unusually downfield shifted. The unusual downfield shifts arise due to deshielding effect caused by delocalized π-electron ring currents of the proximal aromatics rings of residues Y82, F86, F91 and W124 ( Fig. 3c ).
Molecular architecture and RNA binding activity of PiCypA.
(a) β-barrel like fold containing eight β-strands and two helices forming cap at either ends (b) aromatic (cyan) and aliphatic (magenta) forming hydrophobic core (c) Residues showing unusual chemical shifts (Magenta) in NMR spectra and aromatic residues (cyan) responsible for it. (d) Western blot of PiCypA protein probed with anti-his antibody (upper panel) as loading control. For this 1 μg of BSA and increasing concentration of PiCypA (1, 2 and 4 μg) proteins were dot-blotted on precharged PVDF membrane. An identical blot was probed with a 13 mer radio labeled RNA oligo (5′-AUAGCCUCAACCG-3′) and developed. (e) Isothermal titration calorimetry indicating hyperbolic nature for PiCypA binding to the 13 mer RNA (f) Selected regions from the superimposed 2D [15N, 1H] HSQC spectra of free and complexed PiCypA shown in red and blue, respectively. The resonance assignments of residues with large chemical shift changes upon RNA binding (1:1 molar ratio) are shown with arrows. (g) Combined chemical shift changes of PiCypA upon RNA binding in 1:1 molar ratio. Red line indicates one standard deviation for the mean chemical shift change of the dataset. Combined chemical shift changes were calculated by taking the square root of the sum of the squares of the chemical shift changes for the backbone 1HN and 15N resonances after scaling the 15N shifts by 0.2 (h) Mapped RNA binding region (R1, R2 and R3) on the crystal structure of PiCypA.
As PiCypA shares 62% sequence homology with human cyclophilin A, we investigated two characteristic features of cyclophilin-like fold containing proteins, PPIase (peptidyl prolyl cis-trans isomerase) domain architecture and cyclosporin A (CsA) binding activity. PiCypA carries a PPIase domain with all conserved residues (R58, F63, M64, Q67, A104, F116, W124, L125 and H129). NMR titrations with CsA have shown that PiCypA has same conserved CsA binding surface (H57, R58, I60, Q66, G67, G68, H95 and I117). Along with these residues, three more residues that are mutated from the human counterpart (C55T, C64M and C118T) completes CsA interacting region ( Supplementary fig. S3 a–c ).
To explore the effect of high salt on the structure of PiCypA we performed salt titration experiments. In the 2D [15N, 1H] HSQC spectra except for minor changes in chemical shifts of surface exposed residues T55, S80 and K85 no significant backbone amide chemical shift perturbations (CSPs) were observed confirming that structural integrity of the protein is maintained even at 600 mM NaCl22.
PiCypA specifically binds to RNA
We observed a novel RNA binding property of PiCypA when the protein was spotted on a charged PVDF membrane in increasing concentration. PiCypA showed strong RNA binding as shown in autoradiograph. Bovine serum albumin (BSA) was used as a negative control ( Fig. 3d ).
To investigate specificity of RNA binding, PiCypA was titrated against different RNA constructs and we obtained quantitative thermodynamic parameters of binding. The selected RNA sequences were of different length and sequences (1. 5′-GCU GCG-3′; 2. 5′-CGU GCG-3′; 3. 5′-AUA GCC UCA ACC G-3′; 4. 5′-GCU GCU GCG-3′). No RNA binding was observed for 6 mer or 13 mer poly-A RNA sequence. A 13 mer RNA sequence (AUAGCCUCAACCG) was the strongest binder to the protein. The binding constant (Kd) of 22.5 μM indicates moderate to strong binding with ΔH and ΔS values of −118.4 Kcal/mol and −369 cal/mol/deg respectively. The observed isotherms indicated a moderate to strong nature of binding. Therefore, corresponding stoichiometry of the binding could not be determined as the observed ‘c value’ of isotherm was 0.2. As the observed ‘c value’ is outside of the desired range of 1 < c < 1000 a hyperbolic isotherm as compared to sigmoidal isotherm was observed that does not allow correct determination of stoichiometry31,32 ( Fig. 3e ).
13 mer RNA (AUAGCCUCAACCG) was titrated against U-15N labeled PiCypA at pH 6.5 and shifts in the backbone 15N and 1HN resonances in 2D [15N, 1H] HSQC spectra were monitored. A gradual shift in the resonances was observed for all the residues that showed changes indicating a fast exchange (τex < μs) between free and RNA bound PiCypA ( Fig. 3f ). No significant changes in chemical shifts were observed after 1:1 molar ratio of protein and RNA indicating stoichiometry of 1:1 in the complex hence indicating one RNA molecule binding to one PiCypA molecule. The residues showing the high cumulative backbone CSPs are most likely to be involved in the RNA binding. The residues showing significant CSPs are clustered into three major regions of the polypeptide chain. These regions are R1 (L65, G68, T71, G76, T77, G78, G79 and R80), R2 (M103, A104, N105, G107 and T110) and R3 (W124, L125, D126, G127, K128, H129 and V130) along with two individual residues T55 and F115 ( Fig. 3g ). We mapped these residues on the crystal structure of PiCypA. These three regions are clustered around the groove that comprises of aromatic as well as putative hydrogen bonding residues, making it more likely to interact with nucleic acid bases ( Fig. 3h ).
Interaction of PiCypA with other stress induced proteins and their expression level
We performed yeast-2-hybrid assays to find putative interacting protein partners from P. indica cDNA library. A total of 15 interacting proteins of PiCypA were obtained and their putative functions are mentioned in Supplementary Table S2 . Out of these 3 interacting proteins were found to be abiotic stress induced as reported earlier9,33,34,35,36. The expression level of these genes namely phosphoglycerate kinase (PGK), heat shock protein 6 (HSP6) and eIF-4A were found to be higher in transgenic lines as compared to WT ( Fig. 4a–c ) suggesting that PiCypA might be playing vital role in maintenance of mRNA integrity. We also performed semi-quantitative PCR to check the expression level of PGK, HSP6 and eIF-4A mRNAs ( Fig. 4d ). The high expression level of these stress-induced genes together with PiCypA provides increased tolerance to osmotic stress.
Expression level of tobacco homologous genes of PiCypA interacting partners (PGK; phosphoglycerate kinase, HSP6; heat shock protein 6, eIF-4A; DEAD box helicase).
(a), (b) & (c) Real time expression analysis of PGK, HSP6 and eIF-4A, respectively (where WT-N is normal WT plant and WT-S is WT plant with stress treatment in NaCl). (d) Semi-quantitative PCR of these genes PGK [1], HSP6 [2], eIF-4A [3] and actin [4] as control. The gel is cropped from four gels and all four gels are shown in supplementary figure S5.
Discussion
Soil salinity problems have hampered cultivation of many crop plants in large parts of the world. Unregulated large-scale irrigation programs and poor soil management are major reasons for its increased risk of spreading over the years. High salinity stress conditions affect the cellular homeostasis and cause disturbance in water potential and ion distribution. This disturbance in water and ion potential leads to molecular damage, growth inhibition and even death of plants. The molecular chaperones from plants such as heat shock proteins (HSPs) and cyclophilins have been shown to play role in abiotic stress tolerance35,37,38. Cyclophilins have endogenous conserved cytosolic peptidyl-prolyl isomerase (PPiase) activity, which catalyzes the inter-conversion of the cis and trans conformation of peptide bonds of proline residues in polypeptides. This inter-conversion is a rate-limiting step during protein folding39,40. Cyclophilins have been reported to play important role in different cellular functions such as inhibition of cell proliferation, induction of apoptosis and suppression of migratory/invasive capacity41. Interestingly, in silico analysis have predicted Cyclophilin to be an abiotic stress responsive gene16. In salt tolerant model plant Thellungiella, the expression level of cyclophilin was found to be higher44. Role of CYP20–3 (At3g62030) gene from Arabidopsis has been reported in light and other stress conditions16. Another gene from rice OsCyP20–2 (LOC_Os5g01270), showed tolerance against various abiotic stresses16. In the case of yeast cyclophilins, it has been also reported that CPR1 gene help in countering multiple abiotic stresses when overexpressed in E. coli and Yeast16. The cyclophilin protects the cells against oxidative induced death42 and expression levels were found to be elevated after oxygen stress43. However, the role of cyclophilin from P. indica, a salinity tolerant fungus, in stress condition has not been described so far. Here we have dissected the structure-function of PiCypA in osmotic stress tolerance in tobacco plants and demonstrated its novel RNA-binding property.
The PiCypA overexpressing transgenic tobacco plants exhibited better growth and survival ability as compared to normal WT and VC plants under osmotic stress. This study suggests that PiCypA may play a crucial role in stress protection presumably through its active role in maintenance of protein structure during stress. The overexpression of PiCypA in tobacco plants leads to improvement in the antioxidant machinery and accumulation of osmoprotectant like proline, which cope up with the damage caused by elevated ROS during the stress. The harmful effects of increased ROS and improved antioxidant machinery in transgenic rice under stress have also been reported earlier8,21,45. PiCypA expressing T2 transgenic plants also showed normal growth and increased osmotic stress tolerance indicating the functional introduction of the trait, which is stable in T2 generation of transgenic tobacco plants. Interestingly, we have found several interacting partners of PiCypA from P. indica and plant homologues of few of them have been reported in abiotic stress response previously such as PGK, HSP6 and eIF-4A9,33,34,35,36. The expression levels of these genes were significantly higher in PiCypA overexpressing lines as compared to normal WT plants.
Although symbiotic relation of P. indica fungus is known to provide salt stress to plants still molecular mechanism of such action is currently unknown. To understand the molecular mechanism we determined the atomic-resolution structure of a cyclophilin A-like protein from P. indica and observed novel RNA binding property. Crystal structure of PiCypA revealed a conserved canonical cyclophilin like fold comprising two α-helices (P33 – K44 and M139 – V148) and eight β-strands (V9 – I15, Q18 – L27, H57 – I60, F63 – G67, G99 – M103, F115 – T118, F132 – E137 and V159 – T166) in (β1-2)-α1-(β3-5)-α2-β8 architecture. In addition to usual cyclophilin like properties we also observed a novel RNA binding property in PiCypA. As CsA titration data indicates ( Supplementary fig. S3a–c ), perturbed residues upon RNA interaction are shared with those forming CsA interacting region (M103, A104, W124, K128, H129 and V130). All residues involved in RNA interaction are conserved in cyclophilins from different organisms throughout the evolution. PiCypA like AtCyp59 has specificity for GU rich sequences indicates a possibility of its role in stability of mRNA transcripts as ~70% of cellular transcripts carry UG-rich region in their 3′-UTR as well as exon-intron junctions as discussed earlier46. Other Cylophilins like AtCyp59 from Arabidopsis thaliana and hCyP33 from human have been reported to bind RNA but they contain an additional RNA recognition motif (RRM)46,47. Though PiCypA does not have any canonical RRM domain it is still able to bind RNA provides an interesting insight into a new class of RNA binding motif. Possibly PiCypA has evolved and got rid of RRM domain but RNA binding property remains conserved like other RNA binding cyclophilins46. PiCypA might be involved in stabilization of RNAs and possibly protecting the RNA integrity by chaperone-mediated process. This opens up window for further exploration for identification of target RNAs that are interacting with PiCypA.
In this study we have reported the structure of a protein from P. indica fungus at atomic-resolution. We have also demonstrated for the first time a direct evidence of countering osmotic stress tolerance in plant by genetic modification using a P. indica gene. A novel RNA binding activity of PiCypA was also observed that might be part of machinery involved in RNA stabilization during stress conditions. RNA binding activity of PiCypA generates new avenues for involvement of cyclophilins in various mechanisms involving RNA stability and other processes involving transcripts and hnRNPs. These findings open up new areas for investigating the cellular machinery in plants and fungus to counteract salinity stress tolerance and have a great potential for crop improvement.
Methods
Identification and cloning of cyclophilin A-like gene of P. indica
Identification of candidate genes responding to osmotic stress was done as per the method previously described30. One of the clones, cyclophylin A-like protein (accession number GQ214003), was selected for further studies. The gene cloned into pBSK vector was amplified by using PCR Primers (forward: 5′ CTCGAGCATATGTCCCAGCCCAACGTCTACTTTG 3′ and reverse: 5′-GAATTCTTAGACAGTGCCAGACGCAGTAATCTTG 3′). The PCR product (495 bp) was cloned into the pGEMT-easy vector (Promega) and sequenced using the T7 and SP6 primers respectively. Subsequent subcloning was done in pET28a vector (Novagen) using the NdeI and EcoRI restriction sites, generating the pET28a-PiCypA construct.
Expression and purification of PiCypA
The native PiCypA was expressed in E. coli BL21 (DE3) codon plus cell in LB medium. For NMR samples preparation, E. coli BL21 (DE3) codon plus cells were grown in 15NH4Cl and 13C-Glucose labeled media. Cell cultures were grown at 37 °C till OD600 ~ 0.8, induced with Isopropyl β-D-thiogalactoside (IPTG) overnight at 18 °C and then harvested by centrifugation. Soluble fraction of lysates was transferred onto a 5 ml Ni-NTA affinity beads (QIAGEN) on a gravi column (GE Healthcare), pre-equilibrated with lysis and wash buffer. Elutes containing PiCypA were buffer exchanged and concentrated against 50 mM NaCl, 20 mM Tris-Cl buffer pH 6.5. Purified protein was then subjected to Thrombin (Amersham Biosciences) digestion to remove the N-terminal His-tag. Traces of thrombin were removed by passing the reaction mixture through a Benzamidine column (GE Healthcare). Further purification of PiCypA was carried out by size-exclusion chromatography by HiLoad 16/60 Superdex 75 preparative column (GE healthcare). Purified fraction was concentrated to ~1 mM (18 mg/ml) of protein concentration using 3 kDa cut-off Centricon (Millipore).
Plant transformation
The PiCypA from pGEMT digested by XhoI and EcoRI and subcloned into pRT101 and from this the complete cassette (35S CAM promoter + cyclophilin ORF+ NOS terminator) subcloned into pCAMBIA1301 vector. The pCAMBIA1301-PiCypA also carries the HYG (Hygromycin) gene as a selectable marker. Tobacco (Nicotiana tabacum cv. tabbacci) leaf disks were transformed through a procedure9 with Agrobacterium tumefaciens (LBA4404) containing pCAMBIA1301-PiCypA. Putative T0 transgenic plants were regenerated from the callus in the presence of Hygromycin and further screened by using PCR and β-glucuronidase (GUS) assay48. The seeds from these plants, i.e., T0 seeds, were germinated on Hygromycin containing medium to select for all of the transgenic T1 seedlings. GUS activity was visualized in the leaf tissue and fluorescence emission intensity was measured using 1 mM 4-methyl-umbelliferyl-β-D-glucuronide as substrate48.
Leaf disk assay
Leaf disks of 1.0-cm diameter were excised from healthy and fully expanded tobacco leaves of similar age from transgenic and WT plants (60 days old). The disks were floated in a 4-ml solution of 200 mM NaCl and water (experimental control) for 72 h49 and then used for measuring chlorophyll a and b spectrophotometrically after extraction in 80% acetone50. The experiments were repeated at least three times with different transgenic lines.
Genomic DNA PCR and Southern analysis
Genomic DNA was isolated by grinding leaf tissue followed by extraction using the CTAB (N-acetyl-N,N,N trimethylammonium bromide) method51. PCR was performed using the gene specific primers PiCypA-F1 (5′ CTCGAGCATATGTCCCAGCCCAACGTCTACTTTG 3′) starting from the translation initiation site and PiCypA-R1 (5′-GAATTCTTAGACAGTGCCAGACGCAGTAATCTTG 3′). For Southern analysis, 20 μg of genomic DNA from PCR-positive tobacco lines was digested with HindIII, electrophoresed and blotted on Hybond N membranes (Amersham Pharmacia). Radiolabeled 495 bp complete ORF of PiCypA cDNA was used as a probe. Hybridization and washing were carried out at 55°C52.
Real time PCR, immunoblotting and in vitro RNA binding assay
Total RNA was extracted from 200 mg of young leaves of tobacco transgenic lines using the RNeasy total RNA isolation kit (Qiagen, Germany). RT-PCR was performed using total RNA pre-treated with RNase-free DNase I. 5 μg aliquot of total RNA from each sample was used in a 10 ml reverse transcription reaction system with oligo-dT as primer (Takara, Japan). 1 μl aliquot of reverse transcription reaction mixture after the completion of cDNA synthesis was used as the template for real time PCR amplification. Total soluble proteins from the tissue were separated by 12% SDS-PAGE and analysis was performed by using PiCypA specific primary antibodies (1:5,000 dilutions) and alkaline phosphatase conjugated secondary antibodies52. The in vitro RNA binding assay was done using the method described previously53. For this 1 μg of BSA and increasing concentration of PiCypA (1, 2 and 4 μg) proteins were dot-blotted on precharged PVDF membrane and probed with 13 mer α or γ 32P radio-labeled RNA oligo (5′-AUAGCCUCAACCG-3′).
Root growth assay
Seeds of the T0 transgenic tobacco plants were selected by germinating them on Murashige and Skoog54 medium in the presence of hygromycin. These seedlings representing the T1 generation were tested for their sensitivity of root growth to osmotic stress. Individual seedlings were selected for uniformity and their initial root lengths were recorded.
Biochemical and physiological parameters
Quantitation of H2O2, lipid peroxidation, electrolyte leakage, antioxidant enzyme assays and proline estimation, were performed using the method described previously55. Relative water content (RWC) was calculated following the method of Barrs and Weatherley56.
Crystallization, data collection and processing
Purified PiCypA protein was used for crystallization. After screening and optimization diffraction quality crystals belonging to crystal space group C2221 were obtained. The condition of crystallization and data collection has been reported earlier30. Diffraction data were collected with Cu K-radiation (λ = 1.5418 Å) using a Rigaku FR-E+ SuperBright microfocus rotating-anode dual-wavelength (Cu and Cr) X-ray generator equipped with R-AXIS IV++ detectors. The crystals diffracted to 1.97 Å resolution limit. The data sets were indexed, integrated and scaled using MOSFLM57 and ccp4 suite58. Phasing was obtained by molecular replacement59 using human Cyclophilin A Structure (Protein Data Bank ID code 3K0N) as a search model. Model building, phasing and refinement were performed using AutoBuild, phenix-refine and RefMac software on Phenix and ccp4 suits59. Detailed crystallographic statistics are listed in Supplementary Table S1 .
NMR spectroscopy
Spectra measurements were carried out at 30 °C on Bruker Avance III spectrometers equipped with 5 mm cryogenic triple resonance TCI probe-head, operating at field strengths of 500 and 700 MHz. Standard set of triple resonance was used to obtain sequence-specific backbone and side-chain resonance assignments. Details of resonance assignments have been reported earlier22. Temperature calibration was performed using a 100% d4-methanol sample60. Backbone dynamics of PiCypA was studied using 15N relaxation data (R1, R2 and 15N-{1H} nOe) obtained from 2D [15N, 1H] NMR spectroscopy61,62,63. The relaxation data were analyzed using the MODELFREE approach64,65,66.
All titration experiments were performed by adding NaCl, Cyclosporin A or RNA in equal molar steps and 2D [15N, 1H] HSQC spectrum was measured at each molar ratio till no further chemical shift changes were observed. The cumulative backbone chemical shift perturbation (CSP) was calculated using the formula:

where Δδ is the change in backbone amide peaks chemical-shift of PiCypA in the 2D [15N, 1H]-HSQC spectra upon complex formation. Topspin 2.1 (Bruker AG) was used for processing of all spectra and data analysis was performed with CARA67.
Isothermal titration calorimetry
Isothermal titration calorimetry (ITC) experiments were performed by injecting 1.2 mM RNA in the cell containing ~50 μM of PiCypA on MicroCal ITC200 (GE Healthcare). PiCypA was in 10 mM NaCl, 10 mM phosphate buffer (pH 6.5) and RNA was reconstituted in nuclease free water. Data analysis was carried out on Origin 8 software (OriginLab).
Yeast-2-hybrid assay
A Gal4 based two-hybrid system was used as described by the manufacturer (Clontech, USA; http://www.clontech.com/). The coding region of the PiCypA (495 bp) was amplified by PCR with primers harboring restriction sites and cloned in-frame into the NdeI and EcoRI sites of the binding domain vector pGBKT7. The resulting vector pGBKT7-PiCypA was co-transformed with P. indica AD library into yeast strain AH109 harboring two reporter genes (HIS3 and β-galactosidase) by the lithium acetate method68.
References
Wang, W. X., Vinocur, B., Shoseyov, O. & Altman, A. Biotechnology of plant osmotic stress tolerance: physiological and molecular considerations. In: S. Sorvari et al., eds., Int. Symp. On In Vitro Culture and Horticultural Breeding. Acta Hort. 560, 285–292 (2001).
Mahajan, S. & Tuteja, N. Cold, salinity and drought stresses: An overview. Arch. Biochem. Biophys. 444, 139–158 (2005).
Tuteja, N. Mechanisms of high salinity tolerance in plants. Methods Enzymol. 428, 419–438 (2007).
Golldack, D., Lüking, I. & Yang, O. Plant tolerance to drought and salinity: stress regulating transcription factors and their functional significance in the cellular transcriptional network. Plant Cell Rep. 30, 1383–1391(2011).
Parida, K. A. & Das, A. B. Salt tolerance and salinity effects on plant: a review. Ecotox. Environ. Safe. 60, 324–349 (2005).
Flowers, T. J. Improving crop salt tolerance. J. Exp. Bot. 55, 307–319 (2004).
Ashraf, M. & Akram, N. A. Improving salinity tolerance of plants through conventional breeding and genetic engineering: An analytical comparison. Biotechnology Adv. 27, 744–752 (2009).
Gill, S. S. & Tuteja, N. Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol. Biochem. 48, 909–939 (2010).
Mishra, N. S., Pham, X. H., Sopory, S. K. & Tuteja, N. Pea DNA helicase 45 overexpression in tobacco confers high salinity tolerance without affecting yield. Proc. Natl. Acad. Sci. USA 102, 509–514 (2005).
Dang, H. Q., Tran, N. Q., Gill, S. S., Tuteja, R. & Tuteja, N. A single subunit MCM6 from pea promotes salinity stress tolerance without affecting yield. Plant Mol. Biol. 76, 19–34 (2011).
Verma, S. et al. Piriformospora indica gen. et sp. nov., a new root-colonizing fungus. Mycologia 90, 896–903 (1998).
Baltruschat, H. et al. Salt tolerance of barley induced by the root endophyte Piriformospora indica is associated with a strong increase in antioxidants. New Phytol. 180, 501–510 (2008).
Singh, L. P., Gill, S. S. & Tuteja, N. Unraveling the role of fungal symbionts in plant abiotic stress tolerance. Plant Signal. Behav. 6, 175–191 (2011).
Yadav, V. et al. A Phosphate transporter from the root endophytic fungus Piriformospora indica plays a role in Phosphate transport to the host plant. J. Biol. Chem. 285, 26532–26544 (2010).
Waller, F. et al. The endophytic fungus Piriformospora indica reprograms barley to salt-stress tolerance, disease resistance and higher yield. Proc. Natl Acad. Sci. USA 102, 13386–13391(2005).
Trivedi, D. K., Yadav, S., Vaid, N. & Tuteja, N. Genome wide analysis of cyclophilin gene family from rice and Arabidopsis and its comparison with yeast. Plant Signal. Behav. 7, 1–14 (2012).
Godoy, A. V., Lazzaro, A. S., Casalongue, C. A. & Segundo, B. S. Expression of a Solanum tuberosum cyclophilin gene is regulated by fungal infection and abiotic stress conditions. Plant Sci. 152, 123–134 (2000).
Kong, H. Y., Lee, S. C. & Hwang, B. K. Expression of pepper cyclophilin gene is differentially regulated during the pathogen infection and abiotic stress conditions. Physiol. Mol. Plant Path. 59, 189–199 (2001).
Sekhar, K., Priyanka, B., Reddy, V. D. & Rao, K. V. Isolation and characterization of a pigeonpea cyclophilin (CcCYP) gene and its over-expression in Arabidopsis confers multiple abiotic stress tolerance. Plant Cell Environ. 33, 1324–1338 (2010).
Zhu, C. et al. Overexpression of a cotton cyclophilin gene (GhCyp1) in transgenic tobacco plants confers dual tolerance to salt stress and Pseudomonas syringae pv. tabaci infection. Plant Physiol. Biochem. 49, 1264–1271 (2011).
Ruan, S. L. et al. Proteomic identification of OsCYP2, a rice cyclophilin that confers salt tolerance in rice (Oryza sativa L.) seedlings when overexpressed. BMC Plant Biol. 11, 34 (2011).
Trivedi, D. K., Bhatt, H., Johri, A. K., Tuteja, N. & Bhavesh, N. S. Sequence-specific H, C and N NMR assignments of Cyclophilin A like protein from Piriformospora indica involved in salt stress tolerance. Biomol. NMR Assign.; 10.1007/s12104-012-9404-z (2012).
Seki, M., Kamei, A., Yamaguchi-Shinozaki, K. & Shinozaki, K. Molecular responses to drought, salinity and frost: common and different paths for plant protection. Curr. Opin. Biotechnol. 14, 194–199 (2003).
Urano, K., Kurihara, Y., Seki, M. & Shinozaki, K. ‘Omics’ analyses of regulatory networks in plant abiotic stress responses. Curr. Opin. Plant Biol. 13, 132–138 (2010).
Lorkovic, Z. J. Role of plant RNA-binding proteins in development, stress response and genome organization. Trends Plant Sci. 14, 229–236 (2009).
Gomez, J. et al. gene induced by the plant hormone abscisic acid in response to water stress encodes a glycine-rich protein. Nature 334, 262–264 (1988).
Vartanian, N., Marcotte, L. & Giraudat, J. Drought Rhizogenesis in Arabidopsis thaliana (Differential Responses of Hormonal Mutants). Plant Physiol. 104, 761–767 (1994).
Gendra, E., Moreno, A., Alba, M. M. & Pages, M. Interaction of the plant glycine-rich RNA-binding protein MA16 with a novel nucleolar DEAD-box RNA helicase protein from Zea mays. Plant J. 38, 875–886 (2004).
Aneeta, N., Mishra, N. S., Tuteja, N. & Sopory, S. K. Salinity and ABA induced up-regulation and light mediated modulation of mRNA encoding glycine rich RNA-binding protein from Sorghum bicolour. Biochem. Biophys. Res. Commun. 296, 1063–1068 (2002).
Bhatt, H. et al. Cloning, purification, crystallization and preliminary X-ray crystallographic analysis of a cyclophilin A-like protein from Piriformospora indica. Acta Crystallogr. F68, 709–712 (2012).
Grossoehme, N. E., Spuches, A. M. & Wilcox, D. E. Application of isothermal titration calorimetry in bioinorganic chemistry. J. Biol. Inorg. Chem. 15, 1183–1191 (2010).
Turnbull, W. B. & Daranas, A. H. On the value of c: can low affinity systems be studied by isothermal titration calorimetry. J. Am. Chem. Soc. 125, 14859–14866 (2003).
Gu, C. et al. Reference gene selection for quantitative real-time PCR in Chrysanthemum subjected to biotic and abiotic stress. Mol. Biotechnol. 49, 192–197 (2011).
Kim, S. R. & An, G. Rice chloroplast-localized heat shock protein 70, OsHsp70CP1, is essential for chloroplast development under high-temperature conditions. J. Plant Physiol. 170, 854–863 (2013).
Khurana, N., Chauhan, H. & Khurana, P. Wheat chloroplast targeted sHSP26 promoter confers heat and abiotic stress inducible expression in transgenic Arabidopsis plants. PLoS One 8, e54418 (2013).
Vashisht, A. A. & Tuteja, N. Cold stress-induced pea DNA helicase 47 is homologous to eIF4A and inhibited by DNA-interacting ligands. Arch. Biochem. Biophys. 440, 79–90 (2005).
Wang, W., Vinocur, B., Shoseyov, O. & Altman, A. Role of plant heat shock proteins and molecular chaperones in the abiotic stress response. Trends in Plant Sci. 9, 244–252 (2004).
Trivedi, D. K., Ansari, M. W. & Tuteja, N. Multiple abiotic stress responsive rice cyclophilin: (OsCYP-25) mediates a wide range of cellular responses. Comm. Intg. Biology 6(5), eLocation ID, e25260 (2013).
Brandts, J. F., Halvorson, H. R. & Brennan, M. Consideration of the possibility that the slow step in protein denaturation reactions is due to cis-trans isomerism of proline residues. Biochemistry 14, 4953–4963 (1975).
Ishikawa, Y. & Bachinger, H. P. An additional function of the rough Endoplasmic Reticulum protein complex Prolyl 3-Hydroxylase 1 Cartilage associated protein Cyclophilin B: The CXXXC motif reveals disulfide isomerase activity in vitro. J. Bio. Chem.; 10.1074/jbc.M113.498063 (2013 Sep 16).
Li, Z., Min, W. & Gou, J. Knockdown of cyclophilin A reverses paclitaxel resistance in human endometrial cancer cells via suppression of MAPK kinase pathways. Cancer Chem. and Pharmaco.; 10.1007/s00280-013-2285-8 (2013 Sep 14)
Doyle, V., Virji, S. & Crompton, M. Evidence that cyclophilin-A protects cells against oxidative stress. Biochem. J. 341, 127–132 (1999).
Navarette Santos, A., Korber, S., Kullertz, G., Fischer, G. & Fischer, B. Oxygen stress increases prolyl cis/trans isomerase activity and expression of cyclophilin 18 in rabbit blastocysts. Biol. Reprod. 62, 1–7 (2000).
Gong, Z. et al. A DEAD Box RNA helicase is essential for mRNA export and important for development and stress responses in Arabidopsiss. Plant Cell 17, 256–267 (2005).
Tuteja, N., Sahoo, R. K., Garg, B. & Tuteja, R. OsSUV3 dual helicase functions in salinity stress tolerance by maintaining photosynthesis and antioxidant machinery in rice (Oryza sativa L. cv. IR64). Plant J. 10.1111/tpj.12277 (2013 Jun 28).
Bannikova, O. et al. Identification of RNA targets for the nuclear multidomain cyclophilin atCyp59 and their effect on PPIase activity. Nucl. Acids Res. 41, 1783–1796 (2013).
Mi, H., Kops, O., Zimmermann, E., Jäschke, A. & Tropschug, M. A nuclear RNA-binding cyclophilin in human T cells. FEBS Letters 398, 201–205 (1996).
Jefferson, R. A. Assaying chimeric genes in plants: The GUS gene fusion system. Plant Mol. Biol. Rep. 5, 387–405 (1987).
Fan, L., Zheng, S. & Wang, X. Antisense suppression of phospholipase D alpha retards abscisic acid- and ethylene-promoted senescence of postharvest Arabidopsis leaves. Plant Cell 9, 2183–2196 (1997).
Lichtenthaler, H. K. Chlorophyll and carotenoids pigments of photosynthetic biomembranes. Methods Enzymol. 148, 350–366 (1987).
Murray, M. G. & Thompson, W. F. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 8, 4321–4325 (1980).
Pham, X. H., Reddy, M. K., Ehtesham, N. Z., Matta, B. & Tuteja, N. A DNA helicase from Pisum sativum is homologous to translation initiation factor and stimulates topoisomerase I activity. Plant J 24, 219–229 (2000).
Shankar, Pradhan, A. & Tuteja, R. Isolation and characterization of Plasmodium falciparum UAP56 homolog: evidence for the coupling of RNA binding and splicing activity by site-directed mutations. Arch. Biochem. Biophys. 478, 143–153 (2008).
Murashige, T. & Skoog, F. A revised medium for rapid growth and bioassays with tobacco cultures. Physiol. Plant 15, 473–479 (1962).
Garg, B., Jaiswal, J. P., Mishra, S., Tripathi, B. N. & Prasad, M. A comprehensive study on dehydration-induced antioxidative responses during germination of Indian bread wheat (Triticum aestivum L. em Thell) cultivars collected from different agro climatic zones. Physiol. Mol. Biol. Plants 18, 217–228 (2012).
Barrs, H. D. & Weatherly, P. E. A re-examination of the relative turgidity technique for estimating water deficits in leaves. Aust. J. Biol. Sci. 15, 413–428 (1962).
Leslie, A. G. W. Recent changes to the MOSFLM package for processing film and image plate data. Joint CCP4/ESF–EAMCB Newsletter on Protein Crystallography 26, 27–33 (1992).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Cryst. D67, 235–242 (2011).
Navaza, J. A. MoRe: an automated package for molecular replacement. Acta Cryst. A50, 157–163 (1994).
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystal. D58, 1948–1954 (2002).
Findeisen, M., Brand, T. & Berger, S. A 1H-NMR thermometer suitable for cryoprobes. Mag. Reson. Chem. 45, 175–178 (2006).
Kay, L. E., Torchia, D. A. & Bax, A. Backbone dynamics of proteins as studied by 15N inverse detected heteronuclear NMR Spectroscopy: Application to Staphylococcal Nuclease. Biochemistry 28, 8972–8979 (1989).
Skelton, N. J. et al. Practical Aspects of Two-Dimensional Proton-Detected 15N Spin Relaxation Measurements. J. Magn. Reson. B102, 253–264 (1993).
Farrow, N. A. et al. Backbone dynamics of a free and phosphopeptide-complexed Src homology 2 domain studied by 15N NMR relaxation. Biochemistry 33, 5984–6003 (1994).
Lipari, G. & Szabo, A. Model-free approach to the interpretation of nuclear magnetic resonance relaxation in macromolecules 1. Theory and range of validity. J. Am. Chem. Soc. 104, 4546–59 (1982).
Lipari, G. & Szabo, A. Model-free approach to the interpretation of nuclear magnetic resonance relaxation in macromolecules 2. Analysis of experimental results. J. Am. Chem. Soc. 104, 4559–4570 (1982).
Keller, R. The computer aided resonance assignment tutorial, 1st edn. CANTINA Verlag. ISBN 3-85600-112-113 (2004).
Finlayson, S. D., Fleming, C., Berry, D. R. & Johnston, J. R. An improved lithium acetate method for yeast transformation. Biotechnol. Tech. 5, 13–18 (1991).
Acknowledgements
This study is supported by grants to NSB and NT from Department of Science and Technology (DST), Government of India (GOI) and ICGEB core funds. X-ray diffraction data were collected at the National Institute of Immunology (NII), New Delhi. We thank DBT, GOI for providing financial support for the 500 MHz and 700 MHz NMR spectrometers at the ICGEB, New Delhi and NII, New Delhi. HB is a recipient of Council for Scientific and Industrial Research (CSIR) senior research fellowship. DKT is a recipient of DBT Ph.D. fellowship. We also thankful to Dipul Biswas for help in Plant tissue culture work.
Author information
Authors and Affiliations
Contributions
N.S.B. and N.T. designed research; D.K.T. and H.B. performed research; D.K.T., H.B., R.K.P., B.G., A.K.J., R.T., N.S.B. and N.T. analyzed data; and D.K.T., H.B., R.T., N.S.B. and N.T. wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Data deposition: The atomic coordinate and structure factors have been deposited in the Protein Data Bank, www.pdb.org [PDB ID 4EYV]. Sequence-specific NMR resonance assignments have been deposited in the BioMagResBank database, www.bmrb.wisc.edu [BMRB accession number 18312].
Electronic supplementary material
Supplementary Information
Supplementary file
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial- NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Trivedi, D., Bhatt, H., Pal, R. et al. Structure of RNA-interacting Cyclophilin A-like protein from Piriformospora indica that provides salinity-stress tolerance in plants. Sci Rep 3, 3001 (2013). https://doi.org/10.1038/srep03001
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep03001
This article is cited by
-
A high-throughput RNA-Seq approach to elucidate the transcriptional response of Piriformospora indica to high salt stress
Scientific Reports (2021)
-
Comparative transcriptome profiling of rice colonized with beneficial endophyte, Piriformospora indica, under high salinity environment
Molecular Biology Reports (2020)
-
The amazing potential of fungi: 50 ways we can exploit fungi industrially
Fungal Diversity (2019)
-
Transcriptional responses of soybean roots to colonization with the root endophytic fungus Piriformospora indica reveals altered phenylpropanoid and secondary metabolism
Scientific Reports (2018)
-
Genome of Paulownia (Paulownia fortunei) illuminates the related transcripts, miRNA and proteins for salt resistance
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.