Abstract
Most past studies of the biophysical regulation of stem cell differentiation have focused on initial lineage commitment or proximal differentiation events. It would be valuable to understand whether biophysical inputs also influence distal endpoints more closely associated with physiological function, such as subtype specification in neuronal differentiation. To explore this question, we cultured adult neural stem cells (NSCs) on variable stiffness ECMs under conditions that promote neuronal fate commitment for extended time periods to allow neuronal subtype differentiation. We find that ECM stiffness does not modulate the expression of NeuroD1 and TrkA/B/C or the percentages of pan-neuronal, GABAergic, or glutamatergic neuronal subtypes. Interestingly, however, an ECM stiffness of 700 Pa maximizes expression of pan-neuronal markers. These results suggest that a wide range of stiffnesses fully permit pan-neuronal NSC differentiation, that an intermediate stiffness optimizes expression of pan-neuronal genes and that stiffness does not impact commitment to particular neuronal subtypes.
Similar content being viewed by others
Introduction
Stem cells hold great promise for tissue engineering and regenerative medicine due to their ability to differentiate into specialized cell types that may be lost due to disease or injury. Therefore, it is important to understand the microenvironmental cues that regulate stem cell lineage commitment and differentiation. Substrate stiffness has been shown to modulate the differentiation choices of stem cells1,2. For example, adult neural stem cells (NSCs) of the hippocampus can give rise to all three neural cell types – neurons, astrocytes and oligodendrocytes –and we have previously shown that this lineage commitment decision is mechanosensitive, such that NSCs generate more neurons when cultured on soft substrates (<1 kPa) and more astrocytes on hard substrates2,3. Furthermore, exposure of human pluripotent stem cells to soft ECMs early during differentiation increases neuroectodermal specification and the subsequent percentages of neurons obtained4. However, to date, our and others' studies of how mechanical cues regulate stem cell differentiation have largely focused on the initial lineage commitment of multipotent stem cells, such as NSCs committing to any type of neuron or “pan-neuronal” cell and not downstream maturation or subtype specification processes critical to physiological function. While the percentages of pan-neuronal cells may not change with substrate stiffness, pan-neuronal gene expression may. Furthermore, upwards of 100 neuronal subtypes have been identified to date5 and the hippocampus in particular is populated predominantly with GABAergic and glutamatergic neurons6. Because neuronal subtype specification indicates the neurotransmitter(s) a neuron secretes and this in turn dictates synaptic specificity, neuronal maturation leading to subtype specification is crucial for proper neural network function and connectivity. Thus, understanding whether mechanical properties influence pan-neuronal maturation and neuronal subtype differentiation is important for our understanding of organismal and central nervous system development, as well as the generation of therapeutic neuronal subtypes.
Here we study the effect of substrate stiffness on pan-neuronal differentiation and neuronal subtype specification by extending the differentiation time and supplying neuronal maturation factors. Under conditions that drive pan-neuronal differentiation, we find that all substrate stiffnesses permit and do not bias pan-neuronal lineage commitment, as evidenced by similar NeuroD1 gene expression levels and similar percentages of pan-neuronal, TUJ1+ cells. However, an intermediate substrate stiffness close to that of native brain tissue (~700 Pa) increases gene expression of TUJ1 and of pan-neuronal maturation genes P75-NGFR and p21, indicating an optimal stiffness for pan-neuronal maturation. Furthermore, we observe no differences in the percentages of GABA and VGlut1-immunostained neurons and in gene expression of neurotrophic factors TrkA, TrkB and TrkC, known to modulate subtype specification, indicating neuronal subtype specification is not affected by substrate stiffness. In light of recent work demonstrating the ability of stiffness to bias lineage commitment to neurons over other cell types2,3,4,7,8, this work suggests that altering microenvironmental stiffness may remain compatible with the production of many neuronal subtypes and at intermediate stiffness perhaps even promote downstream neuronal maturation and thus contribute to future biomedical applications.
Results
Pan-neuronal TUJ1 gene expression but not pan-neuron generation is optimal on 700 Pa substrates
Our previous work showed that substrate stiffness could instruct NSC lineage commitment toward neurons on soft substrates and astrocytes on stiff substrates in “mixed media conditions” that promote differentiation into a combination of neurons and glia2,3. We therefore asked whether pan-neuronal lineage commitment and subsequent neuronal subtype lineage specification are also regulated by substrate stiffness under conditions that exclusively induce differentiation into neurons. NSCs were cultured on laminin-coated polyacrylamide hydrogels for 6 days in medium that promotes neuronal differentiation (1 μM all-trans retinoic acid, 5 μM forskolin, no serum) adapted from published studies9,10. Gene expression of TUJ1, a pan-neuronal cytoskeletal marker, was found to be optimal on 700 Pa substrates, about 5 fold higher than on 100 and 75000 Pa substrates (Figure 1A). However, gene expression is a population-average measurement and higher neuronal marker gene expression could result from greater neuronal maturation in some cells rather than differences in the actual percentage of neurons generated. Therefore, 100, 700 and 75000 Pa substrates were chosen for their demonstrated differences in TUJ1 gene expression and after 6 days in neuronal differentiation conditions, NSCs were fixed and immunostained against TUJ1. The percentage of TUJ1 positive cells was approximately 40% on all substrate stiffnesses (Figure 1B–C). Thus, the peak in TUJ1 expression on 700 Pa substrates is due to increased TUJ1 expression per cell rather than an increase in the number of TUJ1-positive cells.
Neuronal gene expression but not the generation of neurons is optimal on an intermediate stiffness near that of brain tissue.
Neural stem cells were cultured on polyacrylamide gels of different stiffnesses for 6 days. (A) TUJ1 gene expression normalized to ribosomal 18S, measured by quantitative reverse transcription polymerase chain reaction, is highest near 700 Pa. (B) The percentage of TUJ1+ neurons is invariant with substrate stiffness. (C) Immunofluorescence images corresponding to (B). TUJ1 is red, DAPI is blue. Error bars are 95% confidence intervals, n = 3. Means of multiple groups were compared by analysis of variance, Tukey-Kramer post hoc (ANOVA-TK), points with distinct group letters are statistically different with p < 0.05.
Neuronal subtype specification is not sensitive to substrate stiffness
As described, substrate stiffness does not affect the percentage of TUJ1 positive neurons after 6 days of differentiation (Figure 1B). However, given the differences observed in TUJ1 gene expression, it is possible that while initial pan-neuronal lineage commitment is not influenced by the substrate, subsequent neuronal subtype specification, marked by the expression of neurotransmitters, is affected. In the hippocampus, neurons predominantly express inhibitory γ-aminobutyric acid (GABA) or excitatory glutamate neurotransmitters. We therefore extended the differentiation length to 11 days, when subtype specification is largely complete10. After the first 6 days of differentiation, in addition to retinoic acid and forskolin, 20 ng/mL BDNF was added as a neurotrophin/maturation signal10. Cells were then immunostained for TUJ1, GABA and the vesicular glutamate transporter 1 (VGlut1, a marker of glutamatergic neurons). We did not observe tyrosine hydroxylase (dopaminergic) or choline acetyltransferase (cholinergic) positive cells (data not shown). The percentages of TUJ1 positive cells on day 11 remained similar to day 6, approximately 40%, with no statistically significant variation as a function of substrate stiffness (Figure 2). Consistent with previous work10, the percentages of GABAergic and glutamatergic neurons were relatively low (<5%). Importantly, however, these levels were also insensitive to substrate stiffness, suggesting the observed increase in TUJ1 gene expression at day 6 on 700 Pa substrates does not lead to or correlate with changes in subtype specification.
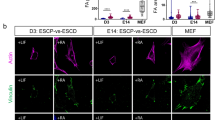
Neuronal subtype specification is not sensitive to substrate stiffness.
Neural stem cells were differentiated for 11 days on polyacrylamide gels of different stiffnesses and immunostained for TUJ1 (green), GABA (red), or VGlut1 (red), which are markers of neurons, GABAergic neurons, or glutamatergic neurons, respectively. DAPI is shown as blue. Inset images are outlined in white. Error bars are 95% confidence intervals, n = 6. Means compared by analysis of variance, Tukey-Kramer post hoc (ANOVA-TK), p > 0.05 for all comparisons.
Time-course of neurotrophic gene expression reveals both mechano-sensitive and -insensitive genes
While the percentages of neurons and neuronal subtypes do not depend on substrate stiffness, TUJ1 gene expression is optimal on 700 Pa substrates (Fig. 1A). To gain further molecular insight into these observations, we performed a time-course of QRT-PCR measurements of neurotrophic genes known to be responsible for different aspects of neuronal differentiation and lineage commitment.
Because similar percentages of TUJ1 positive neurons were observed across all substrate stiffnesses (Figures 1B and 2), we measured the expression of NeuroD1, a key transcription factor that drives neuronal lineage commitment10,11 and whose expression peaks sharply 1 day after induction of neuronal differentiation and then declines10. Given that NeuroD1 expression temporally precedes and may causally drive neuronal lineage commitment and that we do not observe changes in lineage commitment with substrate stiffness, we expected that NeuroD1 expression would also be mechano-insensitive. Indeed, our measurements of NeuroD1 gene expression at day 1 revealed that its expression was similar on all substrate stiffnesses. This supports our observation that neuronal lineage commitment in unipotent (pro-neurogenic) medium conditions is not affected by substrate stiffness.
Analogously, to further investigate the observation that the percentages of GABAergic and glutamatergic neurons were also invariant with substrate stiffness, we asked whether the expression of neurotrophin receptors that transduce maturation and subtype specification signals were also mechano-insensitive. The tyrosine kinase neurotrophin receptors TrkA, B and C, which are responsive to neurotrophic factors including BDNF, initiate expression within the first day of neuronal differentiation and maintain detectable levels throughout neuronal subtype specification10. Therefore, TrkA/B/C mRNA expression was quantified by QRT-PCR after 1, 6 and 11 days of differentiation. No differences in expression were observed as a function of substrate stiffness after 1 day (Figure 3). Expression of TrkA/B/C remained invariant with substrate stiffness through day 6 and 11, with a slight decrease in TrkC expression on 100 Pa relative to harder substrates at day 11. c-fos, an immediate early gene responsive to BDNF via TrkA/B/C signaling, was also expressed at similar levels with respect to substrate stiffness, consistent with TrkA/B/C expression.
Only specific neuronal genes are optimally expressed on intermediate substrate stiffnesses.
Neural stem cells were cultured on polyacrylamide gels of different stiffnesses for the indicated number of days. Gene expression, normalized to ribosomal 18S, was measured by quantitative reverse transcription polymerase chain reaction. Error bars are 95% confidence intervals, n = 3. Means compared by analysis of variance, Tukey-Kramer post hoc (ANOVA-TK), comparison bars represent p < 0.05. Note log-scale y-axes.
While NeuroD1 and TrkA/B/C gene expression were invariant with substrate stiffness, reflecting the similar percentages of neurons and neuronal subtypes on all stiffnesses, TUJ1 gene expression was optimal on 700 Pa substrates (Figure 1A). This finding suggests substrate stiffness exerts an effect on neuronal maturation while not affecting pan-neuronal and neuronal subtype lineage commitment. We thus asked whether the TUJ1 differences observed at day 6 (Figure 1A) were due to faster neuronal differentiation kinetics at 700 Pa, or if this substrate stiffness increases neuronal gene expression in the long term. After 11 days, TUJ1 expression remained optimal on 700 Pa substrates, suggesting this effect may not simply be an isolated observation at a single time point, nor an acceleration of neuronal maturation, but is rather a persistent effect that results in higher long-term TUJ1 levels. We next asked if this effect is isolated to this gene or is a more general effect involving multiple genes related to neuronal maturation but not necessarily subtype commitment decisions. One such gene may be the neurotrophin receptor p75-NGFR, which unlike TrkA/B/C was previously shown not to mediate BDNF (nor neurotrophic factor 3 or NGF) effects on c-fos upregulation or neuronal maturation in hippocampal aNSCs10. While it is thus not yet clear what role p75-NGFR plays in aNSC neuronal differentiation, the p75-NGFR ligand NGF has been shown to induce p21 activity to promote exit from the cell cycle12. Since halting mitosis is an inherent step in neuronal maturation, we investigated both of these genes.
Interestingly, p75NGFR was strongly upregulated on 700 Pa substrates after 1 and 11 days of differentiation. Consistent with this finding and with the dynamics of TUJ1 gene expression, p21, a cyclin-dependent kinase inhibitor whose expression typically signifies cell cycle exit13,14,15 and correlates with the generation of post-mitotic neurons16, was also upregulated on 700 Pa substrates at day 6 and 11. As a whole, this expression analysis of neuronal genes supports a model in which substrate stiffness is permissive of pan-neuronal as well as neuronal subtype lineage commitment but does not modulate their percentages, while intermediate stiffnesses promote pan-neuronal maturation.
Discussion
This work shows that an intermediate substrate stiffness (700 Pa) promotes long-term pan-neuronal gene expression without affecting neuronal lineage commitment or neuronal subtype specification. Consistent with these results, NeuroD1 and TrkA/B/C gene expression, which transduce retinoic acid and BDNF signals to affect lineage and subtype commitment, are not altered at early, intermediate and late time points during neuronal differentiation, while expression of the pan-neuronal genes TUJ1, p75-NGFR and p21 are optimally expressed on 700 Pa substrates. Finally, all substrate stiffnesses (100, 700 and 75000 Pa) allow the generation of both GABAergic and glutamatergic hippocampal subtypes.
We had previously found that under conditions permissive to NSC differentiation into astrocytes, neurons and oligodendrocytes, decreasing substrate stiffness increased the percentage of neurons while decreasing the percentage of astrocytes generated2,3. However, when soluble conditions were changed to induce primarily neuronal differentiation as in this study (i.e. unipotent conditions), neither the percentage of total neurons nor neuronal subtype specification was sensitive to substrate stiffness. However, substrate stiffness did interestingly exert a biphasic effect on pan-neuronal expression of genes such as TUJ1, p75-NGFR and p21, with intermediate stiffnesses near 700 Pa promoting optimal expression. These results indicate the existence of multiple, distinct layers of control over stem cell differentiation, in which three stages of this process – multipotent lineage commitment, pan-neuronal lineage commitment and subsequent subtype specification and the strength of neuronal differentiation indicated by pan-neuronal gene expression – are each governed by distinct sets of biophysical rules. This type of stage-specific regulation may be present in other stem cell systems. Previous studies of mechano-regulation of differentiation of other stem cell types1,17,18,19 have not experimentally distinguished analogous stages of stem cell differentiation and it may be interesting to assess whether observed effects on stem cell differentiation in these cases are primarily due to initial lineage commitment decisions, differentiation processes along a given lineage, or a combination of both. For example, mesenchymal stem cells (MSCs) – which are derived from many locations including bone marrow, umbilical cord blood, fat and muscle – may yield different subtypes of specialized cells, including brown vs. white adipocytes or osteoblasts vs. osteocytes. One study hints at this type of regulation in MSCs, finding that two cell lines derived from different stages of mesenchymal development responded disparately to matrix signals20. In our study, biophysical regulation of neuronal differentiation from a single starting NSC population appears to modulate pan-neuronal maturation but not subtype lineage commitment decisions. Future work may identify differences in connectivity of signaling pathways regulating lineage commitment and neuronal maturation to mechanosensing mechanisms.
Despite differences in the mechanoregulation of NSCs in mixed differentiation (astrocytes, neurons, oligodendrocytes) vs. unipotent neurogenic conditions, 700 Pa is in both cases a key value for stiffness-based NSC regulation. In the mixed differentiation case, NSCs dramatically switch between low to high neuronal lineage commitment at 700 Pa2. In the neuronal differentiation case, optimal TUJ1, p75-NGFR and p21 gene expression is observed also at 700 Pa. Common mechanotransductive mechanisms or cytoskeletal properties of neural cell types may sensitize cells specifically at this stiffness range, which is near that of their native brain tissue microenvironment21,22,23.
Finally, while NSC neuronal lineage commitment and subtype specification seem largely mechano-insensitive, TUJ1, P75-NGFR and p21 gene expression were upregulated on 700 Pa substrates. p75-NGFR upregulation on day 1 may induce p21 expression through autocrine or paracrine NGF signaling12. However, how p75-NGFR may affect neuronal differentiation remains an open question in the field. NGF alone does not promote neuronal maturation, c-fos expression, or subtype specification10. Thus it may serve a more complex function. For example, p75-NGFR has low affinity for NGF relative to TrkA/B/C and thus it has been hypothesized that p75-NGFR acts by increasing the local concentration of NGF to increase Trk signaling24 rather than by directly inducing signaling itself. Additionally, the presence of other less-well characterized ligands of p75-NGFR may be absent in this system and exert substrate-stiffness dependent effects on neuronal differentiation25,26,27. If biological roles for these ligands are identified in the future, additional work can assess their ability to mediate mechanosensitive behavior.
These observations extend our understanding of NSC mechanotransduction and illustrate the complexity of how cells integrate both biochemical and biophysical microenvironmental signals to produce distinct cellular behaviors. Furthermore, a wide range of microenvironmental stiffnesses is permissive of lineage commitment into neuronal subtypes and stiffness may in fact promote neuronal maturation at 700 Pa, near brain tissue stiffness. These results motivate the consideration of biophysical parameters in future tissue engineering and regenerative medicine applications. For example, dynamically tuning tissue scaffold stiffness from very soft (100 Pa), during multipotent lineage commitment to boost neuronal lineage commitment, to intermediate (700 Pa), during neuronal maturation to boost neuronal gene expression, may yield the greatest percentage but also developmentally most mature population of neurons without inhibiting neuronal subtype specification. Additionally, future work may focus on identifying common and distinct nodes at which biochemical pathways intersect biophysical transduction networks as well as elucidate mechanisms by which cells regulate their sensitivity to specific microenvironmental stiffness ranges.
Methods
Stem cell culture
Neural stem cells isolated from the hippocampi of adult female Fischer 344 rats were cultured as described previously28. Specifically, standard cultures were grown on tissue culture polystyrene coated with 10 μg/ml poly-ornithine (Sigma, St. Louis, MO) and 5 μg/ml mouse laminin (Life Technologies, Carlsbad, CA), in Dulbecco's modified Eagle medium (DMEM)/F-12 (1:1, Life Technologies) with N2 supplement (Life Technologies) and 20 ng/ml recombinant human basic fibroblast growth factor (Peprotech, Rocky Hill, NJ). Glass substrates were coated with 20 μg/ml poly-ornithine and 10 μg/ml mouse laminin. Differentiation was induced on day 0 in DMEM:F12 with N2 supplement by the addition of 1 μM retinoic acid (Enzo, Farmingdale, NY) and 5 μM forskolin (Enzo). Half-media changes were performed daily. On day 6, 20 ng/mL brain-derived neurotrophic factor (BDNF, Peprotech) was added to promote neuronal maturation. Cells were fixed and stained or isolated in Trizol (Life Technologies) on day 11.
Immunofluorescence staining
Cells were fixed with 4% paraformaldehyde for 10 minutes, then blocked and permeabilized in 2% goat serum (Sigma) and 0.3% Triton X-100 (Calbiochem, San Diego, CA) in pH 7.4 phosphate buffered solution at room temperature. Samples were incubated for 36 hours at 4°C with the following primary antibodies: rabbit anti-tyrosine hydroxylase (1:1000, Pel-Freez, Rogers, AR), rabbit anti-choline acetyltransferase (1:1000, Abcam, Cambridge, MA), rabbit anti-VGlut1 (1:2500, Synaptic Systems, Goettingen, Germany), mouse anti-TUJ1 (1:1000, Sigma), rabbit anti-GABA (1:2500, Sigma). The primary antibody solution was removed and cells were rinsed three times with phosphate buffered saline and incubated for 2 hours with the following secondary antibodies: FITC-conjugated goat anti-rabbit IgG and Cy5-conjugated goat anti-mouse IgG (both 1:250, from Jackson ImmunoResearch Laboratories, West Grove, PA). Nuclei were stained with DAPI (Life Technologies) at 10 μg/ml. 8-bit images were collected through NIS Elements software at 10× and 20× magnification on a Nikon Eclipse TE2000-E microscope with a Photometrics Coolsnap HQ2 camera and exposure settings were chosen to minimize background fluorescence as determined using control samples without primary antibodies and were uniform across conditions. Image processing (combining channels and linear color level adjustments) in Adobe Photoshop were applied identically to all conditions. Filter cubes used were Nikon C-FL FITC (480/40ex 535/50em), Cy3 (545/30ex 610/75em) and Cy5(620/60ex 700/75em) HYQ. Cells were manually scored positive or negative for lineage markers in regularly spaced and rastered fields with conditions blinded to the counter to avoid bias. ~500 cells were counted per sample with n = 6 samples.
Polyacrylamide substrate preparation
Using a protocol similar to that described previously29, polyacrylamide gels (70 μm nominal thickness) were synthesized on 12 mm glass coverslips using solutions composed of varying concentrations of acrylamide monomer and bisacrylamide crosslinker2. 100 μg/ml laminin was linked to the surface through N-Sulfosuccinimidyl-6-(4′-azido-2′-nitrophenylamino) hexanoate (sulfo-SANPAH, Thermo-Fisher, Waltham, MA) chemistry.
Quantitative reverse transcription polymerase chain reaction
Quantitative reverse transcription PCR (QRT-PCR) was used to accurately quantify specific mRNA concentrations in cells. Cells were lysed and frozen in TRIZOL (Life Technologies) and mRNA was extracted and reverse transcribed to cDNA using the ThermoScript™ RT-PCR System for First-Strand cDNA Synthesis (Life Technologies). Equivalent amounts of total RNA were transcribed into cDNA, which was subsequently used as template for each QRT-PCR reaction (Bio-Rad Laboratories iCycler 5, Hercules, CA). To normalize any remaining variations in starting cDNA amounts, each reaction was carried out in duplicate wells with ribosomal 18S detected using Cal-dye TaqMan probes (Biosearch Technologies, Novato, CA) in one well and the gene of interest detected in the other well using SYBR Green (Life Technologies). The delta cycle-threshold method was used to calculate relative mRNA quantities. QRT-PCR reactions were run for each biological sample with n = 3–6 for each condition.
Primer sequences were: rat trkA 5′ (CACTGGGTGGCAGTTCTCTT), rat trkA 3′ (CATGTACTCGAAGACCATGA), rat trkB 5′ (GTTGCTGACCAAACCAATCG), rat trkB 3′ (CATGTACTCAAAGACCATGA), rat trkC 5′ (CTGAAGGATCCCACCTTGGC), rat trkC 3′ (CATGTATTCAAAGACCATGA), rat p75NGFR 5′ (CCCTGCCTGGACAATGTTAC), rat p75NGFR 3′ (CTGGGCACTCTTCACAC), rat p21 5′ (GCCCAAGATCTACCTGAG), rat p21 3′ (GTGGGCACTTCAGGGCTTTC), rat c-fos 5′ (GGGTTTCAACGCGGACTAC), rat c-fos 3′ (GTTGGCACTAGAGACGGA), mouse NeuroD1 5′ (ATGACCAAATCATACAGCGAGAG), mouse NeuroD1 3′ (TCTGCCTCGTGTTCCTCGT).
References
Engler, A. J., Sen, S., Sweeney, H. L. & Discher, D. E. Matrix elasticity directs stem cell lineage specification. Cell 126, 677–689 (2006).
Keung, A. J., de Juan-Pardo, E. M., Schaffer, D. V. & Kumar, S. Rho GTPases mediate the mechanosensitive lineage commitment of neural stem cells. Stem Cells 29, 1886–1897 (2011).
Saha, K. et al. Substrate modulus directs neural stem cell behavior. Biophys J 95, 4426–4438 (2008).
Keung, A. J., Asuri, P., Kumar, S. & Schaffer, D. V. Soft microenvironments promote the early neurogenic differentiation but not self-renewal of human pluripotent stem cells. Integr Biol (Camb) 4, 1049–1058 (2012).
Heiman, M. et al. A translational profiling approach for the molecular characterization of CNS cell types. Cell 135, 738–748 (2008).
Klausberger, T. & Somogyi, P. Neuronal diversity and temporal dynamics: the unity of hippocampal circuit operations. Science 321, 53–57 (2008).
Leipzig, N. D. & Shoichet, M. S. The effect of substrate stiffness on adult neural stem cell behavior. Biomaterials 30, 6867–6878 (2009).
Teixeira, A. I. et al. The promotion of neuronal maturation on soft substrates. Biomaterials 30, 4567–4572 (2009).
Hsieh, J. et al. IGF-I instructs multipotent adult neural progenitor cells to become oligodendrocytes. J Cell Biol 164, 111–122 (2004).
Takahashi, J., Palmer, T. D. & Gage, F. H. Retinoic acid and neurotrophins collaborate to regulate neurogenesis in adult-derived neural stem cell cultures. J Neurobiol 38, 65–81 (1999).
Lee, J. E. et al. Conversion of Xenopus Ectoderm into Neurons by NeuroD, a Basic Helix-Loop-Helix Protein. Science 268, 836–844 (1995).
Yan, G. Z. & Ziff, E. B. NGF regulates the PC12 cell cycle machinery through specific inhibition of the Cdk kinases and induction of cyclin D1. J Neurosci 15, 6200–6212 (1995).
el-Deiry, W. S. et al. WAF1, a potential mediator of p53 tumor suppression. Cell 75, 817–825 (1993).
Harper, J. W., Adami, G. R., Wei, N., Keyomarsi, K. & Elledge, S. J. The p21 Cdk-interacting protein Cip1 is a potent inhibitor of G1 cyclin-dependent kinases. Cell 75, 805–816 (1993).
Polyak, K. et al. Cloning of p27Kip1, a cyclin-dependent kinase inhibitor and a potential mediator of extracellular antimitogenic signals. Cell 78, 59–66 (1994).
Dobashi, Y., Kudoh, T., Matsumine, A., Toyoshima, K. & Akiyama, T. Constitutive overexpression of CDK2 inhibits neuronal differentiation of rat pheochromocytoma PC12 cells. J Biol Chem 270, 23031–23037 (1995).
Adamo, L. et al. Biomechanical forces promote embryonic haematopoiesis. Nature 459, 1131–1135 (2009).
Chowdhury, F. et al. Material properties of the cell dictate stress-induced spreading and differentiation in embryonic stem cells. Nat Mater 9, 82–88 (2010).
Dalby, M. J. et al. The control of human mesenchymal cell differentiation using nanoscale symmetry and disorder. Nat Mater 6, 997–1003 (2007).
Hsiong, S. X., Carampin, P., Kong, H. J., Lee, K. Y. & Mooney, D. J. Differentiation stage alters matrix control of stem cells. J Biomed Mater Res A 85, 145–156 (2008).
Gefen, A., Gefen, N., Zhu, Q., Raghupathi, R. & Margulies, S. S. Age-dependent changes in material properties of the brain and braincase of the rat. J Neurotrauma 20, 1163–1177 (2003).
Sack, I. et al. The impact of aging and gender on brain viscoelasticity. NeuroImage 46, 652–657 (2009).
Unsgaard, G. et al. Intra-operative 3D ultrasound in neurosurgery. Acta Neurochir (Wien) 148, 235–253 (2006).
Barker, P. A. & Shooter, E. M. Disruption of NGF binding to the low affinity neurotrophin receptor p75LNTR reduces NGF binding to TrkA on PC12 cells. Neuron 13, 203–215 (1994).
Higuchi, H., Yamashita, T., Yoshikawa, H. & Tohyama, M. PKA phosphorylates the p75 receptor and regulates its localization to lipid rafts. EMBO J 22, 1790–1800 (2003).
Tcherpakov, M. et al. The p75 neurotrophin receptor interacts with multiple MAGE proteins. J Biol Chem 277, 49101–49104 (2002).
Ye, X. et al. TRAF family proteins interact with the common neurotrophin receptor and modulate apoptosis induction. J Biol Chem 274, 30202–30208 (1999).
Palmer, T. D., Markakis, E. A., Willhoite, A. R., Safar, F. & Gage, F. H. Fibroblast growth factor-2 activates a latent neurogenic program in neural stem cells from diverse regions of the adult CNS. J Neurosci 19, 8487–8497 (1999).
Pelham, R. J., Jr & Wang, Y. Cell locomotion and focal adhesions are regulated by substrate flexibility. Proc Natl Acad Sci U S A 94, 13661–13665 (1997).
Acknowledgements
This work was supported by a National Defense Science and Engineering Graduate Fellowship and a National Science Foundation Graduate Research Fellowship to A. J. Keung. S. Kumar wishes to acknowledge the support of a UC Berkeley Stem Cell Center Seed Grant, the Arnold and Mabel Beckman Young Investigator Award, a PECASE Award from the Army Research Office (W911NF-09-1-0507) and the NIH Director's New Innovator Award (1DP2OD004213), a part of the NIH Roadmap for Medical Research. D. V. Schaffer and S. Kumar wish to acknowledge the support of NIH 1R01NS074831.
Author information
Authors and Affiliations
Contributions
A.J.K.: conception and design, collection and/or assembly of data, data analysis and interpretation and manuscript writing; M.D.: conception and design, collection and/or assembly of data and data analysis and interpretation; D.V.S. and S.K.: financial support, conception and design, data analysis and interpretation, manuscript writing and final approval of manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Keung, A., Dong, M., Schaffer, D. et al. Pan-neuronal maturation but not neuronal subtype differentiation of adult neural stem cells is mechanosensitive. Sci Rep 3, 1817 (2013). https://doi.org/10.1038/srep01817
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep01817
This article is cited by
-
Mechanical properties of murine hippocampal subregions investigated by atomic force microscopy and in vivo magnetic resonance elastography
Scientific Reports (2022)
-
Recent Advancements in Engineering Strategies for Manipulating Neural Stem Cell Behavior
Current Tissue Microenvironment Reports (2020)
-
Titanium dioxide nanoparticles via oral exposure leads to adverse disturbance of gut microecology and locomotor activity in adult mice
Archives of Toxicology (2020)
-
Mechanotransduction in neuronal cell development and functioning
Biophysical Reviews (2019)
-
Ex vivo biomechanical characterization of syringe-needle ejections for intracerebral cell delivery
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.