Abstract
Crop genetic engineering mostly aims at improving environmental stress (biotic and abiotic) tolerance as well as nutritional quality. Empowering a single crop with multiple traits is highly demanding and requires manipulation of more than one gene. However, we report improved drought tolerance and fungal resistance along with the increased iron and polyunsaturated fatty acid content in tomato by expressing a single gene encoding C-5 sterol desaturase (FvC5SD) from an edible fungus Flammulina velutipes. FvC5SD is an iron binding protein involved in ergosterol biosynthesis. Morphological and biochemical analyses indicated ≈23% more epicuticular wax deposition in leaves of transgenic plants that provides an effective waterproof barrier resulting in improved protection from drought and infection by phytopathogenic fungus Sclerotinia sclerotiorum. Furthermore, the transgenic fruits have improved nutritional value attributed to enhanced level of beneficial PUFA and 2-3 fold increase in total iron content. This strategy can be extended to other economically important crops.
Similar content being viewed by others
Introduction
In recent years, development of the transgenic crops with multiple beneficial traits has emerged as an important area in the field of plant biotechnology. The major objectives of genetic engineering of crops include introduction of traits like improved tolerance towards various biotic and abiotic stresses and enhanced nutritional value. Among all the environmental factors limiting crop yield, drought probably is the abiotic stress with the highest impact1. Besides, biotic stress imposed on crops by plant pathogens are a real threat to worldwide agricul-ture. Phytopathogens like S. sclerotiorum cause substantial loss in crop yield each year throughout the world. This fungus has a broad host range resulting in about 95% loss of economically important crops like oil seedrape, bean, tomato and sunflower. Thus, the transgenic strategies that lead to improved tolerance to drought and phytopathogens can significantly contribute to improvement in crop yield.
Plants are the ultimate source of nutrients in human diet. However, all our major food crops lack certain essential nutrients. Malnutrition is a significant public health issue in most of the developing countries where majority of population relies on staple crops, such as rice or wheat, which does not provide the full complement of essential nutrients. For example, iron deficiency is the most widespread micronutrient deficiency, affecting about 2 billion people. Also the essential ω-3 and ω-6 polyunsaturated fatty acids (PUFA) which are known to have diverse roles in metabolism, cardiovascular health, inflammatory responses, blood pressure regulation, etc. cannot be synthesised de-novo and must be supplied in the diet. Thus, it is conceivable that one of the major goals of genetic engineering is to improve essential nutrient content in the staple crops. This includes enhancing the level of bio-available iron and polyunsaturated fatty acids (PUFA).
The epicuticular waxes which form the outermost layer of aerial plant organs are considered to confer resistance to insect herbivory, fungal pathogens and drought2,3,4. The cuticular waxes are complex mixtures of primarily very long chain fatty acids (VLC, > C18), hydrocarbons, alcohols, aldehydes, ketones, triterpenes, sterols and flavonoids5,6. Modification of cuticular wax layer is one of the strategies to improve drought tolerance by reducing transpirational water loss. Earlier, it has been achieved through the over expression of some genes for transcription factors as well as genes involved in wax biosynthesis7,8,9,10,11,12.
Wax biosynthesis genes like CER1 homolog have been shown to be upregulated in transcript profiling of drought responsive genes in tomato13. A feature common to some of the enzymes involved in epicuticular wax biosynthesis ( including OsGL1-2 from rice11, CER1 and WAX2 from Arabidopsis14,15, TaCer1 from wheat16 and GL1 from maize17) is the presence of N-terminal region exhibiting homology to the C-5 sterol desaturase (FA hydroxylase superfamily) besides an uncharacterized wax-C superfamily domain. FA hydroxylase superfamily consists of a large family of integral membrane enzymes such as fatty acyl desaturases, hydroxylases, ketolases, decarbonylases and monooxygenases found in prokaryotes and eukaryotes18 characterized by the conserved histidine rich motifs that form di-iron-binding site essential for catalytic activity. C-5 sterol desaturase (ERG3) is a membrane bound enzyme that catalyzes the introduction of a C-5 double bond into the B ring of Δ7-sterols to produce the corresponding Δ5, 7-sterols. It has been cloned and characterized in many organisms including Saccharomyces cerevisiae19, Arabidopsis thaliana20, Homo sapiens21,22, Candida albicans23 and most recently in alga Chlamydomonas reinhardtii24.
Introduction of multiple beneficial traits in a single crop can be achieved by stacking of more than one gene either by genetic engineering or conventional cross-breeding of genetically modified plants with two different modifications. However, this strategy is time consuming and labor intensive. In this report, we have achieved desirable traits like drought tolerance, resistance to phytopathogenic fungus S. sclerotiorum alongwith enhanced iron and PUFA content by expressing a single gene coding for Δ7-sterol-C5 (6) desaturase (FvC5SD) from edible fungus Flammulina velutipes. The study appears to be of remarkable interest since enhanced wax content has been achieved by expressing a sterol biosynthesis enzyme which is not reported to be involved in the pathway of wax biosyntheis.
Results
Identification and cloning of putative C-5 sterol desaturase (FvC5SD) from Flammulina velutipes
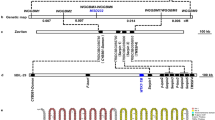
Azam et al isolated a genomic DNA clone from λgt11 genomic library of edible fungus F. velutipes that show a significant homology with C-5 sterol desaturase gene (ERG3) and was named FvC5SD25. Using the sequence information, full length cDNA of FvC5SD was amplified by RT- PCR (reverse transcription PCR) and cloned into T/A cloning vector. The sequencing data revealed an open reading frame (ORF) of 891bp and the deduced amino acid sequence of 296 amino acids ( Fig. S1 ), with a predicted molecular weight of 34.8 kDa. The FvC5SD gene consists of three exons interrupted by two introns.
Multiple alignment of FvC5SD with C-5 sterol desaturase from yeasts, filamentous fungi, plants and mammals ( Fig. 1 ) showed conservation of three histidine rich motifs (HXXXXH, HXXHH and HXXHH). Phylogenetic analysis was performed to establish the evolutionary relationship between C-5 sterol desaturase from various kingdoms ( Fig. S2 ). FvC5SD shows 42% identity to Erg3p of S. cerevisiae, 50% to that of S. pombe, 32% to C. reinhardtii, 34% to A. thaliana and 48% to human C-5 sterol desaturase. FvC5SD was most closely related to sterol desaturase of fungus A. fumigatus (54%) and C. neoformans (53% identity).
C-5 sterol desaturase is conserved across the kingdoms.
Multiple sequence alignment of the deduced amino acid sequence of FvC5SD with C-5 sterol desaturase from other organisms. The amino acid sequences of F. velutipes (JN696291)), Homo sapiens (BAA33729), Mus musculas (O88822) Arabidopsis thaliana (CAA62079), Nicotiana tabacum (AAD20458), Saccharomyces pombe (NP_593135), Saccharomyces cerevisiae (EDN59602) and Candida albicans (XP_713612) are compared. The sequences were aligned using CLUSTAL W software. Accession numbers of sequences are embraced by brackets. Highlighted in yellow is the position of conserved histidine rich motifs.
FvC5SD is a transmembrane enzyme with C-5 sterol desaturase function
In order to establish the function of FvC5SD as C-5 sterol desaturase, biochemical complementation was carried out in S. cerevisiae haploid erg3 knockout strain impaired in ergosterol biosynthesis ( Fig. 2a ). ERG3 encodes C-5 sterol desaturase, which catalyzes the introduction of a C-5 (6) double bond into episterol, a precursor in ergosterol biosynthesis.
FvC5sdp is a transmembrane C-5 sterol desaturase.
(a) Biochemical complementation of erg3 knockout strain of S. cerevisiae with FvC5SD. Total sterol was isolated from WT strain of S. cerevisiae (SC4741), erg3 knockout strain with FvC5SD or empty vector control and analyzed by HPLC. (b) Complementation of cycloheximide hypersensitivity phenotype of erg3 knockout strain of S. cerevisiae with FvC5SD. WT strain of S. cerevisiae (SC4741), erg3 knockout strain with FvC5SD cloned in pGal10HO vector or empty vector were streaked on plates of synthetic minimal medium with and without 0.13 μg/ml of cycloheximide. (c) Kite-Doolittle hydropathy plot analysis (window size of 21) of FvC5sdp predicts presence of four major transmembrane domains. (d) Western blotting of total, soluble and microsomal protein fraction isolated from F. velutipes. Peptide- based antibody against FvC5sdp (raised in rabbit) was used for the analysis. HPLC, High Performance Liquid Chromatography; WT, wild type.
FvC5SD cDNA along with 100 bp upstream to ATG was cloned into galactose inducible vector pGal10 and transformed into the erg3 strain. Ergosterol analysis by HPLC revealed the presence of peaks specific to ergosterol in erg3 knockout expressing FvC5SD which otherwise were absent in case of control with empty vector. Hence, the FvC5SD homology (42% identity) to S. cerevisiae Erg3p was sufficient to complement its function by restoring ergosterol biosynthesis.
S. cerevisiae mutants with defective ergosterol biosynthesis like erg3 have altered membrane sterol composition leading to change in its plasma membrane permeability. This in turn affects the uptake and/or efflux of the drugs like cycloheximide. Cycloheximide hypersensitivity26 is one of the phenotypes exhibited by erg3 knockout strain of S. cerevisiae. We performed phenotypic complementation ( Fig. 2b ) in erg3 knockout strain by expressing cDNA of FvC5SD. The wild type S. cerevisiae strain (SC4741) and erg3 strain expressing FvC5SD were able to grow on plates containing cycloheximide (0.13 μg/ml). However, the erg3 knockout expressing empty vector displayed negligible growth at this concentration of cycloheximide. All the strains were observed to show normal growth on minimal media without cycloheximide. Thus, FvC5SD expression could rescue the cycloheximide hypersensitivity phenotype of the yeast mutant.
The initial prediction based on the hydropathy plot analysis indicates FvC5sdp to be a polytopic membrane protein with as many as four transmembrane helices ( Fig. 2c ). To confirm the membrane association of FvC5sdp, total protein isolated from F. velutipes was subjected to sub-cellular fractionation to separate soluble and microsomal fraction. These fractions were further analyzed by Western blotting with peptide- based antibodies ( Fig. 2d ). FvC5sdp (≈35 kDa) was observed to accumulate primarily in the microsomal fraction.
FvC5SD is an iron regulated gene involved in iron uptake
S. cerevisiae erg3 knockout strain is associated with another phenotype i.e. inability to grow on iron deficient medium. Wild type S. cerevisiae strain and erg3 knockout expressing either empty vector (pGal10) or FvC5SD were spotted on limiting iron medium (LIM) with 100 μM BPA (Bathophenanthroline disulphonic acid, a known iron chelator) in different dilutions. We observed that erg3 knockout with empty vector was unable to grow on LIM. However, erg3 strain expressing FvC5SD could rescue this phenotype by showing growth comparable to wild type on LIM ( Fig. 3a ).
FvC5SD is an iron regulated gene involved in iron uptake.
(a) Spot dilution assay of WT strain of S. cerevisiae (SC4741), erg3 knockout strain with FvC5SD cloned in pGal10HO vector or empty vector on synthetic minimal medium without and with 100μM BPA (limiting iron medium). (b) Northern blot analysis to show iron mediated regulation of FvC5SD transcription. Total RNA was isolated from F. velutipes mycelia grown in medium with or without BPA (100 μM). Graph shows mRNA levels on autoradiograms quantified by densitometry scanning using Fluor-S-(Bio-Rad) software. Data are mean ± SE (n = 3). (c) Comparison of radioactive iron (Fe55Cl3) uptake by S. pombe strain expressing FvC5SD and empty pSL173 vector at different time points. Data are mean ± SE (n = 3). (d) FvC5sdp structure predicted by homology based modelling using ESyPred3D server and threading by LOOP server. The two probable iron binding sites for each structure are also shown. Arrow is used to depict the residue in the iron binding sites which is also part of histidine rich motifs. WT, wild type; BPA, Bathophenanthrolinedisulphonic acid (iron chelator).
In C. albicans, iron deprivation has been shown to regulate ergosterol biosynthesis27,28. We also observed that the FvC5SD expression was transcriptionally regulated by iron in F. velutipes, although no known iron regulatory element (IRE) could be detected by in silico analysis of its promoter. Northern blot analysis showed upregulation of FvC5SD transcript by ≈ 3.0 fold in LIM (with BPA as iron chelator) relative to control condition without BPA ( Fig. 3b ). Inability of erg3 knockout to grow on LIM as well as the upregulation of FvC5SD transcript in LIM suggests that C-5 sterol desaturase may play a significant role in iron uptake in iron deficient conditions. For this purpose, we performed radioactive iron (Fe55Cl3) uptake assay with S. pombe expressing FvC5SD cloned in pSLF173 vector or empty vector ( Fig. 3c ). We observed that the level of uptake increased to ≈ 2.0 fold (after 4 hrs of incubation with Fe55) in case of S. pombe expressing FvC5SD as compared to that with the empty vector.
In order to purify FvC5Sdp, we cloned FvC5SD cDNA in bacterial expression vector pGEX4T-2 and transformed into BL21strain of E. coli for expression. GST- FvC5sdp fusion protein (≈60 kDa) was treated with thrombin to purify FvC5sdp which exhibited the expected molecular weight of ≈34.8 kDa on denaturing SDS-PAGE ( Fig. S3a ). However, purified FvC5sdp exhibited very high molecular weight (>ferritin marker of 440 kDa) under the non-denaturing conditions ( Fig. S3b ). We speculate that the observed high molecular weight could be a consequence of tendency of FvC5sdp to form oligomers in the native state which might result in binding of several atoms of iron.
In order to predict the iron-binding site in FvC5sdp, it was necessary to retrieve information on structure of FvC5sdp. Unfortunately, the structural information on membrane desaturase is scarce owing to technical limitations in obtaining large quantities of purified protein and the intrinsic difficulties in obtaining crystals from membrane proteins. Crystal structures are available only for a few soluble desaturases including the 18:0 Δ9-desaturase29 from Ricinus communis (castor) and a bifunctional desaturase from Hedera helix (ivy)30 which are found in the plastids of the higher plants. Hence, inspite of showing only 17.4% homology to FvC5sdp, we used ivy {Delta}4-16:0-Acp Desaturase (2UW1 chain ‘A’) containing an oxidized (i.e. resting FeIII-FeIII) di-iron active site as template for homology based modeling. ( Fig. 3d upper panel). Two possible metal binding sites (presumably iron) were also predicted for this model. First site is constituted by the residues His 86, His 247 and His 248 and the second site includes the residues Glu 97, His 170 and His174. Among these residues, His 170, His 174, His 247 and His 248 are involved in forming histidine rich motifs. Inspite of showing such low homology to the template, 90.2% of the residues in the predicted structure of FvC5sdp fall in most favored (most allowed) region of the Ramachandran plot31.
‘Threading’ was used as another in silico approach to predict an alternate model for FvC5sdp ( Fig. 3d bottom panel). Protein threading, also known as fold recognition, is a method of protein modeling used to model those proteins which have the same fold as proteins of known structures, but do not have homologous proteins with known structure.The first predicted metal binding site in this model involves His 174, His 228 and Asp 231 whereas the second site involves His 248, His 252 and Glu 293. Among these residues, His 174, His 248 and His 252 are part of histidine rich motifs. Overall 77.7% of the residues fall in most favoured region of Ramachandran plot. The predicted topology of FvC5sdp in the membrane environment (Fig. S4) correlates well with both the models and all the predicted residues involved in iron- binding (which probably form the catalytic centre of the enzyme) lie towards the cytosolic face of the protein.
Characterization of FvC5SD expression
Several known wax biosynthesis enzymes in plants like CER1 consist of domain showing homology to C-5 sterol desaturase. CER1 like protein homolog (AI484218) with sterol desaturase domain has also been identified in tomato. We have chosen tomato (one of the economically important vegetable crops in the Solanaceae family) as a model system for heterologous expression of FvC5SD to examine if it has any effect on quality or quantity of wax.
The ORF of FvC5SD was placed under the control of CaMV35S promoter by replacing the GUSA gene of the binary vector, pBI121 and the resulting construct, pBIFvC5SD, was introduced into tomato by Agrobacterium mediated transformation. Six independent T0 transgenic lines were generated which were confirmed for transgene expression by Northern and Western blot analysis ( Fig. S5 ). Further, we focussed on three independent transgenic lines (D-2, D-11 and D-12) based on different levels of FvC5SD expression for agronomic and biochemical analysis. Leaf surface of the transgenic tomato plants were characterized by darker green colour and glossier surface than the wild type plants ( Fig. 4a left panel). Light microscopy images of the leaves from transgenic plants showed higher epidermal cell density compared to wild type. The epidermal cells of transgenic leaves were smaller in size and arranged compactly with much less intercellular space ( Fig. 4a right panel). However, increase in cell number could compensate for cell size as no significant difference was observed in size of transgenic leaves compared to wild type. Besides this, no other visible change was observed on phenotype and fruit yield of transgenic tomato lines.
FvC5SD expression enhances drought tolerance in transgenics.
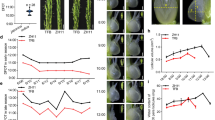
(a) Leaf surface of the transgenic tomato plants had darker green colour and glossier surface than the WT control plants (left). Light microscopy images of leaves from transgenic line (D-2) showed higher epidermal cell density compared to WT (right). (b) Epicuticular wax layers on adaxial leaf surfaces of WT control and transgenic line (D-2) viewed by scanning electron microscopy. Images were taken at two different magnifications of 7000× and 1500×. (c) Composition of leaf cuticular waxes of 4 weeks old WT and transgenic lines as determined by GC-MS. Wax constituents (alkane, alkenes, alcohol, fatty acids and triterpenoids) are expressed in μg/mg of fresh weight of leaves. All values are mean ± SE (n = 3). (d) Carbon chain length distribution of leaf cuticular waxes of 4 weeks old WT and transgenic lines. All values are mean ± SE (n = 3). (e) Chlorophyll leaching assay from leaves of WT control and transgenic line D-2. Value of chlorophyll extracted at each time point is expressed as percentage of total chlorophyll. Data represents the mean ± SE of two independent experiments (n = 3 for each experiment). (f) Transgenic tomato plants expressing FvC5SD showed enhanced drought tolerance. One month old tomato transgenics (three independent transgenic lines D-2, D-11and D-12) and WT plants were subjected to drought stress by withholding the water for 17 days followed by resumption of normal water scheme for recovery. WT and transgenic plants recovered after drought stress are shown in the photographs. WT, wild type; GC-MS, gas chromatography-mass spectrometry.
We compared the wax accumulation on the leaf surfaces of the transgenic and wild type plants by scanning electron microscopy (SEM). SEM study of the tomato leaf surface demonstrated ribbon- like distribution of crystalline wax with much more dense deposition in transgenic leaves than wild type ( Fig. 4b ). To quantify the change in wax deposition on the leaves, detailed chemical analyses of total wax mixtures were conducted on 4 weeks old plants by gas chromatography-mass spectrometry (GC-MS). Plants transformed with FvC5SD showed around 23% more cuticular wax load relative to the wild type. Since wax components were fairly uniform across the transgenic lines (D-2, D-11 and D-12) with no significant statistical difference, pooled data of wax components from these three independent transgenic lines is presented in Fig. 4c & 4d . It is evident that there was increase in accumulation of alkanes by 13.54%, alkenes by 24.2% and alcohol by 33.01%. Fatty acids which are minor content of tomato leaf wax showed the largest percentage increase of about 61% in the transgenics. Amount of Lupeol which was the major triterpenoids detected in leaf wax remained unaltered. Besides alteration of absolute amount of wax compounds, wax composition represented by carbon chain length distribution was also altered by the expression of FvC5SD as shown in Fig. 4d . The alteration in cuticle permeability may be associated with increase or decrease in the wax coverage. To investigate further whether the cuticle properties of the transgenics were altered, chlorophyll leaching assays ( Fig. 4e ) were performed at different times with leaves excised from 4 weeks old plants of wild type control and transgenic line D-2. Results showed that chlorophyll leaching from transgenic leaves was significantly slower as compared to control leaves indicating a decrease in cuticular permeability due to increased wax content.
Drought tolerance of transgenic plants
The most important physiological function of epicuticular wax is to protect the plants from water loss. Therefore, we examined drought tolerance characteristic over one month old plants from wild type and transgenic lines (D-2, D-11 and D-12) by suspending the water for 17 days. Ten days after watering was stopped, all the wild type plants showed wilting while all the transgenic plants still kept their whole plant turgor and showed delayed wilting. After one more week without watering, both wild type and transgenic plants became dehydrated. Upon resumption of the normal watering scheme for 15 days, the transgenic plants recovered much faster and efficiently than the wild type plants ( Fig. 4f ).
Resistance of transgenic plants to S. sclerotiorum infection
Epicuticular wax layer not only acts as an obstacle to drought but also disease by influencing germination and virulence of several plant pathogenic fungi. It acts as the first barrier encountered by invader pathogen that prevents it from coming into direct contact with the underlying epidermal cells and thereby limits infection. Therefore, increased epicuticular wax accumulation in transgenics expressing FvC5SD should lead to enhanced resistance to fungal attack. Since tomato is the natural host of Sclerotinia sclerotiorum, we tested the ability of the fungus to infect leaves of the wild type and transgenic plants. Plants from three different transgenic lines (D-2, D-11 and D-12) were selected for pathogenesis assay. Leaves were considered as diseased when the symptoms of water soaking, browning of tissue and lesion formation appeared. The agar plug containing S. sclerotiorum was applied randomly near the center of the detached leaves. The control set of plants showed typical water soaking and rotting surrounded by chlorotic halo ( Fig. 5a ). The transgenic set showed a slow progress of disease when compared to wild type plants ( Fig. 5b ). Trypan blue staining of infected leaves confirmed that the disease symptoms appeared as a result of spreading of fungal mycelia ( Fig. 5a inset). The fungal hyphal structures in infected leaves were stained blue.
FvC5SD expression in transgenic tomato leads to resistance to infestation by fungus S. sclerotiorum.
(a) The lesion formation in leaves of WT and transgenic lines (D-2, D-11 and D-12) after five days of inoculation. The diseased leaves showed growth of the fungi and lesion formation. Shown in inset is the trypan blue staining of fungal mycelium in one of the infected leaves viewed under light microscope (40× magnification). (b) Time kinetics of disease development in tomato as measured by size of the lesion. Data points represent the mean ±SE of lesion sizes from three leaves. WT, wild type.
FvC5SD expression in tomato enhances total iron content of the transgenics
Use of genetic engineering to increase the iron content of the crops can be achieved by the introduction of genes that code for trace element- binding proteins; overexpression of storage proteins already present and/or increased expression of proteins that are responsible for iron uptake into plants. Since FvC5sdp is an iron-binding protein with a possible role in iron uptake, we estimated the total iron content of tomato fruits from wild type control and FvC5SD expressing transgenic lines (D-3, D-7, D-2, D-11 and D-12) by atomic absorption spectroscopy. Interestingly an increase in total iron content of about 2–3 fold was observed in transgenic tomato fruits compared to that from control plants ( Fig. 6a ).
FvC5SD expression enhances the nutritional value of transgenics.
(a) Total iron content in WT control and transgenic lines (D-3, D-7, D-2, D-11 and D-12) as determined by atomic absorption spectroscopy. Values are mean ± SE (n = 3). (b) Graph showing ratio of C-18 saturated: monounsaturated fatty acids as analyzed by GC-MS based total lipid profiling of tomato fruits from WT and transgenic line D-2. (c) Comparative account of various essential polyunsaturated fatty acids (PUFA) in tomato fruit from WT and transgenic line D-2 as determined by GC-MS. Values are mean ± SE (n = 3). WT, wild type; GC-MS, gas chromatography-mass spectrometry.
FvC5SD expression in tomato enhances polyunsaturated fatty acid content of transgenics
Total lipid profiling of transgenic tomato fruits (from D-2 transgenic line) by GC-MS showed an interesting change in the fatty acid content when compared with wild type fruits. An increased ratio of monounsaturated fatty acid (e.g. oleic acid) to saturated fatty acid (e.g. stearic acid) was observed in the transgenic fruits ( Fig. 6b ). Besides, the essential polyunsaturated fatty acid (PUFA) like linoleic acid (9, 12 octadecadienoic acid), alpha linolenic (9,12,15 octadecatrienoic acid), ω-3,6,9-octadecatrienoic acid and 11, 14 eicosadienoic acid ( Fig. 6c ) were increased by approximately 1.5 fold , 5 fold, 1.5 fold and 2.5 fold respectively compared to the non transgenic tomatoes. This increase further added to the nutritional value of the transgenics as PUFAs have well recognised role in health and nutrition.
Discussion
The increased amount of wax deposition in transgenic tomato expressing FvC5SD, conferring it enhanced drought tolerance and pathogen resistance could be attributed to the presence of C-5 sterol desaturase domain which is present in many enzymes of wax biosynthesis like CER1 of Arabidopsis14. CER1 like protein homolog (AI484218) has also been reported in tomato but has not characterized as yet. The multiple alignment of FvC5sdp with other enzymes involved in epicuticular wax biosynthesis including CER1 revealed little identity ( Fig. S6 ). However, histidine-rich motifs forming di-iron-binding site which are essential for catalytic activity were conserved in all of them. These motifs might allow FvC5sdp to perform similar catalytic reaction like CER1. Besides, FvC5sdp is also a transmembrane protein that localizes predominantly in ER membrane like wax biosynthesis enzyme CER132. If FvC5sdp is considered to be directly involved in wax biosynthesis, then it might be assumed that wax-C superfamily domain (present at C-terminus of wax biosynthesis enzymes like CER1) possibly does not play any significant role in the wax biosynthesis in tomato as FvC5sdp lacks wax-C domain. It might provide some unknown function which needs to be studied further.
Alternatively, instead of direct involvement in wax biosynthesis, FvC5sdp can possibly have an indirect affect on wax biosynthesis. Changes in the sterol desaturation pattern caused due to expression of FvC5SD in tomato could lead to feedback regulation of wax biosynthesis genes resulting in increased wax accumulation.
CER1 in Arabidopsis was proposed to be an aldehyde decarbonylase converting aldehydes to alkanes and its over expression leads to dramatic increase in the production of alkanes but other wax components remains unaffected12,14. However, FvC5SD expression leads to increase in all the wax components including alkanes, alkenes, alcohol and fatty acids which could be due to the wide substrate specificity of the fungal FvC5SD enzyme.
FvC5sdp is an iron-binding protein and its expression lead to enhanced total iron content in transgenic tomato. FvC5sdp forms high molecular weight oligomers in the native state which might contribute to binding of several atoms of Fe resulting in an increased iron level in transgenic fruits.
Increase in level of polyunsaturated fatty acids is surprising since FvC5SD is known to create desaturation in sterols. Therefore, it is speculated that FvC5SD might be performing some non-specific desaturation of fatty acids due to relaxed substrate affinity since both fatty acid desaturase and sterol desaturase have catalytic centre consisting of conserved histidine boxes in common. Stearic acid could be one of the possible substrates for FvC5sdp since ratio of stearic acid/oleic acid was observed to decrease in transgenic fruits relative to wild type.
C-5 sterol desaturase is known to be involved in phytosterol and brassinosteroid biosynthesis in plants33, so it was obvious for us to examine by GC-MS if the transgenic tomato lines expressing FvC5SD had altered level of phytosterols and brassinosteroids. However in comparision to the wild type, no significant changes were observed in the levels of phytosterols and brassinosteroids (data not shown). This could be due to the reason that this enzyme might not be involved in rate limiting step.
It can be concluded that genetic modification of leaf cuticular waxes by expressing C-5 sterol desaturase has a great potential for crop improvement by enhancing drought tolerance and fungal resistance which are the primary causes of reduction in crop yield. Besides, expression of FvC5SD in these crops can also prove to be a potentially useful strategy to increase the total iron content and essential polyunsaturated fatty acids, hence adding additional dietary benefits to the transgenics. It is conceivable that similar strategy could also be extended to other important staple crops.
Methods
Plant material and transgenic plants
To generate transgenic tomato plants, cotyledons from 2-week-old seedlings of Solanum lycopersicum (cv Pusa ruby) were used34. Briefly, seeds were sterilized using 4% commercial bleach and germinated on Murashige and Skoog (MS) medium35. The binary Ti vector pBI121 was used for transformation. The GUS gene of the binary vector was replaced with the FvC5SD gene at the XbaI and SacI restriction sites to gain the new expression construct FvC5SD-pBI121 which was electroporated into Agrobacterium tumefaciens strain EHA105. After 2 weeks of germination, the cotyledons were cut and co-cultivated for 30 min with A. tumefaciens. The cotyledons were then collected for selection on MS plates containing 50 mg/l kanamycin. The regenerated plantlets were transferred to rooting medium. Transgenic seeds were germinated in MS medium containing 50 mg/l kanamycin to get the progeny plants.
Strains, media and growth condition
Flammulina velutipes (ATCC13547) was maintained on media consisting of 5.0% Dextrose, 1.0% peptone, 0.1% KH2PO4, 0.05% MgSO4.7H2O and 1% Malt Extract at 23°C for 15-30 days36. S. sclerotiorum was maintained on potato/dextrose/agar (PDA, 20% potato, 2% dextrose and 1.5% agar) slants. S. pombe was grown aerobically at 30°C in either rich media (YES) or selective synthetic medium (EMM)37. S. cerevisiae was maintained in rich YPD or YPG medium (with 2% galactose or 2% glucose) or in synthetic minimal SD medium. The media was made iron limiting by addition of 100 μM concentration of Bathophenanthrolinedisulphonic acid (BPS). Genetic transformation of yeast was carried out by the alkaline cation method38. The strain and plasmids used in the study are listed in Table S1 .
Scanning electron microscopy (SEM)
For SEM examination, 0.5-cm segments were prepared from the appropriate region of the leaves from one month old tomato plants. The samples were air dried for 72 hrs at room temperature, then mounted onto a copper holder with double adhesive carbon tape, sputter coated with gold for 3 min and examined under an electron microscope (Zeiss).
Chlorophyll leaching assay
Epidermal permeability was measured using a chlorophyll extraction assay. Two or three leaves of same size were collected from 1 month old tomato plants kept in dark for 16 hrs and immersed in 50 ml tubes with 10 ml of 80% ethanol such that all the leaf samples just floated. Tubes were protected from light and agitated gently on a rotator platform at 50 rpm. Aliquots of 1.0 ml were taken out for chlorophyll quantification and poured back to the same tube at every time point. The amount of chlorophyll extracted into the solution was quantified using a spectrophotometer (Beckman, Fullerton, CA, USA) and calculated from light absorption at wavelength of 647 and 664 nm39. Chlorophyll extracted at each time point was expressed as a percentage of total chlorophyll extracted after 48 h of immersion. Data were obtained from two independent experiments, each of which was replicated thrice.
Epicuticular wax extraction and gas chromatography-mass spectrometry analysis (GC-MS analysis)
Leaves from one month old tomato plants were immersed in 30 ml chloroform for 30 s at room temperature. The same leaves were then re-extracted with chloroform at 60°C for 20 s and the two chloroform extracts containing wax were pooled. Chloroform was evaporated and five microgram of n-tetracosane (C24) was added to each sample as an internal standard. To the dried residue 100 μl of derivatization reagent (80 μl BFSTA+ 20 μl TMCS) was added and incubated at 65°C for 1 hr. GC-MS analysis was performed with 1 μl of the sample in split mode on Shimadzu GCMS-QP 2010 plus. The mass spectrometer was tuned according to the manufacturer's recommendations. GC was performed on an Rtx5MS- 30 m column with 0.25-mm ID and 0.25 μm df (Restek). The injection temperature was set to 300°C, the interface temperature to 300°C and the ion source adjusted to 250°C. Helium was used as the carrier gas at a flow rate of 1 ml min−1. The analysis was performed using the following temperature program: 1 min of isothermal heating at 100°C followed by heating at 300°C for 20 mins. Mass spectra were recorded at 2 scan sec−1 with a scanning range of 40 to 850 m/z. Quantification was based on peak areas and normalization based on the internal standard.
Tomato fruit total lipid extraction for GC-MS analysis
Lyophilized fruit tissue (500 mg) was crushed to powder and transferred to a glass vial. 10 μl of 5α -cholest 7en-3β-ol (1 mg/ml stock) was added as an internal standard. To it, 3.75 ml of CHCl3: methanol (1:2) was added and vortexed vigorously. To it 1.25 ml of CHCl3 was added followed by addition of 1.25 ml of dH2O and vortexed well. Bottom organic phase was transferred to a fresh vial and was allowed to evaporate completely at 35°C. To the dried residue, 500 μl of 6% methanolic KOH (w/v) was added and incubated at 85°C for ½ −1 hrs. To it, half the volume i.e. 250 μl of dH2O and then equal volume i.e. 750 μl of n-heptane was added and vortexed well. It was allowed to stand for sometime till the layers get separated. Upper phase was transferred to a fresh vial. Above step was repeated twice. Heptane was allowed to evaporate completely (16–24 hrs). To the dried residue, 100 μl of derivatization reagent (80 μl BFSTA+20 μl TMCS) was added and incubated at 65°C for 1 hr and injected to GC-MS instrument. GC-MS analysis was performed with 1 μl of the sample in the split mode on Shimadzu GCMS-QP 2010 plus under the similar conditions as described above.
Biochemical complementation
FvC5SD cDNA along with 100 bp upstream to ATG was cloned in Hind III restriction site of S. cerevisiae expression vector pGal10-HO under galactose inducible promoter. The construct was transformed in erg3 knockout of S. cerevisiae and transformants were selected on SD minimal media without uracil. Total intracellular sterols were extracted by alcoholic KOH method as reported by Breivik and Owades with slight modifications40. For HPLC analysis of ergosterol, C18 ultra sphere 5 μcolumn (250 × 4.6 mm) was used with methanol and water as solvent (99.7: 0.3 v/v). In the isocratic elution mode flow rate was set to1 ml/min. at RT and U.V detector was used a 281.5 nm. Column temperature was set to 21±0.2°C. Ergosterol (Sigma) was used as standard for analysis.
Atomic absorption spectroscopy for determination of total iron content
Samples (2.0 gm) were dried by lyophilisation and 20 ml of acid mixture ( = conc. nitric acid: conc. perchloric acid :: 4:1) was added to it and evaporated to near dryness on a hot plate. The above procedure was repeated for samples until the fumes and sediment became white. After cooling, 5 ml of conc. hydrochloric acid was poured on the sediment. The samples were boiled for 5 minutes and allowed to dry and cool. After cooling, the sediments were quantitatively transferred in 100 ml volumetric flask dissolving the sediments into sterile water. Then these were filtered twice and kept in polypropylene bottles. The concentration of iron was determined by flame atomic absorption spectrometry using Spectra AA 55 spectrophotometer (Varian, USA; provided by Institution of Environmental Studies and Wetland Management, Kolkata).
Iron uptake assay
Iron uptake assay was performed as described earlier with slight modifications41. Cells were grown in EMM medium to O.D of 2.0 at 600 nm and harvested by centrifugation at 2,770 g. The cells were than washed three times with sterilized distilled water and resuspended in assay buffer (10 mM trisodium citrate- pH 6.5, 5% glucose containing1 μM Fe55Cl3) followed by incubation at 30°C with continuous agitation. At different time intervals, 1.0 ml aliquot of the culture was taken and added to 5.0 ml ice cold 0.25 M EDTA- pH6.5 and mixed to quench any free extracellular radioactive iron. The cells were then harvested by vacuum filtration through a glass fibre filter (WhatmanGF/C25 mm) and washed thrice with 5.0 ml ice cold 0.25 M EDTA- pH 6.5 and twice with ice cold sterilized distilled water. The filters were finally dried and 5.0 ml emulsifier scale scintillation fluid was added. All samples were counted in tritium channel of a Minaxi Tri- carb 400 series Scintillation counter (Canberra Packard).
Pathogenesis assay with S. sclerotiorum and trypan blue staining
S. sclerotiorum infection was carried out by the mycelium agar disc method on detached leaf as described before42. Mycelial agar plugs of 3.0 mm diameter punched from growing margins of a 4-day-old S. sclerotiorum culture was applied on the adaxial surface of the leaves. Leaves were kept under 16-h photoperiod and 100% humidity. The disease symptoms were observed every 24 hrs, over a period of 1 week. This experiment was repeated three times under similar conditions.
For trypan blue staining, infected leaves were stained with 0.05% Trypan blue for 45 min and washed with PBS. All samples were viewed under light microscopy at 40× magnification.
Bioinformatics analysis
ESyPred3D server43 was used for comparative homology modelling and i-TASSER44 loop prediction server was used for threading. Quality of model was checked by PROCHECK analysis45. CHED server was used for prediction of soft metal binding sites in the models.
References
Boyer, J. S. Plant productivity and environment. Science 218, 443–448 (1982).
Jordan, W. R. et al. Environmental physiology of sorghum. Epicuticular wax load and cuticular transpiration. Crop Sci. 24, 1168–1173 (1984).
Jenks, M. A. et al. Chemically induced cuticle mutation affecting epidermal conductance to water vapour and disease susceptibility in Sorghum bicolor (L.) Moench. Plant Physiol. 105, 1239–1245 (1994).
Eigenbrode, S. D. & Espelie, K. E. Effects of plant epicuticular lipids on insect herbivores. Annu. Rev. Entomol. 40, 117–142 (1995)
Kolattukudy, P. E. in Cutin, suberin and waxes, The biochemistry of plants Vol. 4. (eds Stumpf, P. K., Conn, E. E.) 571–645 (NewYork: Academic Press, 1980).
Walton, T. J. in Waxes, cutin and suberin, Methods in plant biochemistry lipids, membranes and aspects of photobiology (eds Harwood, J. L., Bowyer, J. R.) 105–158 (Academic, San Diego, 1990).
Aharoni, A. et al. The SHINE clade of AP2 domain transcription factors activates wax biosynthesis, alters cuticle properties and confers drought tolerance when over expressed in Arabidopsis. Plant Cell 16, 2463–2480 (2004).
Broun, P. et al. WIN1, a transcriptional activator of epidermal wax accumulation in Arabidopsis. Proc. Natl. Acad. Sci. USA 101, 4706–4711 (2004).
Zhang, J. Y. et al. Overexpression of WXP1, a putative Medicago truncatula AP2 domain-containing transcription factor gene, increases cuticular wax accumulation and enhances drought tolerance in transgenic alfalfa (Medicago sativa). Plant J. 42, 689–707 (2005).
Zhang, J. Y. et al. Heterologous expression of two Medicago truncatula putative ERF transcription factor genes, WXP1 and WXP2, in Arabidopsis led to increased leaf wax accumulation and improved drought tolerance, but differential response in freezing tolerance. Plant Mol. Biol. 64, 265–278 (2007).
Islam, M. A. et al. Characterization of Glossy1-homologous genes in rice involved in leaf wax accumulation and drought resistance. Plant Mol. Biol. 70, 443–456 (2009).
Bourdenx, B. et al. Overexpression of Arabidopsis ECERIFERUM1 promotes wax very-long-chain alkane biosynthesis and influences plant response to biotic and abiotic stresses. Plant Physiol. 156, 29–45 (2011).
Gong, P. et al. Transcriptional profiles of drought-responsive genes in modulating transcription signal transduction and biochemical pathways in tomato. J. Exp. Bot. 61, 3563–3575 (2010).
Aarts, M. G. M. et al. Molecular characterization of the CER1 gene of Arabidopsis involved in epicuticular wax biosynthesis and pollen fertility. Plant Cell 7, 2115–2127 (1995).
Chen, X. et al. Cloning and characterization of the WAX2 gene of Arabidopsis involved in cuticle membrane and wax production. Plant Cell 15, 1170–1185 (2003).
Hu, X. et al. cDNA cloning and expression analysis of a putative decarbonylase TaCer1 from wheat (Triticum aestivum L.). Acta. Physiol. Plant 31, 1111–1118 (2009).
Sturaro, M. et al. Cloning and characterization of GLOSSY1, a maize gene involved in cuticle membrane and wax production. Plant Physiol. 138, 478–489 (2005).
Shanklin, J. & Cahoon, E. B. Desaturation and related modifications of fatty acids. Annual Review of Plant Physiol. & Plant Mol. Biol. 49, 611–641 (1998).
Arthington, B. A. et al. Cloning, disruption and sequence of the gene encoding yeast C-5 sterol desaturase. Gene 102, 39–44 (1991).
Gachotte, D. et al. Isolation and characterization of an Arabidopsis thaliana cDNA encoding a Δ7-sterol-C-5-desaturase by functional complementation of a defective yeast mutant. Plant J. 9, 391–398 (1996).
Husselstein, T. et al. Delta7-sterol-C5-desaturase: molecular characterization of wild-type and mutant alleles. Plant Mol. Biol. 39, 891–906 (1999).
Matsushima, M. et al. Molecular cloning and mapping of a human cDNA (SC5DL) encoding a protein homologous to fungal sterol-C5-desaturase. Cytogenet. Cell Genet. 74, 252–254 (1996).
Geber, A. et al. Deletion of the Candida glabrataERG3 and ERG11 genes: effect on cell viability, cell growth, sterol composition and antifungal susceptibility. Antimicrob. Agents Chemother. 39, 2708–2717 (1995).
Brumfield, K. M. Functional characterization of the Chlamydomonas reinhardtii ERG3 ortholog, a gene involved in the biosynthesis of ergosterol. PloS One 5, e8659 (2010).
Azam, M. et al. Cloning and characterization of 5′-flanking region of oxalate decarboxylase gene from F. velutipes. Biochem. J. 367, 67–75 (2002).
Dudley, A. M. et al. A global view of pleiotropy and phenotypically derived gene function in yeast. Mol. Syst. Biol. 1, 0001 (2005).
Lan, C. Y. et al. Regulatory networks affected by iron availability in Candida albicans Mol. Microbiol. 53, 1451–1469 (2004).
Prasad, T. et al. Unexpected link between iron and drug resistance of Candida spp.: iron depletion enhances fluidity and drug diffusion, leading to drug-susceptible cells. Antimicrob. Agents. Chemother. 50, 3597–3606 (2006).
Moche, M. et al. Azide and acetate complexes plus two iron-depleted crystal structures of the di-iron enzyme delta-9- stearoyl-acyl carrier protein desaturase. Implications for oxygen activation and catalytic intermediates. J. Biol. Chem. 278, 25072–25080 (2003).
Whittle, E. et al. A multifunctional acyl-acyl carrier protein desaturase from Hedera helix L. (English Ivy) can synthesize 16- and 18-carbon monoene and diene products. J. Biol. Chem. 280, 28169–28176 (2005).
Ramachandran, G. N., Ramakrishnan, C. & Sasisekharan, V. Stereochemistry of polypeptide chain configurations. J.Mol. Biol. 7, 95–9 (1963).
Kamigaki, A. Suppression of peroxisome biogenesis factor 10 reduces cuticular wax accumulation by disrupting the ER network in Arabidopsis thaliana. Plant Cell Physiol. 50, 2034–2046 (2009).
Sunghwa, C. et al. The Arabidopsisdwf7/ste1 mutant is defective in the Δ7 Sterol C-5 desaturation step leading to brassinosteroid biosynthesis. Plant Cell 11, 207–221 (1999).
Fillati, J. J. et al. Efficient transfer of a C-glyphosphate tolerance gene into tomato using a binary Agrobacterium tumifaciens vector. Biotechnology 5, 726–730 (1987).
Murashige, T. & Skoog, F. A revised medium for rapid growth and bioassays with tobacco tissue culture. Physiol. Plant. 15, 473–497 (1962).
Mehta, A. & Datta, A. Oxalate decarboxylase from Collybia velutipes: Purification, characterization and cDNA cloning. J. Biol. Chem. 266, 23548–23553 (1991).
Maundrell, K. Thiamine-repressible expression vectors pREP and pRIP for fission yeast. Gene 123, 127–130 (1993).
Okazaki, K. et al.High-frequency transformation method and library transducing vectors for cloning mammalian cDNAs by trans-complementation of Schizosaccharomyces pombe. Nucleic Acids Res. 18, 6485–6489 (1990).
Lolle, S. J. et al. Developmental regulation of cell interactions in the Arabidopsis fiddlehead-1 mutant: a role for the epidermal cell wall and cuticle. Dev. Biol. 189, 311–321 (1997).
Breivik, N. & Owades, L. Spectrophotometric semimicrodetermination of ergosterol in yeast. Agricult. & Food Chem 5, 360–363 (1957).
Marvin, M. E., Mason, R. P. & Cashmore, A. M. The CaCTR1 gene is required for high-affinity iron uptake and is transcriptionally controlled by a copper-sensing transactivator encoded by CaMAC1. Microbiology 150, 2197–2208 (2004).
Kesarwani, M. et al. Oxalate decarboxylase from Collybia velutipes : molecular cloning and its overexpression to confer resistance to fungal infection in transgenic tobacco and tomato. J. Biol. Chem. 275, 7230–7238 (2000).
Lambert, C. et al. ESyPred3D: Prediction of proteins 3D structures. Bioinformatics 18, 1250–1256 (2002).
Roy, A., Kucukural, A. & Zhang, Y. I-TASSER: a unified platform for automated protein structure and function prediction. Nat. Protocols 5, 725–738 (2010).
Laskowski, R. A. et al. PROCHECK: a program to check the stereochemical quality of protein structures. J. App. Cryst. 26, 283–291 (1993).
Acknowledgements
We thank Mr. Ajai Kumar and Ruchita Pal, AIRF, JNU for technical help in performing GC-MS and electron microscopy respectively. Prof. Suneel Mukherjee (ICGEB) is acknowledged for critical reading of the manuscript. AK acknowledges CSIR for fellowship.
Author information
Authors and Affiliations
Contributions
AK performed all the experiments. AD supervised the entire project. AK and AD analyzed the data and wrote the paper. MK advised GC-MS, discussed the data and critically read the manuscript. SC, NC and MA critically read the manuscript.
Electronic supplementary material
Supplementary Information
Supplementary information
Rights and permissions
Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Kamthan, A., Kamthan, M., Azam, M. et al. Expression of a fungal sterol desaturase improves tomato drought tolerance, pathogen resistance and nutritional quality. Sci Rep 2, 951 (2012). https://doi.org/10.1038/srep00951
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00951
This article is cited by
-
Characterization and functional analyses of wheat TaPR1 genes in response to stripe rust fungal infection
Scientific Reports (2023)
-
Drought Tolerance Strategies in Plants: A Mechanistic Approach
Journal of Plant Growth Regulation (2021)
-
Sclerotinia stem rot in tomato: a review on biology, pathogenicity, disease management and future research priorities
Journal of Plant Diseases and Protection (2021)
-
The role of sterols in plant response to abiotic stress
Phytochemistry Reviews (2020)
-
Transcriptomic analysis of Eruca vesicaria subs. sativa lines with contrasting tolerance to polyethylene glycol-simulated drought stress
BMC Plant Biology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.