Abstract
We report the first mountain hare (Lepus timidus) transcriptome, produced by de novo assembly of RNA-sequencing reads. Data were obtained from eight specimens sampled in two localities, Alps and Ireland. The mountain hare tends to be replaced by the invading European hare (Lepus europaeus) in their numerous contact zones where the species hybridize, which affects their gene pool to a yet unquantified degree. We characterize and annotate the mountain hare transcriptome, detect polymorphism in the two analysed populations and use previously published data on the European hare (three specimens, representing the European lineage of the species) to identify 4 672 putative diagnostic sites between the species. A subset of 85 random independent SNPs was successfully validated using PCR and Sanger sequencing. These valuable genomic resources can be used to design tools to assess population status and monitor hybridization between species.
Design Type(s) | individual genetic characteristics comparison design • strain comparison design • population dynamics analysis objective |
Measurement Type(s) | transcription profiling assay |
Technology Type(s) | RNA sequencing |
Factor Type(s) | geographic location |
Sample Characteristic(s) | Lepus timidus • liver • Republic of Ireland • Switzerland |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
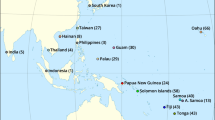
The mountain hare (Lepus timidus) is an Arcto-alpine species that was the most common and widely distributed hare species across Europe during the last glacial periods1. Nowadays, the mountain hare is distributed from Fennoscandia to Eastern Siberia, but also occurs in isolated/refuge populations (e.g., Ireland, Scotland, the Alps, Poland, the Baltics and Japan), and in places where it has been introduced (Iceland, England, Faroe Islands and New Zealand) (see Fig. 1). Even though they are a popular game species and abundant within its range, mountain hares have sharply declined in some regions, particularly in areas of contact with the European hare (Lepus europaeus), where the latter tends to invade and replace the range of the former1–4. Mountain and European hares share extensive natural and human-induced contact zones in Western Europe, from the British Isles to Scandinavia and Central Europe (Fig. 1). Climate change is predicted to affect lagomorphs extensively5,6 and, in particular, to accelerate the replacement of mountain hares by European hares in the contact zones, such as the Alps, Sweden or Ireland7,8. The two species may hybridize when in contact, resulting in some genetic introgression9–13, with potential effects on local adaptation14.
Approximate distributions of the mountain hare, Lepus timidus, and the European hare, L. europaeus, in Eurasia with indication of the areas of contact and of broad geographic overlap between the species (distribution ranges were adapted from IUCN Spatial Data Resources; IUCN 201651). Circles indicate the mountain hare sampling locations for this work (open circle—Ireland; closed circle—Alps).
Even though the mountain hare and other hare species have been the subject of several population genetics studies, these have been mostly based on a few markers10,15–17. Therefore, permanent genomic resources provide fundamental information to develop monitoring tools to evaluate population status and implement protective policies. In this work, we use high-throughput RNA sequencing to: i) generate genomic resources for the mountain hare; and, ii) use published data on the European hare18 to pinpoint candidate fixed differences between the species that can be used to build genotyping tools to monitor gene exchange in the contact zones. We here present the first mountain hare transcriptome, and the most complete among the currently available European Lepus transcriptomes.
Methods
A summary of the methodological workflow is shown in the flowchart of Fig. 2.
Flowchart of the RNA-sequencing setup and data analysis steps. Commands used in the analytical steps shown in bold are detailed in Table 1 (available online only).
Sampling procedure and locations
Specimens from the Alps (see Fig. 1) were sampled during regular permit hunting in Grisons, Switzerland. Specimens from Ireland (see Fig. 1) were captured from the wild in Borris-in-Ossory, by the Irish Coursing Club (ICC) for scientific research purposes under National Parks & Wildlife (NPWS) licence No. C 337/2012 issued by the Department of Arts, Heritage and the Gaeltacht (dated 31/10/2012). Irish hares were dispatched humanely and in accordance with the licence conditions by means of lethal injection administered by Mr William Fitzgerald, Veterinary Laboratory Service Follow (MVB MVM CertCSM), from the Department of Agriculture, Food and the Marine, Regional Veterinary Laboratory, Hebron Road, Kilkenny, R95 TX39. Total RNA was isolated from 8 individuals.
RNA extraction
Liver tissue was freshly collected, immediately preserved in RNAlater and then stored at −80 °C until RNA extraction. Prior to extraction, frozen samples were ground in liquid nitrogen with a ceramic mortar and pestle. Mortar and pestle were washed prior to extraction using a 6-step wash that includes the following washing reagents in order: 70% ethanol, tap water, 10% bleach, milli-Q water, RNase away (Thermo Fisher Scientific) and finishing with molecular grade H2O. RNA extraction was performed using RNeasy Mini Kit according to manufacturer instructions.
RNA sequencing library preparation
The SureSelect Strand-Specific RNA Library Prep for Illumina Multiplexed Sequencing (Agilent Technologies) kit was used to prepare cDNA libraries for all samples. Library sizes were estimated using a Bioanalyzer 2,100 and quantified using KAPA Library quantification kit (KAPA BIOSYSTEMS). Equal molar concentrations of each library were pooled together for sequencing.
Sequence data processing and de novo transcriptome assembly
A detailed description of tools and commands used in the data analysis is shown in Table 1 (available online only). A first quality evaluation of obtained sequence reads (Data Citation 1) was performed with FastQC v0.11.519. After read quality inspection, adapters were removed and quality trimming performed using TRIMMOMATIC v0.3620, with instructions to remove the first ten bases, Illumina adapters, reads below 25 bp long and bases in the ends of reads with quality below 10, and to perform a 4-base sliding window trimming and cutting fragments with an average quality below 10. Trimmed-read quality was rechecked with FastQC (Data Citation 2). A de novo transcriptome assembly was then performed using all properly paired reads from the eight individuals in the dataset using TRINITY v2.2.021, establishing RF as read orientation for a strand-specific assembly. In addition, as a complementary resource, de novo transcriptome assemblies for each of the two sampling localities were also performed. Transrate v1.0.322 was used to evaluate assembly quality and completeness and to remove possible chimeras and poorly supported contigs. Cleaned reads were mapped back to the produced assembly and only the well-supported contigs were retained (Transrate optimal cut-off >0.024). In order to remove redundancy produced by using multi-sample data to perform the assembly, all contigs were clustered using CD-HIT-EST v4.6.423 with a 95% similarity threshold. Open reading frames were predicted with TransDecoder v3.0.024 to remove possible contaminants such as non-coding RNA and DNA contamination. The final filtered transcriptome comprised contigs with predicted open reading frame and/or rabbit (Oryctolagus cuniculus) or pfam annotation. Filtered transcriptome as well as raw assemblies are available in Figshare (Data Citation 2).
Transcriptome annotation
Transcriptome annotation was performed adapting the protocol of Trinotate v3.0.124, using i) Conditional Reciprocal Best BLAST (crb-blast) v0.6.625 against the rabbit transcriptome reference (release 86) and Swiss-Prot database26; ii) protein domain identification by HMMER v3.1b227 onto the PFAM database28; iii) protein signal peptide through signal v 4.129; iv) transmembrane domain prediction using tmHMM v2.030; and v) eggNOG31, GO32and Kegg33 databases annotation. Annotation information was incorporated into an xlsx database (Data Citation 2).
SNP inference
SNP calling was performed separately for mountain hares (Data Citation 1) and European hares (Data Citation 3, from Amoutzias et al.18). The three European hare specimens represent the European lineage of the species18. First, reads from all the individuals were mapped to the filtered mountain hare de novo transcriptome with bwa-mem v0.7.1534 with default parameters and read group information added to each sequencing lane-sample pair. The resulting alignments were converted to a binary file (bam format), sorted and submitted to fixmate step using SAMtools v1.3.135. Duplicate reads were removed using Picard v1.140 (http://broadinstitute.github.io/picard) with the option MarkDuplicates. Realignment and recalibration was performed with Genome Analysis Toolkit v3.6-036. Finally, SNP call was carried out using Reads2snp v2.0.6437 using a threshold of 20 for site and mapping qualities, the paralog filter, a minimum coverage of 10X and a genotype probability >0.95. The resulting VCF file was deposited in Figshare (Data Citation 2). Only SNPs represented in all sampled specimens were retained.
Differentiation, admixture and Gene Ontology enrichment analysis
A set of random 5 502 SNPs, selected from independent contigs in order to reduce the linkage probability, was identified with VCFtools v0.1.1438. PGDSpyder v2.1.1.039 was used to convert this file to the required file formats. Partitions of genetic diversity in the dataset were investigated with a Principal Components Analysis, using PLINK v1.90b3.4540 and ggplot2 R package41 to plot the results. Additionally, the data were analysed using the admixture model implemented in STRUCTURE 2.3.442, with three replicate runs with 1 million steps after a burn-in period of 200 000, and K=2. Results were plotted using CLUMPACK43. Gene Ontology enrichment analyses were performed for the collection of contigs/genes with fixed differences between mountain and European hare samples, and between mountain hare sampling localities. The analysis was based on the rabbit proteome annotations and performed with g:Profiler34, applying the g:SCS multiple test correction and the ‘best per parent group’ hierarchical filter. The background set of genes was reduced to contigs with SNP information.
Independent SNP genotyping
A random set of 110 SNPs, inferred as potentially diagnostic between L. timidus and L. europaeus, was selected for independent validation using Sanger sequencing. DNA was extracted from two of the previously analysed mountain hare samples (one Alpine, Sample_3112, and one Irish, Sample_3103) and two other European hare specimens (sampled in Clermont-Ferrand—Sample—1569—Font-Romeu, Pyrenees—Sample—1550—in France during the regular hunting season). DNA extraction was performed using JETQUICK Tissue DNA Purification kit (Genomed). PCR primers were designed to be anchored in a single exon (taking into account intron-exon boundaries from the European rabbit reference genome) and to amplify a portion of 110 independent contigs containing at least one putative diagnostic SNP. The Primer sets were designed using the Scrimer pipeline44, which depends on Primer345 to design and set the primer conditions. A third internal sequencing primer was designed. PCRs were performed using QIAGEN Multiplex PCR Master Mix (Qiagen) and the following thermal cycling profile: initial denaturation at 95 °C for 15', 35 cycles of denaturation at 95 °C for 30'', annealing at 60–67 °C for 20'' and elongation at 72 °C for 30'', and a final extension step at 72 °C for 5'. PCR products were visually inspected under UV-light after electrophoresis in agarose gels stained with GelRed (Biotium), purified with Exonuclease I (New England Biolabs) and FastAP Thermosensitive Alkaline Phosphatase (Thermo Scientific), and sequenced using internal or, in a few cases, PCR primers in a ABI 3130xl genetic analyzer.
Code availability
Analyses in this work were performed with freely available open access tools mainly using command line versions (Table 1 (available online only)). Parameters are described in the methods section and software versions and commands used are detailed in Table 1 (available online only).
Data Records
Forty-eight raw FASTQ files were submitted to NCBI Sequence Read Archive, with accession number SRP095715 (Data Citation 1 and Tables 2 and 3). FASTQ files were divided in two sets, corresponding to the sampling localities (Ltim_Ireland and Ltim_Alps), and by biosample-specimen (SAMN06186748-3101, SAMN06186761-3102, SAMN06186762-3103 and SAMN06186763-3105; SAMN06186727-3112, SAMN06186728-3113, SAMN06186729-3114 and SAMN06186738-3116). In each biosample, six files were submitted, corresponding to three different Illumina HiSeq sequencing lanes and two read directions. Pre/post-cleaning FASTQC base quality pdf report (FASTQC.pdf) can be accessed in Figshare (Data Citation 2). This dataset is the core of this work and has not been released or analysed previously.
Trinity raw assemblies (Ltimidus_Trinity.fasta, LtimidusIreland_Trinity.fasta and LtimidusAlps_Trinity.fasta) were deposited on Figshare (Data Citation 2 and Table 4). The curated transcriptome assembly fasta files (LtimidusTranscriptome.cds.fasta and LtimidusTranscriptome.pep.fasta) and the annotated database file (LtimidusTranscriptome.xlsx) can also be found in Figshare (Data Citation 2).
The European hare data used here (Data Citation 3) was previously published by Amoutzias et al.18 (NCBI Sequence Read Archive, accession number SRP055741, samples SRR1823098, SRR1863103 and SRR1863605).
Mapping statistics (Table 5), SNP call VCF file (LtimVsLeur.vcf) and population/species diagnostic SNPs tables (Supplementary Tables 1) were deposited in Figshare (Data Citation 2).
Technical Validation
RNA integrity
The quality and quantity of each RNA sample was assessed using the 260/280 and 260/230 absorbance ratios estimated by an IMPLEN P330 NanoPhotometer and RNA Integrity Number (RIN) and concentration (μgμl−1) with a Bioanalyzer 2,100 (Agilent Technologies). All samples had RIN values above 8.
RNA-Seq data quality
The Illumina HiSeq run produced a total raw output of 103 941 215 100 bp paired-end reads (207 882 430 total reads). Adapter removal and quality trimming decreased this number to 201 569 448 reads (97%) (Table 4). Final analysed reads passed the minimum quality parameters as established by FastQC.
Transcriptome assembly curation, annotation and quality
Cleaned reads were assembled into 272 183 contigs with a mean length of 594 bp and a N50 length of 839 bp (Table 4). After assembly curation with Transrate optimal cut-off >0.024, clustering with a 95% similarity threshold and open reading frame prediction, were retained 25 868 transcripts with a mean length of 842 bp and a N50 length of 1 182 (Table 4).
Annotation using a conditional reciprocal best blast hit approach results in 16 772 (65%) annotated transcripts, of which 13 641 were annotated to the rabbit transcriptome and 15 955 to the Swiss-Prot database (Fig. 3). In order to reduce the number of non-annotated transcripts, the less stringent unidirectional blast hit was added to the database. Hits were recovered for 25 549 transcripts (99%) (Fig. 3).
Number of transcripts annotated with different combinations of methods and databases: all transcripts; transcripts annotated with crb-blast against rabbit transcriptome; transcripts annotated with a unidirectional BLASTx against rabbit transcriptome; transcripts annotated with crb-blast against the Swiss-Prot database; and transcripts annotated with a unidirectional BLASTx against the Swiss-Prot database.
The mountain hare transcriptome produced in this study represents an important improvement compared to the currently available transcriptomic resources for European Lepus—L. granatensis46 and L. europaeus18 transcriptomes—as it performs better on several assembly statistics, such as reference coverage (42 versus 32% in L. granatensis and 40% in L. europaeus; using the rabbit transcriptome as reference).
Genetic variation, differentiation and gene ontology enrichment
In total, 218 057 526 reads (63%) were mapped to the filtered transcriptome—136 511 846 mountain hare reads (68%) and 81 545 680 European hare reads (57%) (see statistics in Table 5). After filtering, 159 629 high-quality SNPs were inferred, of which 41 182 (26%) were sequenced in all eleven specimens. A summary of polymorphic, shared and fixed SNPs is shown in Fig. 3. 4 672 putative species-diagnostic SNPs (considered when species presented alternative fixed alleles) were inferred (Data Citation 2,Supplementary Tables 1, also deposited in Figshare). The diagnostic power of our SNP set could be strongly reduced if any of the sequenced specimens was admixed (namely from the Alps, where the species overlap). We therefore conducted a Principal Component Analysis and a Bayesian Assignment analysis to assess our ability to separate the species. The results suggest that the analysed mountain and European hares are well differentiated with our SNP set, and only possible limited levels of admixture were found for Sample—3116 (Fig. 4). An extra table of putative species-diagnostic SNPs excluding that individual was therefore produced (Data Citation 2, Supplementary Table 4, also deposited in Figshare). 25 269 SNPs were inferred in the mountain hare, of which 12 548 and 18 591 were polymorphic in the Irish and Alpine samples respectively, and 126 were fixed between sampling localities (Data Citation 2,Supplementary Tables 5, deposited in Figshare). The ‘membrane part’ gene ontology term was found enriched in the collection of genes with fixed differences between the Irish and Alpine mountain hare samples, while terms ‘lipid metabolic process’, ‘small molecule catabolic process’, ‘extracellular space and acyl-CoA dehydrogenase activity’ were found enriched in genes with fixed differences between samples of the two species. Note however that even though the background gene set was controlled for, RNA-sequencing data does not provide an unbiased sample of information across different genes and these results may represent tissue-related functions.
(a) Relative proportion of the 41 182 SNPs mapped to the mountain hare transcriptome, summarized as polymorphic within each species and fixed or shared between L. timidus (mountain hare) and L. europaeus (European hare). The proportion is shown considering the complete L. timidus dataset (i) and only the Irish (ii) and Alpine (iii) populations. (b) STRUCTURE analysis to evaluate cluster membership and admixture proportions. Individuals are sorted by population and species. Mountain hare populations are shown in blue and European hare individuals in orange. (c) Principal Component Analysis (PCA) plot using one SNP per contig. The first principal component (PC1) splits species and the second (PC2) the sampled populations.
SNP validation
Independent SNP genotyping was performed for a random subset of 110 putative species-diagnostic SNPs from different contigs. Technical validation was considered successful for SNPs showing the expected alternative alleles, being one fixed in L. timidus (note that the sequenced L. europaeus specimens differed from the RNA-sequencing). PCR amplification was successful for 96 of the 110 target contigs (87%), 88 amplicons were successfully sequenced in both species (92%), and concordance between sequences and expected SNPs was obtained for 85 of the sequenced fragments (97%). This represents an overall validation success of 77%, which compares to studies using similar approaches47–49 (Data Citation 2; see Supplementary Table 8 for full genotyping results, and Supplementary Table 9 with the list of all primers, both deposited in Figshare). The reported accuracy of technical validation is conservative, as it is reduced by technical issues in PCR amplification and sequencing, and potential intraspecific polymorphism in the European hare (given the use of two different samples for validation), in addition to real false positives. From the validated SNPs, 73 confirmed alternate alleles in the species, but their diagnostic utility should be tested with larger population sampling.
Usage Notes
These genomic resources (which greatly extend previously available marker sets; e.g.50) will be useful for a variety of studies, particularly in the characterization of genetic diversity in mountain hare populations and on the development of hybridization monitoring tools. Note that SNPs were here inferred from an uneven and small species sample, and therefore any diagnostic genotyping assay built from this data should be first tested with adequate sample sizes from pure parental populations of the species, before being applied to hybrid zones.
Additional Information
How to cite this article: Marques, J. P. et al. Mountain hare transcriptome and diagnostic markers as resources to monitor hybridization with European hares. Sci. Data 4:170178 doi: 10.1038/sdata.2017.178 (2017).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
References
Thulin, C. G. The distribution of mountain hares Lepus timidus in Europe: A challenge from brown hares L. europaeus? Mamm. Rev. 33, 29–42 (2003).
Reid, N. & Montgomery, W. I. Is naturalisation of the brown hare in Ireland a threat to the endemic Irish hare? Biol. Environ. 107, 129–138 (2007).
Reid, N. European hare (Lepus europaeus) invasion ecology: Implication for the conservation of the endemic Irish hare (Lepus timidus hibernicus). Biol. Invasions 13, 559–569 (2011).
Caravaggi, A., Montgomery, W. I. & Reid, N. Range expansion and comparative habitat use of insular, congeneric lagomorphs: invasive European hares Lepus europaeus and endemic Irish hares Lepus timidus hibernicus. Biol. Invasions 17, 687–698 (2015).
Leach, K., Kelly, R., Cameron, A., Montgomery, W. I. & Reid, N. Expertly validated models and phylogenetically-controlled analysis suggests responses to climate change are related to species traits in the order Lagomorpha. PLoS ONE 10, e0122267 (2015).
Leach, K., Montgomery, W. I. & Reid, N. Biogeography, macroecology and species’ traits mediate competitive interactions in the order Lagomorpha. Mammal Review 45, 88–102 (2015).
Acevedo, P., Jiménez-Valverde, A., Melo-Ferreira, J., Real, R. & Alves, P. C. Parapatric species and the implications for climate change studies: A case study on hares in Europe. Glob. Chang. Biol. 18, 1509–1519 (2012).
Caravaggi, A. et al. Niche overlap of mountain hare subspecies and the vulnerability of their ranges to invasion by the European hare; the (bad) luck of the Irish. Biol. Invasions 1–20, 10.1007/s10530-016-1330-z (2016).
Thulin, C. G., Jaarola, M. & Tegelstrom, H. The occurrence of mountain hare mitochondrial DNA in wild brown hares. Mol. Ecol. 6, 463–467 (1997).
Thulin, C. G., Fang, M. & Averianov, A. O. Introgression from Lepus europaeus to L. timidus in Russia revealed by mitochondrial single nucleotide polymorphisms and nuclear microsatellites. Hereditas 143, 68–76 (2006).
Suchentrunk, F. et al. Introgressive hybridization in wild living mountain hares (L. timidus varronis) and brown hares (L. europaeus) and morphological consequences. Mamm. Biol. 70, 39–40 (2005).
Melo-Ferreira, J., Alves, P. C., Freitas, H., Ferrand, N. & Boursot, P. The genomic legacy from the extinct Lepus timidus to the three hare species of Iberia: Contrast between mtDNA, sex chromosomes and autosomes. Mol. Ecol. 18, 2643–2658 (2009).
Zachos, F. E., Ben Slimen, H., Hackländer, K., Giacometti, M. & Suchentrunk, F. Regional genetic in situ differentiation despite phylogenetic heterogeneity in Alpine mountain hares. J. Zool. 282, 47–53 (2010).
Hughes, M., Reid, N., Montgomery, I. & Prodoehl, P. Verification of hybridisation between introduced European and native Irish hares. North. Irel. Environ. Agency Res. Dev. Ser. Irel. Environ. Agency Res. Dev. Ser. 11–11 (2011).
Hamill, R. M., Doyle, D. & Duke, E. J. Spatial patterns of genetic diversity across European subspecies of the mountain hare, Lepus timidus L. Heredity 97, 355–365 (2006).
Melo-Ferreira, J. et al. Recurrent introgression of mitochondrial DNA among hares (Lepus spp.) revealed by species-tree inference and coalescent simulations. Syst. Biol. 61, 367–381 (2012).
Melo-Ferreira, J. et al. Home-loving boreal hare mitochondria survived several invasions in Iberia: the relative roles of recurrent hybridisation and allele surfing. Heredity 112, 265–273 (2014).
Amoutzias, G. D. et al. SNP identification through transcriptome analysis of the european brown hare (Lepus europaeus): Cellular energetics and mother’s curse. PLoS ONE 11, e0159939 (2016).
Andrews S. FastQC: a quality control tool for high throughput sequence data. Available online at: http://www.bioinformatics.babraham.ac.uk/projects/fastqc. 2010.
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible read trimming tool for Illumina NGS data. Bioinformatics 30, 2114–2120 (2014).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Smith-Unna, R., Boursnell, C., Patro, R., Hibberd, J. M. & Kelly, S. TransRate: Reference-free quality assessment of de novo transcriptome assemblies. Genome Res. 26, 1134–1144 (2016).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: Accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012).
Haas, B. J. et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 8, 1494–1512 (2013).
Aubry, S., Kelly, S., Kümpers, B. M. C., Smith-Unna, R. D. & Hibberd, J. M. Deep Evolutionary Comparison of Gene Expression Identifies Parallel Recruitment of Trans-Factors in Two Independent Origins of C4 Photosynthesis. PLoS Genet. 10 (6) e1004365 (2014).
Boutet, E. et al. UniProtKB/Swiss-Prot, the Manually Annotated Section of the UniProt KnowledgeBase: How to Use the Entry View. Methods Mol. Biol. 1374, 23–54 (2016).
Finn, R. D., Clements, J. & Eddy, S. R. HMMER web server: Interactive sequence similarity searching. Nucleic Acids Res. 39, W29–W37 (2011).
Finn, R. D. et al. Pfam: The protein families database. Nucleic Acids Research 42, D1, D222-D230, (2014).
Petersen, T. N., Brunak, S., von Heijne, G. & Nielsen, H. SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat. Methods 8, 785–786 (2011).
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J. Mol. Biol. 305, 567–580 (2001).
Powell, S. et al. eggNOG v3.0: Orthologous groups covering 1133 organisms at 41 different taxonomic ranges. Nucleic Acids Res. 40 (2012).
Gene Ontology Consortium. The Gene Ontology Consortium. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Kanehisa, M., Goto, S., Sato, Y., Furumichi, M. & Tanabe, M. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res. 40, D109–D114 (2012).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
McKenna, A. H. et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Gayral, P. et al. Reference-Free Population Genomics from Next-Generation Transcriptome Data and the Vertebrate-Invertebrate Gap. PLoS Genet. 9, e1003457 (2013).
Danecek, P. et al. The variant call format and VCF tools. Bioinformatics 27, 2156–2158 (2011).
Lischer, H. E. L. & Excoffier, L. PGDSpider: An automated data conversion tool for connecting population genetics and genomics programs. Bioinformatics 28, 298–299 (2012).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis. Springer-Verlag New York, 2009.
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Kopelman, N. M., Mayzel, J., Jakobsson, M., Rosenberg, N. A. & Mayrose, I. Clumpak: A program for identifying clustering modes and packaging population structure inferences across K. Mol. Ecol. Resour. 15, 1179–1191 (2015).
Mořkovský, L., Pačes, J., Rídl, J. & Reifová, R. Scrimer: Designing primers from transcriptome data. Mol. Ecol. Resour. 15, 1415–1420 (2015).
Untergasser, A. et al. Primer3-new capabilities and interfaces. Nucleic Acids Res. 40, e115 (2012).
Marques, J. P. et al. Range expansion underlies historical introgressive hybridization in the Iberian hare. Sci. Rep. 7, 40788 (2017).
Luo, X. et al. Comparative transcriptomics uncovers alternative splicing and molecular marker development in radish (Raphanus sativus L.). BMC Genomics 18, 505 (2017).
Yu, Y. et al. SNP discovery in the transcriptome of white pacific shrimp Litopenaeus vannamei by next generation sequencing. PLoS ONE 9, e87218 (2014).
Cánovas, A., Rincon, G., Islas-Trejo, A., Wickramasinghe, S. & Medrano, J. F. SNP discovery in the bovine milk transcriptome using RNA-Seq technology. Mamm. Genome 21, 592–598 (2010).
Beugin, M.-P. et al. A single multiplex of twelve microsatellite markers for the simultaneous study of the brown hare (Lepus europaeus) and the mountain hare (Lepus timidus). Ecol. Evol. 7, 3931–3939 (2017).
IUCN. IUCN Red List of Threatened Species. Version 2016-1www.iucnredlist.org (2016).
Data Citations
NCBI Sequence Read Archive SRP095715 (2017)
Marques, J. P. Figshare https://doi.org/10.6084/m9.figshare.c.3682042 (2017)
NCBI Sequence Read Archive SRP055741 (2016)
Acknowledgements
This work was funded by project HybridAdapt with reference FCT-ANR/BIA-EVF/0250/2012, supported by Portuguese National Funds through the Fundação para a Ciência e a Tecnologia, FCT. Additional support was obtained from POPH-QREN funds, from the European Social Fund and FCT (IF/00033/2014/CP1256/CT0005 FCT Investigator grant to JM-F, and SFRH/BD/115089/2016 and PD/BD/108131/2015 PhD grants to JPM and MSF respectively). Instrumentation, laboratory and computational support was provided by CIBIO NEW-GEN sequencing platform, supported by European Union's Seventh Framework Programme for research, technological development and demonstration under grant agreement no. 286431. Laboratory work at the University of Montana was performed in the UM Genomics Core, supported with a grant from the M.J. Murdock Charitable Trust. Transcriptome sequencing was performed through the Vincent J. Coates Genomics Sequencing Laboratory at University of California Berkeley, supported by NIH S10 Instrumentation Grants S10RR029668 and S10RR027303. Support was additionally obtained from FLAD (Luso-American Foundation) travel grant to MSF. We are grateful to René Gadient and local hunters in Grisons for their tremendous effort in collecting samples under Alpine conditions. Laura Claffey issued the licence to take animals for scientific purposes in Ireland (dated 31/10/2012) from the Wildlife Licensing Unit, National Parks & Wildlife Service (NPWS), Department of Arts, Heritage & the Gaeltacht, and Mr Jimi Conroy, the local NPWS Conservation Officer consented to sampling within his jurisdiction. We thank the Irish Coursing Club (ICC) for providing hares, and in particular Mr D.J. Histon, Chief Executive and Secretary, for supporting the research. Thanks also to Mr William Fitzgerald, Department of Agriculture, Food and the Marine for dispatching animals in a humane manner.
Author information
Authors and Affiliations
Contributions
J.P.M. and J.M.-F. conceived the study. J.M.-F., P.C.A., N.R., W.I.M., K.H. and H.J. organized and performed the sampling. C.C. and M.S.F. performed laboratory work at the University of Montana under the supervision of J.M.G. L.F. performed laboratory work at CIBIO under the supervision of J.M.-F. J.P.M. analysed the data. J.P.M. and J.M.-F. wrote the paper with contributions from M.S.F. and L.F. All authors read, revised and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
ISA-Tab metadata
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/ The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files made available in this article.
About this article
Cite this article
Marques, J., Ferreira, M., Farelo, L. et al. Mountain hare transcriptome and diagnostic markers as resources to monitor hybridization with European hares. Sci Data 4, 170178 (2017). https://doi.org/10.1038/sdata.2017.178
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2017.178
This article is cited by
-
A new HaCV-EBHSV recombinant lagovirus circulating in European brown hares (Lepus europaeus) from Catalonia, Spain
Scientific Reports (2024)
-
Mountain- and brown hare genetic polymorphisms to survey local adaptations and conservation status of the heath hare (Lepus timidus sylvaticus, Nilsson 1831)
Scientific Data (2022)
-
Bidirectional hybridisation and introgression between introduced European brown hare, Lepus europaeus and the endemic Irish hare, L. timidus hibernicus
Conservation Genetics (2022)