Abstract
Segmentation of anatomical structures is valuable in a variety of tasks, including 3D visualization, surgical planning, and quantitative image analysis. Manual segmentation is time-consuming and deals with intra and inter-observer variability. To develop a deep-learning approach for the fully automated segmentation of the inner ear in MRI, a 3D U-net was trained on 944 MRI scans with manually segmented inner ears as reference standard. The model was validated on an independent, multicentric dataset consisting of 177 MRI scans from three different centers. The model was also evaluated on a clinical validation set containing eight MRI scans with severe changes in the morphology of the labyrinth. The 3D U-net model showed precise Dice Similarity Coefficient scores (mean DSC-0.8790) with a high True Positive Rate (91.5%) and low False Discovery Rate and False Negative Rates (14.8% and 8.49% respectively) across images from three different centers. The model proved to perform well with a DSC of 0.8768 on the clinical validation dataset. The proposed auto-segmentation model is equivalent to human readers and is a reliable, consistent, and efficient method for inner ear segmentation, which can be used in a variety of clinical applications such as surgical planning and quantitative image analysis.
Similar content being viewed by others
Introduction
The inner ear, also known as the labyrinth, is a complex structure located in the temporal bone. It roughly consists of the cochlea, the vestibule, and the semi-circular canals. Understanding changes and variations within these structures can help diagnose and predict a number of conditions1, such as inflammatory and neoplastic processes. Technological developments in imaging techniques have allowed (neuro)radiologists to evaluate the human labyrinth, with recent advances increasing the level of detail1. Moreover, applications of artificial intelligence and the quantitative assessment of medical images for the non-invasive exploration of anatomical structures and the classification of diseases have remarkably increased in recent years2.
The process of the automated extraction and analysis of large amounts of quantitative information from medical images is known as radiomics3,4. A recent study investigated the value of radiomics for diagnosis of Meniere’s disease (MD), an inner ear disorder characterized by episodic vertigo spells, hearing loss, and tinnitus5. Other labyrinthine disorders such as sensorineural hearing loss might benefit from quantitative image analysis as well6.
Image segmentation is a critical step to work towards fully automated diagnostic tools for inner ear disorders. Manual segmentation requires experienced readers, is time-consuming and prone to intra-and inter-observer variability7,8,9. Over the past years, several automatic and semi-automatic inner ear segmentation methods were proposed for both MRI and CT imaging10,11,12,13,14, including region-growing, thresholding and edge detection15, model-based10,14, atlas-based12,13 and machine-learning techniques11. The inner ear’s small and complex structure makes segmentation challenging, especially in MR imaging due to non-homogenous image intensities10,11.
Recent work proposed a statistic shape model (SSM) for inner ear segmentation in MR images (13). However, the proposed methodology present a high computational burden, both in term of time and cost. Another recently published segmentation model showed very good agreement between an atlas-based segmentation and the manual gold standard12, yet requires manual intervention. Additionally, segmentation performance of atlas-based methods decreases for complex structures with variable shape and size16.
Recent studies have demonstrated the successful application of deep learning techniques for detection, segmentation, and classification tasks in the medical field17. Among deep learning techniques, the U-Net architecture is a specific type of convolutional neural network (CNN) consisting of multilayer neural networks. These networks have been implemented successfully, especially for auto-segmentation in medical images18,19. Although U-Net based deep learning approaches do exist for segmentation of the inner ear18,20, they lack the incorporation of anatomical variations, pathological situations or missing anatomical structures, which are part of daily clinical practice. Hence, there is currently no fully automated, generic segmentation method for the inner ear to meet the growing demand for the developments in 3D visualization and quantitative image assessments.
Therefore, this study’s objective was to develop a deep-learning approach for the automatic segmentation of the inner ear in clinical MR images, focusing on the robustness of the method in varying clinical situations, and to evaluate its performance and generalizability with manual segmentation as reference.
Materials and methods
Ethical considerations
This study was performed in accordance with the guidelines outlined by Dutch and Belgian legislation. MRI scans were collected and fully anonymized by the local investigators of four centers. Ethics committee of University Hospital Antwerp approved the study (Approval number—17/09/093) and written informed consent was obtained from the participants. The other centers waived the ethics approval due to the retrospective nature and full anonymization of the data according to the Medical Research involving Human Subjects act (WMO).
Automatic segmentation workflow
The workflow applied in this study consisted of four steps and is illustrated in Fig. 1. Each step of the workflow is detailed in the following paragraphs. Additional details on the model architecture, training, validation and testing can be found in Supplementary Sect. 1 of the Supplementary Information.
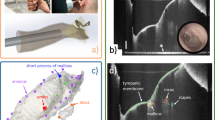
The workflow of autosegmentation of the inner ear in this study, graphically presented in four steps. (A) The image acquisition from four different centers divided into training, validation and an independent test set. (B) Manual segmentation of the labyrinth and pre-processing steps consisting of isotropic voxel resampling, intensity rescaling and center cropping. (C) Extending the data set (data augmentation) by flipping and rotating the input images and training of the model. (D) Validation and testing the model on an independent test cohort.
Training dataset
A total of 1203 images of patients who underwent an MRI scan of the cerebellopontine angle for diverse neuro-otological indications in the period of December 2015 to April 2019 in Maastricht Medical University center (center A) were collected and fully anonymized. All high resolution T2-weighted images were acquired in 1.5 and 3 T (T) MRI scanners, from different vendors with a variety of high-resolution T2-weigted sequences (3D cochlea, DRIVE, SPC_TRA_ISO), with local optimized protocols. MRI scans of the cerebellopontine angle were included if they allowed labyrinth visualization with at least a portion of the labyrinth recognizable and suitable for manual segmentation. MRI scans, which did not allow a clear manual segmentation, were excluded from this study. In total, 259 MRI images were excluded due to unsuitable sequences (DWI, T1, SURVEY MST), poor quality, or skewed MR images. The final training dataset included MRI scans of 944 cases (Table 1).
Validation and test dataset
The validation dataset included MRI scans of 99 cases collected from Maastricht University Medical Center + (center A) in the period from 2005 to 2015 (Table 1). MRI scans collected from 3 different centers, University Hospital Antwerp (center B), Viecure Hospital Venlo (center C), and Apeldoorn dizziness center (center D) during the period of 2005–2017 (Table 1) were used as an independent Test dataset. Both validation and test dataset consisted of T2-weighted MR images of the cerebellopontine angle of patients with uni- or bilateral definitive Meniere’s disease and idiopathic asymmetric sensorineural hearing loss.
Manual segmentation
A team of six readers was trained by the second author (MvdL), an experienced clinician and researcher in inner ear imaging, to manually segment the labyrinth on both sides in 3D Slicer 4.8.119. Manual segmentation was facilitated by intensity-based thresholding and region-growing algorithms. The original MRI scans and the manually segmented masks were visualized by 3D maximum intensity projections as shown in Fig. 2. This provided an overview of the manually segmented results which allowed for thorough quality assessment. All segmentations were curated by the experienced reader (MvdL), where any missing or incorrectly segmented masks were re-segmented by the experienced reader (MvdL). The final segmentation results served as the ground truth for training the CNN. The independent validation and test datasets were segmented and curated by the experienced reader (MvdL).The resulting manual segmentations on the test dataset was used as the ground truth (reference standard).
Pre-processing
In order to generate homogeneous MRI volumes as input for the model, the following pre-processing steps were performed. Firstly, all volumes were resampled by B-spline interpolation to an isotropic voxel size of 0.45 mm. Secondly, the intensities of the MRI volumes were normalized to range [0–1] using the minimum and maximum intensity of each volume. Lastly, since the model’s architecture required inputs of the same dimensions, a center crop of 256 × 256 × 64 pixels was obtained from the pre-processed volumes. This crop size was large enough to contain contextual information of the inner ear. Images smaller than 256 × 256 pixels in the transversal plane and 64 pixels in slice direction were padded with zeros. More details can be found in Supplementary Information Sect. 2.
Model architecture
The model’s architecture is based on a classical 3D U-net21, as illustrated in Fig. 3a,b. It comprises an encoder, a decoder block and skip connections. The encoder network is a contracting path with convolution layers, which extracts high-level features, decreasing the spatial resolution at each layer. The decoder network is an expanding path, which increases the spatial resolution by up-sampling and uses the feature information to segment the pixels corresponding to the Region of interest. Skip connections, between encoder and decoder, allow retrieval of fine details, which might be lost during spatial down-sampling.
(a) The proposed 3D U-Net based architecture used in the study. MRI volumes, at multiple scales, were provided as input to the encoder network. The decoder network outputs a score to classify each voxel as inner ear or not. Notations in blue text (a × a × a × b) highlight the spatial resolution (a × a × a) and the feature map count (b). X block repetitions, IN instance normalization, Conv convolution kernel, ReLU rectified linear unit, 3 × 3 × 3 the size of the 3D CNN kernels. (b) Components of Attention Gating Block. The block receives as inputs, the up-sampled output feature map at each scale in the decoder and the feature map from each scale of the encoder. Attention coefficients generated, scale the input feature maps from the encoder.
The model’s architecture was adapted with attention gates, as the relevant features of the inner ear showed large shape variability and were very small compared to the surrounding structures22.
The attention gates highlight the regions that correspond to inner ear and suppress the regions that correspond to background. The highlighted features are propagated by the skip connections from the deep stages of contracting paths to the expanding paths. More specifically, attention Gates are used to propagate the important spatial information corresponding to inner ear from the encoding to the decoding part of the model. As shown in the Fig. 3b, the input feature maps from the encoder part of the network are scaled by the attention coefficients generated by the Attention Gates, thereby outputting the features relevant to the inner ear. The scaled features are then concatenated with the up-sampled output feature maps at each level in the decoder part of the network.
Since different components of inner ear are more easily accessible at different scales, we additionally input the same volume at three different scales along the encoder path, which has been previously described as an input image pyramid by Oktay et al.22.
Other network parameter changes included an increase in number of convolutional filters from 16 to 128 in the encoder network. Each Maxpooling layer reduced the image spatial resolution by a factor of two. Along the decoder path, transposed convolutions were used for up-sampling which increased the image size by a factor of two at each layer. All the convolutional blocks included 3D convolutions23, ReLu activation24 and Instance Normalization25.
Training, validation and testing
The model was trained with the pre-processed volumes and their corresponding ground truth labels of the training dataset. Randomly selected input volumes were augmented by vertical flipping or rotation during training. The network weights were initialized by using the He-normal initialization method26 and updated by using the Adam optimizer27 at an initial learning rate of 1e−4.
Since the number of positive voxels (i.e. part of the inner ear) and the negative voxels were highly imbalanced, Tversky loss28 was used as an objective loss function while training the model, which penalized false negatives more than false positives at a false positive penalty score (β) of 0.3 and a false negative penalty score (α) of 0.7. This approach emphasizes learning features corresponding to the positive voxels. The loss was calculated in a mini batch of two images per iteration and at the end of each epoch, Tversky loss was calculated on the model’s predictions on the validation dataset to ensure validation loss convergence (i.e., decrease in validation loss).
The final model’s performance was evaluated on the multicentric, independent test dataset.
Outcome measurements
The main outcomes of this study were the Dice similarity coefficient (DSC), true positive rate (TPR), false positive rate (FPR), false negative rate (FNR) and false discovery rate (FDR).
As a secondary outcome a subjective evaluation of clinical validation was performed by the second author (MvdL) in consensus with an experienced neuroradiologist (A.A.Postma). Towards clinical implementation, it is critical that a deep learning model is able to segment the inner ear under all conditions, including those that might alter the shape of the inner ear (e.g., by pathology). Therefore, eight MR images, with their corresponding masks, were selected by the second author (MvdL) in which the signal intensities of the inner ear were altered either by pathology or post-therapeutic changes. These scans were left out from the training dataset and were used for clinical validation of the performance of the model.
Qualitative assessment: in silico clinical study
An in silico clinical study was performed to make a qualitative comparison between manual and model-generated segmentations for 50 MRI volumes randomly selected from the test cohort. An in-house developed software was used to display pairs of segmentations (automated vs manual), at randomized screen positions (left or right) blinded to the participants, overlaid on MRI images, as shown in Fig. 4. The software allowed for scrolling through all image slices and adjustment of window level settings. We enrolled 7 participants (3 computer scientists working in the field of medical imaging and 4 radiologists with an average experience of 2.5 years). For each image, the participants were asked to select their preferred segmentation. For each participant, the qualitative preference score was defined as the percentage of cases with preferred automated segmentation.
Results
The final training dataset included MRI scans of 944 cases (489 men, 455 women, aged 41–74; mean age 57 ± 15). The final validation dataset included MRI scans of 99 cases from center A (69 men, 30 woman, aged 43–69; mean age 56 ± 13) and 177 cases from centers B, C and D (79 men, 59 women, aged 45–74; mean age 59 ± 14).
Segmentation performance
The segmentation accuracy was evaluated against the ground truth by assessing the DSC. DSC measures the overlap between the reference and the model’s output. The overall average metrics of segmentation accuracy, DSC, TPR, FNR, FDR and FPR are summarized in Table 2. Figure 5 shows a comparison between ground truth volume and predicted true positive volume on the validation and test dataset. Figure 6 shows the distribution of DSCs on validation and test dataset. The correlation between the true positive volume and ground truth volume was also investigated. In Fig. 7a,b, agreements for ground truth volume and predicted volume are graphically displayed by Bland–Altman plots. Figure 8a,b shows an example of a well predicted and poorly predicted segmentation.
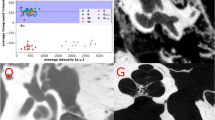
The quantitative analysis showing linear correlations between the ground truth volume and the predicted true positive volume for the validation (plot in blue) and the test sets (plots in orange). The plot of center D shows 2 clear outliers which do not fit the trendline. This suggests under-segmentation of the inner ear in 2 cases belonging to the test cohort from Center D.
Distribution of DSC on the validation (blue curve) and the test dataset (orange curve). The distribution corresponding to Center C and D shows outliers (DSC < 0.7) which means less overlap between ground truth and predicted segmentation. The distribution also shows that the majority of the predictions have DSC between 0.8 to 1.0.
(a) Bland–Altman plot for inner ear volume of the entire test cohort showing percentage difference between predicted volume (PV) and ground truth volume (GTV) as a function of average of Ground Truth and Predicted Volume. The solid line shows the mean difference and the dotted line shows the limits of agreement. PV Predicted volume of inner ear, GTV ground truth volume of inner ear. The plot shows five clear outliers (red dots) with three cases which were under-segmented by 20%, 40% and 60% and two cases which were over-segmented by 40% and 60% respectively. The plot also shows the relationship between the DSC metrics and the level of under/over segmentation percentage. The outliers correspond to the DSC ≤ 0.80. (b) Bland–Altman plot for inner ear volume of the entire test cohort showing percentage difference between predicted volume (PV) and ground truth volume (GTV) as a function of average of ground truth and predicted volume after excluding the outliers shown in Fig. 5A (DSC ≤ 0.80). The solid line shows the mean difference and the dotted line shows the limits of agreement. PV Predicted volume of inner ear, GTV ground truth volume of inner ear. The plot shows that the model, on an average tends to over-segment by 9%.
(a) Example of a well predicted segmentation. The first row denotes the ground truth segmentation. The second row contains the model’s segmentation. (1a) Ground truth, axial plane. (1b) Ground truth, sagittal plane. (2c) Ground truth, coronal plane. (2a) Predicted mask, axial plane. (2b) Predicted mask, sagittal plane. (2c) Predicted mask, coronal plane. DSC: 0.92, ground truth volume: 465.37 mm3, true positive volume: 445.32 mm3, true positive Rate: 95.69%, false negative rate: 4.3%. False discovery rate: 11.7%. (b) Example of a poor segmentation. The first row denotes the ground truth segmentation. The second row contains the model’s segmentation. (1a) Ground truth, axial plane. (1b) Ground truth, sagittal plane. (2c) Ground truth, coronal plane. (2a) Predicted mask, axial plane. (2b) Predicted mask, sagittal plane. (2c) Predicted mask, coronal plane. DSC: 0.48, ground truth volume: 406.05 mm3, true positive volume: 137.96 mm3, true positive rate: 33.97%, false negative rate: 66.02%. False discovery rate: 1.5%.
Performance on clinical validation dataset
On the held-out clinical validation dataset, the model achieved an average DSC of 0.876, TPR of 87.86%, FDR of 15.2% and FNR of 12.13%. The automated segmentations on this dataset are included in Supplementary Information. It includes labyrinths in which parts of the semi-circular canals, the vestibule or the cochlea were missing or not properly displayed. As an example, an MRI scan with a vestibular schwannoma (a tumorous process growing from the vestibular nerve) was included in Fig. 9.
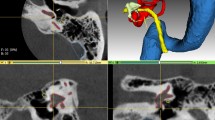
(a) Example of one of the clinical validation MRI scans in the axial and coronal plane. This case shows the presence of a vestibular schwannoma after a translabyrinthine resection on the right side. Therefore, the right semi-circular canals and vestibule are not segmented. DSC: 0.8973, ground truth volume: 316.11 mm3, true positive volume: 294.69 mm3, true positive rate: 93.22%, false negative rate: 6.77%. False discovery rate: 7.3%. (b) The 3D volume rendering of the ground truth and the predicted mask. The semi-circular canals and the vestibule of the right inner ear were not displayed on MRI. The model has correctly not segmented the semi-circular canals and the vestibule. AD auriculum dextra, AS auriculum sinistra.
Qualitative assessment: in silico clinical study
On average, the participants preferred the automated segmentation in 67% of the cases. A paired one-sided t-test for the hypothesis that this average score is greater than 50% was significant (p = 9.82474e−17), indicating that expert users preferred the segmentations generated by the proposed model over the manual segmentations.
Discussion
In this work, a first proof-of-concept of an artificial intelligence based model for the fully automatic segmentation of the inner ear on MRI was demonstrated and validated.
The proposed model showed high performance, with a mean DSC of 0.87 between the manual and the automated segmentation validated across images from three different centers. The mean TPR of 91.5% implies accurate segmentation of the inner ear without significant over or under segmentation as indicated by the low FDR and FNR metrics (14.8% and 8.49% respectively).
The in silico based qualitative analysis showed that on an average, the expert users (radiologists and computer scientists) are more likely to prefer model generated segmentations over manual segmentations. The Bland–Altman plot (Fig. 6) shows 5 outliers. The fact that the slice thicknesses of those scans were high (mean slice thickness—1.1 mm) compared to the mean slice thickness of the training cohort (0.32 mm) could explain the miss-segmentations for these cases. Also, the three scans from the Center D contained either moving artifacts or had tight margins around the labyrinth, which might explain the lower limits of agreements. All the MRI scans of the Center C had noticeably more hyperintense areas at the apex of the pas petrosa compared to the other centers. Although this might have ‘challenged’ the model, it does not explain why only two out of 21 scans had lower DSCs.
A prior study, that used deep learning to facilitate the auto-segmentation of the inner ear, compared the performance of a 3D Fully Connected Network (FCN) to a 2D-FCN20. The study reported an overall DSC of 0.66 and 0.58 when using 3D-FCN and 2D-FCN, respectively. Another recent study reported a high DSC of 0.95 using SSMs based level set10. However, their model was evaluated on a small dataset (10 cases out of 23 cases were held out for testing) and no independent validation was performed. Directly comparing the present approach with the already published methods in terms of DSC is not possible due to differences in datasets. Nevertheless, it is worth noting that our presented method achieves state of the art performance, which can be ascribed to the robust deep learning approach combined with a wide and varied dataset, both for training and validation, an aspect often neglected in similar studies.
There are several important strengths of this study. First of all, the model was trained on a diverse set of MR images of the cerebellopontine region. Although all MR images of the training dataset were collected in one center, they were acquired over a wide time span (2015–2019) and include different acquisition and reconstruction protocols29. Next to this, the training dataset was manually segmented by five independent readers. Therefore, the model learned to eliminate noise in the manually segmented labels caused by inter-reader variability. These methodological aspects resulted in a model that is well generalizable, which is reflected in the high-validation performance. Past studies have shown high inter-reader and intra-reader variability on medical image segmentation tasks30,31. Our method’s consistency (i.e., no segmentation variability) alleviates this issue. Additionally, the interaction time was approximately 10 min per case for manual segmentation by an experienced reader compared to only 6.5 s for the automatic segmentation.
One of the most important strengths of this study is the evaluation on the MR images containing deviant morphological shapes and decreased signal intensities of the labyrinth caused by cerebellopontine pathology. On this held-out clinical validation dataset, the model proved to generalize well with an average DSC and TPR of 0.8768 and 87.86% respectively. So far, previous auto-segmentation studies have trained their models on normal ears or small datasets (13–15). To the best of our knowledge, our study is the first to assess generalizability with respect to pathologies.
Limitations
Several limitations of this study should be noted. First of all, the most important limitation is the lack of a gold standard for manual segmentations from highly experienced neuroradiologists. Due to the extent of the segmentation process, manual segmentation of approximately 1500 labyrinths by one or more senior radiologist was not feasible. Therefore, in this study the authors chose to work with independent readers who were trained and supervised by an experienced clinical researcher in inner ear imaging (MvdL) to generate a first proof of concept. This could have induced noise in the manual segmentations. Also, the intra- and inter-observer variability of the segmentation team was not evaluated. Although manual segmentation was performed under strict supervision of the second author and a curating process was performed to detect incorrectly segmented masks, the quality of the manual segmentation could not be fully guaranteed. Since the manually segmented masks were considered as the reference standard for the evaluation of the model, lower DSC scores might have indicated better automated segmentation compared to manual segmentation.
Nevertheless, efforts have been made to contain this limitation by training a deep learning architecture with large number of parameters and applying Early Stopping to prevent overfitting on the noise in the manual segmentation. Previous studies have proved that overparameterized networks are more robust against noisy labels when Early Stopping is applied32.
Given the very small are occupied by the inner ear in the whole MRI volume, the performance of our model might be further improved by applying bounding box detection33 or shape identification34 prior to automated segmentation especially for abnormal cases.
Secondly, poor generalizability is the most common problems pertaining to deep learning models29. In this study, attempts were made to prevent overfitting by training the model on a large dataset from one center and testing its generalizability by holding out 3 independent validation cohorts. Although the overall DSC scores were markedly high, the model performed poorly and failed to generalize in five cases out of 177 (3 cases from center C and 2 cases from center D had DSC < 0.70). This situation could have been mitigated by training the model on all of the centers. This would have made the training dataset more diverse (e.g., in terms of image acquisition and reconstruction) and the model’s performance could have been evaluated by cross-validation techniques (i.e., holding out 20–30% of the data from each center for a single validation test data). However, this would degrade the credibility of the generalizability of the model due to concerns regarding overfitting.
Lastly, the model was trained and evaluated on datasets that included only Dutch and Belgian population. The generalizability of the model on MRI images from an international cohort is currently unexplored.
Clinical implications and future perspectives
The future clinical advantages of automated 3D image segmentation of the inner ear are versatile. Image segmentation can be used for 3D visualization, allowing a better understanding of the spatial relations and morphological changes within the inner ear, assisting radiologists in the diagnostic process and providing tools for surgical planning35 or learning purposes36. Previous studies have proven the usability of auto-segmentation for pre-operative planning of cochlear implant surgery using CT imaging37 and for the diagnosis of adolescent idiopathic scoliosis using MRI imaging11. Our model proved to be efficient on MRI imaging. However, the proposed methodology can be easily leveraged for similar auto-segmentation applications on different imaging modalities.
Nowadays, quantitative analysis of the inner ear is gaining more importance. Techniques like radiomics6, volumetric assessment of fluid compartments in the labyrinth12,38 and the analysis of the morphoanatomy for the vestibular system11 are used to aid diagnosis of vestibular diseases. Radiomics refers to the process of the automated extraction and analysis of large amounts of quantitative features from medical images. These features are sometimes not perceptual for the human eye and might contain information that reflects underlying tissue heterogeneity and pathophysiology4,39. Quantitative image features involve descriptors of shape, size, volume, intensity distributions and texture heterogeneity patterns39.
A histological feature strongly associated with Meniere’s disease is endolymphatic hydrops (EH), a distention of the endolymphatic compartment in the inner ear40. In conventional MRI, the endolymphatic compartment cannot be distinguished from the perilymphatic compartment, and thus, EH is not depicted41. The differences found in radiomic features between MD and controls could hypothetically be explained by the different composition of the fluids in the labyrinth, causing a different distribution of signal intensities5. Possibly, EH is captured in the quantitative image features due to damage to or morphological changes to the endolymphatic space. Since Meniere’s disease is still a clinical diagnosis challenge42, discovering distinctive image features might benefit the diagnostic trajectory of MD. Another study showed that cochlea CT image features can be useful biomarkers for predicting sensorineural hearing loss in patient with head and neck cancers which received chemoradiation therapy6. Different machine learning methods were used for feature selection, classification and prediction. The advantage of using machine learning in combination with radiomics is that the analysis of the labyrinth could be done autonomously in the future5. However, for both studies, setting a Region Of Interest (ROI) by manual segmentation was necessary. The fully automated segmentation of the inner ear contributes to efficient research on quantitative image analysis of the inner ear.
Next to analyses on conventional MRI and CT imaging, the volumetric assessment of fluid compartments in the labyrinth is also promising for vestibular research38. Contrast-enhanced MR imaging allows the in vivo confirmation and quantification of endolymphatic hydrops12,43.
Several studies investigated the value of the 3D volumetric assessment of the endolymphatic space (ELS) to better monitor EH in vivo, for example in therapeutic trials in Meniere’s disease, and to better compare the ELS in patients with different otological diseases38,44,45. However, the 3D reconstruction were all rendered semi-automatic. Due to this time-consuming process, the applications for volumetric assessment are yet more scientifically than clinically relevant. A recent study proposed atlas-based segmentation for the volume-based quantification of the fluid spaces of the inner ear12. Which created fast, standardized (auto)segmentation. Further research is necessary to explore the option of the proposed U-net model can be leveraged for contrast-enhanced imaging as well, to facilitate volumetric assessment of the ELS in clinic.
Auto-segmentation in its current form, is a step towards fully automated diagnostic tools for inner ear disorders.
Conclusion
In this study, a working first proof-of-concept is demonstrated regarding the fully automatic segmentation of the inner ear using deep learning. Overall, the proposed auto-segmentation model is equivalent to manual segmentation and is a reliable, consistent, and efficient method for inner ear segmentation which can be used in a variety of clinical applications, such as 3D visualization, surgical planning and quantitative image analysis. Auto-segmentation of the inner ear in its current form, might open doors towards automated diagnostic tools for inner ear disorders.
Abbreviations
- DSC:
-
Dice similarity coefficient
- TPR:
-
True positive rate
- FNR:
-
False negative rate
- FPR:
-
False positive rate
- FDR:
-
False discovery rate
References
Pyykkö, I., Zou, J., Gürkov, R., Naganawa, S. & Nakashima, T. Imaging of temporal bone. Adv. Otorhinolaryngol. https://doi.org/10.1159/000490268 (2019).
Sollini, M., Antunovic, L., Chiti, A. & Kirienko, M. Towards clinical application of image mining: A systematic review on artificial intelligence and radiomics. Eur. J. Nucl. Med. Mol. Imaging https://doi.org/10.1007/s00259-019-04372-x (2019).
Kumar, V. et al. Radiomics: The process and the challenges. Magn. Reson. Imaging 30, 1234–1248 (2012).
Gillies, R. J., Kinahan, P. E. & Hricak, H. Radiomics: Images are more than pictures, they are data. Radiology https://doi.org/10.1148/radiol.2015151169 (2016).
van den Burg, E. L. et al. An exploratory study to detect ménière’s disease in conventional MRI scans using radiomics. Front. Neurol. 7, 190 (2016).
Abdollahi, H. et al. Cochlea CT radiomics predicts chemoradiotherapy induced sensorineural hearing loss in head and neck cancer patients: A machine learning and multi-variable modelling study. Phys. Med. 45, 198–204 (2018).
Nogovitsyn, N. et al. Testing a deep convolutional neural network for automated hippocampus segmentation in a longitudinal sample of healthy participants. Neuroimage 197, 589–597 (2019).
Men, K. et al. Fully automatic and robust segmentation of the clinical target volume for radiotherapy of breast cancer using big data and deep learning. Phys. Med. 50, 13–19 (2018).
Akkus, Z., Galimzianova, A., Hoogi, A., Rubin, D. L. & Erickson, B. J. Deep learning for brain MRI segmentation: State of the art and future directions. J. Digit. Imaging 30, 449–459 (2017).
Zhu, S. et al. 3D automatic MRI level set segmentation of inner ear based on statistical shape models prior. In Proceedings—2017 10th International Congress on Image and Signal Processing, BioMedical Engineering and Informatics, CISP-BMEI 2017 (2018). https://doi.org/10.1109/CISP-BMEI.2017.8301973.
Shi, L. et al. Automatic MRI segmentation and morphoanatomy analysis of the vestibular system in adolescent idiopathic scoliosis. Neuroimage https://doi.org/10.1016/j.neuroimage.2010.04.002 (2011).
Kirsch, V., Nejatbakhshesfahani, F., Ahmadi, S. A., Dieterich, M. & Ertl-Wagner, B. A probabilistic atlas of the human inner ear’s bony labyrinth enables reliable atlas-based segmentation of the total fluid space. J. Neurol. https://doi.org/10.1007/s00415-019-09488-6 (2019).
Powell, K. A. et al. Atlas-based segmentation of temporal bone anatomy. Int. J. Comput. Assist. Radiol. Surg. https://doi.org/10.1007/s11548-017-1658-6 (2017).
Reda, F. A., Noble, J. H., Labadie, R. F. & Dawant, B. M. An artifact-robust, shape library-based algorithm for automatic segmentation of inner ear anatomy in post-cochlear-implantation CT. In Medical Imaging 2014: Image Processing (2014). https://doi.org/10.1117/12.2043260.
Todd, C., Kirillov, M., Tarabichi, M., Naghdy, F. & Naghdy, G. An analysis of medical image processing methods for segmentation of the inner ear. In Proceedings of the IADIS International Conference Computer Graphics, Visualization, Computer Vision and Image Processing 2009, CGVCVIP 2009. Part of the IADIS MCCSIS 2009 (2009).
Sharma, N. et al. Automated medical image segmentation techniques. J. Med. Phys. https://doi.org/10.4103/0971-6203.58777 (2010).
Litjens, G. et al. A survey on deep learning in medical image analysis. Med. Image Anal. 42, 60–88 (2017).
Fauser, J. et al. Toward an automatic preoperative pipeline for image-guided temporal bone surgery. Int. J. Comput. Assist. Radiol. Surg. https://doi.org/10.1007/s11548-019-01937-x (2019).
Fedorov, A. et al. 3D Slicer as an image computing platform for the Quantitative Imaging Network. Magn. Reson. Imaging 30, 1323–1341 (2012).
Gong, Z., Li, X., Zhou, L. & Zhang, H. A 3D fully convolutional network based semantic segmentation for ear computed tomography images. In Proceedings—2018 11th International Congress on Image and Signal Processing, BioMedical Engineering and Informatics, CISP-BMEI 2018 (2019). https://doi.org/10.1109/CISP-BMEI.2018.8633242.
Çiçek, Ö., Abdulkadir, A., Lienkamp, S. S., Brox, T. & Ronneberger, O. 3D U-net: Learning dense volumetric segmentation from sparse annotation. In Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics) (2016). https://doi.org/10.1007/978-3-319-46723-8_49.
Oktay, O., et al. Attention U-Net: learning where to look for the Pancreas (2018). https://arxiv.org/abs/1804.03999v3
Ji, S., Xu, W., Yang, M., Yu, K. 3D convolutional neural networks for human action recognition. IEEE Trans. Pattern Anal. Mach. Intell. 35(1), 221–231 (2013). https://doi.org/10.1109/TPAMI.2012.59.
Agarap, A. F. Deep Learning using Rectified Linear Units (ReLU). https://arxiv.org/abs/1803.08375 (2019).
Ulyanov, D., Vedaldi, A., Lempitsky, V. Instance normalization: the missing ingredient for fast stylization. https://arxiv.org/abs/1607.08022 (2017).
He, K., Zhang, X., Ren, S. & Sun, J. Delving deep into rectifiers: Surpassing human-level performance on imagenet classification. In Proceedings of the IEEE International Conference on Computer Vision. https://doi.org/10.1109/ICCV.2015.123 (2015).
Kingma, D. P. & Ba, J. L. Adam: A method for stochastic optimization. In 3rd International Conference on Learning Representations, ICLR 2015—Conference Track Proceedings (2015).
Abraham, N. & Khan, N. M. A novel focal tversky loss function with improved attention u-net for lesion segmentation. In Proceedings—International Symposium on Biomedical Imaging (2019). https://doi.org/10.1109/ISBI.2019.8759329.
Caruana, R., Lawrence, S. & Giles, L. Overfitting in neural nets: Backpropagation, conjugate gradient, and early stopping. In Advances in Neural Information Processing Systems (2001).
Norman, B., Pedoia, V. & Majumdar, S. Use of 2D U-net convolutional neural networks for automated cartilage and meniscus segmentation of knee MR imaging data to determine relaxometry and morphometry. Radiology 288, 177–185 (2018).
Yasaka, K., Akai, H., Abe, O. & Kiryu, S. Deep learning with convolutional neural network for differentiation of liver masses at dynamic contrast-enhanced CT: A preliminary study. Radiology 286, 887–896 (2018).
Li, M., Soltanolkotabi, M. & Oymak, S. Gradient Descent with Early Stopping is Provably Robust to Label Noise for Overparameterized Neural Networks. arXiV (2019).
Zhuang, X. et al. Evaluation of algorithms for multi-modality whole heart segmentation: An open-access grand challenge. Med. Image Anal. 58, 101537 (2019).
Yang, G. et al. Simultaneous left atrium anatomy and scar segmentations via deep learning in multiview information with attention. Future. Gener. Comput. Syst. 107, 215–228 (2020).
Heutink, F. et al. Multi-scale deep learning framework for cochlea localization, segmentation and analysis on clinical ultra-high-resolution CT images. Comput. Methods Progr. Biomed. 191, 105387 (2020).
Ferreira, A., Gentil, F. & Tavares, J. M. R. S. Segmentation algorithms for ear image data towards biomechanical studies. Comput. Methods Biomech. Biomed. Eng. https://doi.org/10.1080/10255842.2012.723700 (2014).
Noble, J. H., Labadie, R. F., Majdani, O. & Dawant, B. M. Automatic segmentation of intracochlear anatomy in conventional CT. IEEE Trans. Biomed. Eng. https://doi.org/10.1109/TBME.2011.2160262 (2011).
Gürkov, R. et al. MR volumetric assessment of endolymphatic hydrops. Eur. Radiol. https://doi.org/10.1007/s00330-014-3414-4 (2015).
Lambin, P. et al. Radiomics: Extracting more information from medical images using advanced feature analysis. Eur. J. Cancer 48, 441–446 (2012).
Merchant, S. N., Adams, J. C. & Nadol, J. B. Pathophysiology of Ménière’s syndrome: Are symptoms caused by endolymphatic hydrops?. Otol. Neurotol. https://doi.org/10.1097/00129492-200501000-00013 (2005).
Lingam, R. K., Connor, S. E. J., Casselman, J. W. & Beale, T. MRI in otology: Applications in cholesteatoma and Ménière’s disease. Clin. Radiol. https://doi.org/10.1016/j.crad.2017.09.002 (2018).
Lopez-Escamez, J. A. et al. Diagnostic criteria for Menière’s disease. J. Vestib. Res. Equilib. Orientat. https://doi.org/10.3233/VES-150549 (2015).
Naganawa, S. & Nakashima, T. Visualization of endolymphatic hydrops with MR imaging in patients with Ménière’s disease and related pathologies: Current status of its methods and clinical significance. Jpn. J. Radiol. https://doi.org/10.1007/s11604-014-0290-4 (2014).
Homann, G. et al. Semi-quantitative vs. volumetric determination of endolymphatic space in Menière’s disease using endolymphatic hydrops 3T-HR-MRI after intravenous gadolinium injection. PLoS ONE https://doi.org/10.1371/journal.pone.0120357 (2015).
Inui, H., Sakamoto, T., Ito, T. & Kitahara, T. Magnetic resonance-based volumetric measurement of the endolymphatic space in patients with Meniere’s disease and other endolymphatic hydrops-related diseases. Auris Nasus Larynx https://doi.org/10.1016/j.anl.2018.11.008 (2019).
Acknowledgements
The authors would like to thank the segmentation team, consisting of the following medical students: J.G.V Muijsenberg, D.M.F. Hoeben, L. Mulder, S.V. Feitsma and Y. L. Leroux, for their contributions to this work. The authors would like to thank Fabio Bottari for the revision of the manuscript. Furthermore, the authors acknowledge financial support from MUMC+ and VieCurie Venlo, ERC advanced grant (ERC-ADG-2015, n° 694812)—Hypoximmuno and the European Program H2020 (PREDICT-ITN-n° 766276), Interreg V-A Euregio Meuse-Rhine (“Euradiomics”).
Author information
Authors and Affiliations
Contributions
A.V.: Conceptualization, methodology, software, validation, formal analysis, investigation, writing—original draft, writing—review & editing, resources, visualization M.F.J.A.L.: formal analysis, investigation, data curation, writing—original draft, writing—review & editing, visualization, R.T.H.L.: Supervision, writing—review & editing, M.H.: Conceptualization, supervision F.Z.: Validation, B.M.: Review & editing, S.P.: Validation, A.A.P.: Resources, data curation, review & editing, validation, T.D.B.: Resources, review & editing, M.A.L.B.: Resources, review & editing, S.H.: Validation, review & editing, P.F.M.D.: Resources, review & editing, V.R.: Writing—resources, review & editing, H.C.W.: Writing—review & editing, W.V.: Conceptualization, supervision, investigation, visualization, S.W.: Conceptualization, supervision, R.B.: Supervision, project administration, funding acquisition, P.L.: Supervision, project administration, funding acquisition.
Corresponding author
Ethics declarations
Competing interests
M.F.J.A van der Lubbe, Marc van Hoof, Sergey Primakov, A.A. Postma, T.D. Bruintjes, M.A.L. Bilderbeek , Hammer Sebastiaan, P.F.M. Dammeijer, V. van Rompaey and R. van de Berg have no competing interests Fadila Zerka, Akshayaa Vaidyanathan, and Benjamin Miraglio are salaried employees of Oncoradiomics SA. Dr Philippe Lambin reports, within and outside the submitted work, grants/sponsored research agreements from Varian medical, Oncoradiomics, ptTheragnostic/DNAmito, Health Innovation Ventures. He received an advisor/presenter fee and/or reimbursement of travel costs/external grant writing fee and/or in kind manpower contribution from Oncoradiomics, BHV, Merck, Varian, Elekta, ptTheragnostic and Convert pharmaceuticals. Dr Lambin has shares in the company Oncoradiomics SA, Convert pharmaceuticals SA and The Medical Cloud Company SPRL and is co-inventor of two issued patents with royalties on radiomics (PCT/NL2014/050248, PCT/NL2014/050728) licensed to Oncoradiomics and one issue patent on mtDNA (PCT/EP2014/059089) licensed to ptTheragnostic/DNAmito, three non-patented invention (softwares) licensed to ptTheragnostic/DNAmito, Oncoradiomics and Health Innovation Ventures and three non-issues, non licensed patents on Deep Learning-Radiomics and LSRT (N2024482, N2024889, N2024889. Ralph T.H. Leijenaar has shares in the company Oncoradiomics and is co-inventor of an issued patent with royalties on radiomics (PCT/NL2014/050728) licensed to Oncoradiomics. Sean Walsh and Wim Vos have shares in the company Oncoradiomics. Henry C. Woodruff has minority shares in the company Oncoradiomics.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vaidyanathan, A., van der Lubbe, M.F.J.A., Leijenaar, R.T.H. et al. Deep learning for the fully automated segmentation of the inner ear on MRI. Sci Rep 11, 2885 (2021). https://doi.org/10.1038/s41598-021-82289-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-82289-y
This article is cited by
-
AI-based MRI auto-segmentation of brain tumor in rodents, a multicenter study
Acta Neuropathologica Communications (2023)
-
75% radiation dose reduction using deep learning reconstruction on low-dose chest CT
BMC Medical Imaging (2023)
-
Towards fully automated inner ear analysis with deep-learning-based joint segmentation and landmark detection framework
Scientific Reports (2023)
-
Teeth and prostheses detection in dental panoramic X-rays using CNN-based object detector and a priori knowledge-based algorithm
Scientific Reports (2023)
-
Deep learning-based pancreas volume assessment in individuals with type 1 diabetes
BMC Medical Imaging (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.