Abstract
As CO2 levels in Earth’s atmosphere and oceans steadily rise, varying organismal responses may produce ecological losers and winners. Increased ocean CO2 can enhance seagrass productivity and thermal tolerance, providing some compensation for climate warming. However, the metabolic shifts driving the positive response to elevated CO2 by these important ecosystem engineers remain unknown. We analyzed whole-plant performance and metabolic profiles of two geographically distinct eelgrass (Zostera marina L.) populations in response to CO2 enrichment. In addition to enhancing overall plant size, growth and survival, CO2 enrichment increased the abundance of Calvin Cycle and nitrogen assimilation metabolites while suppressing the abundance of stress-related metabolites. Overall metabolome differences between populations suggest that some eelgrass phenotypes may be better suited than others to cope with an increasingly hot and sour sea. Our results suggest that seagrass populations will respond variably, but overall positively, to increasing CO2 concentrations, generating negative feedbacks to climate change.
Similar content being viewed by others
Introduction
In addition to being a climate warming greenhouse gas, nearly half of the anthropogenically released carbon dioxide (CO2) is absorbed by the ocean, eliciting both negative and positive organismal responses to the process known as Ocean Acidification1,2. Benthic calcifiers, including corals and mollusks, are expected to respond negatively as calcification becomes energetically more expensive in an acidified ocean3. However, CO2 is also a potentially limiting substrate for photosynthesis in both terrestrial and aquatic ecosystems. Terrestrial plants grown under CO2 enrichment often show increased rates of carbon uptake, sucrose formation, nitrogen and water use efficiencies, dark respiration and growth4. Although many marine autotrophs exhibit very little response to CO2 fertilization5, some nitrogen-fixing cyanobacteria6, coccolithophores7, chlorophytes such as Caulerpa spp. and Ulva spp.8,9, and especially seagrasses10,11,12 exhibit increased rates of photosynthesis, growth and biomass production under CO2 enrichment.
Seagrasses are well recognized as important ecosystem engineers13 and Blue Carbon reservoirs14, but their populations are increasingly threatened by anthropogenic degradation of water quality and climate warming15. Experimental16 and theoretical17 evidence indicates that enhanced photosynthesis stimulated by rising CO2 availability can offset the effects of thermal stress for eelgrass (Zostera marina L.); a problem that appears to be increasing with climate warming18,19,20. Eelgrass, the most widely distributed seagrass species in temperate marine environments of the Northern Hemisphere, persists in geographically isolated populations spanning different thermal environments from the subarctic Bering Sea to the seasonally warm waters of the mid-Atlantic Bight and Mediterranean21. This distribution lends eelgrass to be the most extensively studied seagrass species in terms of ecology, physiology and genetics, making it a unique model organism for exploring the impacts of climate change on seagrass ecosystems. In addition, seagrass meadows are among the most productive aquatic habitats in terms of carbon burial22, suggesting that enhanced seagrass productivity under increasing CO2 conditions may exert a negative feedback on climate change.
Eelgrass exhibits a wide range of morphological variation throughout its geographic distribution that has been related to local environmental conditions, including substrate type, water depth, temperature, light, nutrient availability and water flow23 and inherent genetic variation24. These circumstances suggest populations may have had numerous opportunities for genetic adaptation to different environments, making eelgrass an ideal species for exploring the impacts of climate change across geographic regions. However, the degree of morphological and physiological plasticity within geographically isolated populations remains unknown. Heat stress in European populations of Z. marina and Z. noltii appears to be mediated primarily by its effect on sucrose metabolism25. Consequently, photosynthetic stimulation resulting from CO2 enrichment increases sucrose formation and should reduce the effects of thermal stress. Prolonged exposure to elevated CO2 quantitatively enhances leaf photosynthesis, shoot survival, growth and flowering of eelgrass populations from climates characterized by a narrow thermal range (predominantly cool)16,26 and a wide thermal range that include stressfully warm summers11, thereby demonstrating that eelgrass productivity and thermal tolerance in the modern-day ocean are strongly mediated by CO2 availability.
Here we evaluated the physiological and metabolome responses of two distinct eelgrass populations from Puget Sound, Washington and Chesapeake Bay, Virginia, USA that represent contrasting thermal environments. These populations were subjected to an experimental gradient of five CO2 conditions in an outdoor facility under naturally varying temperature and insolation for one year. Accordingly, we compared the plants physiological processes in response to the environment and characterized the metabolome of the samples at the end of the 1-year exposure period. The metabolome of an organism is considered its chemical phenotype27, as it is the first component responding to external stressors28. The metabolome consists of thousands of low molecular weight metabolites (typically < 800 Da) such as amino acids, organic acids, sugars and phenolic compounds derived from primary and secondary cellular metabolism. This study explored the hypothesis that increased CO2 availability would stimulate carbon fixation pathways and reduce the biosynthesis of stress-related compounds. Differential responses among populations may help identify heritable traits that facilitate adaptation of eelgrass to a changing climate and improve our predictive capacity for restoring and conserving these important ecosystem engineers.
Morphology and whole plant performance
At the start of this experiment, eelgrass (Zostera marina L.) shoots from Puget Sound (Dumas Bay, WA, USA) were significantly longer and wider, reaching up to 1 m in length and 0.3–0.5 cm in width, than eelgrass from the Chesapeake region (South Bay, VA, USA) that grow to about 30 cm in length and 0.1–0.5 cm in width (Fig. 1). CO2 enrichment yielded strong positive effects on individual shoot size, vegetative shoot numbers (shown as % survival) and sucrose content of both populations during the 12-month experiment (Fig. 2). However, plants from South Bay, VA (SBV) showed larger increases in size compared to those from Dumas Bay, WA (DBW), especially during summer and early fall 2013 (Fig. 2A,B and Table S1). The stimulating effects of CO2 enrichment on plant size of both populations decreased during the winter period of low light availability and low temperature (Fig. 2A,B white symbols and lines).
Heat maps showing the responses of South Bay VA and Dumas Bay WA eelgrass populations to [CO2] and time. (A,B) Percent plant size. (C,D) Percent shoot survival. (E,F) Relative growth rate. (G,H) Leaf sucrose concentration. Tick marks on the left vertical axis of each plot indicate the mean [CO2] for each treatment. White symbols on each plot represent the monthly slope of the response variable vs. log [CO2] derived from linear regression analysis. Error bars represent ± 1 SE of the regression slope. South Bay data adapted from11.
High [CO2] stimulated vegetative shoot survival in both eelgrass populations into the fall of 2013 and especially through the winter of 2013–2014 (Fig. 2C,D). However, shoot numbers decreased under ambient [CO2] for both eelgrass populations during the summer period of warm (>25 °C) water temperature. Shoot losses continued under ambient [CO2] as water temperature dropped throughout the fall 2013 and into the winter of 2014. The effect of CO2 on shoot survival, as indicated by the slope of percent survival vs. log [CO2], was highest from winter (January) to spring (May) 2014 for SBV plants and from early (March) to late (May) spring 2014 for DBW plants (Fig. 2C,D white symbols and lines, Table S2). At the end of the experiment (May 2014), shoot numbers in the highest CO2 treatment doubled through vegetative proliferation but less than half originally transplanted shoots survived under ambient [CO2].
Although plant size and therefore absolute growth rate increased with [CO2], there was no CO2 effect on relative growth rate (Fig. 2E,F). Despite CO2 enrichment, both SBV and DBW populations increased growth rates during summer and early fall of 2013, even when water temperature exceeded the 25 °C threshold for eelgrass heat stress18,29,30. Growth rates of both populations declined during winter period of low light availability and temperature in all CO2 treatments (Fig. 2E,F) but recovered as temperature and light availability increased during spring 2014. The monthly trends of relative shoot growth (but not absolute growth) were significantly different between populations at different time points. For example, in late summer (August) and late winter (January) SBV showed higher slopes but DBW slopes were higher in fall (November) and winter (December), (Fig. 2E,F, white symbols and lines, Table S3).
Leaf sugar concentrations peaked in all CO2 treatments for both eelgrass populations in late winter (Fig. 2G,H), when temperature (Fig. S1) and growth were low. Sucrose concentrations of SBV leaves were higher than those from DBW during the whole experiment in high [CO2]. The monthly slopes of leaf sucrose concentration vs. log [CO2] were significantly different between eelgrass populations, SBV presented higher slopes (Fig. 2G,H, white symbols and lines, Table S4) in late summer (August), fall (November) and early winter (December).
Metabolomic Response of Eelgrass
Comparison between populations at high and low CO2
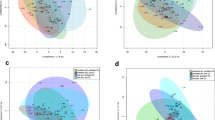
Permutational multivariate analysis of variance (PERMANOVA) of the entire metabolomic fingerprints, including both populations and CO2, showed overall significant differences between the SBV and DBW populations after 1-year growth in the experimental aquaria (Table S5). However, the interaction between CO2 treatment and population was not statistically significant (p = 0.077) (Table S5), suggesting that both populations showed similar responses to elevated CO2. Individual one-way ANOVAs for each metabolite detected significant differences (p < 0.05) between SBV and DBW plants in the abundance of some primary metabolites (Glycolysis – Krebs – Calvin) across CO2 treatments (Tables S6 and S7). Principal component analysis (PCA) including the whole eelgrass metabolomic fingerprints clearly clustered the two populations along the first Principal Component Axis 1(PC1) (Fig. 3A), with CO2 treatments separated along the PC2 showing differences between plants growing at different [CO2].
Although CO2 enhanced growth, survival and thermal tolerance of both eelgrass populations, DBW contained higher abundances of photorespiratory and stress-related compounds in the shikimate pathway than SBV regardless of CO2 treatments (Fig. 4a,b, Tables S6, S7). Higher abundance of dehydroshikimate (Fig. 4a,b, Tables S6, S7) observed in DBW plants, relative to SBV, may indicate up-regulation of metabolic flux through the shikimate pathway31 leading to the synthesis of polyphenols. Stress conditions such as high light and pathogens32, and low CO233 increase the biosynthesis of phenolic compounds in seagrasses. Further, the shikimic intermediates are known to respond to oxidative stress and copper pollution in some macrophytes34,35. On the other hand, SBV plants had higher abundances of α-ketoglutaric acid (TCA Cycle) across CO2 treatments (Fig. 4a,b, Tables S6, S7). Studies have reported accumulation of α-ketoglutaric acid under oxidative stress in Z. marina36 and rice37 and have been suggested as a mitigation mechanisms to manage stressful events limiting the accumulation of pyruvate36.
Representation of the main metabolic pathways of Z. marina from South Bay VA and Dumas Bay WA in response to high and low CO2 concentrations. Only identified metabolites are represented in the schema. Significant changes in any of the metabolite comparisons are represented in bold typeface. Colored boxes below metabolite names represent the result of each of the comparisons after one-way ANOVA. Each letter within each box represent a different comparison: (a) South Bay vs. Dumas Bay plants growing at high CO2 (823 μmol CO2 kg−1SW). (b) South Bay vs. Dumas Bay plants growing at low CO2 (107 μmol CO2 kg−1SW). For a and b, blue and orange colors indicate higher relative abundance in Dumas Bay and South Bay plants, respectively. (c) Highest vs. ambient CO2 conditions (2121 vs 55 μmol CO2 kg−1SW) plants from South Bay (d) High vs. low CO2 conditions (823 vs 107 μmol CO2 kg−1 SW) plants from Dumas Bay. For c and d, blue and orange color indicate higher relative abundance of metabolites in plants growing at high CO2 (2121 μmol CO2 kg−1 SW in c, 823 μmol CO2 kg−1 SW in d) and ambient or low CO2 (55 μmol CO2 kg−1 SW in c, pH 107 μmol CO2 kg−1 SW in d), respectively.
At high [CO2], proline and serine were more abundant in DBW eelgrass than in SBV (Fig. 4a, Table S6). Proline is known to aid stress tolerance by acting as a metal chelator, by providing antioxidative defense and as a signaling molecule38,39 to control mitochondrial functions and, other developmental processes, and activate gene expression that may facilitate plant recovery from stress40. Serine has also been related to stress tolerance (e.g., low temperature and elevated salinity in Arabidopsis thaliana41 and references therein) and is synthesized (i) through the photorespiratory glycolate pathway, (ii) from Calvin Cycle intermediates (the “phosphorylated” pathway) and/or (iii) the glycerate pathway via cytosolic glycolysis42. However, high [CO2] is known to decrease photorespiration in eelgrass43, suggesting that the elevated abundance of serine observed here were likely being driven by non-photorespiratory pathways.
Metabolites involved in biotic/abiotic stress responses were elevated in both populations at low [CO2] (107 μmol CO2·Kg−1 SW). However, the abundance of the photorespiratory metabolites glycerate, glycerate 3-P and succinate semialdehyde (GABA shunt) were higher in DBW leaves than in SBV leaves (Fig. 4b, Table S7). The increase in succinate semialdehyde abundance under low [CO2] in DBW could represent a potential stress response as the GABA shunt may help prevent the accumulation of reactive oxygen intermediates31,32,44,45. SBV plants growing under low [CO2] (107 μmol CO2·Kg−1 SW) had higher abundance of proline and the sugar alcohol myo-inositol (Fig. 4b, Table S7), the latter are known to generate protein stabilizing osmolytes, such as di-myo-inositol phosphate25 that may help protect this population from heat stress25.
South Bay comparison across CO2 treatments
The SBV plants were grown across a CO2 concentration gradient as part of a related experiment10, enabling us to examine their metabolomic responses to different [CO2] in some detail. From the over 5,000 detected metabolic features, 455 (9%) increased with [CO2] and 408 (8.1%) decreased in response to [CO2]. To date, we identified 131 of the 863 responsive metabolites. Experimental CO2 enrichment elevated the concentration of intermediates associated with carbon fixation and amino acid synthesis, as well as sucrose, the latter which is consistent with our prior experimental findings11,16. PERMANOVA of the metabolomic fingerprints of SBV plants showed clear statistical differences between CO2 treatments (p < 0.05, Table S8). PCA distinctly clustered SBV plants growing at the highest CO2 treatment (2121 μmol CO2·Kg−1 SW) from the rest along the first principal component axis (PC1), which explains over 30% of the total variability (Fig. 3B), suggesting that plants growing at the highest CO2 levels experienced the largest metabolic changes. On the other hand, plants growing at ambient CO2 were separated from the other CO2 treatments along the second principal component axis (PC2). This trend was also observed with the Euclidian distances between the metabolomes of plants growing at higher CO2 levels (2121, 823, 370, and 107 CO2·Kg−1 SW) vs. plants growing at ambient [CO2]. The most dramatic overall metabolome change was detected between the highest (2121 μmol CO2·Kg−1 SW) and the ambient (55 μmol CO2·Kg−1 SW) [CO2] (Fig. 5), consistent with our PCA (Fig. 3B) and with the negative log-linear relationship between [CO2] and whole plant performance (Fig. 2). SBV plants grown under high [CO2] had higher abundance of glutamate (Fig. 4c, Table S9) which is involved in N assimilation46. In addition, enhancement of gluconate 6-P in SBV plants under high [CO2] (Fig. 4c, Table S9) suggests activation of the pentose phosphate pathway47 that leads to the synthesis of aromatic amino acids such as phenylalanine; another critical compound in protein synthesis as well as the formation of cell wall components, including lignin48. Univariate analyses showed that SBV plants exposed to ambient [CO2] produced higher abundance of TCA cycle intermediates (Fig. 4c) such as citrate, α-ketoglutarate, pyruvate, and GABA (Table S9). However, we found no differences in leaf dark respiration rates across different [CO2] treatments or between eelgrass populations when exposed to different temperatures (Fig. S2), suggesting that the increases of TCA Cycle metabolites in plants under ambient CO2 may have been diverted to other metabolic pathways (e.g. Shikimate) rather than enhancing respiratory ATP production. Although depriving the plant of potential energy for growth, such diversion leads to the synthesis of secondary compounds with diverse physiological roles, such as cell signaling, production of stress-related compounds and the formation of metabolites associated with the biosynthesis of polyphenols49. Using a small range of CO2 conditions, Cymodocea nodosa revealed up-regulation of genes coding for respiratory metabolism, increasing energetic demand for biosynthesis and stress-related processes under similar ambient [CO2] (pH 7.8/[CO2] 43 μmol Kg−1 SW)50. An accurate quantification of this diversion of respiratory intermediates to different pathways may serve as a good proxy for calculating the energetic cost of the physiological stress response to growth and reproductive output.
Dumas Bay comparison between high and low CO2
For DBW plants, a low number of replicates in some CO2 treatments did not enable metabolomics analyses across all treatments. However, PERMANOVA did not find overall metabolomic differences between plants growing at low [CO2] (107 μmol Kg−1 SW) and plants under high [CO2] (823 μmol Kg−1 SW) (Table S8). The two clustered in different regions along PC1 (Fig. 3C) but according to individual ANOVAs of the metabolomic fingerprints detected significant changes between CO2 treatments (Tables S9 and S10). Under low [CO2] (107 μmol CO2·Kg−1 SW), DBW plants contained higher levels of α-ketoglutarate, succinate, glutamate and glycerate 3-P (Fig. 4d, Table S10) suggesting activation of the GABA shunt as a way to mitigate stress36. On the other hand, high [CO2] (823 μmol CO2·Kg−1 SW) stimulated the abundance of shikimate and proline (Fig. 4d, Table S10) the latter may have increased the thermal tolerance of this population.
Conclusion
Our results revealed that eelgrass populations from two very different thermal environments both exhibited enhanced photosynthetic energy capture, sucrose formation and growth under CO2 enrichment that increased thermal tolerance, potentially counteracting some climate warming impacts on this foundational species. Both populations exhibited positive whole plant responses to CO2 in terms of leaf sucrose, leaf growth, and shoot numbers, but metabolite profiles hint at important genetic differences between these populations. Metabolomic analyses suggest that stress diverts the flow of photosynthates from growth and energy (ATP) production to non-anabolic intermediates that may help (i) elucidate important mechanisms responsible for stress tolerance and (ii) quantify the energetic cost of the stress response.
Although the differences in metabolite pools observed here in response to different [CO2] point to shifts in the activities of metabolic pathways leading to whole plant responses to potential climate forcing, we note that metabolite pool sizes alone are insufficient to fully understand the physiological basis for whole-plant responses to climate-driven environmental change. In addition to making more detailed analyses of metabolite change over time, analyses of changes in the proteome and transcriptome will be necessary to fully understand key genomic functions and metabolic pathways, and those analyses are currently under way. However, the metabolite profiles generated here, in combination with analysis of whole-plant performance, provide a force multiplier for translating ‘omic’ approaches into a predictive understanding of the physiological response of seagrasses to an increasingly hot and sour sea, and the potential for populations to adapt to new environments. Such knowledge will help predict earth system interactions in the context of global cycles and help inform best practices for seagrass restoration.
Methods
Plant collection
Eelgrass shoots with at least 5 rhizomatous internodes and roots were collected by hand from populations in (i) South Bay, Virginia, near the southern limit of their distribution in the Western Atlantic that experience a large seasonal temperature range and (ii) Dumas Bay in southern Puget Sound, WA that are less vulnerable to summertime thermal stress (Fig. S1). Shoots were carefully uprooted by hand, washed free of all sediment and transported to the experimental growth facility at the Virginia Aquarium & Marine Science Center, Virginia Beach, VA. Shoots from Dumas Bay, WA were packed in paper towels moistened with seawater, shipped overnight to VA and immediately transplanted into the experimental facility.
Experimental facility
Eelgrass shoots were grown in fully replicated aquaria using the climate change experimental facility at the Virginia Aquarium & Marine Science Center11. The experimental system consisted of 20 outdoor aquaria plumbed with running water (10 turnovers/day) pumped from the adjacent Owls Creek estuary (Fig S3). Shoots were transplanted into rectangular fiberglass-reinforced plastic containers (0.04 m3 volume, 0.075 m2 surface area) filled with clean intertidal sand and placed in each aquarium. CO2 concentrations were manipulated using beverage-grade CO2 delivered through pH-controlled CO2 bubblers to maintain aquaria at five CO2 concentrations ranging from ambient (55 μmol CO2 Kg−1 SW, pH ~8.0) to 2121 μmol CO2 Kg−1 SW (pH 6) that encompasses a 200-year projection for CO2 availability and yielded a 3-fold gradient in light-saturated photosynthesis for the duration of the experiment10,51.
Whole plant performance
Shoot counts, size, growth, and sucrose content of leaf tissues were measured each month to track the temporal changes across the CO2 treatment range as previously described11.
Metabolic profiling
Tissue collection, storage and processing
One leaf sample (2nd youngest leaf) was collected monthly at random from each plastic container (three plastic containers for SBV and one for DBW in every aquarium). Epiphytes were removed by gently scraping each leaf with a clean razor blade, followed by a brief rinse in 0.2 μm-filtered seawater. The clean leaves were patted dry with a tissue, flash frozen in liquid nitrogen and stored at −80 °C. Samples collected in May 2014 used for the metabolomics analyses performed here were lyophilized for at least 48 h and powdered using a ball mill. The powdered samples were incubated in methanol/deionized water (4/1 v/v) at 10 °C on an orbital shaker (1 h), followed by gentle sonication for 2 min using a Branson ultrasonic cleaner (40 kHz). The extracts were centrifuged and the supernatants transferred to pre-combusted (450 °C for 8 h) amber glass vials for metabolite analysis. Three solvent-only vials were prepared using only methanol/deionized water (no plant material) processed as above.
GC-MS analyses of plant tissues
50 μL of eelgrass extract from each sample was dried and subsequently derivatized in two different steps52. First, compounds were derivatized to a trimethylsilyl ester form using methoxyamine in pyridine solution (30 mg/mL). 20 μL of methoxyamine solution was added to each dried extract and samples were incubated at 37 °C for 90 min in a Thermomixer operating at 1,200 rpm. Later, amine, carboxyl and hydroxyl groups were derivatized using 80 μL of MSTFA (N-Methyl-N-(trimethylsilyl) trifluoroacetamide), subsequently incubated at 37 °C for 30 min at 1,200 rpm. After derivatization, all extracts were vortexed for 10 s and centrifuged at 2,750 × g for 5 minutes and supernatants were used for GC-MS analyses.
GC-MS analyses were performed using an Agilent GC 7890A equipped with an HP-5MS column (30 m × 0.25 mm × 0.25 μm; Agilent Technologies) coupled to a MSD 5975C mass spectrometer (Agilent Technologies, Santa Clara, CA). The injection port temperature was 250 °C. Injection volume was set at 10 μL and split-less. The column was maintained at 60 °C for 1 min and then increased at a rate of 10 °C min−1 to 325 °C during the following 26.5 min and held for 10 min. Experimental blanks from the solvent-only vials were injected every 15 samples and a mixture of fatty acid methyl esters (FAMEs; C8-C28) was analyzed at the beginning of the sequence to calculate retention indices (RI).
Chromatograms were deconvoluted and aligned according to the RI that were calculated from the FAMEs mixture run at the beginning of the sequence. Metabolite identification was conducted by matching MS spectra and RI to an updated version of FiehnLib53. Assigned metabolites were subsequently validated using fragmentation spectra from NIST14 GC-MS library. Parameters used in the Metabolite detector are shown in Table S11. Metabolite matching information in GC-MS is shown in Table S12 and more details as previously described52.
LC-MS analyses of plant tissues
LC-MS analyses were performed using a Vanquish ultra-high pressure liquid chromatography system (UHPLC) coupled to an LTQ Orbitrap Velos mass spectrometer equipped with heated electrospray ionization (HESI) source (Thermo Fisher Scientific, Waltham, Massachusetts, USA). Chromatography was performed with a Hypersil gold C18 reversed-phase column (150 × 2.1 mm, 3 μ particle size; Thermo Scientific, Waltham, Massachusetts, USA) operating at 30° C. Mobile phases consisted of 0.1% formic acid in water (A) and 0.1% formic acid in acetonitrile/water (90:10) (B). The injection volume was 5 μL and flow rate was constant at 0.3 mL min−1. The elution gradient started at 90% A (10% B) constant for 5 min and then linearly changed to 10% A (90% B) during the following 15 min. Those conditions were held for 2 min before the initial conditions were recovered during the subsequent 2 min and the column was washed and stabilized for 11 min. All samples were injected in both negative (−) and positive (+) ionization modes. The MS operated at a resolution of 60,000 in Fourier Transform Mass Spectrometry (FTMS) full-scan mode measuring a mass range of 50 to1000 m/z. Experimental blanks from the solvent-only vials were injected every 15 samples. Details on the MS was previously described54.
LC-MS negative and positive chromatograms were separately processed with MZmine 2.2655. Chromatograms were baseline corrected, deconvoluted, aligned and metabolic features were assigned to metabolites according to retention time (RT) and exact mass of standard compounds included in our in-house library (second level identification according to Sumner et al.56,57). The parameters used for the extraction of the metabolic fingerprints are given in Table S13. Metabolite matching information in LC-MS is shown in Table S14.
Statistical analysis
Whole plant performance
Within-aquarium replicate measures of each performance property were combined each month to generate statistically independent means for each aquarium (without error), resulting in statistically independent replicate measurements for each CO2 treatment each month. Consequently, statistical significance of treatment effects was determined using a repeated-measures ANCOVA implemented in the mixed model analysis of the linear mixed model component of IBM SPSS Statistics 22 using population and month as the fixed factors (within subjects) and log CO2 as the covariate (between subjects). The time series observations were treated as repeated subjects for each measured parameter. All error terms were expressed as standard errors unless otherwise noted.
Metabolomics
The final metabolomic dataset was composed of two categorical factors (Population and CO2 treatment) and 5757 continuous variables (metabolomic features), including 133 metabolites identified by our LC-MS and GC-MS compound libraries. Full factorial PERMANOVAs (Population + CO2 + Population × CO2) were performed to test for overall metabolomic differences between populations and CO2 levels. Both populations were exposed to all five [CO2] levels however for the metabolomics analyses the DBW population had a low number of replicates in some CO2 treatments constraining the metabolomics analyses across all treatments. Therefore, we only used two levels of CO2 for both populations (823 and 107 μmol CO2·Kg−1 SW) to be consistent with both populations for the full PERMANOVA model. Additional PERMANOVAs were performed to test for overall differences for CO2 treatments within each eelgrass population. All PERMANOVAs were computed using the Euclidean distance and 10,000 permutations. In addition, one-way ANOVAs were performed on individual metabolic features contrasting (i) DBW vs. SBV plants at high [CO2] (823 μmol CO2 kg−1 SW), (ii) DBW vs. SBV plants at low [CO2] (107 μmol CO2 kg−1 SW), (iii) highest [CO2] (2121 μmol CO2 kgSW−1) vs. ambient [CO2] (55 μmol CO2 kgSW−1) for SBV plants, and (iv) high [CO2] (823 μmol CO2 kgSW−1) vs. low [CO2] (107 μmol CO2 kgSW−1) for DBW plants.
The datasets (SBV + DBW, SBV alone, and DBW alone) were subsequently subjected to principal component analysis (PCA) to visualize the overall metabolomic variability of the study cases along the two axes explaining most variability (principal component (PC) 1 and PC2).
The Euclidean distances (metabolomic distances) between plants growing at ambient CO2 (55 μmol CO2· kg−1SW) and plants at higher CO2 concentrations (2121, 823, 370, and 107 µmol CO2 kg−1SW) were calculated using the entire metabolomic fingerprints. Those distances were posteriorly submitted to one-way ANOVAs followed by Tukey HSD post-hoc test to understand which [CO2] had a higher response in comparison to ambient [CO2]. Pearson correlation coefficients (r) between CO2 gradient and individual metabolic features for SBV plants were calculated.
All statistical analyses for metabolomics were performed in R version 3.6.158. PERMANOVAs were performed using the adonis function from the package “vegan”59. One-way ANOVAs and Euclidian distances were calculated with the functions aov and dist, respectively, included in the package “stats”58. Tuskey HSD post hoc tests were conducted using the HSD.test function from the agricolae package60. All PCAs were performed using the function PCA found in 2323 “FactoMineR” package61. For PCAs, missing data were previously imputed using the function imputePCA from “missMDA” package62.
Data availability
Whole-plant metabolic data are available from BCO-DMO (Principal Investigator Richard Zimmerman, Project: Impact of Climate Warming and Ocean Carbonation on Eelgrass (Zostera marina L.). Metabolomics data are also available in the Supporting Information for this manuscript.
References
Doney, S. C., Fabry, V. J., Feely, R. A. & Kleypas, J. A. Ocean acidification: the other CO2 problem. Annual Review of Marine Science 1, 23, https://doi.org/10.1146/annurev.marine.010908.163834 (2009).
Penuelas, J. & Sardans, J. Ecological metabolomics. Chemistry and Ecology 25, 305–309 (2009).
Kleypas, J. et al. Impacts of ocean acidification on coral reefs and other marine calcifiers: a guide for future research. Report of a workshop held 18–20 April 2005 St. Petersburg, FL sponsored by NSF, NOAA, and the U.S. Geological Survey 18, 88 (2006).
Leakey, A. D. et al. Elevated CO2 effects on plant carbon, nitrogen, and water relations: six important lessons from FACE. Journal of Experimental Botany 60, 2859–2876 (2009).
Mackey, K. R., Morris, J. J., Morel, F. M. & Kranz, S. A. Response of photosynthesis to ocean acidification. Oceanography 28, 74–91 (2015).
Hutchins, D. et al. CO2 control of Trichodesmium N2 fixation, photosynthesis, growth rates, and elemental ratios: Implications for past, present, and future ocean biogeochemistry. Limnology and Oceanography 52, 1293–1304 (2007).
Rivero-Calle, S., Gnanadesikan, A., Del Castillo, C. E., Balch, W. M. & Guikema, S. D. Multidecadal increase in North Atlantic coccolithophores and the potential role of rising CO2. Science 350, 1533–1537 (2015).
Beer, S. Mechanisms of inorganic carbon acquisition in marine microalgae (with special reference to the Chlorophyta). Progress in Phycological Research 10, 179–179 (1994).
Hall-Spencer, J. M. et al. Volcanic carbon dioxide vents show ecosystem effects of ocean acidification. Nature 454, 96 (2008).
Invers, O., Zimmerman, R., Alberte, R., Perez, M. & Romero, J. Inorganic carbon sources for seagrass photosynthesis: an experimental evaluation for bicarbonate use in temperate species. The Journal of Experimental Marine Biology and Ecology 265, 203–217 (2001).
Zimmerman, R. C. et al. Experimental impacts of climate warming and ocean carbonation on eelgrass Zostera marina. Marine Ecology Progress Series 566, 1–15 (2017).
Jiang, Z. J., Huang, X. P. & Zhang, J. P. Effects of CO2 Enrichment on Photosynthesis, Growth, and Biochemical Composition of Seagrass Thalassia hemprichii (Ehrenb.) Aschers. Journal of Integrative Plant Biology 52, 904–913, https://doi.org/10.1111/j.1744-7909.2010.00991.x (2010).
Jones, C. G., Lawton, J. H. & Shachak, M. in Ecosystem management 130-147 (Springer, 1994).
Macreadie, P. I., Serrano, O., Maher, D. T., Duarte, C. M. & Beardall, J. Addressing calcium carbonate cycling in blue carbon accounting. Limnology and Oceanography Letters 2, 195–201 (2017).
Orth, R. J. et al. A global crisis for seagrass ecosystems. Bioscience 56, 987–996 (2006).
Palacios, S. L. & Zimmerman, R. C. Response of eelgrass Zostera marina to CO2 enrichment: possible impacts of climate change and potential for remediation of coastal habitats. Marine Ecology Progress Series 344, 1 (2007).
Zimmerman, R. C., Hill, V. J. & Gallegos, C. L. Predicting effects of ocean warming, acidification, and water quality on Chesapeake region eelgrass. Limnology and Oceanography 60, 1781–1804, https://doi.org/10.1002/lno.10139 (2015).
Moore, K. A. & Jarvis, J. C. Environmental Factors Affecting Recent Summertime Eelgrass Diebacks in the Lower Chesapeake Bay: Implications for Long-term Persistence. Journal of Coastal Research, 135-147 (2008).
Winters, G., Nelle, P., Fricke, B., Rauch, G. & Reusch, T. B. Effects of a simulated heat wave on photophysiology and gene expression of high- and low-latitude populations of Zostera marina. Marine Ecology Progress Series 435, 83–95 (2011).
Thom, R., Southard, S. & Borde, A. Climate-linked mechanisms driving spatial and temporal variation in eelgrass (Zostera marina L.) Growth and assemblage structure in Pacific Northwest estuaries, USA. Journal of Coastal Research 68, 1–11 (2014).
Moore, K. & Short, F. In Seagrasses: Biology, Ecology and Conservation (eds Anthony W.D. Larkum, Robert J. Orth, & Carlos M. Duarte) Ch. 16, 361-386 (Springer, 2006).
Mcleod, E. et al. A blueprint for blue carbon: toward an improved understanding of the role of vegetated coastal habitats in sequestering CO2. Frontiers in Ecology and the Environment 9, 552–560 (2011).
Holmer, M., Baden, S., Boström, C. & Moksnes, P.-O. Regional variation in eelgrass (Zostera marina) morphology, production and stable sulfur isotopic composition along the Baltic Sea and Skagerrak coasts. Aquatic Botany 91, 303–310 (2009).
Hughes, A. R., Stachowicz, J. J. & Williams, S. L. Morphological and physiological variation among seagrass (Zostera marina) genotypes. Oecologia 159, 725–733 (2009).
Gu, J. et al. Identifying core features of adaptive metabolic mechanisms for chronic heat stress attenuation contributing to systems robustness. Integrative Biology 4, 480–493 (2012).
Zimmerman, R. C., Kohrs, D. G., Steller, D. L. & Alberte, R. S. Impacts of CO2 enrichment on productivity and light requirements of eelgrass. Plant Physiology 115, 599–607 (1997).
Fiehn, O. Metabolomics – the link between genotypes and phenotypes. Plant Molecular Biology 48, 155–171, https://doi.org/10.1023/a:1013713905833 (2002).
Gargallo-Garriga, A. et al. Root exudate metabolomes change under drought and show limited capacity for recovery. Scientific Reports 8, 12696, https://doi.org/10.1038/s41598-018-30150-0 (2018).
Zimmerman, R. C., Smith, R. D. & Alberte, R. S. Thermal acclimation and whole-plant carbon balance in Zostera marina L. (eelgrass). Journal of Experimental Marine Biology and Ecology 130, 93–109, https://doi.org/10.1016/0022-0981(89)90197-4 (1989).
Thayer, G. W., Wolfe, D. A. & Williams, R. B. The Impact of Man on Seagrass Systems. American Scientist, 288-296 (1975).
Singh, S. A. & Christendat, D. Structure of Arabidopsis Dehydroquinate Dehydratase-Shikimate Dehydrogenase and Implications for Metabolic Channeling in the Shikimate Pathway. Biochemistry 45, 7787–7796, https://doi.org/10.1021/bi060366+ (2006).
Vergeer, L. H. T., Aarts, T. L. & de Groot, J. D. The ‘wasting disease’ and the effect of abiotic factors (light intensity, temperature, salinity) and infection with Labyrinthula zosterae on the phenolic content of Zostera marina shoots. Aquatic Botany 52, 35–44, https://doi.org/10.1016/0304-3770(95)00480-N (1995).
Arnold, T. et al. Ocean acidification and the loss of phenolic substances in marine plants. PLoS One 7, e35107, https://doi.org/10.1371/journal.pone.0035107 (2012).
Zou, H.-X. et al. Behavior of the Edible Seaweed Sargassum fusiforme to Copper Pollution: Short-Term Acclimation and Long-Term Adaptation. Plos One 9, e101960, https://doi.org/10.1371/journal.pone.0101960 (2014).
Kumari, P., Reddy, C. R. K. & Jha, B. Methyl Jasmonate-Induced Lipidomic and Biochemical Alterations in the Intertidal Macroalga Gracilaria dura (Gracilariaceae, Rhodophyta). Plant and Cell Physiology 56, 1877–1889, https://doi.org/10.1093/pcp/pcv115 (2015).
Hasler-Sheetal, H., Fragner, L., Holmer, M. & Weckwerth, W. Diurnal effects of anoxia on the metabolome of the seagrass Zostera marina. Metabolomics 11, 1208–1218, https://doi.org/10.1007/s11306-015-0776-9 (2015).
Miro, B. & Ismail, A. Tolerance of anaerobic conditions caused by flooding during germination and early growth in rice (Oryza sativa L.). Frontiers in Plant Science 4, https://doi.org/10.3389/fpls.2013.00269 (2013).
Verbruggen, N. & Hermans, C. Proline accumulation in plants: a review. Amino Acids 35, 753–759, https://doi.org/10.1007/s00726-008-0061-6 (2008).
Hayat, S. et al. Role of proline under changing environments. Plant Signaling & Behavior 7, 1456–1466, https://doi.org/10.4161/psb.21949 (2012).
Szabados, L. & Savouré, A. Proline: a multifunctional amino acid. Trends in Plant Science 15, 89–97, https://doi.org/10.1016/j.tplants.2009.11.009 (2010).
Ho, C.-L. & Saito, K. Molecular biology of the plastidic phosphorylated serine biosynthetic pathway in Arabidopsis thaliana. Amino Acids 20, 243–259, https://doi.org/10.1007/s007260170042 (2001).
Bourguignon J, Rebéillé F & Douce R. in Plant amino acids: Biochemistry & biotechnology (ed Singh BK) 111–146 (Marcell Dekker, Inc, 1998).
Celebi, B. Potential impacts of Climate Change on photochemistry in Zostera marina L. PhD thesis, Old Dominion University, Norfolk, VA, (2016).
Shelp, B. J., Bown, A. W. & McLean, M. D. Metabolism and functions of gamma-aminobutyric acid. Trends in Plant Science 4, 446–452, https://doi.org/10.1016/S1360-1385(99)01486-7 (1999).
Bouché, N., Fait, A., Bouchez, D., Møller, S. G. & Fromm, H. Mitochondrial succinic-semialdehyde dehydrogenase of the γ-aminobutyrate shunt is required to restrict levels of reactive oxygen intermediates in plants. Proceedings of the National Academy of Sciences 100, 6843–6848, https://doi.org/10.1073/pnas.1037532100 (2003).
Forde, B. G. & Lea, P. J. Glutamate in plants: metabolism, regulation, and signalling. Journal of Experimental Botany 58, 2339–2358, https://doi.org/10.1093/jxb/erm121 (2007).
Tabita, F. R. & McFadden, B. A. Regulation of ribulose-1,5-diphosphate carboxylase by 6-phospho-D-gluconate. Biochemical and Biophysical Research Communications 48, 1153–1159, https://doi.org/10.1016/0006-291X(72)90831-5 (1972).
Bonawitz, N. D. & Chapple, C. The genetics of lignin biosynthesis: connecting genotype to phenotype. Annual review of genetics 44, 337–363, https://doi.org/10.1146/annurev-genet-102209-163508 (2010).
Weaver, L. M. & Herrmann, K. M. Dynamics of the shikimate pathway in plants. Trends in Plant Science 2, 346–351, https://doi.org/10.1016/S1360-1385(97)84622-5 (1997).
Ruocco, M. et al. Genomewide transcriptional reprogramming in the seagrass Cymodocea nodosa under experimental ocean acidification. Molecular Ecology, n/a-n/a, https://doi.org/10.1111/mec.14204 (2017).
Cottingham, K. L., Lennon, J. T. & Brown, B. L. Knowing when to draw the line: designing more informative ecological experiments. Frontiers in Ecology and the Environment 3, 145–152 (2005).
Kim, Y.-M. et al. Diel metabolomics analysis of a hot spring chlorophototrophic microbial mat leads to new hypotheses of community member metabolisms. Frontiers in Microbiology 6, 209 (2015).
Kind, T. et al. FiehnLib: mass spectral and retention index libraries for metabolomics based on quadrupole and time-of-flight gas chromatography/mass spectrometry. Analytical Chemistry 81, 10038–10048 (2009).
Rivas-Ubach, A. et al. Similar local, but different systemic, metabolomic responses of closely related pine subspecies to folivory by caterpillars of the processionary moth. Plant Biology 18, 484–494, https://doi.org/10.1111/plb.12422 (2016).
Pluskal, T., Castillo, S., Villar-Briones, A. & Orešič, M. MZmine 2: modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC bioinformatics 11, 395 (2010).
Sumner, L. W. et al. Proposed minimum reporting standards for chemical analysis. Metabolomics 3, 211–221 (2007).
Myers, O. D., Sumner, S. J., Li, S., Barnes, S. & Du, X. One Step Forward for Reducing False Positive and False Negative Compound Identifications from Mass Spectrometry Metabolomics Data: New Algorithms for Constructing Extracted Ion Chromatograms and Detecting Chromatographic Peaks. Analytical Chemistry 89, 8696–8703, https://doi.org/10.1021/acs.analchem.7b00947 (2017).
Team, R. C. R Core Team (2019). R: A language and environment for statistical computing. R Found. Stat. Comput. Vienna, Austria. URL http://www.R-project. org/, page R Foundation for Statistical Computing (2019).
Oksanen, J., et al. Vegan: Community Ecology Package. R package version 2.3.2. R package version 2.3.2. (2013).
Mendiburu, F. agricolae: Statistical Procedures for Agricultural Research. R package version 1.2-3. (2015).
Husson, F., Josse, J., Le, S. & Mazet, J. Multivariate exploratory data analysis and data mining. R Package (2016).
Josse, J. & Husson, F. missMDA: a package for handling missing values in multivariate data analysis. Journal of Statistical Software 70, 1–31 (2016).
Acknowledgements
We thank V. Hill, D. Ruble, B. Celebi and M. Jinuntuya for maintenance of the eelgrass experiments and assistance with collection of the leaf samples. Mark Swingle, Sean Bourgeois and Charles Jourdant of the Virginia Aquarium provided operational maintenance of the experimental system. Financial support for the experimental portion of this research was provided by grants from the National Science Foundation (OCE-1061823 and OCE-1635403) to R.C.Z. Metabolomics analyses and interpretation were conducted under the Laboratory Directed Research and Development (LDRD) Program at the Pacific Northwest National Laboratory (PNNL), a multi-program national laboratory operated by Battelle for the U.S. Department of Energy (DOE) under Contract DE-AC05–76RL01830. A portion of this research was also performed under the Facilities Integrating Collaborations for User Science (FICUS) initiative (#50358) and used resources at the Environmental Molecular Sciences Laboratory (EMSL), which is a DOE Office of Science User Facility. EMSL is sponsored by the DOE Office of Biological and Environmental Research and operated under contract DE-AC05-76RL01830.
Author information
Authors and Affiliations
Contributions
R.C.Z., N.D.W. and C.Z.S. designed research. R.C.Z. and C.Z.S. maintained the experimental system, conducted growth measurements and collected all leaf samples. L.J.K. conducted sample processing. A.R.U. led the metabolomics sample analyses and data compilation. All of the authors participated in writing the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zayas-Santiago, C.C., Rivas-Ubach, A., Kuo, LJ. et al. Metabolic Profiling Reveals Biochemical Pathways Responsible for Eelgrass Response to Elevated CO2 and Temperature. Sci Rep 10, 4693 (2020). https://doi.org/10.1038/s41598-020-61684-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-61684-x
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.