Abstract
Hyperprogressive disease (HPD), an unexpected acceleration of tumor growth kinetics, is described in cancer patients treated with anti-PD-1/anti-PD-L1 agents. Here, our aim was to take into consideration the host and explore whether single nucleotide polymorphisms (SNPs) in key genes involved in immune response might predispose to HPD. DNA was extracted from blood-samples from 98 patients treated under CPI monotherapy. Four candidate genes (PD-1, PD-L1, IDO1 and VEGFR2) and 15 potential SNPs were selected. The TGKR (ratio of the slope of tumor growth before treatment and the slope of tumor growth on treatment) was calculated. Hyperprogression was defined as a TGKR≥2. TGKR calculation was feasible for 80 patients (82%). HPD was observed for 11 patients (14%) and was associated with shorter overall survival (P = 0.003). In univariate analysis, HPD was significantly associated with age ≥70 y (P = 0.025), immune-related toxicity (P = 0.016), VEGFR2 rs1870377 A/T or A/A (P = 0.005), PD-L1 rs2282055 G/T or G/G (P = 0.024) and PD-L1 rs2227981 G/A or A/A (P = 0.024). Multivariate analysis confirmed the correlation between HPD and age ≥70 y (P = 0.006), VEGFR2 rs1870377 A/T or A/A (P = 0.007) and PD-L1 rs2282055 G/T or G/G (P = 0.018). Immunogenetics could become integral predictive factors for CPI-based immunotherapy.
Similar content being viewed by others
Introduction
Checkpoint inhibitors (CPIs) including compounds targeting PD-1/PD-L1 axes have brought significant improvements in terms of overall survival in several types of advanced cancers1,2,3,4,5,6. A single response profile, such as pseudo-progression, is observed under CPIs7. Among these typically-related response profiles under CPIs is hyperprogressive disease (HPD) which was defined as an unanticipated and paradoxical acceleration of the tumor growth7,8. The incidence of HPD is variable according to the way it is defined and ranges between 4 and 29%7. Though such acceleration of the tumor growth kinetic was also observed with other agents (chemotherapy9, tyrosine kinase inhibitors10), the intensity and the frequency of the phenomenon appears to be higher with checkpoint inhibitors used alone7. A single response profile, such as pseudo-progression, is observed under CPIs7. Among these typically-related response prfiles under CPIs is hyperprogressive disease (HPD), which has been defined as an unanticipated and paradoxical acceleration of tumor growth7,8. The incidence of HPD is variable according to the way it is defined and ranges between 4 to 29%7. Although this acceleration of tumor growth kinetics was also observed with other agents (chemotherapy9, tyrosine kinase inhibitors10), the intensity and frequency of the phenomenon appears to be higher with checkpoint inhibitors used alone7. HPD may be associated with a worsening of the outcome11. Different physiopathological hypotheses have been tested to explain phenomena such as tumoral genomics variations12,13. Indeed, CPI has been shown to hasten tumor growth in a mouse model with a relative lack of PD-1 expression14. As HPD was observed in several malignant tumor types, a role for the host variations has been advocated13,15,16. Indeed, allelic variations of HLA class I genes have been shown to impact clinical outcome under CPI17. However, dedicated germinal immunogenetics studies remains rare in the context of CPI-based treatment18. To better elucidate the potential relationship between host immunogenetics and CPI treatment outcome and particularly HPD, we correlated the outcome of patients treated with CPI and selected polymorphisms described in four key genes: PD-1 (Programmed Cell Death 1 gene, 2q37.3), PD-L1 (Programmed Death Ligand 1 gene, 9p24.1), IDO1 (Indoleamine 2,3-Dioxygenase 1 gene, 8p11.21) and VEGFR2 (Vascular Endothelial Growth Factor Receptor 2 gene, 4q12).
Results
Patient characteristics and outcome
Patient baseline characteristics are given in Table 1. All patients were treated for an advanced malignancy. Non-small cell lung cancer (NSCLC) (n = 48) was the largest subgroup followed mainly by head and neck squamous cell carcinoma (HNSCC) (n = 16), renal cell carcinoma (RCC) (n = 14) and melanoma (n = 13). Importantly, all patients were treated by CPI monotherapy alone (anti-PD-1 or anti-PD-L1), with a majority of anti-PD1 (87%). Median age was 68 (range: 32–85), 65 were males (66%) and 70 were smokers (83%). Sixty-six patients had received previous irradiation (69%). The SNP genotype, gene information and genotype frequency are shown in Table 2.
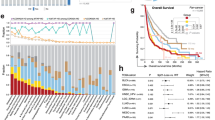
Median follow-up was 13.3 months (95% confidence interval [CI]; 10.6 months to 15.4 months). Median irPFS was 16.8 months (95% confidence interval [CI]; 10.2 months to NA) and median OS was not reached. Twelve-month OS and 12-month PFS were 80% (95% confidence interval [CI], 72% to 90%) and 47% (95% confidence interval [CI]; 5% to 60%), respectively.
Fifteen patients experienced grade 3–4 IrAEs (15.5%), 67 grade 1–2 IrAEs (68.25%) and 16 patients had no IrAE (16.25%). Overall response was complete for 8 patients (8%), partial for 43 patients (44%), stable disease for 28 patients (28.5%) and progressive disease for 19 patients (19.5%). TGKR could be calculated for 80 patients (15 patients had CPI as first line for advanced disease; pre-baseline scanner was not available for 3 patients). HPD was observed in 11 patients (14%). HPD was correlated with shorter OS (Fig. 1) compared with non-HPD patients (P = 0.003).
HPD predictive factors
In univariate analysis (Table 3), HPD was significantly associated with age ≥70 years (25% versus 6%; P = 0.025), immune-related toxicity grade ≥3 (38.5% versus 9.5%; P = 0.016), VEGFR2 rs1870377 A/T or A/A (26% versus 4%; P = 0.005), PD-L1 rs2282055 G/T or G/G (23% versus 2.5%; P = 0.024) and PD-L1 rs2227981 G/G (4.5% versus 23.5%; P = 0.024). HPD was not significantly correlated with lactate dehydrogenase (LDH) blood levels at baseline (p = 0.055). Similarly, the neutrophil-to-lymphocyte ratio (NLR) was not linked to HPD (p = 0.936). Also, tumor burden was not associated with HPD (p = 0.732). Multivariate analysis revealed an independen t association between HPD and age ≥ 70 years (OR = 14.42; 95% confidence interval [CI]; 2 to 100; P = 0.006), rs1870377 T/A or A/A, and VEGFR2 (OR = 15.36; 95% confidence interval [CI]; 1.92 to 119; P = 0.007) and rs2282055 T/G or G/G, PDL1 (OR = 17.73; 95% confidence interval [CI]; 11.55 to 227; P = 0.01).
A risk score was calculated by logistic regression and integrated the 3 independent variables (age, rs2282055, rs1870377) for predicting HPD. The risk for HPD was optimally estimated (OR = 18.34; 95% confidence interval [CI]; 3.38 to 99.58; P <0.001) (Table 4).
Discussion
We observed HPD in 14% of treated patients by CPI, a figure in the range of figures reported in independent series7. We identified older age as a predictive variable for HPD in accord with previously reported series11. However, this point is controversial and observations have been reported in recent studies by Kim et al.19 and Ferrara et al.9 showingno association between HPD and age. These discrepancies may be due to the different evaluation methods used to evaluate HPD as well as to the retrospective nature of these studies. In agreement with others19, we noted that patients with HPD had higher baseline LDH levels but which did not reach statistical significance in our hands. Our negative finding contrasts with that of Kim and coworkers19 reporting that patients with HPD had baseline NLR values higher than those of patients without HPD. This discrepancy can be explained by the retrospective nature of both studies and also by the relatively small number of patients. Clearly, prospective studies based on a larger set of patients would be more likely to provide firmer conclusions regard of this possible association between baseline NLR and the risk to developing HPD under CPI. To the best of our knowledge, the present study is the first cohort that explores the link between host gene polymorphisms and HPD under CPI. Our data highlight two germinal variations with rs2282055 (PD-L1) and rs1870377 (VEGFR2) having a significant and independent influence on the occurrence of HPD.
The group of patients with rs2282055 (PD-L1) G allele, either homozygous or heterozygous, was found to be significantly associated with a higher risk of developing HPD in comparison with T/T genotype, the locus being located on chromosome 9p24.1. When expressed on tumor cells, this gene down-regulates the activation of T effector cells through a key mechanism responsible for immune response evasion20. However, the real impact of tumor PD-L1 expression on treatment outcome under CPI remains controversial21. The regulation of tumoral and non-tumoral PD-L1 expression is a complex phenomenon and is influenced by multiple molecular pathways22,23,24. rs2282055 (PD-L1) is associated with 10 other SNP all inserted in different introns of the PD-L1 gene25. It has been shown that introns may have a direct or indirect influence on mRNA expression: GTEX portal (https://gtexportal.org/home/) indicates that rs2282055 is associated with down-regulated expression of PD-L1 (CD274 gene) in brain tissue while it is overexpressed in the pancreas, suggesting that rs2282055 may impact PD-L1 expression differently in different tissues. rs2282055 (PD-L1) was recently evaluated for its association with survival of patients not treated by CPI26. In this latter study, the impact of rs2282055 (PD-L1) polymorphism on survival was found to be non-significant, thus suggesting a non-prognostic role of this polymorphism. Since PD-L1 expression was not available in our cohort, we could not examine potential links between this rs and the level of expression of PD-L1 protein In conclusion, it can be suggested that rs2282055 (PD-L1) may interfere with CPI-HPD development, while the underlying mechanism remains to be elucidated.
VEGFR2 is a gene encoding for vascular endothelial growth factor receptor 2 expressed on both endothelial cells and various immune cells27,28. VEGFR2 is a key regulator of tumor angiogenesis and tumor microenvironment by mainly promoting a high level of Tregs and by reducing the ability of T effector cells to penetrate the tumor cell bed29. Of note, rs1870377 (KDR, VEGFR2, NM_002253.3:c.1416A>T) induces a missense substitution Q472H in the fifth (out of seven) extracellular Ig-like motifs that has been shown to increase VEGF-A binding and activity inducing increased microvessel density in tumor tissue of patients with non-small cell lung cancer30. In our series, carriers of rs1870377 (VEGFR2) with any A genotype were more prone to develop HPD. Thus, VEGFR2 substitution Q472H may play a potential role in increased tumor size due to increased angiogenesis and microvessel development in these patients. It is thus conceivable that the impact of VEGFR2 on tumor and its microenvironment may differ according to the allelic inheritance of the host with an influence on HPD development under CPI.
Collectively, one can formulate a working hypothesis with HPD occurring in a subset of patients harboring unfavorable alleles which modulate the expression of different genes inducing tumor progression under CPI. It was interesting to identify key immunology-linked genes like PD-L1 and VEGFR2 gene variants using this approach. The present reported results remain challenging in clinical practice with particular attention given to the fact that most allelic variations are present at relatively low frequencies. However, this study contains a number of limitations which do not allow drawing definitive conclusion: the sample size is relatively small (11 HPD cases) and patients received two different classes of PD-1 and PD-L1 CPI. TGKR was not assessable for first-line treated patients. The study covered different histological types and some patients had been more or less heavily pretreated. According to the meta-analysis by Kim and coworkers31, the histological type of the tumor is not predictive value for the occurrence of HPD. However, it has been reported that renal cell carcinoma (RCC) patients may be at a lesser risk of HPD11,32. Of note, our cohort was also enriched with long-responding patients as all patients alive and treated with CPI in the department were asked their consent to dedicated blood sampling for the study. This explains the high response rate reported in our series (52%). Above all, the study remains original leading to identification of potential host-linked biomarkers for HPD prediction. Interestingly, it was possible to establish a powerful (OR = 18.34; 95% confidence interval [CI]; 3.38 to 99.58; P <0.001) predictive score combining host characteristics such as age and germinal gene polymorphisms. Evaluating the risk of HPD by testing host immunogenetics must remain probabilistic in nature and may differ according to ethnic population, thus limiting extrapolation of the present study outside the Caucasian population. Efforts to expand other candidate genes and their polymorphisms are currently ongoing in larger prospective cohorts. Particular attention should be paid to allelic variations of HLA class I genes.
Finally, our results support the notion of a genetic susceptibility potentially impacting the development of HPD in a Caucasian population. In a broader perspective, it is hoped that the present data can stimulate further studies integrating both somatic and germinal variability aimed at satisfying the still unmet need for faithful predictive biomarkers to ensure enhanced management of cancer therapy by CPI.
Patients and Methods
Study design and patients
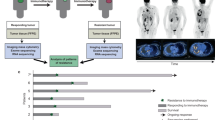
This is a retrospective study covering the period April to August 2018. All data were retrieved from the clinical database of the Centre Antoine Lacassagne (Nice, France). Tumor responses were evaluated after monotherapy according to RECIST 1.1 criteria (complete response (CR), partial response (PR), stable disease (SD), and progressive disease (PD)). Objective response was evaluated as previously published33,34,35. Immune-related adverse events (irAEs) were evaluated according to National Cancer Institute Common Terminology Criteria for Adverse Events (NCI-CTCAE V5). Pre-baseline, baseline, and initial imaging results were recorded and were to calculate the TGKR (ratio of the slope of tumor growth before treatment and the slope of tumor growth during treatment), as previously reported8. The sum of the largest diameter of target lesions at baseline indicated the tumor burden at baseline. HPD was defined as a TGKR ≥2. Written informed consent was systemically obtained before collecting a study-dedicated blood sample. Patient characteristics, at baseline, also included age, gender, histology, smoker status, lactate dehydrogenase (LDH), neutrophil-to-lymphocyte (NLR) and tumor burden.
SNP selection and genotyping
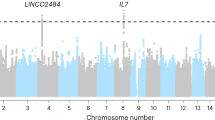
Seventeen SNPs of PD-1 (rs10204525; rs11568821; rs22727981), PD-L1 (rs2282055; rs2297136; rs2297137; rs4143815; rs10815225; rs822339), IDO1 (rs3739319; rs3808606; rs373931; rs9657182; rs34820341) and VEGFR2 (rs2305948; rs1870377; rs2071559) were selected according to their functional and/or clinical relevance. Genomic DNA was extracted from a blood sample using the commercially-available Maxwell® 16 LEV Blood DNA Kit (#AS1290, Promega). The assay to screen the 17 SNPs was created by using Assay Design Suite v2.0 (AGENA Bioscience online software) with the “Genotyping Design”option. We had created the assay to screen the 17 SNPs. Data were verified and compatible with DNA controls polymorphism for 15 SNPs; the remaining 2 SNPs had been eliminated (PD-L1 rs822339 and IDO1 rs34820341) because incompatible with DNA control polymorphism (https://www.coriell.org/1/NIGMS/Collections/CEPH-Resources). For 15 SNPs minor allele frequency was ≥5% in Caucasians according to SNPpedia (http://www.snppedia.com) and the Ensemble database (http://www.Ensembl.org). All tested SNPs were in Hardy-Weinberg equilibrium (Table 2).
Statistical considerations
The link between the 15 SNPs and clinico-radiological parameters and CPI response according to ir-RECIST35 criteria and irAEs was examined. Statistical comparisons were performed using χ2 test or Fisher’s exact test for categorical data and Student’s test or Wilcoxon’s test for continuous variables. Immune-related progression-free survival (irPFS) and Overall Survival (OS) were respectively calculated from the baseline CT scan to progression (according to ir-RECIST criteria) or death and presented graphically using the Kaplan-Meier method. All variables significant at the 5% level in both univariate and multivariate logistic regression models were included. Co-linearity between all variables of the initial multivariate model was evaluated. The choice of the final model was made by performing a backward stepwise selection model. A fitted score for each participant by logistic regression was used to define two risk groups of patients (low or high risk of HPD). The optimal number of risk groups for predictive models was obtained using the Younden method36. Statistical analyses were performed using R version 3.5.0 on Windows®.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee (French National Commission for Informatics and Liberties N°17010).
Informed consent
All patients provided written informed consent before enrollment.
Change history
12 June 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Bellmunt, J. et al. Pembrolizumab as Second-Line Therapy for Advanced Urothelial Carcinoma. N. Engl. J. Med. 376, 1015–1026, https://doi.org/10.1056/NEJMoa1613683 (2017).
Ferris, R. L. et al. Nivolumab for Recurrent Squamous-Cell Carcinoma of the Head and Neck. N. Engl. J. Med. 375, 1856–1867, https://doi.org/10.1056/NEJMoa1602252 (2016).
Herbst, R. S. et al. Pembrolizumab versus docetaxel for previously treated, PD-L1-positive, advanced non-small-cell lung cancer (KEYNOTE-010): a randomised controlled trial. Lancet 387, 1540–1550, https://doi.org/10.1016/S0140-6736(15)01281-7 (2016).
Horn, L. et al. Nivolumab Versus Docetaxel in Previously Treated Patients With Advanced Non-Small-Cell Lung Cancer: Two-Year Outcomes From Two Randomized, Open-Label, Phase III Trials (CheckMate 017 and CheckMate 057). J. Clin. Oncol. 35, 3924–3933, https://doi.org/10.1200/JCO.2017.74.3062 (2017).
Motzer, R. J. et al. Nivolumab versus Everolimus in Advanced Renal-Cell Carcinoma. N. Engl. J. Med. 373, 1803–1813, https://doi.org/10.1056/NEJMoa1510665 (2015).
Robert, C. et al. Pembrolizumab versus Ipilimumab in Advanced Melanoma. N. Engl. J. Med. 372, 2521–2532, https://doi.org/10.1056/NEJMoa1503093 (2015).
Borcoman, E. et al. Novel patterns of response under immunotherapy. Ann. Oncol. 30, 385–396, https://doi.org/10.1093/annonc/mdz003 (2019).
Saada-Bouzid, E. et al. Hyperprogression during anti-PD-1/PD-L1 therapy in patients with recurrent and/or metastatic head and neck squamous cell carcinoma. Ann. Oncol. 28, 1605–1611, https://doi.org/10.1093/annonc/mdx178 (2017).
Ferrara, R. et al. Hyperprogressive Disease in Patients With Advanced Non-Small Cell Lung Cancer Treated With PD-1/PD-L1 Inhibitors or With Single-Agent Chemotherapy. JAMA. Oncol. 4, 1543–1552, https://doi.org/10.1001/jamaoncol.2018.3676 (2018).
Mellema, W. W., Burgers, S. A. & Smit, E. F. Tumor flare after start of RAF inhibition in KRAS mutated NSCLC: a case report. Lung Cancer 87, 201–203, https://doi.org/10.1016/j.lungcan.2014.11.014 (2015).
Champiat, S. et al. Hyperprogressive Disease Is a New Pattern of Progression in Cancer Patients Treated by Anti-PD-1/PD-L1. Clin. Cancer Res. 23, 1920–1928, https://doi.org/10.1158/1078-0432.CCR-16-1741 (2017).
Kato, S. et al. Hyperprogressors after Immunotherapy: Analysis of Genomic Alterations Associated with Accelerated Growth Rate. Clin. Cancer Res. 23, 4242–4250, https://doi.org/10.1158/1078-0432.CCR-16-3133 (2017).
Liu, X. S. & Mardis, E. R. Applications of Immunogenomics to Cancer. Cell 168, 600–612, https://doi.org/10.1016/j.cell.2017.01.014 (2017).
Wartewig, T. et al. PD-1 is a haploinsufficient suppressor of T cell lymphomagenesis. Nat. 552, 121–125, https://doi.org/10.1038/nature24649 (2017).
Fairfax, B. P. et al. Innate immune activity conditions the effect of regulatory variants upon monocyte gene expression. Sci. 343, 1246949, https://doi.org/10.1126/science.1246949 (2014).
Sun, C., Mezzadra, R. & Schumacher, T. N. Regulation and Function of the PD-L1 Checkpoint. Immun. 48, 434–452, https://doi.org/10.1016/j.immuni.2018.03.014 (2018).
Chowell, D. et al. Patient HLA class I genotype influences cancer response to checkpoint blockade immunotherapy. Sci. 359, 582–587, https://doi.org/10.1126/science.aao4572 (2018).
Nomizo, T. et al. Clinical Impact of Single Nucleotide Polymorphism in PD-L1 on Response to Nivolumab for Advanced Non-Small-Cell Lung Cancer Patients. Sci. Rep. 7, 45124, https://doi.org/10.1038/srep45124 (2017).
Kim, Y. et al. Comprehensive Clinical and Genetic Characterization of Hyperprogression Based on Volumetry in Advanced Non-Small Cell Lung Cancer Treated With Immune Checkpoint Inhibitor. J. Thorac. Oncol. 14, 1608–1618, https://doi.org/10.1016/j.jtho.2019.05.033 (2019).
Wang, Y. et al. Regulation of PD-L1: Emerging Routes for Targeting Tumor Immune Evasion. Front. Pharmacol. 9, 536, https://doi.org/10.3389/fphar.2018.00536 (2018).
Coelho, M. A. et al. Oncogenic RAS Signaling Promotes Tumor Immunoresistance by Stabilizing PD-L1 mRNA. Immun. 47, 1083–1099 e1086, https://doi.org/10.1016/j.immuni.2017.11.016 (2017).
Topalian, S. L., Drake, C. G. & Pardoll, D. M. Immune checkpoint blockade: a common denominator approach to cancer therapy. Cancer Cell 27, 450–461, https://doi.org/10.1016/j.ccell.2015.03.001 (2015).
Ben Nasr, M. et al. PD-L1 genetic overexpression or pharmacological restoration in hematopoietic stem and progenitor cells reverses autoimmune diabetes. Sci Transl Med 9, https://doi.org/10.1126/scitranslmed.aam7543 (2017).
Sun, L. O. et al. Spatiotemporal Control of CNS Myelination by Oligodendrocyte Programmed Cell Death through the TFEB-PUMA Axis. Cell 175, 1811–1826 e1821, https://doi.org/10.1016/j.cell.2018.10.044 (2018).
Ward, L. D. & Kellis, M. HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res. 40, D930–934, https://doi.org/10.1093/nar/gkr917 (2012).
Yoon, S. et al. Prognostic relevance of genetic variants involved in immune checkpoints in patients with colorectal cancer. J. Cancer Res. Clin. Oncol. 142, 1775–1780, https://doi.org/10.1007/s00432-016-2196-2 (2016).
Shibuya, M. Vascular Endothelial Growth Factor (VEGF) and Its Receptor (VEGFR) Signaling in Angiogenesis: A Crucial Target for Anti- and Pro-Angiogenic Therapies. Genes. Cancer 2, 1097–1105, https://doi.org/10.1177/1947601911423031 (2011).
Miettinen, M., Rikala, M. S., Rys, J., Lasota, J. & Wang, Z. F. Vascular endothelial growth factor receptor 2 as a marker for malignant vascular tumors and mesothelioma: an immunohistochemical study of 262 vascular endothelial and 1640 nonvascular tumors. Am. J. Surg. Pathol. 36, 629–639, https://doi.org/10.1097/PAS.0b013e318243555b (2012).
Zhu, P., Hu, C., Hui, K. & Jiang, X. The role and significance of VEGFR2(+) regulatory T cells in tumor immunity. Onco Targets Ther. 10, 4315–4319, https://doi.org/10.2147/OTT.S142085 (2017).
Glubb, D. M. et al. Novel functional germline variants in the VEGF receptor 2 gene and their effect on gene expression and microvessel density in lung cancer. Clin. Cancer Res. 17, 5257–5267, https://doi.org/10.1158/1078-0432.CCR-11-0379 (2011).
Kim, J. Y. et al. Hyperprogressive Disease during Anti-PD-1 (PDCD1) / PD-L1 (CD274) Therapy: A Systematic Review and Meta-Analysis. Cancers (Basel) 11, https://doi.org/10.3390/cancers11111699 (2019).
Hwang, I., Park, I., Yoon, S. K. & Lee, J. L. Hyperprogressive Disease in Patients With Urothelial Carcinoma or Renal Cell Carcinoma Treated With PD-1/PD-L1 Inhibitors. Clin Genitourin Cancer, https://doi.org/10.1016/j.clgc.2019.09.009 (2019).
Nishino, M. et al. Developing a common language for tumor response to immunotherapy: immune-related response criteria using unidimensional measurements. Clin. Cancer Res. 19, 3936–3943, https://doi.org/10.1158/1078-0432.CCR-13-0895 (2013).
Seymour, L. et al. iRECIST: guidelines for response criteria for use in trials testing immunotherapeutics. Lancet Oncol. 18, e143–e152, https://doi.org/10.1016/S1470-2045(17)30074-8 (2017).
Wolchok, J. D. et al. Guidelines for the evaluation of immune therapy activity in solid tumors: immune-related response criteria. Clin. Cancer Res. 15, 7412–7420, https://doi.org/10.1158/1078-0432.CCR-09-1624 (2009).
López-Ratón, M., Rodríguez-Álvarez, M. X., Cadarso-Suárez, C. & Gude-Sampedro, F. OptimalCutpoints: an R package for selecting optimal cutpoints in diagnostic tests. J. Stat. Softw. 61, 1–36 (2014).
Acknowledgements
The authors acknowledge support from Centre Antoine Lacassagne, Oncopharmacology unit team, University Côte d’Azur, France.
Author information
Authors and Affiliations
Contributions
G.M. and E.S.B. conceived the research idea. J.Ga. and N.E. performed the data analysis. S.R. collected the data. S.R., J.Ga., P.B., D.G., D.B., N.E., F.P., J.Gu., G.M. and E.S.B. participated in the writing and are involved in critical revision of this manuscript for important intellectual content. All authors approved the final manuscript as submitted and agree to be accountable for all aspects of the work.
Corresponding author
Ethics declarations
Competing interests
Gérard Milano is a member of an advisory board at B.M.S., M.S.D. and Merck. Fréderic Peyrade is a member of an advisory board at M.S.D. and Merck. Delphine Borchiellini is a member of an advisory board at M.S.D., Pfizer, Astra-Zeneca, Roche, B.M.S. Joel Guigay is a member of an advisory board at Merck. The remaining authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Refae, S., Gal, J., Brest, P. et al. Hyperprogression under Immune Checkpoint Inhibitor: a potential role for germinal immunogenetics. Sci Rep 10, 3565 (2020). https://doi.org/10.1038/s41598-020-60437-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-60437-0
This article is cited by
-
Association of PD-1 and PDL-1 gene polymorphisms with colorectal cancer risk and prognosis
Molecular Biology Reports (2022)
-
Mechanisms of hyperprogressive disease after immune checkpoint inhibitor therapy: what we (don’t) know
Journal of Experimental & Clinical Cancer Research (2020)
-
Hyperprogressive disease and its clinical impact in patients with recurrent and/or metastatic head and neck squamous cell carcinoma treated with immune-checkpoint inhibitors: Korean cancer study group HN 18–12
Journal of Cancer Research and Clinical Oncology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.