Abstract
The general model presented here for the formation of chromatin domains, LADs and TADs, is primarily based on the 3-D structures of the corresponding DNA sequences, the GC-poor and GC-rich isochores. Indeed, the low-heterogeneity GC-poor isochores locally are intrinsically stiff and curved because of the presence of interspersed oligo-Adenines. In contrast, the high-heterogeneity GC-rich isochores are in the shape of peaks characterized by increasing levels of GC and of interspersed oligo-Guanines. In LADs, oligo-Adenines induce local nucleosome depletions leading to structures that are well suited for the attachment to (and embedding in) the lamina. In TADs, the gradients of GC and of oligo-Guanines are responsible for a decreasing nucleosome density, decreasing supercoiling and increasing accessibility. This “moulding step” shapes the “primary TADs” into loops that lack self-interactions, being CTCF/cohesin-free structures. The cohesin complex then binds to the tips of “primary TADs” and slides down the loops, thanks to Nipbl, an essential factor for loading cohesin and for stimulating its ATPase activity and its translocation. This “extruding step” leads to closer contacts and to self-interactions in the loops and stops at the CTCF binding sites located at the base of the loops that are thus closed and insulated.
Similar content being viewed by others
Introduction
Fifty years ago, well before any sequencing technique was available, a new approach, later called compositional genomics, was developed in order to understand the organization of the mammalian genome. This approach was based on the “short-sequence designs”1 of DNA molecules. Since short sequences are responsible for (1) the fine structure of the double helix; and (2) the interactions of DNA with proteins, compositional genomics is, in fact, based on genome structure and function.
The original investigation2 concerned two satellite DNAs, that appeared in CsCl density gradient ultracentrifugation experiments as unresolved shoulders on the “main band” DNAs of mouse and guinea pig, respectively, and that were completely separated by ultracentrifugation in Cs2SO4/Ag+ gradients. The separation was possible because of the differential binding of silver ions on the short repeats of the satellite DNAs and led to the conclusion that satellites DNA are “conformationally slightly different” from main-band DNA, the differences being related to their particular nucleotide sequences2. This point was confirmed by the different compositions of the short-sequences of the two satellites specifically split by DNAses3. When the Cs2SO4/Ag+ was applied to the bovine genome, the standard mammalian genome at that time, not only several cryptic satellite DNAs were revealed, but, much more importantly, the “main band” DNA molecules were resolved into three families or “major components”4. This was the first evidence that, neglecting satellites, the genome consisted of large DNA regions that differed in the frequency of short-sequence binding sites for silver ions and of short-sequences split by DNAses. The “major components” of mammalian DNAs were shown to be “fairly homogeneous” sequences originally estimated as ≥0.3 Mb in size5,6 that were called isochores7. Subsequent investigations showed that isochores from the human genome (and other mammalian genomes) belong to five families, L1, L2, H1, H2, H3, characterized by increasing GC levels, increasing compositional heterogeneity, increasing gene density and by DNA sizes ≥0.2 Mb. These results were confirmed when sequences from the human genome became available8,9,10 (for a review, see ref.11).
While the functional importance of isochores was already evident a long time ago because of their correlations with all genome properties tested and led to their definition as “a fundamental level of genome organization”12, the basic reason for their existence is not yet understood12, even if links with DNA and chromatin structure were predicted13 and later shown. Indeed, the GC-rich, gene-rich, and the GC-poor, gene-poor isochores (the “genome core” and the “genome desert”, see Supplementary Table S1 and Fig. S1) were shown to correspond at interphase to “open” and “closed” chromatins (later called A and B compartments14) that were centrally and peripherally located, respectively, in the nucleus15.
Recent investigations16 showed that maps of GC-rich and GC-poor isochores of all human and mouse chromosomes match maps of TADs, the Topologically Associating Domains, 0.2–2 Mb in size17, and maps of LADs, the Lamina Associated Domains, ~0.5 Mb medium size18 (that are characterized by histone modificator H3K27me3) respectively (see the Nomenclature section in Materials and Methods). In fact, the average size, ~0.9 Mb, of human isochores10, is almost identical to that, ~0.88 Mb, of 91% of the mouse chromatin domains19, and both isochores and chromatin domains are evolutionarily conserved in mammals11,19. A work still in press showed that the isochores of Drosophila20 also correspond to TADs21.
It should be noted that the current investigations on chromatin based on the Hi-C approach have not taken into account earlier observations on isochores that could provide critical insights. This was done here by studying isochores not only in their composition but also in their topology, and led to the discovery that isochores are characterized by 3-D structures that play a crucial role in the “moulding” of TADs and LADs.
Results and Discussion
A working hypothesis
The starting point of this work was the consideration that the match between isochores and chromatin domains16 suggested a possible similarity between the 3-D structures of the low-heterogeneity GC-poor and the high-heterogeneity GC-rich isochores on the one hand, and the compositionally even structures of LADs and the loops of TADs, on the other. This working hypothesis was tested on human chromosome 21, which comprises isochores from all the five families and, being the smallest human chromosome, allows a more expanded graphical presentation of data. This was done by comparing the previous results obtained by the compositional approach used so far with the new results derived from a topological approach. The previous results basically comprised (1) the compositional profile of the DNA sequence as obtained through non-overlapping 100Kb windows (Fig. 1A; see Materials and Methods for the choice of this window size); and (2) the assemblies of the 100Kb sequences into isochores using a fixed window (Fig. 1B) or a sliding-window approach (Fig. 1C). The new results, in contrast, rely on a point-by-point plot of the GC levels of 100Kb DNA segments.
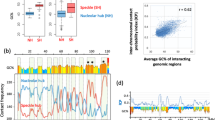
Compositional and isochore profiles of human chromosome 21. (A) Compositional profile of release hg38 as seen through non-overlapping 100-Kb windows46. DNA sequences from isochore families L1 to H3 are represented in different colors, deep blue, light blue, yellow, orange, red, respectively. The left-side ordinate values are the minima GC values between the isochore families (see Supplementary Table S1). (B) Isochore profile (release hg38) using a non-overlapping 100Kb window and the isoPlotter program46 with “fixed” isochore borders (see Supplementary Table S1). (C) Isochore profile (from the matched b37 assembly47) using a sliding window approach with “fixed” isochore borders (see Supplementary Table S1). This profile is slightly more extended on the centromeric (left) side than those of (A,B). Permissions to publish (A–C) were obtained from the copyright owners. (D) GC levels of 100 Kb windows are presented as a point-by-point plot. This figure shows that, except for the H1 (a) and L2+ peaks (b to f), that happen to be more abundant in chromosome 21 compared to other chromosomes, individual isochores from the H1 to H3 families are in the form of peaks that do not appear in a clear way in the standard presentation of compositional profiles of chromosomes (A). Black, blue and red lines, as well as double lines (X,Y), separate regions 1 to 6.
The 3-D structure of isochores
The compositional point-by-point profile (split into several regions) of chromosome 21 (see Fig. 1D and its enlarged version, Fig. 2) revealed: (1) several single peaks (a to f) located in regions 1 and 3, as well as a compositionally low-heterogeneity L1 isochore present in region 2 (these features are also seen in Fig. 1A–C); (2) the isochores from H1 (region 4) and H2 (region 5) families that consist of multiple-peaks that emerge from common compositional bases; (3) a series of extremely sharp peaks ranging from 40–45% to 55–60% GC (region 6); these peaks, previously visualized (see Fig. 1B,C) as a series of alternating H2 and H3 isochores (because the compositional approach used involved merging into isochores 100Kb segments comprised within the ranges of Supplementary Table S1) should now be called “H3 peaks” since they cannot be strictly defined as H3 isochores not being “fairly homogeneous” sequences.
Figure 1D as displayed at a higher magnification. Thick horizontal lines correspond to minima GC values between isochore families. The color code is as in Fig. 1. (A) Region 1 comprises a (rare) H1 single peak (a) and two L2 peaks (b,c). Region 3 comprises three L2 peaks (d–f); peak d may be seen as a (rare) L1 single peak if the “extended” GC-range of isochores is used (see Supplementary Table S1). (B) Regions 4 and 5 comprise the multi-peak H1 and H2 isochores respectively. (C) Region 6 comprises H3 peaks, region 2 corresponds to a compositionally even L1 isochore.
It so happens that, in contrast with other chromosomes, chromosome 21 barely comprises homogeneous L2 sequences (shown in Supplementary Fig. S2 and in ref.16) that define an L2− sub-family, as distinct from an L2+ sub-family formed by the L2 single peaks of regions 1 and 3 (these peaks are more frequent in chromosome 21 than in other chromosomes). Indeed, only one example of an L2− isochore is present in chromosome 21, the Y “valley” isochore (see Figs 1D and 2). Moreover, upon close inspection, the peaks of regions 4 to 6 were seen to correspond to the minute peaks of Fig. 1C. This observation is very relevant in that it allows generalizing the present results to all human chromosomes as investigated by the sliding window approach.
In conclusion, the point-by-point approach has revealed (1) a low-heterogeneity L1 sequence; (2) several L2+ single peaks (in addition to a rare H1 single peak; see legend of Fig. 2); (3) the multi-peak structures of H1 and H2 isochores; and (4) the H3 peaks.
Oligo-nucleotides and isochores
The compositional and topological properties of isochores will now be considered in more detail. Using previous data22, one can assess not only the levels of AAA/TTT, GGG/CCC, but also those of “A/T-only” and “G/C-only” trinucleotides, in isochore families. The results for L1 and H3 isochores, presented in Supplementary Table S2, indicate that specific sets of trinucleotides have widely different distributions in different isochore families. Indeed, in L1 isochores (35.5% GC) the “A/T-only” trinucleotides represent 26% of the sequences, with AAA/TTT corresponding to 9% and consisting of a series of blocks that may extend to tetra- to octa-A’s23. In contrast, the L2+, H1, H2 and H3 peaks are characterized by gradients of increasing GC and increasing frequencies of oligo-G’s. In the “classical” H3 isochores (55% GC), “G/C-only” tri-nucleotides represent 15.4% of the sequences with GGG/CCC corresponding to 6.2% and extending to tetra- to octa-Gs23.
One should now consider that the interspersed oligo-A’s and oligo-G’s are intrinsically stiff for different structural reasons24 and that, specifically, oligo-A’s are intrinsically curved25. Two different classes of isochores can, therefore, be distinguished from a topological viewpoint: (1) the L1 (and, probably L2−) isochores characterized by low compositional heterogeneity that are locally stiff because of the presence of interspersed oligo A’s; and (2) the L2+ and H3 single-peaks and the H1/H2 multi-peaks that are characterized by a high compositional heterogeneity, by increasing densities of interspersed oligo-G’s (as well as of CpG’s and CpG islands26), and by increasing densities of oligo-A’s in the troughs of H3 peaks. In conclusion, the A- and G-oligonucleotides are responsible for specific topological differences in GC-poor and GC-rich isochores. Expectedly, “A/T-only” and “G/C-only” sequences also contribute to such topological differences23.
Isochores and nucleosomes
A crucial point now is to consider that the features just described that are responsible for the topology of GC-poor and GC-rich isochores, namely the presence of interspersed oligo-As and oligo-Gs (as well as of CpGs and CpG islands) are strongly inhibitory to nucleosome formation24,25,26. It should also be noted that the 3-D structures of GC-poor and GC-rich isochores appear after nucleosomes dissociate from the parental DNA strands and before they re-associate (along with neo-synthesized nucleosomes) on the daughter strands (see Supplementary Fig. S3A and refs27,28).
Isochores and chromatin domains
Yet another, critical, way to classify isochores can be based on their correspondence with chromatin domains. This also leads to visualize the same two groups of isochores. Indeed, the L1 and L2− isochores (that include some “valley” isochores such as X and Y of Figs 1D and 2) correspond to LADs, while the single-peak and the multi-peak isochores of the other families correspond to interLADs (single-loop and multi-loop TADs, respectively; see Supplementary Fig. S2). In the case of L2 isochores, the existence of two sub-families L2− and L2+ is confirmed by their correspondence with LADs and interLADs (i.e., TADs), respectively (see Supplementary Fig. S2).
A model for the formation of chromatin domains
The results presented so far allow building models for the formation of LADs and TADs. In the case of LADs (Fig. 3A), the presence of interspersed oligo-A’s that causes local DNA stiffness and bending also causes local nucleosome depletions that induce LADs to assume a structure which is well suited to attach to (and even embed in) the lamina29. This structure can also be visualized as the result of a “moulding”of LADs by the short-sequence designs of L1 and L2− isochores.
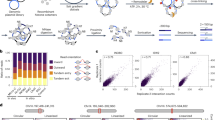
The proposed models for the formation of LADs and TADs. (A) A representation of a chromatin fibre corresponding to a GC-poor isochore (a blue bar with yellow oligo-Adenines blocks is delimited by CTCF binding sites, green boxes). Topologically, the structure of the chromatin fibre is modeled by intrinsically stiff and curved oligo-Adenines that cause local bending, twisting and nucleosome depletions; this chromatin structure favors its 3-D physical adaptation to the lamina and may be considered as the result of a “moulding” of LADs by the DNA sequences. (B) A compositional representation of a chromatin fibre corresponding to a GC-rich peak; the yellow to red color gradient indicates the corresponding gradients of GC, oligo-Guanines, CpG’s and CpG islands that alternate with the AT gradients of the oligo-Adenines present in the troughs23 both gradients being responsible for the decreasing nucleosome density, decreasing supercoiling and increasing accessibility as well as for the looping; this “moulding step” by the DNA sequence is followed by: (C) the binding of CTCF and cohesin to the peaks of the loops and by a “sliding/extrusion step” down the loops (helped by the cohesin ATP-ase) which ends when the CTCF/cohesin complex reaches the CTCF binding sites at the base of the loops; this process leads to approaching the two branches of the loop, creating self-interactions and closing and insulating the loop. The broken arrow indicates that the depletion of CTCF or cohesin reverses TADs into “primary TADs”, that still are transcriptionally functional31,32,33.
In the case of TADs, their formation appears to take place through a two-step mechanism. In a critical first step, the increasing oligo-Gs, CpGs and CpG islands of the peaks are responsible for a decreasing nucleosome density25,26, a decreasing supercoiling30 (and an increasing nuclease accessibility), that constrain chromatin to fold into loops, the tips of the loops corresponding to the highest GC levels and to the lowest nucleosome density. Incidentally, the nucleosome-depleted troughs between the peaks due to the increasing presence of oligo-Adenines help the folding of the chromatin fibre in the valleys between the H3 peaks. This initial “moulding step”, which may involve essentially equal-size chromatin branches (as suggested by the symmetry of GC peaks; see Figs 1D and 2), leads to CTCF/cohesin-free “primary TADs” (Fig. 3B), that probably are already functional since cohesin and CTCF depletions are not followed by any remarkable alteration of transcription as recently shown31,32,33. Interestingly, a wider nucleosome spacing and higher flexibility of the loops compared to boundary regions was proposed by the “insulation/attraction model”17.
Two points should now be raised about the “moulding step”. The first one concerns which one of the two factors, the 3-D structure of isochores or the nucleosome density, is primarily responsible for moulding. The demonstration (see Supplementary Fig. S3B) that TADs are not very different in fibroblasts and in spermatozoa34 (in which case nucleosomes are replaced by protamines) provides an answer in favor of the first possibility. The second point is that the existence of “primary TADs” helps to solve the notorious open problem of the energy source required by the extrusion models35,36, that involve pulling megabase-size chromatin fibres through the CTCF/cohesin ring. Indeed, the “marginal” energy burden proposed in a very recent article37 essentially rests on indirect evidence, namely on the fact that an ATP hydrolysis dependent molecular motor was recently identified in condensin and shown to use 2ATP per sec per condensin38.
In a second step (Fig. 3C), one may consider that the CTCF/cohesin ring complex binds to the peaks of the loops and starts an “extrusion step” down the “primary TADs”. In this connection, it is of great interest to consider the recent proposal39 that Nipbl, an essential factor for loading cohesin on the loops, may also stimulate cohesin translocation down the loops. The ATPase activity associated with cohesin sub-units Smc1 and Smc3, also stimulated by Nipbl40, would provide the energy necessary for sliding, an energy obviously far below the level required by “active extrusion”. This sliding step leads to closer contacts between the two branches of the loops and to self-interactions and stops at the CTCF binding sites, that are located at the base of the loops, thus closing and insulating them. Incidentally, the possibility that extrusion could also be mediated by a single cohesin ring sliding over the top of a preformed chromatin loop was taken into consideration previously41.
Conclusions
The present investigations led to (1) the discovery of the 3D topology of isochores; (2) the discovery that this topology is due to short sequences, such as oligo-Adenines and oligo-Guanines, as well as to a number of other short sequences23; and (3) the discovery that the 3D topology of isochores is responsible for the “moulding step” of chromatin domains. Incidentally, the roots of the first two conclusions go very back in time2,3.
These conclusions represent a paradigm shift compared to current models that only rest on architectural proteins, such as CTCF and cohesin, neglecting, as also remarked by other authors42, the role played by DNA. Indeed, they introduce a primary “moulding step” based on isochore topology which precedes the “extrusion step” and makes it easier in terms of the energy required.
These investigations, that only dealt with the evolutionary conserved features of chromatin architecture, also led to a new vision of the “genomic code”, a definition which was originally coined43,44 (see also ref.11 for a review) for the compositional correlations that hold between coding and contiguous non-coding sequences and among the three codon positions. In fact, the “genomic code” should now be visualized (G. Bernardi, paper in preparation) as (1) a short-sequence code for the 3D topology of isochores and the moulding of chromatin domains; (2) a pervasive code since it not only concerns the ~98% of non-coding sequences, leaving little or no room for “junk DNA” (see ref.45 for a review), but also constrains the genetic code in terms of composition of coding sequences, codon usage and aminoacid choice11.
Materials and Methods
The compositional profile displayed in Fig. 1A (from ref.46) concerns DNA sequences derived from release hg 38. Non-overlapping 100 Kb windows were used because 100Kb is a plateau value under which the composition of DNA segments shows an increasing variance with decreasing size, due to the contribution of different specific sequences, such as interspersed repeated sequences10. Figure 1B shows an isochore profile obtained by using a sliding window approach16. Figure 1B,C are from the matched b37 assembly of ref.47; these profiles are slightly more extended on the centromeric (left) side than that of Fig. 1A.
Nomenclature
Although TADs comprise, by definition, all the topologically associating domains, in the present context TADs indicate the chromatin domains other than LADs. The main reason for this choice is that the mechanisms of formation of the two sets of domains are different, even if based on the same DNA properties, 3-D structure of isochores and nucleosome binding.
Data Availability
All data generated or analyzed during this study are included in this published article (and its Supplementary Information files).
References
Hudson, A. P., Cuny, G., Cortadas, J., Haschemeyer, E. V. & Bernardi, G. An analysis of fish genomes by density gradient centrifugation. Eur. J. Biochem. 112, 203–210 (1980).
Corneo, G., Ginelli, E., Soave, C. & Bernardi, G. Isolation and characterization of mouse and guinea pig satellite deoxyribonucleic acids. Biochemistry 7, 4373–4379 (1968).
Bernardi, G., Ehrlich, S. D. & Thiery, J. P. The specificity of Deoxyribonucleases and their use in nucleotide sequence studies. Nature New Biology 246, 36–40 (1973).
Filipski, J., Macaya, G. & Bernardi, G. An analysis of the bovine genome by Cs2SO4-Ag density gradient centrifugation. J Mol Biol 80, 177–197 (1973).
Thiery, J. P., Macaya, G. & Bernardi, G. An analysis of eukaryotic genomes by density gradient centrifugation. J. Mol. Biol. 108, 219–235 (1976).
Macaya, G., Thiery, J. P. & Bernardi, G. An approach to the organization of eukaryotic genomes at a macromolecular level. J Mol Biol 108, 237–254 (1976).
Cuny, G., Soriano, P., Macaya, G. & Bernardi, G. The major components of the mouse and human genomes. 1. Preparation, basic properties and compositional heterogeneity. Eur. J. Biochem. 115, 227–233 (1981).
Saccone, S., Pavlicek, A., Federico, C., Paces, J. & Bernardi, G. Genes, isochores and bands in human chromosomes 21 and 22. Chromosome Res. 9, 533–539 (2001).
Bernardi, G. Misunderstandings about isochores. Part 1. Gene 276, 3–13 (2001).
Costantini, M., Clay, O., Auletta, F. & Bernardi, G. An isochore map of human chromosomes. Genome Res 16, 536–541 (2006).
Bernardi, G. Structural and Evolutionary Genomics. Natural Selection in Genome Evolution. Elsevier, Amsterdam. 2004 This out-of-print book is freely available at, www.giorgiobernardi.eu (reprinted in 2005).
Eyre-Walker, A. & Hurst, L. The evolution of isochores. Nature Rev Genet 2, 549–555 (2001).
Bernardi, G. et al. The mosaic genome of warm-blooded vertebrates. Science 228, 953–958 (1985).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Saccone, S., Federico, C. & Bernardi, G. Localization of the gene-richest and the gene-poorest isochores in the interphase nuclei of mammals and birds. Gene 300, 169–78 (2002).
Jabbari, K., Bernardi, G. An isochore framework underlies chromatin architecture. PloS One, https://doi.org/10.1371/journal.pone.0168023 (2017).
Dixon, J. R., Gorkin, D. U. & Ren, B. Chromatin domains: the unit of chromosome organization. Mol Cell 62, 668–680 (2016).
van Steensel, B. & Belmont, A. Lamina-Associated Domains: links with Chromosome Architecture, Heterochromatin, and Gene Repression. Cell 169, 780–791 (2017).
Dixon, J. R., Selvaraj, S., Yue, F., Kim, A. & Li, Y. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Jabbari, K. & Bernardi, G. The distribution of genes in the Drosophila genome. Gene 247, 287–292 (2000).
Labena, A. A., Guo, H., Dong, C., Li, L. & Guo, F. The topologically associated domains (TADs) of a chromatin correlated with isochores organization of a genome. Current Bioinformatics 13 (in press, 2018).
Costantini, M. & Bernardi, G. The short-sequence designs of isochores from the human genome. Proc Natl Acad Sci USA 105, 13971–13976 (2008).
Lamolle, G., Sabbia, V., Musto, H. & Bernardi, G. The short sequence design of DNA and its involvement in the 3-D structure of the genome. Sci. Rep. (in press).
Struhl, K. & Segal, E. Determinants of nucleosome positioning. Nature Struct and Mol Biol 20, 267–273 (2013).
Barbic, A., Zimmer, D. P. & Crothers, D. M. Structural origins of adenine-tract bending. PNAS 100, 2369–2373 (2003).
Fenouil, R. et al. CpG islands and GC content dictate nucleosome depletion in a transcription-independent manner at mammalian promoters. Genome Res 22, 2399–2408 (2012).
Tagami, H., Ray-Gallet, D., Almouzni, G. & Nakatani, Y. Histone H3.1 and H3.3 complexes mediate nucleosome assembly pathways dependent or independent of DNA synthesis. Cell 116, 51–61 (2004).
Allison, L. A., DNA replication and telomere maintenance in Fundamental Molecular Biology 135 Wiley (2011).
Kind, J. et al. Genome-wide Maps of Nuclear Lamina Interactions in Single Human Cells. Cell 163, 134–147 (2015).
Naughton, C. et al. Transcription forms and remodels supercoiling domains unfolding large-scale chromatin structures. Nat Struct Mol Biol 20, 387–395 (2013).
Schwarzer, W. et al. Two independent modes of chromatin organization revealed by cohesin removal. Nature 551, 51–56 (2017).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320 (2017).
Nora, E. P. et al. Targeted degradation of CTCF decouples local insulation of chromosome domains from genomic compartmentalization. Cell 169, 930–944 (2017).
Battulin, N. et al. Comparison of the three-dimensional organization of sperm and fibroblast genomes using the Hi-C approach. Genome Biology 16, 77–91 (2015).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. CellReports 15, 2038–2049 (2016).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc Natl Acad Sci USA 112, 6456–6465 (2015).
Fudenberg, G., Abdennur, N., Imakaev, M., Goloborodko, A., Mirny, L. A. Emerging evidence of chromosome folding by loop extrusion. Cold Spring Harb Symp Quant Biol (2018).
Terakawa, T. et al. The condensin complex is a mechanochemical motor that translocates along DNA. Science 358, 672–676 (2017).
Rhodes, S., Mazza, D., Nasmyth, K. & Uphoff, S. Scc2/Nipbl hops between chromosomal cohesin rings after loading. eLife 6, 30000–30020 (2017).
Murayama, Y. & Uhlmann, F. Biochemical reconstitution of topological DNA binding by the cohesin ring. Nature 505, 367–371 (2014).
Merkenschlager, M. & Nora, E. P. CTCF and cohesin in genome folding and transcriptional gene regulation. Annu Rev Genomics Hum Genet 17, 17–43 (2016).
Todolli, S., Perez, P. J., Clauvelin, N. & Olson, W. K. Contributions of sequence to the higher-order structures of DNA. Biophys J 112, 416–426 (2017).
Bernardi, G. Le Génome des Vertébrés: Organisation, Fonction et Evolution. Biofutur 94, 43–46 (1990).
Bernardi, G. & Bernardi, G. Compositional properties of nuclear genes from cold-blooded vertebrates. J Mol Evol 33, 57–67 (1991).
Palazzo, A. F. & Gregory, T. R. The case for junk DNA. PLOS Genetics 10, 1–8 (2014).
Cozzi, P., Milanesi, L. & Bernardi, G. Segmenting the human genome into isochores. Evol Bio Informa 11, 253–261 (2015).
Rao, S. S. P. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Acknowledgements
The author thanks Paolo Ascenzi for hospitality, Giacomo Bernardi (Santa Cruz, California, USA), Oliver Clay (Medellin, Colombia), Paolo Cozzi (Milan, Italy) and, especially, Kamel Jabbari (Cologne, Germany) for critical reading, comments and discussions. Caterina Nuvoli and Marta Ritucci provided excellent technical help. This research was supported by the Kimura Prize for Molecular Evolution and Evolutionary Genomics conferred to the author (Tokyo, June 2016).
Author information
Authors and Affiliations
Contributions
Giorgio Bernardi wrote the manuscript and prepared the figures.
Corresponding author
Ethics declarations
Competing Interests
The author declares no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bernardi, G. The formation of chromatin domains involves a primary step based on the 3-D structure of DNA. Sci Rep 8, 17821 (2018). https://doi.org/10.1038/s41598-018-35851-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-35851-0
This article is cited by
-
Mechanical Pressure Driving Proteoglycan Expression in Mammographic Density: a Self-perpetuating Cycle?
Journal of Mammary Gland Biology and Neoplasia (2021)
-
DNA sequence-dependent chromatin architecture and nuclear hubs formation
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.